Abstract
Multiprotein bridging factor 1 (MBF1) is an evolutionarily conserved transcriptional co-activator in archaea and eukaryotes that has been demonstrated previously to play an important role in various types of stress response. In this study, a full-length MBF1 cDNA sequence (VvMBF1) was isolated from grape (Vitis labrusca × V. vinifera) and was found to be up-regulated in the leaves of grape plants following both drought and abscisic acid (ABA) treatments. Furthermore, constitutive expression of VvMBF1 in Arabidopsis thaliana enhanced drought stress tolerance in transgenic plants. To gain further insight into the role of VvMBF1 in drought resistance, we analyzed various physiological parameters related to stress response in transgenic Arabidopsis lines and found that transgenic plants were better able to prevent water loss under stress conditions than wild-type (WT) plants. This was likely due to an increase in the sensitivity of stomata to ABA, which is a well-known signaling molecule in plant drought response. In addition, dehydration stress yielded less cell damage to transgenic plants than WT plants. We also found that VvMBF1-expressing transgenic lines exhibited up-regulation of two drought-responsive genes that are known to function in the ABA-dependent drought-response pathway. Taken together, these results reveal that VvMBF1 is likely involved in drought-responsiveness in grape, and confers increased drought tolerance in transgenic plants, possibly through an ABA-dependent signal transduction pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drought stress is one of the most important environmental factors limiting crop growth and geographical distribution, and can have a devastating impact on production (Zhu 2002). Since plants are sessile organisms, they have evolved multifaceted mechanisms to respond and adapt to drought conditions, including morphological, physiological, biochemical and molecular adaptations. When exposed to drought stress, plants produce high levels of the hormone abscisic acid (ABA), which induces stomatal closure as well as the expression of various genes. Indeed, while it has been shown that exogenous application of ABA can mimic such drought stress responses (Bray 1997; Shinozaki and Yamaguchi-Shinozaki 1997), not all drought stress-induced genes are responsive to exogenous ABA treatment. In line with this, it has been suggested that both ABA-independent and ABA-dependent signal transduction cascades are associated with drought stress in plants (Ingram and Bartels 1996; Shinozaki and Yamaguchi-Shinozaki 1996; Leung and Giraudat 1998).
Multiprotein bridging factor 1 (MBF1) has been suggested to play an important role in ABA-mediated stress response in plants (Tsuda et al. 2004; Kim et al. 2007; Arce et al. 2010). This protein is an evolutionarily conserved transcriptional co-activator that enhances the transcription of its target genes by bridging transcription factors and TATA-box-binding protein (Li et al. 1994; Takemaru et al. 1997). Co-activators are a class of transcription factor capable of interconnecting a regulating DNA-binding protein with a component of the basal transcription machinery, thus allowing transcriptional activation to proceed. In yeast and animals, MBF1 has been found to play a role in the regulation of diverse processes, including endothelial cell differentiation, hormone-regulated lipid metabolism, central nervous system development, and histidine metabolism (Takemaru et al. 1998; Brendel et al. 2002; Busk et al. 2003; Liu et al. 2003) respectively.
Arabidopsis thaliana contains three distinct homologs encoding MBF1, all of which can complement MBF1 deficiency in yeast (Tsuda et al. 2004). While the expression of AtMBF1a and AtMBF1b are developmentally regulated (Tsuda and Yamazaki 2004), that of AtMBF1c is up-regulated in response to heat, hydrogen peroxide (H2O2), dehydration, salinity, pathogen infection and application of the plant hormones ABA or salicylic acid (Tsuda and Yamazaki 2004; Suzuki et al. 2005, 2008, 2011; Arce et al. 2010). Functionally, AtMBF1a appears to be involved in stress tolerance as well as in ethylene and glucose signaling (Kim et al. 2007), while AtMBF1c plays a role in leaf cell cycle and expansion (Toji et al. 2009). Furthermore, the AtMBF1 genes have also been suggested to play a role in the ABA-dependent inhibition of germination (Mauro et al. 2012). Various MBF1 genes have also been isolated and characterized from other plant species. For example, the expression of MBF1 genes from potato, tomato, Retama raetam, tobacco and wheat have all been found to be induced by various types of abiotic stress, biotic stress and/or hormone treatment (Godoy et al. 2001; Arce et al. 2006; Zegzouti et al. 1999; Pnueli et al. 2002; Rizhsky et al. 2002; Zhang et al. 2009). In fact, it appears that virtually all plant MBF1 genes that have been studied to date may be important for biotic and abiotic stress response.
Although grape is one of the most widely cultivated and economically important fruit crops in the world, its susceptibility to drought limits its use in areas where irrigation is not readily available. Therefore, improvement of drought tolerance in this crop may result in an expansion of its production regions, thus contributing to the maintenance of a sustainable grape industry. Since MBF1 homologs have yet to be isolated from grape, we sought to clone such a gene (VvMBF1) and characterize its response to both drought stress and ABA treatment. In addition, we also endeavored to provide insight into the mechanism by which VvMBF1 is able to elicit stress tolerance through its heterologous expression in Arabidopsis. These findings not only further our understanding of the role of MBF1 genes in drought tolerance, but may also provide a foundation for the potential development of drought resistant crops in the future.
Materials and methods
Plant materials
The grape variety ‘Kyoho’ (Vitis labrusca × V. vinifera) used in this study was grown in the greenhouse at Northwest Agriculture & Forestry University, Yangling, Shaanxi, China (34°200′N, 108°240′E). Two-year-old ‘Kyoho’ grape seedlings were used for gene cloning and expression analyses following drought and ABA treatments. Arabidopsis thaliana Columbia-0 (WT) plants were grown in growth chambers under intense light (150 μmol m−2 s−1) at 21 °C, with 70 % relative humidity and a day length of 16 h.
Cloning and sequence analysis of VvMBF1
Total RNA was extracted from the leaves of 2-year-old ‘Kyoho’ seedlings, which were grown under conditions detailed above, using the E.Z.N.A.® Plant RNA Kit (Omega Bio-tek, Norcross, GA, USA) following the manufacturer’s protocol, with residual DNA removed using DNase I (Promega, Madison, WI, USA). First-strand cDNA was synthesized using 500 ng of total RNA as template with PrimeScript™RTase (TaKaRa Biotechnology, Dalian, China) according to the manufacturer’s instructions. The full-length VvMBF1 cDNA was obtained using the BD SMA RT™ RACE cDNA Amplification kit (Clontech, Palo Alto, CA, USA). Gene-specific primers GSP1 (5′-CCG CCA AGA AGG ACG AGA AAG CTG TC-3′) and GSP2 (5′-TTT CCT CGA AGT TTC ACT CCA AGA GCC C-3′) were designed for 5′ and 3′ RACE amplification based on a known partial VvMBF1 cDNA sequence (GenBank accession no. FG269343). The resulting PCR products were cloned and sequenced, and full-length cDNA was obtained through the assembly of the 5′ and 3′ sequences. Subsequently, the full VvMBF1 open reading frame (ORF) was cloned into the pGEM-Teasy vector (Promega, Madison, WI, USA) using primers F (5′-ATG GCA GGA GTC GGA C-3′) and R (5′-TTA TTT CTT TCC TCG AAG TT-3′). This plasmid was re-sequenced for confirmation purposes and was termed pGEM-Teasy-VvMBF1. Amino acid sequences of homologous MBF1 proteins from other plant species were obtained from NCBI using Blastp (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Multiple sequence alignment of deduced protein sequences and phylogenetic analysis were carried out with MEGA 5.0 software using the neighbor-joining (NJ) method.
Drought and ABA treatment in grape
Two-year-old ‘Kyoho’ grape seedlings were subjected to drought stress treatment by withholding water when the third to fifth leaves were fully expanded. Leaves were harvested 24, 48, 72, 96, 120, 144 and 168 h post-treatment. Drought-stressed plants were then re-watered to soil saturation and leaves were collected 24 h after re-watering. Well-watered seedlings were used as a control. ABA treatment was performed by spraying leaves with 300 µM ABA, followed by sampling 1, 6, 12, 24 and 48 h post-treatment. Leaves sprayed with sterile distilled water were used as a negative control. At every time point of each treatment, six leaves from six separate plants were combined to form one sample. All treatments were performed in triplicate. Plant samples were immediately frozen in liquid nitrogen and stored at −80 °C for subsequent RNA isolation and expression analysis.
Generation and selection of transgenic Arabidopsis expressing VvMBF1
The coding sequence of VvMBF1 (with BamHI and KpnI sites added to its 5′ and 3′ ends, respectively) was amplified from pGEM-Teasy-VvMBF1 using gene-specific primers F1 (5′-AAA GGA TCC ATG GCA GGA GTC GGA C-3′; BamHI site underlined) and R1 (5′-CGC GGT ACC TTA TTT CTT TCC TCG AAG TT-3′; KpnI site underlined), and was inserted immediately downstream of the CaMV 35S promoter in the plant over-expression vector, pCambia2300 (Cambia, Brisbane, QLD, Australia). The resulting vector was transferred into Agrobacterium tumefaciens strain EHA105 and Arabidopsis plants were transformed using the floral dip method (Clough and Bent 1998). T0 seeds were harvested and sown on MS medium (Murashige and Skoog 1962) supplemented with 50 mg l−1 kanamycin. Quantitative real-time RT-PCR was used to select the three lines (L7, L20, and L21) with the highest VvMBF1 expression levels from 52 independent lines, from which T3 homozygous lines were generated and subsequently utilized for all further experiments.
Expression of VvMBF1 and drought-responsive genes in Arabidopsis following drought treatment
In order to provide further evidence that the VvMBF1 gene is responsive to drought, homozygous T3 lines, as well as wild-type (WT) plants, were grown on MS medium for 10 days, after which time they were transferred to liquid MS medium or liquid MS medium supplemented with 300 mM mannitol (commonly used to mimic drought stress). After 3 days, seedlings were sampled for expression of VvMBF1 and other drought-responsive genes (AtRD22, AtRD29B, AtRD29A and AtERD1) using quantitative real-time RT-PCR assays.
Germination of Arabidopsis seeds under mannitol stress
For germination of Arabidopsis seeds, approximately 100 seeds from each of the three selected T3 homozygous lines, as well as WT plants, were vernalized for 3 day at 4 °C, surface-sterilized based on standard protocols (Weigel and Glazebrook 2002), and then sown on MS medium or MS medium supplemented with 300 mM mannitol. The percentage of germinated seeds was calculated based on the number of seedlings that had reached the cotyledon stage at 2 weeks (Saleki et al. 1993). All germination assays were performed in triplicate.
Drought stress treatment of Arabidopsis plants
Ten-day-old transgenic and WT seedlings grown on MS medium plates were transferred to pots filled with humus, and were watered well for 7 days. During the next 7 days, the seedlings were not watered, after which point it was visually apparent that they were suffering from drought stress. Following drought treatment, plants were re-watered. Phenotypes before and after treatment were monitored and photographed using a digital camera (Canon, Tokyo, Japan). Survival rates were also examined at the end of the experiment.
Measurements of water loss rate, electrolyte leakage, malondialdehyde and proline contents
To determine the rate of water loss, leaves (approximately 0.2 g) were harvested from 3-week-old transgenic and WT seedlings, and were placed on dry filter paper. Fresh weights (FW) were measured every 10 min for 50 min to determine the rate of water loss. Electrolyte leakage (EL), malondialdehyde (MDA) and proline content, as well as the accumulation of reactive oxygen species (ROS: O2 − and H2O2), were measured at the end of the water loss experiment.
For determination of EL, leaves were placed in deionized water, shaken on a gyratory shaker at 100 rpm for 1 h at room temperature, and the conductivities of the resulting solutions (C1) were determined. Subsequently, the leaves were boiled for 10 min in deionized water, cooled to room temperature, and the conductivities of the solutions (C2) were re-measured. The C1 to C2 (C1/C2) ratios were then calculated and used as a measure of relative electrolyte leakage from the leaves (Liu et al. 2006).
Malondialdehyde (MDA) content was measured by homogenizing leaves in 10 ml of TCA (trichloroacetic acid) and subsequently centrifuging at 4,000 rpm for 10 min. 2 ml of the resulting supernatants were heated with 2.0 ml of 0.67 % (w/v) TBA (thiobarbituric acid) at 100 °C for 15 min, cooled quickly on ice, and then centrifuged at 4,000 rpm for 5 min, 4 °C. Absorbances were then measured at 532 nm (A532), 600 nm (A600), and 450 nm (A450) using a spectrophotometer (Hitachi Limited, Tokyo, Japan). MDA content was calculated using the equation: C (MDA content) = 6.45 (A532 − A600) − 0.56 × A450 (Heath and Packer 1968).
For measurements of proline content, leaves were homogenized in 5 ml of 3 % sulphosalicylic acid and centrifuged at 4,000 rpm for 5 min, 4 °C. 2 ml of the resulting supernatants were incubated with 2 ml of ninhydrin reagent (2.5 % (w/v) ninhydrin, 60 % (v/v) glacial acetic acid, and 40 % 6 M phosphoric acid) and 2 ml of glacial acetic acid at 100 °C for 30 min. Reactions were terminated by snap cooling in an ice bath, 4 ml of toluene was added, and the mixtures were vortexed then incubated at 23 °C for 24 h. Proline contents were then determined as described previously (Bates et al. 1973).
Detection of O2 − and H2O2
For visualization of O2 − and H2O2, leaves were stained with nitro blue tetrazolium (NBT; Sigma, Steinheim, Germany) and diaminobenzidine (DAB; Sigma), respectively. In the case of O2 −, leaves were immersed in HEPES buffer (pH 7.5) containing 6 mM NBT for 2 h or until blue spots appeared, as described previously (Kim et al. 2011). In the case of H2O2, leaves were placed in 1 mg ml−1 DAB solution until brown spots became visible (Kotchoni et al. 2006). In both instances, stained samples were then de-pigmented in 80 % (v/v) ethanol at 80 °C for 2 h. Images were acquired using a digital camera.
Measurement of ABA content
Seven-day-old seedlings grown on MS medium were transferred onto new media containing basal MS, or MS supplemented with 300 mM mannitol, and grown for 1 week. Seedlings were harvested for ABA extraction and measurement according to the method of Liu and Zhang (2010). ABA was extracted using organic solvents (Liu and Zhang 2010) while HPLC (high performance liquid chromatography (Shimadzu, Kyoto, Japan)) fractionation was carried out using a C18 chromatography column (250 mm × 4.6 mm, 5 μm) with a mobile phase of 30 % of acetonitrile and 70 % 0.02 mol l−1 acetic acid. Aliquots of 20 μl of the extracted solutions were used for detection at 262 nm with a column temperature of 25 °C. ABA content was determined based on chromatographic peak area.
Measurement of stomatal closure in response to ABA treatment
Stomatal closure assays were conducted as described previously (Pei et al. 1997). Rosette leaves from WT plants and three of each T3 homozygous transgenic line were placed on the surface of a solution containing 50 µM CaCl2, 10 mM KCl, 10 mM MES (2-(N-morpholino) ethanesulfonic acid)-Tris, pH 6.15, and were exposed to light for 2 h. ABA (Sigma) was then added to the solution at concentrations of either 1 or 5 µM, respectively, and samples were incubated for a further 2 h. Following ABA treatment, 40 stomatal apertures were measured from each leaf as the ratio of stomatal width to length. All assessments were carried out in triplicate.
Quantitative real-time RT-PCR
Total RNA extractions from the leaves of ‘Kyoho’ grape and Arabidopsis seedlings, as well as first-strand cDNA synthesis, were carried out as described above. Subsequent quantitative real-time PCR analyses were conducted using SYBR green (TaKaRa Biotechnology) on an IQ5 real-time PCR instrument (Bio-Rad, Hercules, CA, USA) with the following thermal profile: 95 °C for 30 s, 40 cycles of 95 °C for 5 s, and 60 °C for 30 s. Following PCR amplification, dissociation curves were generated to verify that only a single product was amplified in each case using the following program: 95 °C for 15 s, followed by a constant increase from 60 to 95 °C. Fragments of the grape Actin1 (GenBank accession no. AY680701) or Arabidopsis Actin2 (GenBank accession no. At3G18780) transcripts were utilized as internal controls, respectively. Primers used for qRT-PCR were designed using Primer 5.0 software and are listed in Supplemental Table S1. Relative expression levels were determined using IQ5 software and the normalized expression method.
Statistical analysis
All experiments were carried out in triplicate as three independent trials. Results were expressed as means and standard errors, and were calculated using Microsoft Excel. Paired t tests were performed using SPSS Statistics 17.0 software (IBM China Company Ltd., Beijing, China) to assess significant differences.
Results
Cloning and sequence analysis of VvMBF1
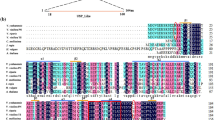
Gel electrophoresis of VvMBF1 RACE products obtained from ‘Kyoho’ grape leaf tissue yielded bands of approximately 510 bp (5′ RACE) and 650 bp (3′ RACE) (Supplemental Figure S1). The full-length assembled cDNA comprised 767 bp, including a 60 bp 5′ untranslated region (UTR), a 429-bp ORF, and a 246 bp 3′ UTR. This sequence encodes a deduced protein 142 amino acids in length and contains a highly conserved MBF1 domain (Supplemental Figure S2). The deduced amino acid sequence of VvMBF1 exhibits a relatively high level of identity (between 77 and 84 %) with other plant MBF1 proteins shown previously to be responsive to various types of stress (Fig. 1). Furthermore, both sequence alignments (Fig. 1a) and phylogenetic analysis (Fig. 1b) indicated that VvMBF1 exhibited higher degree of identity with AtMBF1a and AtMBF1b than with AtMBF1c, and was thus classified as a member of plant group Ι MBF1 proteins (Tsuda and Yamazaki 2004).
Comparison of VvMBF1 with other plant MBF1 proteins. a Multiple alignments of predicted amino acid sequences of VvMBF1 with other plant MBF1 proteins. b Phylogenetic analysis of the deduced VvMBF1 protein sequence with other plant MBF1 proteins. MBF1 proteins utilized in this analysis were as follows: XP_002280992.1 (VvMBF1; V. vinifera a), XP_003634667.1 (putative V. vinifera MBF1-like; V. vinifera a-like), AAK44095.1 (Arabidopsis AtMBF1a; A. thaliana a), AAG41491.1 (Arabidopsis AtMBF1b; A. thaliana b), AEE76905.1 (Arabidopsis AtMBF1c; A. thaliana c), AAD46402.1 (Lycopersicon esculentum), BAB88859 (Nicotiana tabacum), AAL32037.2 (Retama raetam), CAA89698.1 (Ricinus communis), AAF81108.1 (Solanum tuberosum), ACO36694.1 (Triticum aestivum), XP_002331813.1 (Populus trichocarpa), XP_003527342.1 (Glycine max), ADX60234.1 (Oryza sativa), ACF06505.1 (Elaeis guineensis), ACG33346.1 (Zea mays)
VvMBF1 expression is induced by drought and ABA treatment in grape
Quantitative real-time RT-PCR assays of VvMBF1 transcript levels in the leaves of ‘Kyoho’ grape in response to drought treatment indicated that VvMBF1 expression was significantly induced by drought 72–168 h post-treatment, with peak levels of expression occurring 144 h following treatment (Fig. 2a). At its most elevated point, VvMBF1 expression in drought-treated plants was approximately threefold higher than the negative control. Once drought-treated plants were re-watered, VvMBF1 expression declined until transcript levels were once again similar to untreated controls. In the case of ABA treatment, similar assays demonstrated that the expression of VvMBF1 was significantly induced 6 h post-treatment, after which time expression decreased back to levels that were comparable to untreated controls (Fig. 2b).
Quantitative real-time RT-PCR analysis of VvMBF1 expression in ‘Kyoho’ plants exposed to drought and ABA treatments. Relative expression profiles of VvMBF1 following drought (a) and ABA (b) treatments. Blocks represent the mean values ± SD from three independent experiments. Asterisks indicate statistically significant differences (**P < 0.01, Student’s t test) compared to untreated control plants
Validation of transgenic Arabidopsis lines expressing VvMBF1
The role of VvMBF1 in stress tolerance was further investigated via the constitutive expression of VvMBF1 in Arabidopsis. A total of 52 independent transgenic lines were generated and confirmed by quantitative real-time RT-PCR (Supplemental Figure S3). Three of these lines (L7, L20 and L21) were selected for the generation of homozygous T3 lines, which were used for all further analyses based on their high levels of VvMBF1 expression in leaf tissue as evidenced by quantitative real-time RT-PCR (Fig. 3a).
Validation of VvMBF1 expression and drought-responsiveness in transgenic Arabidopsis. a Quantitative real-time RT-PCR analysis of VvMBF1 transcript levels in untreated WT and transgenic lines (L7, L20 and L21). b Quantitative real-time RT-PCR analysis of VvMBF1 transcript levels in WT and transgenic lines following mannitol-induced osmotic stress. Datas represent mean values ± SD from three independent experiments. Asterisks indicate statistical significance (**P < 0.01, Student’s t test) in comparison with WT
Confirmation of the drought-responsiveness of VvMBF1 in these particular transgenic Arabidopsis lines was obtained using quantitative real-time RT-PCR to assay expression levels in seedlings following treatment with mannitol. Results from this experiment demonstrated that the levels of VvMBF1 transcript in plants derived from all three transgenic lines following drought stress were 4–6 times higher than transcript levels in untreated transgenic lines that (Fig. 3b), validating their use throughout the remainder of the study.
Constitutive expression of VvMBF1 in transgenic Arabidopsis enhances drought tolerance
In an attempt to determine whether the expression of VvMBF1 in transgenic Arabidopsis lines provided any benefits in terms of drought tolerance compared to untransformed lines, we initially tested their ability to germinate on growth medium containing mannitol. In the absence of mannitol, nearly 100 % of seeds from both transgenic and WT plants germinated successfully. In contrast, when sown on medium containing mannitol, seeds from transgenic lines exhibited significantly higher germination rates than those of WT (Fig. 4a). Indeed, while the average germination rate of transgenic seeds was almost 90 %, an average of only 37.45 % of WT seeds germinated (Fig. 4b).
Drought stress response of WT and VvMBF1-expressing transgenic Arabidopsis lines. a Representative images of WT and transgenic (L7, L20 and L21) seedlings 2 weeks after seeds were sown on MS basal medium or MS basal medium supplemented with 300 mM mannitol (to induce osmotic stress). b Germination rates of seeds sown on MS basal medium or MS basal medium supplemented with 300 mM mannitol. c Representative images of 2-week-old potted WT and transgenic lines deprived of water for 7 days and then re-watered. d Survival rates of WT and transgenic lines 24 h after re-watering. Datas in graphs represent mean values ± SD from three independent experiments. Asterisks indicate statistical significance (*P < 0.05, **P < 0.01, Student’s t test) of differences between transgenic lines and WT
To garner further evidence that VvMBF1 functions to enhance drought tolerance in transgenic Arabidopsis, both transgenic and WT plants were subjected to water restriction for 7 days, after which time morphological differences became apparent. As shown in Fig. 4c, all WT plants exhibited severe water loss-related symptoms, with considerable wilting following the drought treatment. In contrast, only slight wilting was observed in the majority of transgenic plants. 24 h after re-watering, almost all WT plants had died (8.47 % recovery rate), whereas most of the transgenic plants had resumed normal growth (ranging from 47.72 to 84.14 % recovery rate; Fig. 4d).
Transgenic Arabidopsis lines display distinct physiological changes as a result of drought stress compared to WT
To evaluate the physiological response of transgenic and WT plants to drought stress, leaves from 3-week-old transgenic and WT plants were assessed for water loss following detachment, EL, the contents of MDA, proline and ABA, as well as accumulation of O2 − and H2O2 (which are all important markers of drought tolerance). Measurement of the rate of water loss from the leaves of both transgenic and WT plants at various time points following leaf detachment demonstrated that transgenic lines displayed less water loss than WT lines at every time point (Fig. 5a). Furthermore, while EL and MDA contents were found to be similar in the leaves of well-watered transgenic and WT plants, the contents of both were significantly lower in transgenic lines than WT lines following dehydration treatment (Fig. 5c, e). Similarly, while there were no significant differences between the ABA and proline contents of transgenic and WT lines grown under normal conditions. However, when exposed to dehydration treatment, transgenic lines exhibited significantly higher proline contents, but little difference in ABA content, compared to WT lines (Fig. 5b, d).
Physiological changes associated with drought stress response in WT and VvMBF1-expressing transgenic Arabidopsis plants. a Rate of water loss from detached leaves of WT and transgenic (L7, L20 and L21) plants. b ABA content in the leaves of WT and transgenic seedlings. c Relative electrolyte leakage from detached leaves of WT and transgenic seedlings. d Proline content in the leaves of WT and transgenic seedlings. e MDA content in the leaves of WT and transgenic seedlings. f Histochemical staining assay of O2 − and H2O2 accumulation with nitro blue tetrazolium (NBT) and diaminobenzidine (DAB), respectively, in WT and transgenic leaves following drought stress. In all cases, datas represent mean values ± SD from three independent experiments. Asterisks indicate statistical significance (*P < 0.05, **P < 0.01, Student’s t test) of differences between transgenic and WT plants
As water stress can also lead to increases in reactive oxygen species, such as O2 − and H2O2, which can result in oxidative damage, we also assessed the accumulation of these two chemicals in response to drought treatment in transgenic and WT Arabidopsis. Following dehydration treatment, leaves from WT plants exhibited a deeper staining than transgenic plants in both cases, indicating that transgenic plants expressing VvMBF1 accumulated less O2 − and H2O2 during drought stress (Fig. 5f).
Stomata from transgenic Arabidopsis lines exhibit increased sensitivity to ABA treatment
Since water loss from plants largely depends upon stomatal regulation, which is, in turn, affected by ABA, we compared stomatal apertures from the leaves of transgenic and WT plants treated with different concentrations of ABA. Untreated transgenic and WT lines displayed very similar, and relatively high, width to length ratios signifying fully open stomata. Conversely, following ABA treatment transgenic lines exhibited lower stomatal width to length ratios than WT plants (Fig. 6), indicating that stomata of transgenic lines had undergone a higher degree of closure than those of WT.
Stomatal closure in response to ABA treatment in WT and VvMBF1-expressing transgenic Arabidopsis plants. a Comparison of stomatal apertures in response to different concentrations of exogenous ABA in WT and transgenic (L7, L20 and L21) plants. b Stomatal aperture width to length ratios following treatment with different concentrations of exogenous ABA. Datas represent mean ± SD of three replicates. Asterisks indicate statistical significance (*P < 0.05, **P < 0.01, Student’s t test) of differences between transgenic and WT plants
VvMBF1 expression alters the expression of several drought-responsive genes in transgenic Arabidopsis
To investigate the molecular mechanism underlying the enhanced response of VvMBF1-expressing transgenic Arabidopsis lines to drought stress, the expression of four genes that have been found previously to be drought-responsive (AtRD22, AtRD29B, AtRD29A and AtERD1 from Arabidopsis) were analyzed via quantitative real-time RT-PCR. Compared to WT lines, both AtRD22 and AtRD29B exhibited a significantly higher level of expression in transgenic lines following drought treatment, whereas the expression of AtRD29A and AtERD1 remained unchanged (Fig. 7).
Expression levels of drought-responsive genes in WT and VvMBF1-expressing transgenic plants. Relative gene expression levels were analyzed using quantitative real-time RT-PCR. Blocks represent the mean ± SD of three independent experiments. Asterisks indicate statistical significance (*P < 0.05, **P < 0.01, Student’s t test) of differences between transgenic (L7, L20 and L21) and WT plants
Discussion
Genetic engineering via the transfer of stress-responsive genes has proven to be a powerful tool for enhancing stress resistance in plants (Nakashima et al. 2009; Xu et al. 2013; Cui et al. 2013; Long et al. 2013; Wang et al. 2013). Since virtually all plant MBF1 genes identified to date have been found to be involved in biotic and abiotic stress response, we endeavored to isolate an MBF1 homolog from grape, assess its ability to confer enhanced drought resistance in a transgenic plant system, and characterize its mechanism of function.
Previously, it has been shown that plant MBF1 proteins can be classified into two groups: one comprising AtMBF1a- and AtMBF1b-like sequences (plant group I) and the other consisting of sequences similar to that of AtMBF1c (plant group II; Tsuda and Yamazaki 2004). Unfortunately, distinct functional differences between the two clades have yet to be deciphered and members of both have been implicated in stress response (Zegzouti et al. 1999; Godoy et al. 2001; Pnueli et al. 2002; Rizhsky et al. 2002; Suzuki et al. 2005, 2008, 2011; Arce et al. 2006; Kim et al. 2007; Zhang et al. 2009). Therefore, while our results suggest that VvMBF1 is a member of the group I MBF1 clade (Fig. 1), at present, the significance of this finding remains unclear.
Quantitative real-time PCR analysis of VvMBF1 expression in ‘Kyoho’ grape leaves demonstrated that it was significantly induced by both drought and ABA treatment (Fig. 2), suggesting a putative role in stress response. To further understand the function of this gene, we transformed Arabidopsis with a constitutively expressed VvMBF1 cassette and confirmed its responsiveness to drought treatment (Fig. 3b) prior to additional analyses. In terms of general drought-tolerance, it was found that transgenic seeds exhibited significantly increased levels of seed germination under osmotic stress compared to WT seeds (Fig. 4a, b). Furthermore, transgenic seedlings were much better able to tolerate dehydration than WT plants (Fig. 4c, d). These findings correlate well with results from previous studies of MBF1 genes from other plant species. For example, overexpression or mutation of Arabidopsis AtMBF1 genes resulted in an enhancement or decline, respectively, in tolerance to various forms of biotic and abiotic stress (Kim et al. 2007; Arce et al. 2010; Suzuki et al. 2005, 2008, 2011). Moreover, the expression of MBF1 genes from plants such as potato, tomato, tobacco and wheat, have all been demonstrated to be up-regulated in response to various types of stress (Zegzouti et al. 1999; Godoy et al. 2001; Rizhsky et al. 2002; Zanetti et al. 2003; Matsushita et al. 2002; Arce et al. 2006; Zhang et al. 2009). Therefore, it seems that plant MBF1 genes likely exhibit a broadly conserved function of mediating responses to stress, and thus can be deemed potential targets for the genetic enhancement of crop species.
In an attempt to provide insight into the mechanism driving VvMBF1-induced drought tolerance, we sought to monitor changes in a number of physiological responses that are known to be associated with dehydration stress (Seki et al. 2007). Increases in electrolyte leakage, MDA content, and the production of ROS can indicate drought-induced damage to cell membranes, which are one of the first targets of this type of stress (Bajji et al. 2001; Chen and Murata 2002; Carvalho 2008). Following dehydration treatment, we found that electrolyte leakage, MDA content and ROS production in VvMBF1-expressing transgenic plants were all decreased compared to WT plants (Fig. 5c, e, f), suggesting reduced damage to cell membranes in transgenic plants, and hence, increased drought resistance. In addition to its role in stress response, our results imply that VvMBF1 may also function to maintain redox homeostasis within cells by preventing the oxidative damage caused by increased production of ROS. This scenario would resemble that of a number of other previously characterized MBF1 genes, which have also been implicated in the response to oxidative stress (Jindra et al. 2004; Tsuda and Yamazaki 2004; Arce et al. 2010).
When exposed to drought treatment, both VvMBF1-expressing transgenic lines and WT control plants exhibited elevations in their endogenous ABA levels (Fig. 5b), which is consistent with the fact that drought is known to dramatically stimulate ABA biosynthesis, which in turn plays a significant role in drought stress responses (Basra 1994; Ingram and Bartels 1996; Rock 2000). However, we did not note any significant difference in endogenous ABA levels between VvMBF1-expressing transgenic lines and WT, which suggests that VvMBF1 is likely not involved in the ABA biosynthetic pathway itself.
The ability of a plant to reduce water loss is also an important determining factor of its tolerance to drought. In this study, we found that detached leaves from VvMBF1-expressing transgenic plants displayed less water loss over time than those from WT plants (Fig. 5a). Furthermore, transgenic lines displayed a significantly greater accumulation of proline, which functions as an osmotic regulator and prevents water loss from cells when plants are subjected to dehydration and salt stress (Bais and Ravishankar 2002; Gill and Tuteja 2010; Hussain et al. 2011), than WT plants when grown under drought conditions (Fig. 5d). This suggests that proline may be a contributing factor to the observed dehydration resistance of VvMBF1-expressing transgenic plants.
Water loss is controlled in large part by the regulation of stomatal apertures, which can occur within minutes of experiencing drought stress and involve changes in the activities of various signaling molecules, including ABA (Finkelstein et al. 2002). When a plant is exposed to dehydration, pairs of epidermal guard cells surrounding stomatal pores perceive increased ABA levels, which triggers a reduction in their turgor and causes stomatal closure (Kim et al. 2010). Therefore, we compared the response of stomata to various concentrations of exogenous ABA in VvMBF1-expressing transgenic lines and WT plants, and found that the stomata of transgenic plants exhibited lower width to length ratios than WT plants following ABA treatment (Fig. 6a, b). These findings imply that VvMBF1 plays a role in ABA-mediated stomatal closure, which almost certainly contributes to its function in drought tolerance.
Previously, several genes have been found to respond to drought at the transcriptional level (Ingram and Bartels 1996; Shinozaki and Yamaguchi-Shinozaki 1996; Bray 1997). Of these, AtRD22 and AtRD29B have been found to function in the ABA-dependent pathway, whereas AtRD29A and AtERD1 function in the ABA-independent pathway (Shinozaki and Yamaguchi-Shinozaki 2000). We found that the expression of AtRD22 and AtRD29B, but not AtRD29A and AtERD1, was significantly increased in VvMBF1-expressing transgenic lines compared to WT plants following drought treatment (Fig. 7). Taken together with the fact that VvMBF1 was found to be up-regulated by both exogenous ABA and drought treatments (Fig. 2), our findings suggest that this gene likely functions upstream of other drought-responsive genes in an ABA-dependent pathway. This also appears to be the case for MBF1 genes from Arabidopsis, whereby their functions in stress response have been attributed to roles in ethylene and ABA signal transduction pathways (Arce et al. 2010; Kim et al. 2007; Suzuki et al. 2005; Mauro et al. 2012).
In conclusion, we identified a grape MBF1 gene (VvMBF1), and demonstrated that like its counterparts in many other plant species, it is responsive to both drought and ABA. Analysis of VvMBF1-expressing transgenic plants indicated that VvMBF1 leads to improved drought tolerance through an integrated effect comprising changes in numerous physiological traits, the regulation of stomatal closure, and the up-regulation of other ABA-dependent stress-responsive genes. These results imply that VvMBF1 may confer drought avoidance in plants by an ABA-dependent signal transduction pathway. Since the greatest enhancements in stress tolerance in transgenic plants have been achieved via the transfer of stress-responsive genes that provide an upstream function in a signal transduction pathway (Yao et al. 2012; Liu et al. 2013; Luo et al. 2013), VvMBF1 has the potential to be an excellent target for future crop improvement.
Abbreviations
- ABA:
-
Abscisic acid
- MBF1:
-
Multiprotein bridging factor 1
- H2O2 :
-
Hydrogen peroxide
- RACE:
-
Rapid-amplification of cDNA ends
- GSP:
-
Gene-specific primer
- cDNA:
-
Complementary deoxyribonucleic acid
- RT-PCR:
-
Reverse transcriptase-polymerase chain reaction
- FW:
-
Fresh weights
- EL:
-
Electrolyte leakage
- MDA:
-
Malondialdehyde
- ROS:
-
Reactive oxygen specie
- O2 − :
-
Superoxide anion
- TCA:
-
Trichloroacetic acid
- TBA:
-
Thiobarbituric acid
- NBT:
-
Nitro blue tetrazolium
- DAB:
-
Diaminobenzidine
- HPLC:
-
High performance liquid chromatography
References
Arce DP, Tonón C, Zanetti ME, Godoy AV, Hirose S, Casalongué CA (2006) The potato transcriptional co-activator StMBF1 is up-regulated in response to oxidative stress and interacts with the TATA-box binding protein. J Biochem Mol Biol 39:355–360
Arce DP, Godoy AV, Tsuda K, Yamazaki K, Valle EM, Iglesias MJ, Mauro MFD, Casalongué CA (2010) The analysis of an Arabidopsis triple knock-down mutant reveals functions for MBF1 genes under oxidative stress conditions. J Plant Physiol 167:194–200
Bais HP, Ravishankar GA (2002) Role of polyamines in the ontogeny of plants and their biotechnological applications. Plant Cell, Tissue Organ Cult 69:1–34
Bajji M, Kinet JM, Lutts S (2001) The use of the electrolyte leakage method for assessing cell membrane stability as a water stress tolerance test in durum wheat. Plant Growth Regul 36:61–70
Basra AS (1994) Stress-induced gene expression in plants. In: Bray EA (ed) Alterations in gene expression in response to water deficit. Harwood, USA, pp 1–23
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Bray EA (1997) Plant responses to water deficit. Trends Plant Sci 2:48–54
Brendel C, Gelman L, Auwerx J (2002) Multiprotein bridging factor-1 (MBF-1) is a cofactor for nuclear receptors that regulate lipid metabolism. Mol Endocrinol 16:1367–1377
Busk PK, Wulf-Andersen L, Strøm CC, Enevoldsen M, Thirstrup K, Haunsø S, Sheikh SP (2003) Multiprotein bridging factor 1 cooperates with c-Jun and is necessary for cardiac hypertrophy in vitro. Exp Cell Res 286:102–114
Carvalho MH (2008) Drought stress and reactive oxygen species: production, scavenging and signaling. Plant Signal Behav 3:156–165
Chen TH, Murata N (2002) Enhancement of tolerance of abiotic stress by metabolic engineering of betaines and other compatible solutes. Curr Opin Plant Biol 5:250–257
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cui Y, Xu G, Wang M, Yu Y, Li M, Rocha PSCF, Xia X (2013) Expression of OsMSR3 in Arabidopsis enhances tolerance to cadmium stress. Plant Cell Tiss Organ Cult 113:331–340
Finkelstein RR, Gampala SSL, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14:15–45
Gill SS, Tuteja N (2010) Polyamines and abiotic stress tolerance in plants. Plant Signal Behav 5:26–33
Godoy AV, Zanetti ME, Segundo BS, Casalongué CA (2001) Identification of a putative Solanum tuberosum transcriptional coactivator up-regulated in potato tubers by Fusarium solani f. sp. eumartii infection and wounding. Physiol Plant 112:217–222
Heath RL, Packer L (1968) Photoperoxidation in isolated chloroplasts: I. Kinetics and stoichiometry of fatty acid peroxidation. Arch Biochem Biophys 125:189–198
Hussain SS, Ali M, Ahmad M, Siddique KHM (2011) Polyamines: natural and engineered abiotic and biotic stress tolerance in plants. Biotechnol Adv 29:300–311
Ingram J, Bartels D (1996) The molecular basis of dehydration tolerance in plants. Annu Rev Plant Biol 47:377–403
Jindra M, Gaziova I, Uhlirova M, Okabe M, Hiromi Y, Hirose S (2004) Coactivator MBF1 preserves the redox-dependent AP-1 activity during oxidative stress in Drosophila. EMBO 23:3538–3547
Kim MJ, Lim GH, Kim ES, Ko CB, Yang KY, Jeong JA, Lee MC, Kim CS (2007) Abiotic and biotic stress tolerance in Arabidopsis overexpressing the multiprotein bridging factor 1a (MBF1a) transcriptional coactivator gene. Biochem Biophys Res Commun 354:440–446
Kim TH, Böhmer M, Hu H, Nishimura N, Schroeder JI (2010) Guard cell signal transduction network: advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu Rev Plant Biol 61:561–591
Kim SH, Woo DH, Kim JM, Lee SY, Chung WS, Moon YH (2011) Arabidopsis MKK4 mediates osmotic-stress response via its regulation of MPK3 activity. Biochem Biophys Res Commun 412:150–154
Kotchoni SO, Kuhns C, Ditzer A, Kirch HH, Bartels D (2006) Over-expression of different aldehyde dehydrogenase genes in Arabidopsis thaliana confers tolerance to abiotic stress and protects plants against lipid peroxidation and oxidative stress. Plant, Cell Environ 29:1033–1048
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Annu Rev Plant Biol 49:199–222
Li FQ, Ueda H, Hirose S (1994) Mediators of activation of fushi tarazu gene transcription by BmFTZ-F1. Mol Cell Biol 14:3013–3021
Liu TX, Zhang YP (2010) Determination of ABA content in the seedling of capsicum by HPLC. Guangdong Agric Sci 8:249–250 (In Chinese)
Liu QX, Jindra M, Ueda H, Hiromi Y, Hirose S (2003) Drosophila MBF1 is a co-activator for tracheae defective and contributes to the formation of tracheal and nervous systems. Development 130:719–728
Liu JH, Nada K, Honda C, Kitashiba H, Wen XP, Pang XM, Moriguchi T (2006) Polyamine biosynthesis of apple callus under salt stress: importance of the arginine decarboxylase pathway in stress response. J Exp Bot 57:2589–2599
Liu P, Xu ZS, Lu PP, Hu D, Chen M, Li LC, Ma YZ (2013) A wheat PI4 K gene whose product possesses threonine autophophorylation activity confers tolerance to drought and salt in Arabidopsis. J Exp Bot 64:2915–2927
Long L, Gao W, Xu L, Liu M, Luo X, He X, Yang X, Zhang X, Zhu L (2013) GbMPK3, a mitogen-activated protein kinase from cotton, enhances drought and oxidative stress tolerance in tobacco. Plant Cell Tiss Organ Cult. doi:10.1007/s11240-013-0392-1
Luo X, Bai X, Sun X, Zhu D, Liu B, Ji W, Cai H, Cao L, Wu J, Hu M, Liu X, Tang L, Zhu Y (2013) Expression of wild soybean WRKY20 in Arabidopsis enhances drought tolerance and regulates ABA signalling. J Exp Bot 64:2155–2169
Matsushita Y, Miyakawa O, Deguchi M, Nishiguchi M, Nyunoya H (2002) Cloning of a tobacco cDNA coding for a putative transcriptional coactivator MBF1 that interacts with the tomato mosaic virus movement protein. J Exp Bot 53:1531–1532
Mauro MF, Iglesias MJ, Arce DP, Valle EM, Arnold RB, Tsuda K, Yamazaki K, Casalongué CA, Godoy AV (2012) MBF1 s regulate ABA-dependent germination of Arabidopsis seeds. Plant Signal Behav 7:188–192
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149:88–95
Pei ZM, Kuchitsu K, Ward JM, Schwarz M, Schroeder JI (1997) Differential abscisic acid regulation of guard cell slow anion channels in Arabidopsis wild-type and abi1 and abi2 mutants. Plant Cell 9:409–423
Pnueli L, Hallak-Herr E, Rozenberg M, Cohen M, Goloubinoff P, Kaplan A, Mittler R (2002) Molecular and biochemical mechanisms associated with dormancy and drought tolerance in the desert legume Retama raetam. Plant J 31:319–330
Rizhsky L, Liang H, Mittler R (2002) The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiol 130:1143–1151
Rock CD (2000) Pathways to abscisic acid-regulated gene expression. New Phytol 148:357–396
Saleki R, Young PG, Lefebvre DD (1993) Mutants of Arabidopsis thaliana capable of germination under saline conditions. Plant Physiol 101:839–845
Seki M, Umezawa T, Urano K, Shinozaki K (2007) Regulatory metabolic networks in drought stress responses. Curr Opin Plant Biol 10:296–302
Shinozaki K, Yamaguchi-Shinozaki K (1996) Molecular responses to drought and cold stress. Curr Opin Biotechnol 7:161–167
Shinozaki K, Yamaguchi-Shinozaki K (1997) Gene expression and signal transduction in water-stress response. Plant Physiol 115:327–334
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Suzuki N, Rizhsky L, Liang H, Shuman J, Mittler R (2005) Enhanced tolerance to environmental stress in transgenic plants expressing the transcriptional coactivator multiprotein bridging factor 1c. Plant Physiol 139:1313–1322
Suzuki N, Bajad S, Shuman J, Shulaev V, Mittler R (2008) The transcriptional co-activator MBF1c is a key regulator of the motolerance in Arabidopsis thaliana. J Biol Chem 283:9269–9275
Suzuki N, Sejima H, Tam R, Schlauch K, Mittler R (2011) Identification of the MBF1 heat-response regulon of Arabidopsis thaliana. Plant J 66:844–851
Takemaru K, Li FQ, Ueda H, Hirose S (1997) Multiprotein bridging factor 1 (MBF1) is an evolutionarily conserved transcriptional coactivator that connects a regulatory factor and TATA element-binding protein. Proc Natl Acad Sci USA 94:7251–7256
Takemaru K, Harshima S, Ueda H, Hirose S (1998) Yeast coactivator MBF1 mediates GCN4-dependent transcriptional activation. Mol Cell Biol 18:4971–4976
Toji T, Tsuda K, Yoshizumi T, Ikeda A, Yamaguchi J, Matsui M, Yamazaki K (2009) Arabidopsis MBF1 s control leaf cell cycle and its expansion. Plant Cell Physiol 50:254–264
Tsuda K, Yamazaki K (2004) Structure and expression analysis of three subtypes of Arabidopsis MBF1 genes. Biochim Biophys Acta 1680:1–10
Tsuda K, Tsuji T, Hirose S, Yamazaki K (2004) Three Arabidopsis MBF1 homologs with distinct expression profiles play roles as transcriptional co-activator. Plant Cell Physiol 45:225–231
Wang X, Li Z, Yan F, Khalil R, Ren Z, Yang C, Yang Y, Deng W (2013) ZmSKIP, a homologue of SKIP in maize, is involved in response to abiotic stress in tobacco. Plant Cell Tiss Organ Cult 112:203–216
Weigel D, Glazebrook J (2002) Arabidopsis. Cold Spring Harbor Laboratory, New York, pp 12–13
Xu X, Guo R, Cheng C, Zhang H, Zhang Y, Wang X (2013) Overexpression of ALDH2B8, an aldehyde dehydrogenase gene from grapevine, sustains Arabidopsis growth upon salt stress and protects plants against oxidative stress. Plant Cell Tiss Organ Cult 114:187–196
Yao X, Xiong W, Ye T, Wu Y (2012) Overexpression of the aspartic protease ASPG1 gene confers drought avoidance in Arabidopsis. J Exp Bot 63:2579–2593
Zanetti ME, Blanco FA, Daleo GR, Casalongué CA (2003) Phosphorylation of a member of the MBF1 ranscriptional co-activator family, StMBF1, is stmiulated in potato cell suspensions upon fungal elicitor challenge. J Exp Bot 54:623–632
Zegzouti H, Jones B, Frasse P, Marty C, Maitre B, Latché A, Pech JC, Bouzayen M (1999) Ethylene-regulated gene expression in tomato fruit: characterization of novel ethylene-responsive and ripening-related genes isolated by differential display. Plant J 18:589–600
Zhang Y, Zhang G, Dong YL, Guo J, Huang LL, Kang ZS (2009) Cloning and characterization of a MBF1 transcriptional coactivator factor in wheat induced by stripe rust pathogen. Acta Agronomica Sinica 35:11–17
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31272136), 948 Project from the Ministry of Agriculture of China (2012-S12), as well as the Program for Innovative Research Team of Grape Germplasm Resources and Breeding (2013KCT-25).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yan, Q., Hou, H., Singer, S.D. et al. The grape VvMBF1 gene improves drought stress tolerance in transgenic Arabidopsis thaliana . Plant Cell Tiss Organ Cult 118, 571–582 (2014). https://doi.org/10.1007/s11240-014-0508-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-014-0508-2