Abstract
Some species in Vicia sativa complex, also called aggregate (CVS), have economic and ecological advantages and they are frequently used for pasture, silage, and green manure. The main objective of this study was to explore the secondary gene pool for enhancement of common vetch (Vicia sativa L.) germplasm by genome introgressions via conventional crosses made between cultivated common vetch (Vicia sativa L. subspecies sativa) used as female parent with five other subspecies [amphicarpa (L.), Batt., cordata (Wulfen ex Hoppe) Asch. & Graebner, macrocarpa (Moris) Arcang., nigra (L.) Ehrh., and segetalis (Thuill.)], used as male parents. As indicated with very low seed sets, higher levels of sterility were the rule in hybrids in CVS. Hybrids were confirmed using flower petal color and anthocyanin pigmentation markers. Chromosome count studies revealed that certain crossing-experiments were successfully produced interspecific hybrids. Certain offspring progenies in further generations were also obtained. Our studies clearly indicated that complex CVS and hybrids obtained from these species consisted of 10, 11, 12, 13 and 14 pair of somatic chromosomes. Results indicated that there exist significant incompatibility alleles possibly expressing in the zygote or post-zygotic developmental stages within the CVS. In conclusion, present study revealed that sufficient amount of seeds could be obtained from the crosses in the CVS and this stimulates renewed interest in utilizing the secondary gene pool as a source of genetic variation in breeding programs of common vetch.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Vicia L. is a member of the tribe Fabeae (also referred to as Vicieae/Leguminosae) of the subfamily Papilionoideae which includes Vicia, Lathyrus, Lens and Pisum (Hanelt and Mettin 1989). Pinning down exact numbers of species in genus Vicia L. is difficult due to cytological and morphological differences. Although numbers vary, it is estimated that this genus comprises about 166–210 annual or perennial species naturally grown in Europe, Asia, and North America, temperate regions of South America and tropical Africa (Hanelt and Mettin 1989; Tate and Ennenking 2006; El-Bok et al. 2015; Raveendar et al. 2015; Martin et al. 2018). Common vetch (Vicia sativa L.), also known as tare, is widespread around many parts of world, including the Mediterranean basin, west and central Asia, China, eastern Asia, India, and the USA (Hueze et al. 2011). It is commonly grown as winter forage legumes for its high nutritional value, green manure, pasture, silage and hay. Since common vetch has the ability to grow in wider range of climate and soil conditions it is suitable intercrops for cereals to reduce diseases and help to improve soil properties (Acikgoz 1986; Seymour et al. 2002; Cakmakci et al. 2003, 2006; Chung et al. 2013; Mikic et al. 2014; Chai et al. 2017; Tiryaki et al. 2016; Dong et al. 2017; Geogieva 2018).
The Mediterranean region is accepted as the principal center of diversification, and the highest specific diversity of Vicia is found in Turkey and northwest Asia (Hanelt and Mettin 1989; Frediani et al. 2004; El-Bok et al. 2015; Raveendar et al. 2015; Dong et al. 2017; Martin et al. 2018). The number of polyploid species in the genus is very low, for instance among 87 species studied, ten were found polyploid ones, indicating the polyploidy is almost insignificant in the evolution of the genus Vicia, and the tribe Vicieae (Ladizinsky and Shefer 1982). Maxted (1993) divided the subgenus Vicia into nine sections using morphological key markers. These sections of subgenus Vicia are (1) Atossa (Alef.) Asch. & Graebner, (2) Microcarinae Maxted, (3) Hypechusa (Alef.) Asch. & Graebner, (4) Peregrinae Kupicha, (5) Wiggersia (Alef.) Maxted, (6) Vicia, (7) Narbonensis (Radzhi) Maxted, (8) Bithynicae (B. Fedtsch. ex Radzhi) Maxted and (9) Faba (Mill.) Ledeb. (Martin et al. 2018).
Existence of large structural differences in chromosome numbers and arrangements, great variation in the genome size along with the presence of allogamy, autogamy and even functional cleistogamy make classification more difficult in Vicia (Davis and Plitmann 1970; Hollings and Stace 1978; Zohary and Plitmann 1979; Raveendar et al. 2015). Phenological, morphological, cytological, karyological and cytogenetic studies were performed for taxonomic classification and genetic relationship studies. Extensive karyological studies based on chromosome size, centromeric index and banding patterns, and cytogenetic landmarks based on fluorescence in situ hybridization (FISH) or primed in situ DNA labelling (PRINS) have been established on several species (Navratilova et al. 2003). Biotechnological approaches using DNA and proteins have been used to identify divergence among wild populations in several vetch species (Haider and El-Shanshoury 2000; Shiran and Raina 2001; Arslan et al. 2012; Chung et al. 2013; El-Bok et al. 2014; Kim et al. 2015). Restriction fragment length polymorphism (RFLP), amplified fragment Length polymorphism (AFLP), random amplified polymorphic DNA (RAPD), Start Codon Targeted (SCoT) markers, and microsatellites were used in Vicia species taxonomic classification. However, due to the much closer similarities resulted from inter- and intra-specific hybridization between certain accessions of different Vicia species rather than between accessions within each taxon were unable to assess the relationships between sativa species complex. Treating taxa within the sativa species complex whether species or subspecies seems to be a contentious issue for taxonomists of Vicia (Shiran and Raina 2001; Potokina et al. 1999, 2002; El-Bok et al. 2014; 2015; Raveendar et al. 2015; Chai et al. 2017; Han et al. 2017).
Within the genus Vicia several species are genetically, physiologically and morphologically related to certain species that are known as a Vicia sativa complex (CVS) or sativa complex or sometime they are also referred as sativa aggregate. Members of the annual species of this complex are widespread in the Mediterranean countries and in the Near and Middle East (Hollings and Stace 1978; Zohary and Plitmann 1979; Ladizinsky and Shefer 1982; Hanelt and Mettin 1989; Gil and Cubero, 1993; van de Wouw et al. 2001, 2003; Schifino-Wittmann 2000; El-Shanshoury 2007). However, members of this complex are morphologically, karyologically and ecologically variable and were considered to be in active evolution (El-Bok et al. 2015) making identification difficult and confusing. The confused taxonomic status of the CVS is probably due to hybridization between individuals of different cytotypes (Ladizinsky and Shefer 1982; Schifino-Wittmann 2000; Celiktas et al. 2006; El-Bok et al. 2015). van de Wouw et al. (2001, 2003) proposed six taxa of the sativa complex (CVS) consisting of amphicarpa, cordata, macrocarpa, nigra (syn. = V. angustifolia L.), sativa and segetalis (Schifino-Wittmann 2000; El-Shanshoury 2007). Some members in CVS were considered to be distinct species and were divided into several subspecies or varieties by several other researchers (Ladizinsky and Shefer 1982; Potokina 1997). Some species contain more cytotypes within the same taxon. For instance, Schifino-Wittmann (2000) detected 2n = 12 and 2n = 14 plants of nigra in southern Brazil. Based on molecular data, Shiran and Raina (2001) revealed that seven species (V. sativa L., V. incisa Bieb., V. nigra, V. cordata Wulfen ex Hoppe, V. amphicarpa L., V. macrocarpa (Moris) Bertol and V. angustifola L.) within the CVS shared a common ancestor and suggested that they could be referred as subspecies not species.
Earlier, some cytogenetic and karyological studies were conducted in CVS (Donnelly and Clark 1962; Ladizinsky and Shefer 1982; Yamamoto 1968, 1974; Gil and Cubero 1993; El-Shanshoury 2007). Several experiments revealed that subspecies of CVS consisted of 2n = 10, 12 and 14 chromosome complements. Within single or mixed populations of two, three or four subspecies, different chromosome numbers were observed in the natural habitats (Ladizinsky 1978; Ladizinsky and Temkin 1978; Zohary and Plitmann 1979; Han et al. 2017). A study of Ladizinsky and Shefer (1982) revealed that 2n = 10 cytotypes were found in secondary and artificial habitats while 2n = 12 were found in natural vegetation among dwarf shrubs, or in the maquis. On the other hands, the 2n = 14 cytotypes were found in dry habitats in steppes bordering the Mediterranean vegetation in Israel. Celiktas et al. (2006) reported that somatic chromosome counts of wild population of Vicia sativa L. were consistently 2n = 10, while the cultivar had 2n = 12 in Turkey. The same authors also noted that there were no significant differences between wild and cultivated cytotypes in plant height, seed number or pod number but the wild types had more branches, smaller seeds, and lower germination rates.
Numerous seed banks worldwide hold genetic resources of common vetch. Many common vetch varieties have been developed using the gene bank accessions. However, many unwanted characteristics have been co-transferred from other related gene pools. For instance, the presence of cyanoalanine, favism toxin, vicine and the neurotoxic peptide toxins in the seeds of vetch limits its usefulness in rations for certain animals and for human consumption (Enneking and Wink 2000; Firincioglu et al. 2007; Uzun et al. 2011; Kim et al. 2015). Furthermore, pod-shattering and diseases such as downy mildew (Abd El-Moneim 1993; Ahmed et al. 2000), or lack of winter hardiness (Acikgoz 1982a, b) restrict the use of common vetch accessions for improvement breeding studies. Interspecific hybrids obtained from interspecific crosses of different species such as within CVS may provide several advantages. For instance, some forms of amphicarpa produce two types of pods: aerial and underground (subterraneous pods). These vetches have ability to survive in desert-like environments and produce herbage and pods (Acikgoz 1984; Ladizinsky 2014).
Increasing the observed genetic vulnerability in many crop species have stimulated renewed interest in utilizing the secondary and if possible tertiary gene pool as a source of genetic variation in plant breeding programs. Interspecific introgression to increase the genetic diversity is one of the solutions to the problem of genetic vulnerability and influences of founder effect. This study was undertaken to create interspecific hybrids between common vetch and 5 members within the complex CVS. Genetic variations obtained could be used as a gene pool for obtaining or cloning resistance to various biotic and abiotic stress factors, insect and plant diseases, forage quality, earliness, etc., or transferring them to the cultivated lines (Celiktas et al. 2006; Tiryaki et al. 2016).
Materials and methods
Plant materials
Plant materials used in the present study consisted of six accessions of subspecies within the complex CVS. Subspecies (subsp.) studied were sativa L., amphicarpa (L.), Batt., cordata (Wulfen ex Hoppe) Asch. & Graebner, macrocarpa (Moris) Arcang., nigra (L.) Ehrh., and segetalis (Thuill.). Vicia sativa L. subsp. sativa was represented with “W-1”, (a Turkish line grown for hay and seed production), subsp. cordata was represented with “VIC701”, (an accession originated from Egypt), subsp. segetalis was represented with “VIC420”, (an accession from Greece), subsp. amphicarpa was represented with “VIC724”, (an accession of Ukraine received from Leibniz-Institute of Plant Genetics and Crop Plant Research, Germany), subsp. macrocarpa was represented with “5283”, (an accession from International Center for Agricultural Research in the Dry Areas, Syria, ICARDA), and subsp. nigra was represented with “K-1”, (an accession from Turkey).
Hybridization studies (crosses)
All the Vicia accessions mentioned above were established and were grown in controlled greenhouse conditions using conventional vetch production practices (Acikgoz 1982b). Approximately 8–10 seeds per 30 L pots filled with a field soil were sown in four consecutive times with 15-day intervals in order to synchronize flowering of the parents. A total of 30 pots per accessions were made and after emergence of seedlings, one or two plants were left in each pot. In the crosses, a sativa line, K-1, was used as female parent and other five accessions (amphicarpa, macrocarpa, segetalis, cordata and nigra) were used as male parents. One-way crosses were made in order to detect the real hybrids based on anthocyanin and white petal flower color. Anthocyanin coloration of seedlings and purple flower color are known to be completely dominant to bright green seedlings and white flower color (Donnelly and Clark 1962; Chowdhury et al. 2004). Therefore, hybrid plants were easily detected by anthocyanin coloration at seedling stage and purple flowers during flowering period. Approximately 5–10 seeds from each F1 cross were sown as mentioned above. Mature plants were selfed to obtain F2 progeny of sativa x amphicarpa, sativa x macrocarpa, sativa x segetalis, sativa x cordata and sativa x nigra. Further studies were undertaken to obtain F3 to F7 generations.
Chromosome studies
Chromosome counts were obtained from cells of young root tips of hybrids and their parental lines according to reported protocols (Hollings and Stace 2006). About 10 seeds of each parent and five seeds from each hybrid were germinated on moist filtered paper in Petri dishes. Germination studies were initiated with seeds kept at 4 °C in a refrigerator for 1 week to break dormancy, and followed by incubation at room temperature at 18-24 °C for 2–3 days. During the germination, emerging roots were checked every day to catch root tips with accumulated cells at metaphase. Root tips of 1–2 cm were cut and transferred into clean tubes. These roots were treated with a saturated aqueous solution of α-mono-bromo-naphthalene and incubated at 4 °C for 8 h. Root tips were washed in running tap water on a steel sieve for 5 min and rinsed 3–5 times with tap water, then fixed with 99% glacial acetic acid (v/v) for 30 min. After fixation, root tips were washed twice with 70% ethanol (v/v) and stored at 4 °C in a refrigerator until use.
For staining, refrigerated root tips were rinsed 3 times with plenty of distilled water for 5 min and hydrolyzed at room temperature with 1 N HCl for 20 min followed by rinsing 3 times with plenty of distilled water for 5 min. Root tips were stained using 1% aceto-orcein solution for 2 h. Root tips were removed from the stain solution and incubated at room temperature in distilled water for 10 min. The meristematic areas of the root tips were excited into small pieces using a razor blade and then transferred on microscope slides and gently crushed between slides in a drop of 1% aceto-orcein (w/v) and mounted in 45% acetic acid (v/v). Suitable mitotic plates were observed with a binocular microscope, photographed under an optical microscope type Olympus with BX51/BX52 attached digital camera and stored in a personal computer. The chromosome counts of the subspecies and hybrids studied were confirmed in at least ten cells.
Pollen viability studies
Pollens were collected from parental lines and their hybrids for pollen viability studies. Pollen viability was assessed using the basic fuchsin staining (glycerin, gelatin and crystalline basic fuchsin). Pollen grains were transferred on a microscope slide and washed with 70% ethanol (v/v) on a slide to remove oily substances and the remaining of anther tissues (Trognitz 1991). Remains of alcohol was allowed to evaporate by heating the slide. A drop of melted glycerin jelly with basic fuchsin dissolved in a water bath at 50 °C was put on slide to embed the pollen grains. Embedded pollens were covered with a coverslip. Slides were incubated in a heater at 50 °C for 1–2 min and dried upside down before at least one-day storage. Light microscope type Olympus BX51/BX52 attached digital camera was used to assess the pollen viability. Viable pollen grains appear pink colored and well-rounded while nonviable pollen grains were not stained and looked shrunken. The percentage of viable and unviable pollens was measured by examining approximately 100 pollen grains in two different specimens.
Results and discussion
A total of six hybrids were obtained using a line belonging to subspecies sativa as female parent with the genetic marker of white flower color known governed by a single recessive gene. On the other hands, the subspecies used as male parents had purple flower which is completely dominant to white flower color. Observations revealed that purple flower color was associated with anthocyanin pigmentation in the stems of seedlings and mature plants while white-flowered plants showed none of this pigmentation. These morphological differences were used as selection and confirmation markers for confirmation of successful hybridizations between subspecies sativa and other accessions. With the use of anthocyanin coloration marker, we could identify successful hybrids immediately after seedling emerged by the visualization of reddish-purple color in the stems. We selected and utilized F1 hybrids that contained reddish-purple color in the stems and purple petal colors. Since the presence of anthocyanin in the stems of seedlings and purple color in petal were dominant to light green shoots and white flower color, heterozygote (hybrids) contained these markers while those female sativa plants contained light green shoots and white flower color. Hybrids with heterozygote genotypes produced F2 progenies with a 3:1 ratio of reddish-purple color in the stems with purple petal colors, and light green shoots with white flower color. For instance, when a total of 59 F2 plants obtained from selfed sativa x nigra hybrids were studied, a total of 59 F2 plants consisting of 43 purple and 16 white flower were identified. This segregation was typically a 3:1 segregation and confirmed previous reports that the color of the flower was governed by a single gene (Donnelly 1958; Donnelly and Clark 1962; Chowdhury et al. 2004; Ladizinsky 2014).
Chromosome counts
Chromosome counts were performed in all parents and their hybrids using at least 3 root tips obtained from approximately 10 seeds of each parent and five seeds from each hybrid. For each plant sample at least 10 counts were performed. Vicia sativa subsp. sativa used in all crosses as female parent, contained 2n = 12 chromosomes. Subspecies cordata and segetalis had 2n = 10 chromosomes, macrocarpa and nigra had 2n = 12 chromosomes while amphicarpa had 2n = 14 chromosomes in root tip cells. Chromosome counts in all the F1 hybrids obtained in the hybridization studies revealed that the number of chromosomes was an average of the two parents (Fig. 1). Mitotic chromosome number (2n) of hybrids ranged from 11 to 14. Mitotic chromosome number of hybrid sativa x macrocarpa, sativa x cordata, sativa x segetalis, sativa x amphicarpa and sativa x nigra were found 2n = 12, 2n = 11, 2n = 11, 2n = 13 and 2n = 12, respectively (Fig. 1). Observed somatic chromosome numbers of 10, 11, 12, 13 and 14 within the parental lines and their hybrids indicated that there were three different basic chromosome numbers x = 5, 6 and 7. This indicated that the basic chromosome number was n = 7, from which n = 6 and n = 5 have originated in Vicia. In close agreements with our studies, the same somatic chromosome numbers of Vicia sativa subspecies reported in earlier studies were also counted in the present study (Yamamoto 1968; Hollings and Stace 1974; Ladizinsky 1978; Ladizinsky and Temkin 1978; Zohary and Plitmann 1979).
Analyses showed that parental accessions were diploid, and no aneuploidy and higher ploidy levels were detected in the present study. Results revealed that the shape and length of somatic chromosomes of the complex CVS were different. We noted that there seemed an inverse relationship between chromosome number and the size of chromosome within the complex CVS (Fig. 1). For instance, there were a reduction in chromosome size per se 2n = 14 > 2n = 12 > 2n = 10 for the complex CVS. The differences in number, shape and length of chromosomes indicated the complex CVS have great cytogenetic property that could be an indication of genetic diversity (Potokina et al. 2002; Chung et al. 2013; Kim et al. 2015). Although it is not universally acceptable in plant genomes, it could be stated that the greater genetic distance between parents, the greater the amount of genetic recombination and hybrid vigor could be obtained (Ince et al. 2010; Karaca and Ince 2018a). Higher cytological differences may cause high degree of segmental homology between the parental genomes that may result in forming a segmental alloploid, characterized in meiosis by multivalent associations, producing unbalanced gametes (Ladizinsky and Shefer 1982).
Pollen viability
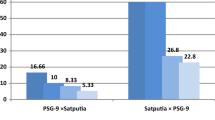
Viable and dead pollens were visualized (Fig. 2) and the viability rates of parents and their hybrids were determined. Pollen viability of the parental lines and accessions ranged from 100% in subspecies macrocarpa and amphicarpa to 44% in nigra. On the other hands, pollen viability of the hybrids ranged from 8.1% in sativa x segetalis hybrid to 0.5% in sativa x nigra hybrid. In general, pollen viability of lines and accessions was found to be very high in all of the parents with the exception in subspecies nigra. Pollen viability of subspecies sativa was 98.1%, macrocarpa 100%, segetalis 99%, nigra 44%, cordata 91%, and amphicarpa 100%. Analysis revealed that sativa x segetalis (8.1%), sativa x cordata (7.2%) and sativa x amphicarpa (7.1%) produced better pollen viability than sativa x nigra (0.5%) and sativa x macrocarpa (3.1%). Based on the study of more than 500 pollen grains, we concluded that when pollen viability rates of parental lines were high, hybrids had higher pollen fertility, thus; higher seed sets were obtained. Although this study does not have any genotypic analysis, using accession data we could state that viability and seed set of interspecific crosses can depend on the genotype of the lines used as parents. Low level of pollen viability in hybrids associated with the low level of seed retention. Zohary and Plitmann (1979) observed just negligible percentage of normally stained pollen in F1 plants of sativa x cordata hybrids. Watanabe and Yamada (1958) also reported very low pollen fertility in sativa x angustifolia hybrids. In the present study, it was observed that high rate of mortality in the pollens could be correlated a low seed set of 1 or 2 seeds per pod in sativa x nigra and in sativa x macrocarpa hybrids. Earlier, Cooper (1958) reported that intra-specific crosses in Vicia sativa were quite fertile, but they could not obtain seeds from 24 hybrid Vicia species. Watanabe and Yamada (1958) reported that sativa x angustifolia hybrids had very low (5%) pollen fertility. Also Cubero (1982) reported that hybridizations between subspecies sativa and macrocarpa, amphicarpa or angustifolia were not successful, there were very low seed sets, poor pollen fertility or no seed germination.
Varying rates of viability and mortality among the hybrids obtained by crosses of 6 subspecies clearly showed that some subspecies could hybridize more easily than others. We speculated that this was probably due to the existence of incompatibility alleles in some subspecies than other (Abdalla 1977). In the present study, we did not use reciprocal crosses because of the selection markers were valuable only crosses in which female parent was sativa. High rate of mortality in the crosses were probably due to incompatibility, which causes poor agronomic qualities of the progeny, distorted segregation or sterility and limited recombination. The low level of pollen viability observed in certain hybrids directly correlated with their infertility. All living hybrids produced mostly single-seeded, rarely two-seeded pods (Fig. 3). For instance, hybrids between sativa x segetalis produced a total of 60 pods, 58 of which consisted of a single seed while two had two seeds. A total of 17 pods consisting of 16 single seeds and one with two-seeds were obtained from hybridization of sativa x cordata. Similarly, 16 pods and 16 seeds in sativa x amphicarpa hybrid, 17 pods and 18 seeds in sativa x nigra hybrid, 34 pods and 42 seeds in sativa x macrocarpa hybrid were obtained. We speculated that the differences in chromosome numbers or chromosome structures probably prevented regular matching in meiosis. This infertility in F1 hybrid plants is consistent with the literature. Many researchers (Donnelly and Clark 1962; Mettin and Hanelt 1964; Zohary and Plitmann 1979; Ladizinsky 1981) reported that pollen viability and seed set were very low in crosses between Vicia species and subspecies at F1 generation. We speculated that crosses between sativa × segetalis, sativa × cordata and sativa × amphicarpa produce homomorphic bivalents that support better survivorship and fertility (Ladizinsky and Shefer 1982).
State of interspecific crosses in generations
Crossing studies at the interspecific level within the complex CVS were difficult, and, therefore, limited number of hybrid seeds were obtained in our experiments. It was speculated that difficulties might have come from segregation distortion, suppression of recombination, and linkage drag since they are often encountered in interspecific crosses (Watanabe and Yamada 1958; Rousi 1961). As indicated with very low seed sets, higher levels of sterility were the rule in hybrids between subspecies sativa x other subspecies of the complex CVS. It was interesting to note that even when some F1 hybrids showed considerable hybrid vigor as observed from plant heights and pod sizes; members of F2 generations were mixtures of etiolated leaves (also some variegated), lethal and some normal types. Watanabe and Yamada (1958) reported crosses between subspecies sativa and subspecies angustifolia, using angustifolia as a male parent. These authors found that pods of hybrids resembled the pods of subspecies angustifolia. In further studies, the authors showed that chromosome pairing at meiosis was very poor, and as a consequence pollen fertility was low. In our study, it was observed that after the F2 generations, fertility of some hybrids started to improve and hybrids held good seed set especially after the F3 generation (Donnelly and Clark 1962; Ladizinsky 1981). Researchers attributed this improvement as fixation of chromosome numbers in advanced generations. Zohary and Plitmann (1979) reported that many crosses between subspecies with sativa did not produce viable hybrids. These researchers found that the rate of seed retention in the F1 of the sativa x cordata hybrids was only 12–16%, whereas it was 90–95% in the parents. They also reported that fertility has improved after the F2 generation. In the present study, improvement of viability and fertility in F3 and F4 generations of some crosses were probably due to the fact that some gametes were eliminated during zygote formation and euploid genomes become advantages in further generations.
Seeds from parental and hybrids were harvested and 20 seeds from each sample were sown to obtain F1 plants. Majority of seed either not germinated, or germinated but did not survive, or did not produce seeds. A total of 27 seeds from sativa x amphicarpa, 61 seeds from sativa x macrocarpa×20 seeds from sativa × segetalis, 27 seeds from sativa x cordata and 59 seeds from sativa x nigra were harvested. Varying number of F2 plants were grown but the highest infertilities were observed in F2 generation. However, fertility rates increased during F3 and F4 generations. A total of 81 lines consisting of subspecies 2 sativa × amphicarpa, 46 sativa × macrocarpa, 9 sativa × segetalis, 11 sativa × cordata and 13 sativa x nigra were developed. Further studies using reciprocal crosses within the complex CVS would produce additional information regarding elimination, recovery and fertilization.
In order to improve the rate of viability and fertility of hybrids obtained from interspecific crosses within the complex CVS, suitable lighting, temperature and other environmental conditions, hormonal treatments, embryo rescue, embryo culture, decapitating styles and grafting the floral parts should be considered (Karaca and Ince 2018b; Karere et al. 2010). However, when the incompatibility is not in the stigma and/or style affects the zygote or post-zygotic development, these techniques would not provide amendments on the fertility ratio. Alternative approaches such as chromosome grafting based translocations via irradiating fragments of donor chromosomes to the recipient genome can be considered (Karere et al. 2010).
Conclusions
The genus Vicia has long been an important subject of scientific research because it contains several species of economic importance and provides interesting features for ecological and genetic aspects for plant scientists and breeders. In the present study, we reported several interesting findings that could be used in the breeding and genetic studies of Vicia species. We concluded that successful interspecific hybridizations within the complex CVS were possible but hybridizations have some bottlenecks. The highest infertilities were observed at F1 and F2 generation while fertility rates increased as generation progress forward and full fertility were obtained during F3 and F4 generations. These findings indicated that interspecific crosses within the complex CVS could be used to manipulate genetic and epigenetic traits involving in processes of hard seed, winter hardiness, and resistance to bacterial and fungal diseases, root knot nematodes, spider mites and aphids. However, higher success in interspecific hybridization within the complex CVS is still difficult, novel approaches such as bridge crosses, mentor pollen application, stigma grafting and reverse breeding approaches would increase the rate of success within the hybridization studies within the complex CVS. Upon obtaining successful genome transfer (hybridization), interspecific hybrids as genetic stocks could be used to introduce some nuclear and/or cytoplasmic genetic traits with desired economic values to breeder lines. Therefore, introgression via interspecific crosses in common vetch not only definitely enhance breeding efforts but also expand wealth of genetic resources.
References
Abd El-Moneim AM (1993) Selection for non-shattering common vetch, Vicia sativa L. Plant Breed 110:168–171
Abdalla MM (1977) Intraspecific unilateral incompatibility in Vicia faba L. Theor Appl Genet 50:227–233
Acikgoz E (1982a) Cold tolerance and its association with seedling morphology and chemical composition in annual forage legumes. II. Vetch (Vicia) species. Plant Breed 88:278–286
Acikgoz E (1982b) Parameters of cold tolerance in common vetch. Euphytica 31:997–1001
Acikgoz E (1984) Herbage and seed yield of subterranean vetch in response to cutting treatments. J Agron Crop Sci 153:260–263
Acikgoz E (1986) Annual forage legumes in the arid and semi-arid regions of Turkey. In: Beck DP, Materon LA (eds) Nitrogen fixation by legumes in Mediterranean agriculture. Martinus Nijhoff Publ, Leiden, pp 47–54. https://doi.org/10.1007/978-94-009-1387-5_6
Ahmed S, Akem C, Abd El Moneim AM (2000) Sources of resistance to downy mildew in narbon (Vicia narbonensis) and common (Vicia sativa) vetches. Genet Resour Crop Evol 47:153–156
Arslan E, Ertugrul K, Ozturk AB (2012) Karyological studies of some species of the genus Vicia L. (Leguminosae) in Turkey. Caryologia 65:106–113
Cakmakci S, Aydinoglu B, Karaca M (2003) Determining relationships among yield and yield components using correlation and path coefficient analyses in summer sown common vetch (Vicia sativa L.) genotypes. Pak J Bot 35:387–400
Cakmakci S, Aydinoglu B, Karaca M (2006) Heritability of yield components in common vetch (Vicia sativa L.). Acta Agric Scand, Sect B-Soil Plant Sci 56(1):54–59. https://doi.org/10.1080/09064710510008531
Celiktas N, Can E, Hatipoglu R, Avci S (2006) Comparison between a wild population and cultivar of common vetch (Vicia sativa L., Fabaceae) on cytological and agronomic characteristics. N Z J Agric Res 49:389–393
Chai X, Dong R, Liu W, Wang Y, Liu Z (2017) Optimizing sample size to assess the genetic diversity in common vetch (Vicia sativa L.) populations using start codon targeted (SCoT) markers. Molecules 22:567. https://doi.org/10.3390/molecules22040567
Chowdhury DMS, Rathjen JM, Tate ME, McDonald D (2004) Genetics of colour traits in common vetch (Vicia sativa L.). Euphytica 136:249–255
Chung JW, Kim TS, Suresh S, Lee SY, Cho GT (2013) Development of 65 novel polymorphic cDNA-SSR markers in Common Vetch (Vicia sativa subsp. sativa) using next generation sequencing. Molecules 18:8376–8392. https://doi.org/10.3390/molecules18078376
Cooper RL (1958) Hybridization in vetch. Thesis, Michigan State University
Cubero I (1982) Interspecific hybridization in Vicia. In: Hawtin G, Webb C (eds) Faba bean improvement, proceedings of the faba bean conference. Martinus Nijhoff Publ, Leiden, pp 91–108
Davis PH, Plitmann U (1970) Vicia L. In: Davis PH (ed) Flora of Turkey. The University Press, Edinburgh, pp 274–325
Dong R, Dong D, Luo D, Zhou Q, Chai X, Zhang J, Xie W, Liu W, Dong Y, Wang Y, Liu Z (2017) Transcriptome analyses reveal candidate pod shattering-associated genes involved in the pod ventral sutures of common vetch (Vicia sativa L.). Front Plant Sci 8:649. https://doi.org/10.3389/fpls.2017.00649
Donnelly ED (1958) Inheritance of white flower color in common vetch, Vicia sativa. Agron J 50:763–764
Donnelly ED, Clark EM (1962) Hybridization of the genus Vicia. Crop Sci 2:141–145
El-Bok S, Zoghlami-Khelil A, Brahim TB, Ouji A, Hassen H, Lamine O, Jabri C, Douggari R, El-Gazzah M (2014) Chromosome number and karyotype analysis of some taxa of Vicia genus (Fabaceae): revision and description. Int J Agric Biol 16:1067–1074
El-Bok S, Zoghlami-Khelil A, Dougari R, Jabri C, Lamine O, El-Gazzah M (2015) Vicia sativa subsp. sativa (Fabaceae): new taxonomic division in Tunisia based on karyological data. Pak J Agric Sci 52:279–283
El-Shanshoury AR (2007) Characterization of infraspecific electrophoretic genetic variation within Vicia sativa subspecies sativa population. J Biol Sci 7:918–924
Enneking D, Wink M (2000) Towards the elimination of antinutritional factors in grain legumes. In: Knight R (ed) Linking research and marketing opportunities for pulses in the 21st century, current plant science and biotechnology in agriculture. Kluwer Academic Publishers, Dordrecht, pp 375–384
Firincioglu HK, Tate M, Unal S, Dogruyol L, Ozcan I (2007) A selection strategy for low toxin vetches (Vicia sativa spp.). Turk J Agric For 31:303–311
Frediani M, Maggini F, Gelati MT, Cremonini R (2004) Repetitive DNA sequences as probes for phylogenetic analysis in Vicia genus. Caryologia 57:379–386
Geogieva N (2018) Suitability of vetch (Vicia sativa L. and V. villosa Roth) cultivars for organic farming conditions. Pak J Bot 50:161–167
Gil J, Cubero JI (1993) Multivariate analysis of the Vicia sativa L. aggregate. Bot J Linn Soc 113:389–400
Haider AS, El-Shanshoury AR (2000) Variability of storage proteins and esterase isozymes in Vicia sativa subspecies. Biol Plant 43:205–209
Han Y, Liu Y, Wang H, Liu X (2017) The evolution of Vicia ramuliflora (Fabaceae) at tetraploid and diploid levels revealed with FISH and RAPD. PLoS ONE 12(1):e0170695. https://doi.org/10.1371/journal.pone.0170695
Hanelt P, Mettin D (1989) Biosystematics of the genus Vicia L. (Leguminosae). Annu Rev Ecol Evol Syst 20:199–223
Hollings E, Stace CA (1974) Karyotype variation and evolution in the Vicia sativa aggregate. New Phytol 73:195–208
Hollings E, Stace CA (1978) Morphological variation in the Vicia sativa L. aggregate. Watsonia 12:1–14
Hollings E, Stace CA (2006) Karyotype variation and evolution in the Vicia sativa L. aggregate. New Phytol 73:195–208
Hueze V, Tran G, Baumont R (2011) Common vetch (Vicia sativa). Feedipedia 12:53–62
Ince AG, Karaca M, Onus AN (2010) A reliable gender diagnostic PCR assay for jojoba (Simmondsia chinensis (Link) Schneider). Genet Resour Crop Evol 57:773. https://doi.org/10.1007/s10722-009-9516-1
Karaca M, Ince AG. (2018a). Identification of heterotic group for genetic and epigenetic studies in maize. In: Proceedings og international congress on agriculture and animal science, 7–9 November, Alanya, Turkey, pp 546–552
Karaca M, Ince AG (2018b) New generation plant breeding methods (Molecular Plant Breeding) some advantages & disadvantage. Res J Agric Sci 11:39–49
Karere GM, Lyons LA, Froenicke L (2010) Enhancing radiation hybrid mapping through whole genome amplification. Hereditas 147:103–112. https://doi.org/10.1111/j.1601-5223.2010.02166.x
Kim TS, Raveendar S, Suresh S, Lee GA, Lee JR, Cho JH, Lee SY, Ma KH, Cho GT, Chung JW (2015) Transcriptome analysis of two Vicia sativa subspecies: mining molecular markers to enhance genomic resources for vetch improvement. Genes 6:1164–1182. https://doi.org/10.3390/genes6041164
Ladizinsky G (1978) Chromosomal polymorphism in wild populations of Vicia sativa. Caryologia 31:233–241
Ladizinsky G (1981) Consequences of hybridization in Vicia sativa aggregate. Heredity 47:431–438
Ladizinsky G (2014) Chromosomal polymorphism in wild populations of Vicia sativa L. Caryologia 31:233–241
Ladizinsky G, Shefer Y (1982) Polyploidy in the Vicia sativa aggregate. New Phytol 91:541–547
Ladizinsky G, Temkin R (1978) The cytogenetic structure of Vicia sativa aggregate. Theor Appl Genet 53:33–42
Martin E, Yildiz HK, Kahraman A, Binzat OK, Eroglu HE (2018) Detailed chromosome measurements and karyotype asymmetry of some Vicia (Fabaceae) taxa from Turkey. Caryologia 71:224–232. https://doi.org/10.1080/00087114.2018.1460058
Maxted N (1993) A phenetic investigation of Vicia L. subgenus Vicia (Leguminosae–Vicieae). Bot J Linn Soc 111:155–182
Mettin D, Hanelt P (1964) Cytosystematische untersuchungen in der artengruppe urn Vicia sativa L. Kulturpflanze 12:163–225
Mikic A, Mihailovic V, Cupina B, Milic D, Katic S, Karagic D, Pataki I, D’Ottavio P, Kraljevic-Balalic M (2014) Forage yield components and classification of common vetch (Vicia sativa L.) cultivars of diverse geographic origin. Grass Forage Sci 69:315–322
Navratilova A, Neumann P, Macas J (2003) Karyotype analysis of four Vicia species using in situ hybridization with repetitive sequences. Ann Bot 91:921–926
Potokina EK (1997) Vicia sativa L. aggregate (Fabaceae) in the flora of the former USSR. Genet Resour Crop Evol 44:199–209
Potokina E, Tomooka N, Vaughan DA, Alexandrova T, Xu R-Q (1999) Phylogeny of Vicia subgenus Vicia (Fabaceae) based on analysis of RAPDs and RFLP of PCR-amplified chloroplast genes. Genet Resour Crop Evol 46:149–161
Potokina E, Blattner F, Alexandrova T, Bachmann K (2002) AFLP diversity in the common vetch (Vicia sativa L.) on the world scale. Theor Appl Genet 105:58–67
Raveendar S, Lee GA, Jeon YA, Lee YJ, Lee JR, Cho GT, Cho JH, Park JH, Ma KH, Chung JW (2015) Cross-amplification of Vicia sativa subsp. sativa microsatellites across 22 other Vicia species. Molecules 20:1543–1550. https://doi.org/10.3390/molecules20011543
Rousi A (1961) Cytotaxonomical studies on Vicia cracca L. and V. tenuifolia Roth. I. Chromosome number and karyotype evolution. Hereditas 47:81–110
Schifino-Wittmann MT (2000) The cytogenetics and evolution of forage legumes from Rio Grande do Sul: a review. Genet Mol Biol 23:989–995
Seymour M, Siddique KHM, Brandon N, Martin L, Jackson E (2002) Response of vetch (Vicia spp.) to plant density in south-western Australia. Aust J Exp Agric 42:1043–1051
Shiran B, Raina SN (2001) Evidence of rapid evolution and incipient speciation in Vicia sativa species complex based on nuclear and organellar RFLPs and PCR analysis. Genet Resour Crop Evol 5:519–532
Tate M, Ennenking D (2006) Common vetch (Vicia sativa ssp. sativa): feed or future food. Grain Legumes 47:16–17
Tiryaki GY, Cil A, Tiryaki I (2016) Revealing seed coat colour variation and their possible association with seed yield parameters in common vetch (Vicia sativa L.). Int J Agron. https://doi.org/10.1155/2016/1804108
Trognitz BR (1991) Comparison of different pollen viability assays to evaluate pollen fertility of potato dihaploids. Euphytica 56:143–148
Uzun A, Gucer S, Acikgoz E (2011) Common vetch (Vicia sativa L.) germplasm: correlations of crude protein and mineral content to seed traits. Plant Foods Hum Nutr 66:254–260
Van de Wouw M, Maxted N, Chabane K, Ford-Lloyd BV (2001) Molecular taxonomy of Vicia ser. Vicia based on amplified fragment length polymorphisms. Plant Syst Evol 229:99–105
Van de Wouw M, Maxted N, Chabane K, Ford-Lloyd BV (2003) A multivariate and cladistic study of Vicia L. ser. Vicia (Fabaceae) based on analysis of morphological characters. Plant Syst Evol 237:19–39
Watanabe K, Yamada T (1958) Studies on the interspecific hybridization of common vetch Vicia sativa, and Yahazuendo, V. angustifolia var. segetalis. Bull Natl Inst Agric Sci Jpn Ser G 15:109–145
Yamamoto K (1968) On the interspecific hybrids between Vicia sativa and V. macrocarpa. Jpn J Breed 18:283–290
Yamamoto K (1974) Hybrid plants between two races of Vicia amphicarpa having 2n = 10 and 2n = 14 chromosomes. Jpn J Breed 24:73–80
Zohary D, Plitmann U (1979) Chromosome polymorphism, hybridization and colonization in the Vicia sativa group (Fabaceae). Plant Syst Evol 31:143–156
Acknowledgements
This study was financially supported in part by the Scientific and Technological Research Council of Turkey (Project No: 214O224).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kartal, G.K., Senbek, G., Karaca, M. et al. Hybridization studies in Vicia sativa complex. Euphytica 216, 29 (2020). https://doi.org/10.1007/s10681-020-2566-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-020-2566-3