Abstract
Camelina (Camelina sativa (L.) Crantz) is a re-emergent oilseed crop that is also becoming important as a model for applied projects based on studies in Arabidopsis thaliana, since the two species are closely related members of the tribe Camelineae of the Brassicaeae. Since camelina can be transformed genetically by floral dip, genetically modified (GM) camelina is being created in many laboratories, and small-scale field trials are already being conducted in the US and Canada. Although camelina does not cross-fertilize Brassica crop species, such as oilseed rape, nothing was known about its ability to cross with other members of the tribe Camelineae, which in addition to arabidopsis includes the widespread weed, shepherd’s purse (Capsella bursa-pastoris). We have tested the ability of camelina to cross with arabidopsis and C. bursa-pastoris, as well as with the more distantly related Cardamine hirsuta, tribe cardamineae. No seeds were produced in crosses with arabidopsis, and a few seeds were obtained in crosses with C. hirsuta, but the embryos aborted at an early stage of development. A few seeds were also obtained in crosses with C. bursa-pastoris, which germinated to produce plants of a phenotype intermediate to that of the parents, but the hybrids were both male and female sterile. Therefore, the likelihood of pollen-mediated gene flow from camelina to these related species is low.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Camelina (Camelina sativa) is an oilseed crop that has been grown in Central Asia and Europe since the Neolithic (Hovsepyan and Willcox 2008; Toulemonde 2010), but during the nineteenth and twentieth centuries it was replaced progressively by more productive species, such as oilseed rape. Camelina has attracted renewed interest in the past few years for several reasons. The oil has attractive nutritional features, since it is particularly rich in omega-3 lipids (Zubr 1997). Also, since camelina grows well under low water conditions (Bramm et al. 1990; Zubr 1997), it is being grown increasingly in dryer areas of Western North America, where it is being promoted primarily as a source of biodiesel.
Camelina is also an interesting model species, particularly for testing potential applications based on studies carried out with arabidopsis. Indeed, Camelina and Arabidopsis are closely related genera belonging to the tribe Camelineae of the family Brassicaceae (Beilstein et al. 2008), and camelina genes have on the order of 90–95 % sequence identity with corresponding arabidopsis genes (Hutcheon et al. 2010; Nguyen et al. 2013). Further, camelina can be transformed genetically by floral dip (Lu and Kang 2008), similarly to arabidopsis and capsella (Bartholmes et al. 2008; Clough and Bent 1998). Also, its life cycle is short, approximately 3 months, and in our experience a single plant in the greenhouse produces approximately 1 g of seed, with an oil content of ~30 to 40 % oil (Zubr 1997), making camelina a particularly useful species for testing potential modifications of seed oil content and composition. This was shown by Sayanova et al. (2011), who overexpressed an acyl-CoA delta-6-desaturase in camelina. Further, strategies based on gene knockdown can also be effective; Kang et al. (2011) used an antisense gene to knock down expression of fatty acid desaturase 2 (FAD2) expression, and Nguyen et al. (2013) used hairpin RNA production to knock down expression of either FAD2 or fatty acid elongase 1 (FAE1).
Although modest amounts of seed and oil can be produced in the greenhouse, it will rapidly become useful to carry out small-scale field trials of genetically modified (GM) camelina in order to produce sufficient quantities of seed for characterization of nutritional and/or industrial properties of the oil and residual cake. Such field trials will be facilitated by the recent release by the Canadian Food Inspection Agency (CFIA) of a plant biology document that describes the general properties of C. sativa (www.inspection.gc.ca/english/plaveg/bio/dir/camelsate.shtml). In view of small-scale field trials, it is particularly important to be able to prevent gene flow from camelina to related species. Hybrids can be created between C. sativa and the two other Camelina species, C. alyssum and C. microcarpa (Séguin-Swartz et al. 2010), while attempts to cross C. sativa with the distantly related crop species Brassica napus (oilseed rape), B. rapa and B. juncea were uniformly unsuccessful (Séguin-Swartz 2008). According to the CFIA biology document, there is no information regarding the potential for gene flow from C. sativa to members of other genera of the tribe Camelineae; such information regarding two particularly widespread, abundant members of the Camelineae, Capsella bursa-pastoris and Arabidopsis thaliana, would be highly pertinent for planning confinement of future GM camelina field trials. To facilitate gene flow studies, we have created a transgenic line of camelina that expresses a DsRed transgene, and have used it to evaluate in the greenhouse the potential for crossing of camelina—as pollen donor—with three wild Brassicaeae that are abundant in Versailles: A. thaliana, C. bursa-pastoris and Cardamine hirsuta. The latter is a more distantly related species, member of the tribe Cardamineae, but falling within the same major lineage of Brassicaceae as C. sativa (lineage 1; Beilstein et al. 2006, 2008).
Materials and methods
Plant materials
Seeds of C. sativa cv Céline were provided by the CAVAC (Coopérative agricole Vendéenne d’approvisionnement, de ventes de céréales et autres produits agricoles, La Roche-sur-Yon, France). Local ecotypes of C. bursa-pastoris, A. thaliana and C. hirsuta were collected on the INRA-Versailles campus, in spring of 2012. C. bursa-pastoris G4 was provided by G. Theissen (Bartholmes et al. 2008). Plants were grown in the greenhouse (16 h day length, 25 °C day and 22 °C night, 40 % relative humidity). For manual crossing, flowers of the female parent were emasculated before anthesis to avoid selfing, and were pollinated manually with the pollen of DsRed-expressing camelina line 1.6.5.
Genetic transformation of camelina using Agrobacterium tumefaciens
The protocol was adapted from standard arabidopsis floral dip techniques. C. sativa cv Céline plants were transformed with Agrobacterium tumefaciens C58C1 bearing pBinGlyRed2 (www.camelinagene.org), in which expression of a DsRed coding sequence is driven by the promoter of Cassava vein mosaic virus (CsVMV) (Verdaguer et al. 1996). Camelina plants were grown 10/pot in 15 cm pots. When plants began to flower, approximately 40 days after planting, the flowers were dipped for 1 min in a solution of an overnight culture of Agrobacterium resuspended in twice the volume of 50 g/l sucrose containing 0.05 % (v/v) Silwet L77 (Lehle Seeds, Round Rock, Texas, USA). After dipping, plants were kept for 24 h under low light conditions, with the flowers covered with wet paper towels to maintain high humidity. Plants were dipped three times at weekly intervals, corresponding to the 3 weeks of flowering. The seeds produced were screened for transformants by observation of DsRed expression in seeds, as visualized under a dissecting microscope with a Nikon LH-M100C-1 light source, using filters to obtain green light at 558 nm. In a typical experiment ~25,000 seeds produced, of which six were genetically transformed. The presence of the transgene was verified by PCR with forward primer F5DsRed (GAGTTCATGCGCTTCAAGGTG) and reverse primer R5DsRed (GGGCTTCTTGGCCATGTAGAT), using the following conditions: 30 s 95 °C, 30 s 55 °C, 1 min 72 °C, 35 cycles.
Flow cytometry
Relative nuclear DNA amounts were determined with a CyFlow Space flow cytometer as described by Marie and Brown (1993). For each plant genotype, three flower bud samples were analyzed.
Results
Transgenic camelina expressing a DsRed transgene for use as pollen donor
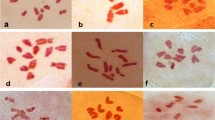
In order to facilitate screening for inter-generic hybrids, we transformed camelina with a DsRed gene controlled by the CsVMV promoter, which was expressed throughout the plant, as was observed for other species (Verdaguer et al. 1996). DsRed expression in embryos allowed simple screening for transformants among the T1 progeny and evaluation of the transgene transmission to further progeny generations (Fig. 1a). The presence of the DsRed transgene in putative transformants was confirmed by PCR.
Using the DsRed marker gene to screen for transgenic T1 individuals and for transgene transmission to progeny. a, c, and e were photographed under white light, while b, d, and f show the same view under green light (558 nm). a and b Progeny of a camelina plant that had been used in Agrobacterium-mediated floral dip transformation, showing a DsRed-positive putative transformant. c and d seeds produced in a C. hirsuta × C. sativa cross. e and f: seeds produced in a C. bursa-pastoris × C. sativa cross
Crosses between camelina and wild Brassicaceae that flower simultaneously with camelina
Three wild Brassicaceae species are particularly abundant in Versailles, and are widespread in temperate regions: A. thaliana, C. bursa-pastoris and C. hirsuta. In order to evaluate whether gene flow from camelina to these species can occur, seeds of all three wild species were collected on the INRA-Versailles campus during the early spring of 2012. All three continue to flower into mid-summer, simultaneously with camelina grown in the field in Versailles. These local ecotypes were used as female parents in crosses in the greenhouse. Since all species concerned are autogamous, it was necessary to emasculate the female parent flowers before applying pollen of DsRed-transgenic camelina of the T1 generation to the stigmas of the female parent.
As shown in Table 1, no seeds were produced from 79 A. thaliana × C. sativa crosses. In contrast, a total of eight seeds were produced from 64 C. hirsuta × C. sativa crosses, of which three were DsRed positive, but the embryos aborted at an early stage of development (Fig. 1d). The DsRed-negative seeds probably include additional hybrids, since the C. sativa parent was hemizygous for the DsRed marker, although we cannot exclude formally that some of them were due to accidental self-pollination. The 95 C. bursa-pastoris × C. sativa crosses yielded 26 seeds, of which eight were DsRed positive (Fig. 1f). In this case, embryo development was more complete, but not entirely normal (Fig. 1f). In control crosses in which flowers of all four species were emasculated and self pollinated, seeds were successfully produced (Table 1).
Morphology of F1 progeny of C. bursa-pastoris × C. sativa
The DsRed-positive putative hybrids were sown, but only those of the C. bursa-pastoris × C. sativa crosses germinated. The morphology of the hybrids is compared to that of the parents in Fig. 2. In the early vegetative stages (Fig. 2a), capsella plants remained in a rosette, while the stem of the camelina plants elongated. The hybrids were intermediate, with stems that elongated, but less than those of camelina. The flowers of capsella and camelina were distinctly different in flower color and size (Fig. 2b). The flowers of the putative hybrids were more like those of capsella, but the stamens failed to elongate, and they produced no pollen. In the reproductive stage, unlike the parents, the hybrids did not self-pollinate, and in early phases the silicules aborted. Later during the reproductive phase, silicules with unique features developed on the hybrids, but they produced no seeds (Fig. 2c). The silicules of capsella were strongly compressed perpendicular to the plane of the septum, those of camelina were nearly isodiametric in cross section, and the silicules of putative hybrids were flattened like those of capsella, but the contour of the silicules of the hybrids was intermediate between the parents. All attempts to backcross the hybrids with pollen from the parental species failed to produce seeds.
Comparison of the morphology of putative C. bursa-pastoris × C. sativa hybrids with that of the parental species. a Plants 3 weeks old, during vegetative growth. b Flowers partly dissected to display floral organs. c: Full-grown green silicules. Maturing seeds are visible in the parental species, but not in the putative hybrid
Flow cytometry of the relative nuclear DNA content of putative hybrids
As shown above, the progeny of the C. bursa-pastoris × C. sativa crosses was morphologically intermediate to the parental species. In order to determine whether the plants were true hybrids, the relative nuclear DNA amounts of the parental species and the hybrids were compared. The four species differ in their ploidy: A. thaliana and C. hirsuta are diploid (Carlson et al. 2009), C. bursa-pastoris is tetraploid (Hurka et al. 2012), and C. sativa is hexaploid (Gehringer et al. 2006; Hutcheon et al. 2010), and this is reflected in their relative nuclear DNA amounts (Fig. 3a). As shown in Fig. 3b, the progeny of the C. bursa-pastoris × C. sativa crosses had nuclear DNA amounts intermediate to those of the parents, and were thus genuine hybrids.
Flow cytometry of nuclear DNA amounts. a The relative DNA amount of 2n and 4n peaks of C. sativa, C. bursa pastoris, C. hirsuta, and A. thaliana. Values are averages from three plants, showing standard error of the means. b The relative DNA amount of 2n and 4n peaks of C. sativa, putative hybrids, and C. bursa-pastoris. Since the three putative hybrids gave DNA amounts that could not be distinguished, the average value shown is that of three samples for each of the three hybrids
Discussion
We have evaluated the ability of DsRed-transgenic camelina to cross-fertilize Versailles ecotypes of the locally most abundant wild Brassicaceae, C. bursa-pastoris, A. thaliana and C. hirsuta. Only C. sativa × C. bursa-pastoris crosses produced a few hybrid plants, and these displayed both male and female sterility. These results suggest that the potential for pollen-mediated gene flow from future field trials in Versailles of GM camelina would be extremely limited. Further, all species considered are autogamous, and in the field self-fertilization predominates (Walsh et al. 2012); as a consequence, pollen competition should reduce even further the probability of cross-pollination by camelina. Thus, standard protocols to prevent pollen- or seed-mediated gene flow from small-scale trial plots would be sufficient.
It is not clear to what extent these results can be generalized. Hurka et al. (2012) proposed that C. bursa-pastoris arose several times by chromosome doubling of an unidentified diploid progenitor species, and as a result C. bursa-pastoris is highly variable. For instance, in the greenhouse, the capsella ecotype G4 from Germany (Bartholmes et al. 2008) is quite different from the Versailles ecotype; G4 has a lengthy rosette phase and highly divided leaves, while the Versailles ecotype has an extremely brief rosette phase and entire leaves. In preliminary attempts to cross G4 with DsRed camelina, no hybrid progeny was obtained. Although this suggests that there are capsella ecotypes that are less able to cross with camelina, this also means that we cannot exclude that there may be ones with increased ability to hybridize.
Although commercial development of GM camelina is perhaps only in the distant future, it is useful to consider potential impacts of future release starting early in the development process, so that any necessary research can be carried out in a timely fashion. A useful tool for considering potential risks, which is based on three fundamental steps, has been described recently (Tepfer et al. 2013). The first step is to define the concerns in the form of risk hypotheses, in which the potential harm is clearly stated. The second is to clarify how the harm could occur, through defining a chain of cause and effect, in the form of a pathway to harm. And the third step is to determine how the current state of knowledge validates or invalidates the steps in the pathway.
We have chosen a hypothetical herbicide-tolerant camelina to illustrate this risk assessment strategy (Fig. 4). The equally hypothetical potential harm considered is that gene flow from the herbicide-tolerant camelina could create herbicide-tolerant capsella, which would become a more difficult weed to eliminate. We have stated the concern as a risk hypothesis in Fig. 4a, and in 4b have detailed the steps by which the potential harm could occur. We have also indicated how the current state of knowledge, including the results presented here, informs the different steps of the pathway to harm. If we consider the steps in the pathway in sequence, steps 1 and 2 clearly were observed in Versailles: camelina and capsella did flower simultaneously, and potential pollinators (honey bees, bumblebees, and hoverflies) were observed visiting the flowers. Our experimental crossing results show that step 3 is unlikely to occur in the field, since only a few C. sativa × C. bursa-pastoris hybrids were observed in hand-pollinated, emasculated flowers. And further, step 4 is also unlikely, since the few hybrids obtained were both male and female sterile. If the breaks in the pathway to harm at steps 3 and 4 are valid with other genotypes and in other environments this suggests that the potential risk is very low, and these same considerations would be valid for other potential risks with GM camelina that could occur via pollen-mediated gene flow to capsella. If there are circumstances where gene flow from camelina to capsella is of concern, it would be appropriate to consider whether there are means to break the pathway to harm in a more definitive manner.
Analysis of a potential risk associated with hypothetical herbicide-tolerant (HT) GM camelina. The example illustrates a three-step risk assessment process giving a central role to problem formulation: statement of a risk hypothesis, development of a pathway to harm, evaluation of how the current state of knowledge supports or invalidates steps in the pathway (Tepfer et al. 2013). a formulation of a risk hypothesis based on concerns that transfer of a gene conferring herbicide tolerance from GM camelina to C. bursa-pastoris could increase the latter’s weediness. b development of a pathway to harm showing the steps that could lead from deployment of the GM camelina to the potential harm being considered. Results supporting or invalidating steps in the pathway are indicated
References
Bartholmes MA, Nutt P, Theissen G (2008) Germline transformation of shepherd’s purse (Capsella bursa-pastoris) by the ‘floral dip’ method as a tool for evolutionary and developmental biology. Gene 409:11–19
Beilstein MA, Al-Shehbaz IA, Mathews S, Kellogg EA (2006) Brassicaceae phylogeny and trichome evolution. Am J Bot 93:607–619
Beilstein MA, Al-Shehbaz IA, Mathews S, Kellogg EA (2008) Brassicaceae phylogeny inferred from phytochrome A and NDHF sequence data: tribes and trichomes revisited. Am J Bot 95:1307–1327
Bramm A, Dambroth M, Schulte-Korne S (1990) Analysis of yield components of linseed, false flax, and poppy. Landbauforschung Voelkenrode 40:107–114
Carlson T, Bleeker W, Hurka H, Elven R, Brochmann C (2009) Biogeography and phylogeny of Cardamine (Brassicaceae). Ann Missouri Bot Gard 96:215–236
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Gehringer A, Friedt W, Lühs W, Snowdon RJ (2006) Genetic mapping of agronomic traits in false flax (Camelina sativa subsp. sativa). Genome 49:1555–1563
Hovsepyan R, Willcox G (2008) The earliest finds of cultivated plants in Armenia: evidence from charred remains and crop processing residues in pisé from the Neolithic settlements of Aratasche and Aknashen. Veg Hist Archeobot 17(suppl. 1):63–71
Hurka H, Friessen N, German DA, Franzke A, Neuffer B (2012) ‘Missing link’ species Capsella orientalis and Capsella thracica elucidate evolution of model plant genus Capsella (Brassicaceae). Mol Ecol 21:1223–1238
Hutcheon C, Ditt RF, Beilstein M, Comai L, Schroeder J, Goldstein E, Shewmaker CK, Nguyen T, De Rocher J, Kiser J (2010) Polyploid genome of Camelina sativa revealed by isolation of fatty acid synthesis genes. BMC Plant Biol 10:233
Kang J, Snapp AR, Lu C (2011) Identification of three genes encoding microsomal oleate desaturases (FAD2) from the oilseed crop Camelina sativa. Plant Physiol Biochem 49:223–229
Lu C, Kang J (2008) Generation of transgenic plants of a potential oilseed crop Camelina sativa by Agrobacterium-mediated transformation. Plant Cell Rep 27:273–278
Marie D, Brown SC (1993) A cytometric exercise in plant DNA histograms, with 2C values for 70 species. Biol Cell 78:41–51
Nguyen T, Collins-Silva JE, Macrander J, Yang W, Nazarenus TJ, Nam J-W, Jaworski JG, Lu C, Scheffler BE, Mockaitis K, Cahoon EB (2013) Camelina seed transcriptome: a tool for meal and oil improvement and translational research. Plant Biotech J. doi: 10.1111/pbi.12068
Sayanova O, Ruiz-Lopez N, Haslam RP, Napier JA (2011) The role of delta-6-desaturase acyl-carrier specificity in the efficient synthesis of long-chain polyunsaturated fatty acids in transgenic plants. Plant Biotech J 11:157–168
Séguin-Swartz G (2008) Hybridization between Camelina sativa (L.) Crantz (false flax) and Brassica napus, B. rapa and B. juncea. Report to the Canadian Food Inspection Agency. July 31, 2008
Séguin-Swartz G, Warwick SI, Gugel RK, Olivier CY, Soroka J, Strelkov SE, Klein-Gebbinck H, Falk KC (2010) Biology of Camelina sativa (L.) Crantz (false flax). Final report to the Canadian Food Inspection Agency. June 7, 2010
Tepfer M, Racovita M, Craig W (2013) Putting problem formulation at the forefront of GMO risk analysis. GM Crops Food 4:1–6
Toulemonde F (2010) Camelina sativa: l’or végétal du Bronze et du Fer. Anthrobotanica 1(1):3–14
Verdaguer B, de Kochki A, Beachy RN, Fauquet C (1996) Isolation and expression in transgenic tobacco and rice plants of the Cassava vein mosaic virus (CVMV) promoter. Plant Mol Biol 31:1129–1139
Walsh KD, Puttick DM, Hills MJ, Yang R-C, Topinka KC, Hall LM (2012) First report of outcrossing rates in camelina (Camelina sativa (L.) Crantz). Can J Plant Sci 92:681–685
Zubr J (1997) Oil-seed crop: Camelina sativa. Ind Crop Prod 6:113–119
Acknowledgments
We thank E. Cahoon for providing pBinGlyRed2, G. Theissen for providing C. bursa-pastoris G4, the CAVAC for providing camelina cv Céline, P. Grillon for growing the plants in the greenhouse, M. Racovita and W. Craig for helpful discussions. This research was supported in part by 3BCAR Institut Carnot project Camelina Oil.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Julié-Galau, S., Bellec, Y., Faure, JD. et al. Evaluation of the potential for interspecific hybridization between Camelina sativa and related wild Brassicaceae in anticipation of field trials of GM camelina. Transgenic Res 23, 67–74 (2014). https://doi.org/10.1007/s11248-013-9722-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-013-9722-7