Abstract
In order to study the properties of a thermostable uricase produced by Microbacterium sp. strain ZZJ4-1, the enzyme was purified by ammonium sulfate precipitation and DEAE-cellulose ion exchange, hydrophobic and molecular sieve chromatography. The molecular mass of the purified enzyme was estimated to be 34 kDa by SDS-PAGE. The enzyme was stable between pH 7.0 and 10.00. The optimal reaction temperature of the enzyme was 30 °C at pH 8.5. The K m and K cat of the enzyme were 0.31 mM and 3.01 s−1, respectively. Fe3+ could enhance the enzyme activity, whereas Ag+, Hg2+, o-phenanthroline and SDS inhibited the activity of the enzyme considerably. After purification, the enzyme was purified 19.7-fold with 31% yield. As compared with uricases from other microbial sources, the purified enzyme showed excellent thermostability and other unique characteristics. The results of this work showed that strains of Microbacterium could be candidates for the production of a thermostable uricase, which has the potential clinical application in measurement of uric acid.
Similar content being viewed by others

Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Uricases (urate oxidoreductase; EC 1.7.3.3), which catalyse the oxidative breakdown of uric acid, belong to a group of enzymes in the purine degradation pathway found in animals (Keilin 1959; Wallrath and Friedman 1991), plants (Montalbini et al. 1997), fungi (Montalbini et al. 1999), yeasts (Adamek et al. 1990; Hongoh et al. 2000; Koyama et al. 1996) and bacteria (Yamamoto et al. 1996). Since Bongaert et al. (Bongaert et al. 1978) found that Bacillus fastidiosus could produce uricase and use uric acid as the only carbon source for growth in 1978, many reports on bacterial and yeast uricases, such as those from Arthrobacter globiformis (Nobutoshi et al. 2000), Bacillus subtilis (Hunag and Wu 2004), Candida utilis (Liu et al. 1994) and Pseudomonas aeruginosa (Ishikawa et al. 2004), have been published.
In the human body, uric acid is a final product of purine catabolism and is excreted out of the body by the kidney. When the level of uric acid in blood increases over the normal value, it can lead to a group of diseases, such as gout (Nakagawa et al. 2006), idiopathic calcium urate nephrolithiasis (Masseoud et al. 2005) and renal failure (Capasso et al. 2005), and it was also reported that a high level of uric acid was related to leukemia in children (Larsen & Loghman-Adham 1996). Consequently, uric acid concentration is an important parameter monitored in urine and blood samples in routine clinical examinations. At present, the colorimetric method that employs uricase and peroxidase is widely accepted as a simple, sensitive and highly specific test for uric acid examination. In this enzymatic system of uric acid analysis, uricase plays an important role:
By determining the amount of peroxide formed in the reaction, the concentration of uric acid can be estimated. For the best result in this examination, the properties of the uricase in this reaction system are important, especially its stability, which will determine the precision of the measurement. At present, most enzymes applied in the clinical test are used in solution, and most proteins, including this enzyme, are relatively unstable when dissolved in aqueous solution. Therefore, for the enzymatic examination application, research was undertaken to search for a thermostable enzyme (Guo et al. 2006; Huang et al. 1998; Zhou et al. 2005).
As mentioned above, there were many uricases that have been isolated from microorganisms, but the thermostability of the published uricases were relatively low (Suzuki et al. 2004). The most thermostable uricase was reported by Suzuki, but it would lose its activity after a short period of treatment at 60 °C (Suzuki et al. 2004). This low stability is a disadvantage in clinical applications.
In a previous study, we isolated a bacterium Microbacterium sp. strain ZZJ4-1 that produced a thermostable uricase. The enzyme was stable at 65 °C and its solution retained its original activity even after storage at 37 °C for 40 days (Zhou et al. 2005). Considering that this enzyme has a potential value in practical application, the present study was undertaken to purify and study the properties of this new enzyme.
Materials and methods
Materials and chemicals
The culture of Microbaterium sp. (strain ZZJ4-1) was maintained in our laboratory (Zhou et al. 2005).
The protein standards for SDS-PAGE were purchased from Invitrogen (Shanghai, China). All other chemicals used were of reagent or molecular biology grade and purchased from Hangzhou Huadong Medicine Group Co., Ltd (Hangzhou, China).
Cultivation conditions
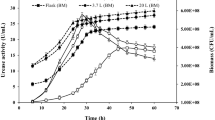
The fermentation medium consisted of 3.0 g of uric acid, 10.0 g of maize milk, 0.5 g of MgSO4 · 7H2O, 0.5 g of KH2PO4, 2.0 g of K2HPO4 · 3H2O, 0.1 g of NaCl, 1.0 l of tap water and the pH was adjusted to 7.5 (Zhou et al. 2005). To cultivate the strain for production of the uricase, a loop of bacteria from a slant was inoculated into a 500 ml flask containing 100 ml liquid medium and incubated at 30 °C for 30 h with a rotary shaker at 120 rev/min.
Enzyme assay and protein measurement
The principle of enzyme measurement was as follows: uricase can catalyse the oxidation of uric acid to form 5-hydroxyisourate and H2O2, which is then measured using a reaction system containing 4-aminoantipyrine, phenol and peroxidase as chromogens. In practical analysis, 0.10 ml enzyme solution was incubated with a mixture of 0.6 ml 0.1 M sodium borate buffer (pH 8.5) containing 2 mM uric acid, 0.15 ml of 30 mM 4-aminoantipyrine, 0.1 ml of 1.5% phenol and 0.05 ml peroxidase (15 U/ml) at 37 °C for 20 min (Masaru 1981). The reaction was stopped by addition of 1.0 ml ethanol and the absorbance at 540 nm was read against the blank in a spectrophotometer. One unit of enzyme was defined as the amount of enzyme that produces 1.0 μmole of H2O2 per minute under the standard assay conditions.
The protein was measured by the Folin-phenol method (Lowry et al. 1951).
Enzyme purification
Unless otherwise stated, all of the enzyme purification processes were performed with 0.1 M phosphate buffer at pH 7.0 (buffer A) at 4 °C.
Cells from 10.0 l of media were harvested by centrifugation and washed twice with 50 mM phosphate buffer (pH 7.5), re-suspended in buffer A and disrupted by ultrasonic oscillation (120 W oscillating for 3 s with 6 s intervals, repeated 100 times). After the cell debris had been separated by centrifugation, solid ammonium sulfate was added to the enzyme solution and the precipitate of the fractions from 55% to 80% saturation was collected by centrifugation (8000 rpm, 40 min, 4 °C). The enzyme was then dissolved in a small amount of buffer (2:1, volume of the buffer/weight of the precipitate), dialyzed against 0.01 M phosphate buffer until the ammonium sulfate was removed.
The dialyzed enzyme solution was put onto a DEAE-Cellulose column (5.5 × 50 cm) previously equilibrated with buffer A and it was then eluted with 2.0 l of buffer A with a linear gradient of 0–1.5 M KCl (flow rate: 2 ml/min). The fractions containing enzyme activity were collected. After the ammonium sulfate had been added to 65% saturation, the enzyme solution was loaded onto a Toyopearl HW-65 column (5.5 × 85 cm) equilibrated previously with buffer A containing 65% ammonium sulfate. The enzyme was eluted with 2.0 l of the same buffer with a linear gradient of ammonium sulfate from 65% to 0% (flow rate: 2 ml/min). When the fractions containing enzyme activity had been collected, the ammonium sulfate was added to get 85% saturation. The precipitate of the enzyme was then collected by centrifugation, dissolved in a small amount of buffer A, dialysed with the same buffer containing 0.1 M KCl and applied to a Sephadex G-75 column (5.5 × 100 cm) equilibrated with the same buffer. The enzyme was then eluted with 2.0 l of the same buffer (flow rate: 2 ml/min) and the fractions containing the highest specific activity were collected for further study.
Characterization of enzyme
To study the effect of pH on the activity of uricase, the enzyme was assayed at different pHs in the range from 4.0 to 11.0 with intervals of 0.5. The following buffers were used: 100 mM citrate for pH 4.0–6.0, 100 mM phosphate for pH 6.0–8.5 and 100 mM borate for pH 8.5–11. The pH stability was studied by incubating the purified enzyme solution in the corresponding buffers in the range from 4.0 to 11.0 at 25 °C for 18 h and measuring the residual activity.
To study the effect of temperature on the uricase activity, the standard enzyme reaction solution was pre-incubated at temperatures of 20–60 °C with 5 °C intervals for 5 min and the enzyme solution was then added and incubated for 20 min at the same temperature to measure its activity. For thermostability testing, the purified uricase solution was incubated at temperatures in the range from 20 °C to 80 °C for 30 min and the remaining activity was then measured.
The apparent K m of the uricase was estimated by the double reciprocal plot method. At different concentrations of uric acid, the enzyme activity was assayed and the Km was calculated by the Lineweaver-Burk plotting according to the Michaelis-Menten equation.
To study the effects of chemicals on the uricase activity, the enzyme solution was pre-incubated with the chemicals for 30 min at room temperature in phosphate buffer and the remaining uricase activity was assayed with the standard reaction system containing the corresponding chemical.
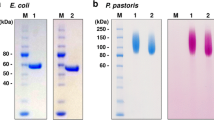
The relative molecular mass of the enzyme was determined by SDS-PAGE with 10% polyacrylamide gels (Laemmli 1970). The purity was determined by the specific activity of the enzyme and PAGE with 10% polyacrylamide gels. The protein was stained with 0.1% Coomassie brilliant blue R 250 in 4:1:5 methanol/acetic acid/water (vol/vol/vol) solution and destained in the same solution without dye.
Results
Purification of the enzyme
In the first step of the purification, ammonium sulfate precipitation was applied. The amount of protein and the enzyme activity of each fraction were measured. The fractions with ammonium sulfate concentrations from 65% to 80% had the highest enzyme specific activity (0.43 U/mg), while fractions from 35–55, 55–65 to 80–85 had the specific activity of 0.02 U/mg, 0.09 U/mg and 0.07 U/mg, respectively. The fractions with concentrations from 65% to 80% were collected, dialyzed and loaded onto a DEAE-cellulose column. The enzyme was eluted with the same buffer containing a linear concentration gradient of KCl from 0 M to 1.5 M. Enzyme activity was found in fractions from 75 to 130 and fractions from 80 to 105 were pooled, solid ammonium sulfate was added to 65% saturation and the solution was loaded onto a Toyopearl HW 65-C column. The enzyme was then eluted with the same buffer with a decreasing linear concentration of ammonium sulfate from 65% to 0%. The enzyme activity was found in the fractions from 100 to 160 and fractions from 110 to 135 were pooled, concentrated and applied to the Sephadex G-75 column. After being washed with buffer A containing 0.1 M NaCl, the enzyme activity was observed in the fractions from 30 to 80. The fractions from 50 to 60 were pooled as purified enzyme for further study.
The purity of the enzyme after each purification step was examined by SDS-PAGE and is shown in Figure 1. The results of the purification process are summarized in Table 1 . During the purification, the enzyme was purified 19.7-fold with a recovery of 31% and the purified enzyme had a specific activity of 5.32 U mg-1.
Molecular mass determination
The purified enzyme showed a single protein band in SDS-PAGE and its molecular mass was estimated to be 34 kDa (Fig. 2).
Optimum reaction temperature and thermostability of the purified enzyme
The purified enzyme was stable at a relative high temperature. As shown in Fig. 3, it was stable at 65 °C and it retained 64% of its original activity even after being treated at 70 °C for 30 min. By comparison between the optimal reaction and stable temperature, it was shown that although the enzyme was stable at 65 °C, its optimum temperature was 30°C, which was relatively low, and it only showed 21% relative activity at 60 °C.
Effects of temperature on uricase activity and stability (△) The relative activity of the purified enzyme, measured in different reaction temperatures. The relative activity of the uricase in 30 °C reaction system was defined as 100%. (•) The percentage of the residual activity, after treatment for 30 min at different temperatures in sodium borate pH 8.5 buffer. The residual activity of uricase after 30 min at 45 °C was defined as 100%
Optimum pH and the stability of the enzyme at different pHs
The activity of the enzyme was measured in different buffers with a pH range from 4.0 to 11.0. The results showed that the enzyme had low activity at pH below 5.5 or over 10.5 and had relatively high activity in the range from 7.0 to 10.0, with the optimal reaction pH at 8.5 (Fig. 4). As shown in Fig. 4, after being incubated in different buffers at 25 °C for 18 h, the uricase was stable in the pH range from 5.5 to 9.5.
Kinetics and effect of chemicals on the activity of the enzyme
The K m of the purified enzyme for uric acid was estimated to be 0.31 mM, which was calculated from the slopes and intercepts of the regression lines of the Lineweaver-Burk plot by determining the enzyme activity at 37 °C and the K cat of the enzyme was estimated to be 3.01 s−1.
The effects of different chemicals on the activity of the enzyme are summarized in Table 2. It was shown that among the metal ions, Li+, Ag+ and Hg+ greatly inhibited the enzyme activity. The strongest suppression was observed in the case of Hg+, which suppressed almost all of the activity of this enzyme. The chelating reagents had different effects; EDTA had no inhibitory effect, whereas o-phenanthroline (OPT) could inhibit the activity of uricase considerably.
Discussion
At present, the uricases from many microorganisms have been studied and some gene sequences of uricases have been cloned and studied. It was shown that uricases belong to a group of enzymes that have the same catalytic character, but a great diversity of molecular structures. Uricases from different sources may have different molecular masses and amino acid sequences. In this study, the molecular mass of the uricase from strain ZZJ4-1 was estimated to be 34 kDa by SDS-PAGE, whereas uricases produced by Candida utilis (Koyama et al. 1996) and Pseudomonas aeruginosa (Ishikawa et al. 2004) had the molecular masses of 34 and 54 kDa, respectively. The apparent K m value of this uricase was 0.31 mM, while uricases from Arthrobacter, Bacillus sp. and Candida sp. had K m values of 75, 75, 46 μM, respectively (Suzuki et al. 2004).
Some relatively thermostable uricases, such as those produced by Arthrobacter globiformis FERM BP-360, Bacillus sp.TB-90 and Candida utilis, have been studied and these were stable at 55 or 60 °C (Suzuki et al. 2004). In this work, the uricase produced by strain ZZJ4-1 showed good thermostability. After incubation at 65 °C for 30 min, the enzyme from strain ZZJ4-1 still retained 98.9% of its original activity, and even after incubation at 70 °C for 30 min, the remaining activity was 64%. Additionally, the pH stability of the enzyme from strain ZZJ4-1 was a little broader than that of the uricases from the above three species (Suzuki et al. 2004).
The effects of metal ions on the enzyme activity from strain ZZJ4-1 were also compared with the above three enzymes (Suzuki et al. 2004). Considering that some uricases require metal ion cofactors for activity (Wu et al. 1989; Chu et al. 1996) and whether or not an enzyme is inhibited by certain metals is an important characteristic, experiments were carried out to examine the effect of metals on the activity of this enzyme. It was shown that the enzyme from strain ZZJ4-1 was not inhibited by Cu2+, Fe3+ or Zn2+, while uricases from the other three strains were strongly inhibited by these ions. Ag+ is a strong inhibitor of the uricases from strain ZZJ4-1, Candida utilis and Bacillus sp. TB-90, while it had no such effect on the uricase from Arthrobacter globiformis FERM BP-360. When EDTA, a chelating reagent, was added to the enzyme solution at a final concentration of 20 mM, the activity of the uricase was only slightly inhibited. When the stronger metal-ion-chelating reagent o-phenanthroline was added to the enzyme solution, the activity of the enzyme was strongly inhibited. This phenomenon indicates that some yet unidentified metal ion is strongly bound in the enzyme and forms part of the uricase structure, which is very important to keep its catalytic activity. This property is also different from the uricase of Arthrobacter globiformis FERM BP-360 (Suzuki et al. 2004).
It was shown that although the uricase from the strain ZZJ4-1 was stable at 65 °C, the enzyme was at its optimal activity at 30 °C and it only showed 21% of its maximal activity at 60 °C. This implied that the molecular structure of enzyme had a reversible change at a temperature between 30 ° and 60 °C, which had a negative effect on its catalysis activity. But considering that most clinical enzymatic examinations are undertaken at 25 to 37°C, this property will not affect its application in clinical examination.
References
Adamek V, Suchova M, Demnerova K et al (1990) Fermentation of Candida utilis for uricase production. J Indu Microbiol 6:85–90
Bongaerts GP, Uitzetter J, Brouns R et al (1978) Uricase of Bacillus fastidiosus. Properties and regulation of synthesis. Biochim Biophys Acta 527:348–358
Capasso G, Jaeger Ph, Robertson WG et al (2005) Uric acid and the kidney: urate transport, S tone disease and progressive renal failure. Curr Pharm Des 11:4153–4159
Chu R, Lin Y, Usuda N, Rao MS et al (1996) Mutational analysis of the putative copperbinding site of rat urate oxidase. Ann NY Acad Sci 804:777–780
Guo K, Ma X, Sun G et al (2006) Expression and characterization of a thermostable sarcosine oxidase (SOX) from Bacillus sp. in Escherichia coli. Appl Microbiol Biotechnol 73:559–566
Hongoh Y, Sasaki T, Ishikawa H (2000) Cloning, sequence analysis and expression in Escherichia coli of the gene encoding a uricase from the yeast-like symbiont of the brown planthopper, Nilaparvata lugens. Insect Biochem Mol Biol 30:173–182
Huang HS, Kabashima T, Ito K et al (1998) Thermostable glycerol kinase from Thermus flavus: cloning, sequencing, and expression of the enzyme gene. Biochim Biophys Acta 1382:186–190
Hunag S, Wu T (2004) Modified colorimetric assay for uricase activity and a screen for mutant Bacillus subtilis uricase genes following StEP mutagenesis. Eur J Biochem 271:517–523
Ishikawa J, Yamashita A, Mikami Y et al (2004) The complete genomic sequence of Nocardia farcinica IFM 10152. Proc Natl Acad Sci USA 101:14925–14930
Keilin J (1959) The biological significance of uric acid and guanine excretion. Biol Rev 34:265–296
Koyama Y, Ichikawa T, Nakano E (1996) Cloning, sequence analysis, and expression in Escherichia coli of the gene encoding the Candida utilis urate oxidase (uricase). J Biochem (Tokyo), 120:969–973
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Larsen G, Loghman-Adham M (1996) Acute renal failure with hyperuricemia as initial presentation of leukemia in children. J Pediatr Hematol Oncol 18:191–194
Liu J, Li G, Liu H et al (1994) Purification and properties of uricase from Candida sp. and its application in uric acid analysis in serum. Appl Biochem Biotechnol 47:57–63
Lowry OH, Rosebrough NJ, Farr AL et al (1951) Protein measurement with the Folin Phenol reagent. J Biol Chem 193:265–275
Masaru P (1981) Purification and some properties of Sarcosine Oxidase from Corynebacterium sp. U-96. J Biochem 89:599–607
Masseoud D, Rott K, Liu-Bryan R et al (2005) Overview of hyperuricaemia and gout. Curr Pharm Des 11:4117–4124
Montalbini P, Aguilar M, Ineda M (1999) Isolation and characterization of uricase from bean leaves and its comparison with uredospore enzyme. Plant Sci 147:139–147
Montalbini P, Redondo J, Caballero JL (1997) Uricase from leaves: its purification and characterization from three different higher plants. Planta 202:277–283
Nakagawa T, Mazzali M, Kang DH et al (2006) Uric acid - a uremic toxin? Blood Purif 24:67–70
Nobutoshi K, Keisuke S, Takao M et al (2000) Determination of uric acid in plasma by closed-loop fia with a coimmobilized enzyme flow cell. Anal Sci 16:1203–1205
Suzuki K, Sakasegawa SI, Misaki H et al (2004) Molecular cloning and expression of uricase gene from Arthrobacter globiformis in Escherichia coli and characterization of the gene product. J Biosci Bioeng 98:153–158
Wallrath LL, Friedman TB (1991) Species differences in the temporal pattern of Drosophila urate oxidase gene expression are attributed to trans-acting regulatory changes. Proc Natl Acad Sci USA 88:5489–5493
Wu XW, Lee CC, Muzny DM et al (1989) Urate oxidase: primary structure and evolutionary implications. Proc Natl Acad Sci USA 86:9412–9416
Yamamoto K, Kojima Y, Kikuchi T et al (1996) Nucleotide sequence of the uricase gene from Bacillus sp. TB-90. J Biochem 119:80–84
Zhou XL, Ma XH, Sun GQ (2005) Isolation of a thermostable uricase-producing bacterium and study on its enzyme production conditions. Process Biochem 40:3749–3753
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kai, L., Ma, XH., Zhou, XL. et al. Purification and characterization of a thermostable uricase from Microbacterium sp. strain ZZJ4-1. World J Microbiol Biotechnol 24, 401–406 (2008). https://doi.org/10.1007/s11274-007-9489-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-007-9489-1