Abstract
The remediation of groundwater contaminated with benzene, toluene, ethylbenzene, and the xylenes (BTEX) typically involves in situ biodegradation. Although the mechanisms of aerobic BTEX biodegradation in laboratory cultures have been well studied, less is known about the microorganisms responsible in mixed culture samples or at contaminated sites. In this study, the microorganisms responsible for in situ degradation within mixed culture samples were investigated using the molecular method stable isotope probing (SIP). For this, m-xylene was utilized as a model BTEX contaminant. Specifically, DNA-based SIP was utilized to identify active m-xylene degraders in microcosms constructed with soil from three sources (a gasoline-contaminated site and two agricultural sites). Replicate microcosms were amended with either labeled (13C) or unlabeled m-xylene, and the extracted DNA samples were ultracentrifuged, fractioned, and subjected to terminal restriction fragment length polymorphism (TRFLP). The dominant m-xylene degraders (responsible for 13C uptake) were determined by comparing relative abundance of TRFLP phylotypes in heavy fractions of labeled m-xylene (13C) amended samples to the controls (from unlabeled m-xylene amended samples). Four phylotypes were identified as the dominant m-xylene degrading species, falling within either the β Proteobacteria or the Bacilli. Of these, two 16S rRNA gene sequences were highly novel, displaying very limited similarity (94% and 90%) to any previously reported 16S rRNA gene sequence. Further, three of these phylotypes fell within genera with limited or no previous links to BTEX degradation, suggesting much information is still to be gained concerning the identity of microorganisms responsible for degradation within mixed culture samples.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Benzene, toluene, ethylbenzene, and p-, o-, and m-xylene (BTEX) are common contaminants of groundwater, primarily because of leaking underground storage tanks. Significant efforts have been directed towards remediating such contaminated sites because of the toxicity associated with these chemicals. Remediation typically occurs through biodegradation, and although many isolates have been obtained, less is known about the microorganisms actually responsible for BTEX transformation in mixed cultures or, indeed, in situ at contaminated sites.
Methods to identify the microorganisms able to transform problematic contaminants have traditionally involved enrichment culture followed by isolations. However, these approaches typically select for organisms able to compete well under laboratory conditions and likely do not reflect those responsible while in the mixed culture of an environmental sample. Although molecular approaches (e.g., denaturing gradient gel electrophoresis, clone libraries, or terminal restriction fragment length polymorphism (TRFLP)) have enabled the identification of community members in a mixed culture sample, these techniques typically do not directly link the ability to degrade a specific contaminant to individual species. However, a method developed recently, called stable isotope probing (SIP), is emerging as a key tool for linking activity or function with identity for mixed community or environmental samples (Radajewski et al. 2000). The method involves addition of a labeled substrate to a mixed community sample and identification of the organisms responsible for label uptake through DNA or RNA extraction and ultracentrifugation (isolates the labeled nucleic acids), followed by 16S rRNA gene sequencing. The technique has previously been used to identify microorganisms able to transform key organic compounds, such as polycyclic aromatic hydrocarbons, RDX, methane, benzene, and toluene (Hutchens et al. 2004; Kasai et al. 2006; Kunapuli et al. 2007; Liou et al. 2008; Luo et al. 2009; Oka et al. 2008; Roh et al. 2009; Singleton et al. 2005; Xie et al., submitted for publication 2009). These reports linked a number of novel microorganisms to biodegradation, indicating that information gained from isolations may not reflect degradation in mixed culture or complex environmental samples.
To obtain a more complete understanding of BTEX degradation in mixed culture, DNA-based SIP was applied here to identify the dominant organisms responsible for the transformation of m-xylene in microcosms constructed from three soils of different origin. To our knowledge, this represents the first report directly linking m-xylene degradation to specific microorganisms in a mixed community sample using SIP. Soils from two agricultural and one gasoline-contaminated site were chosen for this work to provide a better understanding of the diversity of m-xylene degraders as well as the variation in their m-xylene transformation abilities.
2 Materials and Methods
2.1 Soil Incubations and m-Xylene Analyses
Soils were collected from three different locations in Michigan, including a gasoline-contaminated site and two agricultural sites. Agricultural site 1 was previously under alfalfa production, and site 2 was under corn production. Both agricultural soils had received biosolids from a municipal wastewater treatment plant 2–3 years before sample collection. Following sample collection, soils were homogenized, sieved through a 4-mm screen, and stored at 4°C until use. Soil microcosms consisted of phosphate-buffered mineral medium (25 mL), as previously described (Mu and Scow 1994), and soil (3 g) in serum bottles (150 mL). The bottles were sealed with rubber stoppers and aluminum seals. The treatments included the following: sterile controls and unlabeled (6 µL, 99% pure, Sigma Aldrich) and labeled m-xylene (6 µL, methyl-13C2 m-xylene, 99%, Sigma Aldrich) amended samples. The sterile controls were obtained by autoclaving repeatedly (three times in 1 day). All samples exposed to m-xylene (labeled or unlabeled) were prepared in duplicate, and thus the SIP investigation (DNA extraction, ultracentrifugation, fractionation, and TRFLP) occurred in duplicate for each soil. Microcosms were incubated on a horizontal shaker at room temperature (18–22°C), and headspace m-xylene concentrations (200 µL samples) were determined daily with a gas chromatograph (Perkin Elmer) equipped with flame ionization detector and a capillary column (J&W Scientific, DB-624, diameter 0.53 mm). The injector and detector temperatures were set at 200°C, the column temperature was 80°C, and helium was utilized as the carrier gas.
2.2 DNA Extraction and Ultracentrifugation
Following the depletion of m-xylene, DNA was extracted from soil samples of replicate labeled and replicate unlabeled m-xylene amended microcosms with the Powersoil kit (Mobio Laboratories, Carlsbad, CA) following the manufacturer’s instructions. Ultracentrifugation was performed in Quick-Seal polyallomer tubes (13 × 51 mm, 5.1 mL, Beckman Coulter). Approximately 10 µg DNA (quantified with Nanodrop, ND-1000) was added to the tubes along with a Tris–EDTA (pH 8.0)/CsCl solution. The buoyant density (BD) of each sample was determined (model AR200 digital refractometer, Leica Microsystems Inc.) prior to sealing the tubes (cordless quick-seal tube topper, Beckman) and adjusted by adding small volumes of CsCl solution or Tris–EDTA buffer. The tubes were centrifuged within a Sorvall WX 80 Ultra Series Centrifuge (Thermo Scientific) at 178,000×g (20°C) for 48 h in a Stepsaver 70 V6 Vertical Titanium Rotor (8 × 5.1 ml capacity). Following ultracentrifugation, the tubes were placed into a fraction recovery system (Beckman Coulter) for fraction (150 µL) collection. The BD of each fraction was measured. The DNA was purified by the addition of glycogen (20 mg mL−1), water, and ethanol, overnight storage at 4°C, centrifugation, and pellet resuspension in sterile water. The resuspended pellet was stored at −20°C.
2.3 Polymerase Chain Reaction, TRFLP, and 16S rRNA Gene Sequencing
Each ultracentrifugation fraction from DNA extracted from the replicate labeled and replicate unlabeled m-xylene amended microcosms was analyzed by terminal restriction fragment length polymorphism following standard procedures (Liu et al. 1997). The fractions were amplified with 27F-FAM (5′ AGAGTTTGATCMTGGCTCAG, 5′ end-labeled with carboxyfluorescine) and 1492R (5′-GGTTACCTTGTTACGACTT) (Operon Biotechnologies) using the following polymerase chain reaction (PCR) program: 94°C (5 min); 94°C (30 s); 55°C (30 s); 72°C (1.5 min) (30 cycles); 72°C (5 min). Following amplification, PCR products (150 ng) were purified with a QIAquick PCR purification kit (Qiagen Inc.), following the manufacturer’s instructions, and digested with Hae III (New England Biolabs) with a 6-h incubation period. Additional digests (Hha I and Msp I) for TRFLP analyses were included to correlate the TRFLP fragment lengths to the in silico cut sites of the cloned 16S rRNA gene sequences and thus determine the identity of the enriched fragments. DNA fragments were separated by capillary electrophoresis (ABI Prism 3100 Genetic Analyzer, Applied Biosystems) at the Research Technology Support Facility (RTSF) at Michigan State University. Data were analyzed with GeneScan software (Applied Biosystems), and the percent abundance of each fragment was determined as previously described (Yu and Chu 2005).
For 16S rRNA gene sequencing, heavy fraction 13C-DNA was amplified as above, except the forward primer was unlabeled (27F 5′ AGAGTTTGATCMTGGCTCAG), and the final extension period was extended (72°C for 15 min). The PCR products were purified with QIAquick PCR purification kit (Qiagen Inc.) and cloned into Escherichia coli TOP10 vector supplied with a TOPO TA cloning kit (Invitrogen Corporation). E. coli clones were grown on Luria–Bertani medium solidified with 15 g agar L−1 with 50 µg mL−1 ampicillin for 16 h at 37°C. Colonies with inserts were verified by PCR with primers M13 F (5′-TGTAAAACGACGGCCAGT-3′) and M13 R (5′-AACAGCTATGACCATG-3′), plasmids were extracted from the positive clones with a QIAprep miniprep system (Qiagen, Inc.), and the insertions were sequenced at RTSF. The Ribosomal Database Project (Center for Microbial Ecology, Michigan State University) analysis tool “classifier” (Wang et al. 2007b) was utilized to assign taxonomic identity. The partial 16S rRNA gene sequences of organisms linked to m-xylene degradation were deposited with GenBank under accession numbers GU294754, GU294755, GU294756, and GU294757.
3 Results and Discussion
3.1 Comparison of m-Xylene Degradation Rates in the Three Soils
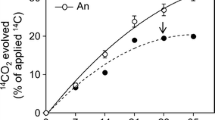
The time required for m-xylene degradation varied between the three soil microcosms (Fig. 1). Degradation occurred rapidly (7 days) in the microcosms constructed from the contaminated soil, whereas in both agricultural soils, m-xylene removal was slower. In agricultural soil 1, approximately 9 days was required, and in agricultural soil 2, degradation was not complete until day 14. This pattern of slower degradation in agricultural soils compared to the contaminated site soil is expected as these microorganisms have likely had no previous exposure to this contaminant. A significant loss occurred in the autoclaved controls, likely due to sorption. However, in all three cases, there was a significant difference in m-xylene concentration between controls and samples, clearly indicating a biological removal mechanism.
3.2 Enriched TRFLP Fragments Following Label Uptake
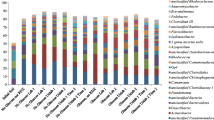
DNA extracts from replicate labeled and replicate unlabeled m-xylene amended soil microcosms were subjected to ultracentrifugation and fractionation, followed by TRFLP on each fraction. Unlabeled samples provided a control for DNA present in heavy fractions because of high guanine cytosine (GC) content rather than label incorporation. In other words, TRFLP fragments present in heavy fractions of samples, but not controls, represented the dominant microorganisms responsible for label uptake, therefore m-xylene degradation. Analyses of TRFLP profiles illustrated the enrichment of different fragments in each soil in the heavy fractions obtained from the 13C m-xylene microcosms but not in the unlabeled controls (Fig. 2). This trend suggests that a different dominant microorganism was responsible for 13C label uptake, thus contaminant degradation, in each soil. One TRFLP fragment (219 bp) was enriched in the microcosms constructed with soil from a gasoline-contaminated site. Similarly, one fragment was enriched from agricultural soil 1 (214 bp), whereas two TRFLP fragments (217 or 229 bp) were enriched in agricultural soil 2. DNA extracted from replicate microcosms for each soil produced similar trends (Fig. 2).
Relative abundance of the four highly enriched TRFLP fragments (a–d) in fractions of three labeled m-xylene amended soil microcosms compared to the controls (unlabeled m-xylene amended soils). Fractions from labeled and unlabeled m-xylene amended replicated microcosm are illustrated by the closed and opened symbols, respectively. Note the y-axis has different scales
Peak TRFLP relative abundance values and the buoyant density at which they occurred differed for each of these fragments. In the contaminated site soil, the maximum relative abundance of fragment 219 bp was 54.5% occurring at 1.725 g mL−1. A similar high TRFLP relative abundance level (67.0%) was observed for the fragment enriched in agricultural soil 1, occurring at a similar buoyant density (1.724 g mL−1). In contrast to this, a lower level of enrichment was noted for agricultural soil 2, likely because two microorganisms appeared responsible. TRFLP fragments 217 and 229 bp had peak relative abundance values of 6.0% and 5.2%, occurring at buoyant density values of 1.733 and 1.736 g mL−1, respectively. These values are similar to those found previously for aerobic benzene (13C labeled) SIP degradation studies (Cupples et al. 2009), which ranged from 25.4% to 84.1% for peak relative abundance occurring between 1.723 and 1.739 g mL−1. These statistics should be useful for others preparing SIP studies to determine the range of fractions to analyze for enriched DNA.
3.3 Identity of m-Xylene Degraders in the Three Soils
To determine the identity of these enriched fragments, approximately ten plasmids were partially sequenced for each soil microcosm. In addition, dominant TRFLP fragments obtained with 13C-enriched heavy fractions with two additional restriction enzymes (Hha I and Msp I) were used to provide a positive identification of the Hae III TRFLP-enriched fragments (219, 214, 217, and 229 bp). The dominant fragments obtained from all TRFLP restriction enzymes were compared to those obtained from in silico digests to determine the 16S rRNA sequence of the enriched fragments. A comparison of TRFLP cut sites and in silico cut sites for each identified phylotype is presented (Table 1). The slight differences (two to three bases) between the measured fragment lengths and those predicted using sequence data have also been noted by others (Clement et al. 1998; Liu et al. 1997; Osborn et al. 2000). The taxonomic identities, as determined by the Ribosomal Database Project (RDP) analysis tool “classifier” (Wang et al. 2007b), of each enriched fragment (cloned 16S rRNA sequence), thus m-xylene degrading microorganism, are presented in Table 2.
3.4 Relevant Genera Characteristics of Identified m-Xylene Degraders
Although others have used SIP to link function with identify for benzene and toluene degradation (Kasai et al. 2006; Kunapuli et al. 2007; Liou et al. 2008; Luo et al. 2009; Oka et al. 2008), to our knowledge, the current study is the first to use SIP to examine in situ m-xylene degraders.
Isolates able to degrade m-xylene include members of the α Proteobacteria (genus Sphingobium) (Chadhain et al. 2007; Kim and Zylstra 1999), β Proteobacteria (genera Alcaligenes and Ralstonia) (Cavalca et al. 2000; Olsen et al. 1994), γ Proteobacteria (genera Pseudomonas and Pseudoxanthomonas) (Assinder and Williams 1990; DiLecce et al. 1997; Duetz et al. 1997; Jeong et al. 2006; Kim et al. 2008; Velazquez et al. 2005, 2006; Williams and Worsey 1976; Worsey and Williams 1975), as well as members of the Actinobacteria (genus Rhodococcus) (Jung and Park 2004; Lee and Cho 2008). Isolates linked to m-xylene degradation from a contaminated site were primarily γ Proteobacteria (Pseudomonas or Stenotrophomonas species) (Hendrickx et al. 2006), whereas clone library analysis revealed that a culture consuming m-xylene was dominated by a β Proteobacteria (Delftia acidovorans), with minor members of the community including species of the Proteobacteria (genera Rhizobium and Mesorirhizobium), Firmicutes (genus Paenibacillus), or Bacteroidetes (genus Sphingobacterium) (Khomenkov et al. 2005). A member of the Firmicutes (Bacillus firmus) was also used to seed a biotrickling filter for xylene removal; however, the xylene isomer was not reported (Liu et al. 2006). In the current study, m-xylene degradation was directly linked to two members of both the β Proteobacteria and Bacilli.
The m-xylene degrader in agricultural soil 1 was classified within the genus Paenibacillus. This genus was separated from members of Bacillus group 3 in 1993 (Ash et al. 1993) and contains more than 80 recognized species (Lee et al. 2008) with Paenibacillus polymyxa as the type species. Bacteria in this genus have been isolated from a wide variety of environments, including desert sand (Jeon et al. 2009), on chestnut leaves (Valverde et al. 2008), in soil (Lee et al. 2008), poultry manure compost (Wang et al. 2007a), warm springs (Saha et al. 2005), Antarctic sediment (Montes et al. 2004), cow feces (Velaquez et al. 2004), blood cultures (Roux and Raoult 2004), and raw and heat-treated milk (Scheldeman et al. 2004). Interestingly, it is an organism in this genus, Paenibacillus larvae, that is the causative agent of the deleterious honey bee disease (American foulbrood) (Genersch 2008). Relevant to the research presented here, there have been several studies linking Paenibacillus species to the degradation of both natural and anthropogenic organic compounds. For example, several have been linked to the degradation of polysaccharides (Khianngam et al. 2009; Lee and Yoon 2008; Park et al. 2007; Velaquez et al. 2004), the dye Indigo carmine (Ramya et al. 2008), dibenzofuran (Iida et al. 2006), the polycyclic aromatic hydrocarbons naphthalene, biphenyl and phenanthrene (Daane et al. 2002), and polychlorinated biphenyls (Sakai et al. 2005). One study did find Paenibacillus as a minor component of a microbial community displaying m-xylene degradation (Khomenkov et al. 2005), although no direct link between these organisms and substrate depletion was provided. Significantly, no sequence in GenBank was highly similar to the 16S rRNA gene sequence of the Paenibacillus strain identified here. The three closest (all 94%) sequences were obtained from a pyrosequencing environmental survey (FJ478656.1 656/691 or 94%), soil iron–manganese nodules (EF492898.1, 657/693 or 94%), and petroleum-contaminated sediments (DQ664166.1, 653/689 or 94%). The data presented here represent the first direct report linking m-xylene degradation to any organism in this genus.
The m-xylene degrader identified in the microcosms constructed from the contaminated site sediment was classified as belonging to the genus Ramlibacter within the class β Proteobacteria. Surprisingly little information exists on microorganisms within this genus. For example, only 38 16S rRNA gene sequences are present for Ramlibacter within the RDP, and of these, only seven belong to isolates. Furthermore, the Web of Science contains only seven entries under the topic search of “Ramlibacter.” The genus was reported only recently (2003) with Ramlibacter tataouinensis (type species) and Ramlibacter henchirensis. These were described as cyst-producing bacteria isolated from subdesert soil in Tunisia (Heulin et al. 2003). Interestingly, both require growth factors and are slow growing under optimal conditions (Heulin et al. 2003), making them ideal targets for SIP. Four other isolates classifying within this genus were obtained in 2007 from paddy soil microcosms (Shrestha et al. 2007). Interestingly, the remaining isolate was obtained from chlorinated solvent-contaminated groundwater (Connon et al. 2005). The three most closely related 16S rRNA gene sequences in GenBank to the m-xylene degrader reported here were from a hydrocarbon-contaminated soil (AM936163.1, 730/735 or 99%) and two different rice field soils (EU589304, 745/755 or 98% and AY360686.1, 744/756 or 98%). The data presented here directly link an organism in the Ramlibacter genus to the degradation of an important environmental contaminant. To our knowledge, this is the first study to directly link an organism in this genus to the degradation of any BTEX compound.
The two microorganisms responsible for m-xylene degradation in agricultural soil 2 was classified within the β Proteobacteria as an unclassified Incertae sedis 5 and within the Firmicutes as a Bacillus species. The 16S rRNA gene sequence of the Incertae sedis 5 illustrated very little similarity (90% identity) to any sequence within Genbank. The three most similar partial 16S rRNA gene sequences originated from paddy soil (AM949517.1, 745/825 or 90%), the rhizosphere of trembling aspen (EF018506.1, 733/821 or 89%), and another soil (EF417657.1, 698/769 or 90%). Although the Proteobacteria are well known for their ability to transform organic contaminants, including m-xylene, the uniqueness of the 16S rRNA gene sequence identified here appears to represent a truly novel m-xylene degrader within the phylum. The three most similar partial 16S rRNA gene sequences in Genbank to the m-xylene degrader classifying as a Bacillus species originated from a clean room (EU071493.1, 711/725 or 98%), urban aersosol (DQ129492.1, 706/723 or 97%), and soil (AY289495.1, 705/723 or 97%). There have been a small number of reports linking Bacillus species to BTEX degradation. These include a soil study that suggested both Bacillus and Actinobacteria populations were affected by exposure to BTEX (Ji et al. 2007). Another study obtained a Bacillus sp. (Bacillus sphaericus 205y) from a group of isolates that could use benzene and/or toluene as their sole carbon source (Hun et al. 2003). Furthermore, B. firmus was used to seed biotrickling filter columns designed for xylene treatment (the isomer was not provided) (Liu et al. 2006). Another study isolated B. sphaericus from a lab-scale biofilter treating an airstream mixture of BTEX and reported that it had a high BTEX degrading activity (Mathur et al. 2007). Others isolated Bacillus species from seawater and deepsea sediment with benzene, toluene, and m-xylene degrading abilities (Wang et al. 2008).
4 Conclusions
DNA-based stable isotope probing was used to investigate the diversity of m-xylene degraders between microcosms constructed with sediment from contaminated and uncontaminated sites. Four phylotypes were identified as the dominant degraders within these microcosms, all falling within the β Proteobacteria or Bacilli. In the contaminated site microcosms, an organism belonging to the genus Ramlibacter (within β Proteobacteria) was identified as the dominant m-xylene degrader, representing the first report linking BTEX degradation to this genus. In agricultural soil 1, the m-xylene degrader was classified within the genus Paenibacillus and illustrated limited similarity (94% 16S rRNA gene identity) to any previously reported microorganism. Interestingly, Paenibacillus represents a genus with previous links to organic compound degradation but not directly to m-xylene degradation. In agricultural soil 2, two dominant m-xylene degraders were identified, one (unclassified Incertae sedis 5 within β Proteobacteria) with very little similarity (90% 16S rRNA gene identity) to any previously reported microorganism. The other was classified as a Bacillus species and thus corroborates previous reports linking BTEX degradation by Bacillus species. In summary, although two of the m-xylene degrading microorganisms identified belonged to a phylum (Proteobacteria) previously linked to BTEX degradation, neither represented commonly reported aerobic BTEX degraders (Pseudomonas, Sphingobium, Ralstonia) within this phylum. The other two were classified with the Bacilli, with one illustrating a unique 16S rRNA sequence. Such data indicate that previous studies involving isolations and clone libraries may not accurately represent the active m-xylene (or BTEX) degraders in all mixed community samples. These data provide an interesting illustration of the use of SIP to determine the role of uncultured microorganisms in environmental processes.
References
Ash, C., Priest, F. G., & Collins, M. D. (1993). Molecular identification of rRNA group 3 Bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie Van Leeuwenhoek, 64, 253–260.
Assinder, S. J., & Williams, P. A. (1990). The TOL plasmids: Determinants of the catabolism of toluene and the xylenes. Adv Microb Physiol, 31, 1–69.
Cavalca, L., Di Gennaro, P., Colombo, M., Andreoni, V., Bernasconi, S., Ronco, I., et al. (2000). Distribution of catabolic pathways in some hydrocarbon-degrading bacteria from a subsurface polluted soil. Res Microbiol, 151, 877–887.
Chadhain, S. M. N., Moritz, E. M., Kim, E., & Zylstra, G. J. (2007). Identification, cloning, and characterization of a multicomponent biphenyl dioxygenase from Sphingobium yanoikuyae B1. J Ind Microbiol Biotechnol, 34, 605–613.
Clement, B. G., Kehl, L. E., DeBord, K. L., & Kitts, C. L. (1998). Terminal restriction fragment patterns (TRFPs), a rapid, PCR-based method for the comparison of complex bacterial communities. J Microbiol Meth, 31, 135–142.
Connon, S. A., Tovanabootr, A., Dolan, M., Vergin, K., Giovannoni, S. J., & Semprini, L. (2005). Bacterial community composition determined by culture-independent and -dependent methods during propane-stimulated bioremediation in trichloroethene-contaminated groundwater. Environ Microbiol, 7, 165–178.
Cupples, A. M., Xie, S., Luo, C., & Sun, W. (2009). Novel toluene, m-xylene and benzene degraders at contaminated and uncontaminated sites identified by SIP. 109th General Meeting. Philadelphia: American Society for Microbiology.
Daane, L. L., Harjono, I., Barns, S. M., Launen, L. A., Palleroni, N. J., & Haggblom, M. M. (2002). PAH-degradation by Paenibacillus spp. and description of Paenibacillus naphthalenovorans sp nov., a naphthalene-degrading bacterium from the rhizosphere of salt marsh plants. Int J Syst Evol Microbiol, 52, 131–139.
DiLecce, C., Accarino, M., Bolognese, F., Galli, E., & Barbieri, P. (1997). Isolation and metabolic characterization of a Pseudomonas stutzeri mutant able to grow on the three isomers of xylene. Appl Environ Microbiol, 63, 3279–3281.
Duetz, W. A., Wind, B., Kamp, M., & vanAndel, J. G. (1997). Effect of growth rate, nutrient limitation and succinate on expression of TOL pathway enzymes in response to m-xylene in chemostat cultures of Pseudomonas putida (pWW0). Microbiology-Uk, 143, 2331–2338.
Genersch, E. (2008). Paenibacillus larvae and American Foulbrood—long since known and still surprising. Journal Fur Verbraucherschutz Und Lebensmittelsicherheit-Journal of Consumer Protection and Food Safety, 3, 429–434.
Hendrickx, B., Junca, H., Vosahlova, J., Lindner, A., Ruegg, I., Bucheli-Witschel, M., et al. (2006). Alternative primer sets for PCR detection of genotypes involved in bacterial aerobic BTEX degradation: Distribution of the genes in BTEX degrading isolates and in subsurface soils of a BTEX contaminated industrial site. J Microbiol Meth, 64, 250–265.
Heulin, T., Barakat, M., Christen, R., Lesourd, M., Sutra, L., De Luca, G., et al. (2003). Ramlibacter tataouinensis gen. nov., sp nov., and Ramlibacter henchirensis sp nov., cyst-producing bacteria isolated from subdesert soil in Tunisia. Int J Syst Evol Microbiol, 53, 589–594.
Hun, C. J., Rahman, R. N. Z. A., Salleh, A. B., & Basri, M. (2003). A newly isolated organic solvent tolerant Bacillus sphaericus 205y producing organic solvent-stable lipase. Biochem Eng J, 15, 147–151.
Hutchens, E., Radajewski, S., Dumont, M. G., McDonald, I. R., & Murrell, J. C. (2004). Analysis of methanotrophic bacteria in Movile Cave by stable isotope probing. Environ Microbiol, 6, 111–120.
Iida, T., Nakamura, K., Izumi, A., Mukouzaka, Y., & Kudo, T. (2006). Isolation and characterization of a gene cluster for dibenzofuran degradation in a new dibenzofuran-utilizing bacterium, Paenibacillus sp strain YK5. Arch Microbiol, 184, 305–315.
Jeon, C. O., Lim, J. M., Lee, S. S., Chung, B. S., Park, D. J., Xu, L. H., et al. (2009). Paenibacillus harenae sp nov., isolated from desert sand in China. Int J Syst Evol Microbiol, 59, 13–17.
Jeong, E., Hirai, M., & Shoda, M. (2006). Removal of p-xylene with Pseudomonas sp. NBM21 in biofilter. J Biosci Bioeng, 102, 281–287.
Ji, S. C., Kim, D., Yoon, J. H., & Lee, C. H. (2007). Metagenomic analysis of BTEX-contaminated forest soil microcosm. J Microbiol Biotechnol, 17, 668–672.
Jung, I. G., & Park, C. H. (2004). Characteristics of Rhodococcus pyridinovorans PYJ-1 for the biodegradation of benzene, toluene, m-xylene (BTX), and their mixtures. J Biosci Bioeng, 97, 429–431.
Kasai, Y., Takahata, Y., Manefield, M., & Watanabe, K. (2006). RNA-based stable isotope probing and isolation of anaerobic benzene-degrading bacteria from gasoline-contaminated groundwater. Appl Environ Microbiol, 72, 3586–3592.
Khianngam, S., Akaracharanya, A., Tanasupawat, S., Lee, K. C., & Lee, J. S. (2009). Paenibacillus thailandensis sp nov and Paenibacillus nanensis sp nov., xylanase-producing bacteria isolated from soil. Int J Syst Evol Microbiol, 59, 564–568.
Khomenkov, V. G., Shevelev, A. B., Zhukov, V. G., Kurlovich, A. E., Zagustina, N. A., & Popov, V. O. (2005). Application of molecular systematics to study of bacterial cultures consuming volatile organic compounds. Appl Biochem Microbiol, 41, 154–161.
Kim, E., & Zylstra, G. J. (1999). Functional analysis of genes involved in biphenyl, naphthalene, phenanthrene, and m-xylene degradation by Sphingomonas yanoikuyae B1. J Ind Microbiol Biotechnol, 23, 294–302.
Kim, J. M., Le, N. T., Chung, B. S., Park, J. H., Bae, J. W., Madsen, E. L., et al. (2008). Influence of soil components on the biodegradation of benzene, toluene, ethylbenzene, and o-, m-, and p-xylenes by the newly isolated bacterium Pseudoxanthomonas spadix BD-a59. Appl Environ Microbiol, 74, 7313–7320.
Kunapuli, U., Lueders, T., & Meckenstock, R. U. (2007). The use of stable isotope probing to identify key iron-reducing microorganisms involved in anaerobic benzene degradation. ISME J, 1, 643–653.
Lee, E. H., & Cho, K. S. (2008). Characterization of cyclohexane and hexane degradation by Rhodococcus sp. EC1. Chemosphere, 71, 1738–1744.
Lee, J. C., & Yoon, K. H. (2008). Paenibacillus woosongensis sp nov., a xylanolytic bacterium isolated from forest soil. Int J Syst Evol Microbiol, 58, 612–616.
Lee, F. L., Tien, C. J., Tai, C. J., Wang, L. T., Liu, Y. C., & Chern, L. L. (2008). Paenibacillus taichungensis sp nov., from soil in Taiwan. Int J Syst Evol Microbiol, 58, 2640–2645.
Liou, J. S. C., DeRito, C. M., & Madsen, E. L. (2008). Field-based and laboratory stable isotope probing surveys of the identities of both aerobic and anaerobic benzene-metabolizing microorganisms in freshwater sediment. Environ Microbiol, 10, 1964–1977.
Liu, W. T., Marsh, T. L., Cheng, H., & Forney, L. J. (1997). Characterization of microbial diversity by determining terminal restriction fragment length polymorphisms of genes encoding 16S rRNA. Appl Environ Microbiol, 63, 4516–4522.
Liu, Q., Babajide, A. E., Zhu, P., & Zou, L. P. (2006). Removal of xylene from waste gases using biotrickling filters. Chem Eng Technol, 29, 320–325.
Luo, C., Xie, S., Sun, W., Li, X., & Cupples, A. M. (2009). Identification of a novel toluene-degrading bacterium from the candidate phylum TM7, as determined by DNA stable isotope probing. Appl Environ Microbiol, 75, 4644–4647.
Mathur, A. K., Majumder, C. B., & Chatterjee, S. (2007). Combined removal of BTEX in air stream by using mixture of sugar cane bagasse, compost and GAC as biofilter media. J Hazard Mater, 148, 64–74.
Montes, M. J., Mercade, E., Bozal, N., & Guinea, J. (2004). Paenibacillus antarcticus sp nov., a novel psychrotolerant organism from the Antarctic environment. Int J Syst Evol Microbiol, 54, 1521–1526.
Mu, D. Y., & Scow, K. M. (1994). Effect of trichloroethylene (TCE) and toluene concentrations on TCE and toluene biodegradation and the population density of TCE and toluene degraders in soil. Appl Environ Microbiol, 60, 2661–2665.
Oka, A. R., Phelps, C. D., McGuinness, L. M., Mumford, A., Young, L. Y., & Kerkhof, L. J. (2008). Identification of critical members in a sulfidogenic benzene-degrading consortium by DNA stable isotope probing. Appl Environ Microbiol, 74, 6476–6480.
Olsen, R. H., Kukor, J. J., & Kaphammer, B. (1994). A novel toluene-3-monooxygenase pathway cloned from Pseudomonas pickettii PKO1. J Bacteriol, 176, 3749–3756.
Osborn, A. M., Moore, E. R. B., & Timmis, K. N. (2000). An evaluation of terminal-restriction fragment length polymorphism (T-RFLP) analysis for the study of microbial community structure and dynamics. Environ Microbiol, 2, 39–50.
Park, M. J., Kim, H. B., An, D. S., Yang, H. C., Oh, S. T., Chung, H. J., et al. (2007). Paenibacillus soli sp nov., a xylanolytic bacterium isolated from soil. Int J Syst Evol Microbiol, 57, 146–150.
Radajewski, S., Ineson, P., Parekh, N. R., & Murrell, J. C. (2000). Stable-isotope probing as a tool in microbial ecology. Nature, 403, 646–649.
Ramya, M., Anusha, B., & Kalavathy, S. (2008). Decolorization and biodegradation of Indigo carmine by a textile soil isolate Paenibacillus larvae. Biodegradation, 19, 283–291.
Roh, H., Yu, C. P., Fuller, M. E., & Chu, K. H. (2009). Identification of hexahydro-1, 3, 5-trinitro-1, 3, 5-triazine-degrading microorganisms via N-15-stable isotope probing. Environ Sci Technol, 43, 2505–2511.
Roux, V., & Raoult, D. (2004). Paenibacillus massiliensis sp nov., Paenibacillus sanguinis sp nov and Paenibacillus timonensis sp nov., isolated from blood cultures. Int J Syst Evol Microbiol, 54, 1049–1054.
Saha, P., Mondal, A. K., Mayilraj, S., Krishnamurthi, S., Bhattacharya, A., & Chakrabarti, T. (2005). Paenibacillus assamensis sp nov., a novel bacterium isolated from a warm spring in Assam, India. Int J Syst Evol Microbiol, 55, 2577–2581.
Sakai, M., Ezaki, S., Suzuki, N., & Kurane, R. (2005). Isolation and characterization of a novel polychlorinated biphenyl-degrading bacterium, Paenibacillus sp KBC101. Appl Microbiol Biotechnol, 68, 111–116.
Scheldeman, P., Goossens, K., Rodriguez-Diaz, M., Pil, A., Goris, J., Herman, L., et al. (2004). Paenibacillus lactis sp nov., isolated from raw and heat-treated milk. Int J Syst Evol Microbiol, 54, 885–891.
Shrestha, P. M., Noll, M., & Liesack, W. (2007). Phylogenetic identity, growth-response time and rRNA operon copy number of soil bacteria indicate different stages of community succession. Environ Microbiol, 9, 2464–2474.
Singleton, D. R., Powell, S. N., Sangaiah, R., Gold, A., Ball, L. M., & Aitken, M. D. (2005). Stable-isotope probing of bacteria capable of degrading salicylate, naphthalene, or phenanthrene in a bioreactor treating contaminated soil. Appl Environ Microbiol, 71, 1202–1209.
Valverde, A., Peix, A., Rivas, R., Velazquez, E., Salazar, S., Santa-Regina, I., et al. (2008). Paenibacillus castaneae sp nov., isolated from the phyllosphere of Castanea sativa Miller. Int J Syst Evol Microbiol, 58, 2560–2564.
Velaquez, E., de Miguel, T., Poza, M., Rivas, R., Rossello-Mora, R., & Villa, T. G. (2004). Paenibacillus favisporus sp nov., a xylanolytic bacterium isolated from cow faeces. Int J Syst Evol Microbiol, 54, 59–64.
Velazquez, F., Parro, V., & de Lorenzo, V. (2005). Inferring the genetic network of m-xylene metabolism through expression profiling of the xyl genes of Pseudomonas putida mt-2. Mol Microbiol, 57, 1557–1569.
Velazquez, F., de Lorenzo, V., & Valls, M. (2006). The m-xylene biodegradation capacity of Pseudomonas putida mt-2 is submitted to adaptation to abiotic stresses: Evidence from expression profiling of xyl genes. Environ Microbiol, 8, 591–602.
Wang, C. M., Shyu, C. L., Ho, S. P., & Chiou, S. H. (2007a). Species diversity and substrate utilization patterns of thermophilic bacterial communities in hot aerobic poultry and cattle manure composts. Microb Ecol, 54, 1–9.
Wang, Q., Garrity, G. M., Tiedje, J. M., & Cole, J. R. (2007b). Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol, 73, 5261–5267.
Wang, L., Qiao, N., Sun, F. Q., & Shao, Z. Z. (2008). Isolation, gene detection and solvent tolerance of benzene, toluene and xylene degrading bacteria from nearshore surface water and Pacific Ocean sediment. Extremophiles, 12, 335–342.
Williams, P. A., & Worsey, M. J. (1976). Ubiquity of plasmids in coding for toluene and xylene metabolism in soil bacteria—evidence for existence of new TOL plasmids. J Bacteriol, 125, 818–828.
Worsey, M. J., & Williams, P. A. (1975). Metabolism of toluene and xylenes by Pseudomonas putida (arvilla) mt-2: Evidence for a new function of TOL plasmid. J Bacteriol, 124, 7–13.
Yu, C. P., & Chu, K. H. (2005). A quantitative assay for linking microbial community function and structure of a naphthalene-degrading microbial consortium. Environ Sci Technol, 39, 9611–9619.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xie, S., Sun, W., Luo, C. et al. Stable Isotope Probing Identifies Novel m-Xylene Degraders in Soil Microcosms from Contaminated and Uncontaminated Sites. Water Air Soil Pollut 212, 113–122 (2010). https://doi.org/10.1007/s11270-010-0326-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11270-010-0326-z