Abstract
A reliable Agrobacterium-mediated transformation and shoot regeneration protocol was developed for breeding lines of commercially important western-shipper cantaloupe and honeydew melons, ‘F39’ and ‘150’, respectively. Different media were tested to select a shoot regeneration system for each of three elite breeding lines ‘F39’, ‘141’ and ‘TMS’. Murashige & Skoog (MS) basal medium supplemented with 1 mg l−1 benzyladenine (BA), 0.26 mg l−1 abscisic acid (ABA) and 0.8 mg l−1 indole-3-acetic acid (IAA) was used for shoot regeneration from cotyledonary explants in ‘F39’ and ‘150’. Kanamycin sensitivity as well as Timentin™ and Clavamox® were evaluated using wild-type ‘F39’ and ‘150’ cotyledons. Kanamycin concentrations of 200 and 150 mg l−1 were chosen as the threshold levels for ‘F39’ and ‘150’, respectively. No significant differences were found between Timentin™ and Clavamox® in ‘F39’; however, Clavamox® reduced the incidence of vitrification and increased the frequency of shoot elongation in ‘150’. A. tumefaciens strain EHA105, harboring pCNL56 carrying neomycin phosphotransferase II (nptII) and gusA reporter genes, was selected to establish a transformation protocol for ‘F39’ and ‘150’. Putative transformants were evaluated using β-glucuronidase (GUS) histochemical assay, polymerase chain reaction (PCR) and Southern blot analyses. Based on these parameters, the transformation efficiency for cantaloupe ‘F39’ was 0.3% and that for honeydew ‘150’ was 0.5%.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Melon (Cucumis melo L.) is a group of high-value crops that include cantaloupe (muskmelon), honeydew and casaba melon. Despite the development of genetic transformation protocols for melon reported in the last 20 years, transformation remains genotype-dependent and efficiencies are relatively low (0–12.5%) (Nuňez-Palenius et al. 2006, 2008). To date, the most successful methods for producing transgenic melon have been achieved in French (Fang and Grumet 1990), Israeli (Nuňez-Palenius et al. 2006) and Asian germplasm (Dong et al. 1991; Wu et al. 2009), which constitute a low percentage of the US consumer market. Successful transformation protocols are needed for improvement of commercial genotypes grown in the US, especially the western shipper cantaloupe (C. melo var. reticulatus) and honeydew (C. melo var. inodorus).
A review of pertinent literature reveals only a few studies on the tissue culture or transformation of western shipper and honeydew melons. At this writing, there were no transformation studies available for honeydew melon; however, a number of studies on shoot regeneration in culture have been reported (Ficcadenti and Rotino 1995; Keng and Hoong 2005; Kintzios and Taravira 1997; Oridate et al. 1992; Orts et al. 1987). In western shipper cantaloupe, one microprojectile mediated transformation (Gonsalves et al. 1994) and two Agrobacterium-mediated transformations (Clough and Hamm 1995; Fuchs et al. 1997) were reported to confer resistance to cucumber mosaic virus (CMV), zucchini yellow mosaic virus (ZYMV), and watermelon mosaic virus (WMV). A report on muskmelon transformation via ovary injection showed changes in the soluble sugar composition in fruits (Fan et al. 2007). Also, C. melo L. cv. Hetao was transformed using the pollen-tube pathway approach, which produced melons having improved firmness and shelf-life (Hao et al. 2011).
A. tumefaciens-mediated transformation has become the method of choice for melon transformation (Nuňez-Palenius et al. 2008). We opted to use Agrobacterium due to reports of low copy gene transfers, which help to reduce the chances for multi-gene triggered silencing in the transgenic plants produced. Unfortunately, previously reported procedures provided insufficient detail for us to replicate.
Plant growth regulators have been used in melon tissue culture to optimize shoot regeneration. Both auxins and cytokinins are known to be essential for bud/shoot induction and the optimal auxin/cytokinin levels, critical for recovery of plants, are often genotype specific. The auxin indole-3-acetic acid (IAA) and cytokinin 6-benzyladenine (BA) are commonly used in organogenesis studies. The synthetic auxin 2,4-dichlorophenoxyacetic acid has been widely used to induce somatic embryogenesis (Akasaka-Kennedy et al. 2004; Gray et al. 1993; Guis et al. 1997; Kintzios et al. 2004; Kintzios and Taravira 1997; Oridate and Oosawa 1986). In addition, 6-(γ,γ-dimethylallylamino)-purine (2iP), gibberellic acid (GA3), kinetin, thidiazuron and α-naphthaleneacetic acid have been used in melon culture (Fang and Grumet 1990; Ficcadenti and Rotino 1995; Guis et al. 2000; Souza et al. 2006; Yadav et al. 1996). The effects of these growth regulators on melon regeneration differed depending on plant genotype, explant type and culture conditions. Other media components also affect melon regeneration efficiency. For example, abscisic acid (ABA) and sucrose have been shown to enhance somatic embryogenesis in melon (Nakagawa et al. 2001). The anti-gibberellin analogue, ancymidol, was reported to promote shoot regeneration from cotyledonary explants of ‘Galia’ melon in combination with a low concentration of BA (0.44 μM), while the addition of GA3 to this medium reduced regeneration frequency 12-fold after 13 d of treatment (Gaba et al. 1996).
We evaluated cotyledonary explant responsiveness for shoot initiation and plant regeneration in elite western shipper cantaloupe and casaba lines, using the six different regeneration media reported to be optimal for other melon genotypes. The best medium for regeneration of western shipper cantaloupe line ‘F39’ was then used to determine the sensitivity of kanamycin selection. Two beta-lactam antibiotics frequently used to remove Agrobacterium from inoculated tissues, Clavamox® and Timentin™ (Gould and Magallens-Cedeno 1998; Cheng et al. 1998), were compared with respect to their influences on shoot regeneration capacity. In optimization tests for transformed shoot production, we also evaluated the effects of light versus dark during co-cultivation. This factor has been neglected in melon transformations but has shown the importance in other plant species, such as Phaseolus acutifolius and Arabidopsis thaliana (Zambre et al. 2003). In addition, two A. tumefaciens strains, EHA105 (Hood et al. 1993) and LBA4404 (Hoekema et al. 1983), were evaluated using β-glucuronidase (GUS) transient expression in our genotypes. Melon regeneration and transformation are still considered to be difficult due to several factors such as strong genotype dependence, a high percentage of polyploids, high rates of ‘escapes’ and aberrant shoot development (Castelblanque et al. 2008; Chovelon et al. 2008; Dong et al. 1991; Akasaka-Kennedy et al. 2004; Wu et al. 2009). The goal of this research was to establish a reliable genetic transformation system in these two elite breeding lines with commercial qualities.
Materials and methods
Plant material
Three melon breeding lines, ‘F39’, ‘141’ and ‘TMS’ were initially selected to test published melon regeneration protocols. ‘F39’ and ‘141’ are inbred lines of western shipper cantaloupe (C. melo var. reticulatus), which produce high quality orange-fleshed fruits with netted surface and have been inbred for over 10 generations at Texas AgriLife Research Center, Weslaco, TX. In addition, ‘F39’ has medium to high resistance to multiple diseases. ‘TMS’ is an elite white-fleshed casaba melon (C. melo var. inodorus) with smooth surface. However, it was replaced with honeydew ‘150’ in later studies due to limited seed availability. ‘150’ is an inbred line of pale green-fleshed honeydew (C. melo var. inodorus) with a smooth rind surface. Preliminary tests showed that the frequency of ‘150’ regeneration was very high on the medium that had been chosen for ‘F39’. Therefore, ‘F39’ and ‘150’ were used to establish the transformation protocol described here.
Explant preparation
Seeds were surface sterilized in a 50% commercial bleach solution (3% sodium hypochlorite) containing a drop of Tween-20 for 30 min and rinsed three times with sterile water. After soaking seeds in sterile water for 4–8 h, the seed coats were removed to expose each embryo. De-coated embryos were sterilized in 70% ethanol for 30 s followed by a 5% commercial bleach solution (0.3% sodium hypochlorite) containing a drop of Tween-20 for 10 min. Embryos were rinsed three times with sterile water and cultured in the dark on a germination medium, consisting of Murashige and Skoog (MS) (1962) salts medium supplemented with 30 g l−1 sucrose and solidified by 5 g l−1 agar, pH 5.7–5.8. Seven days later, cotyledons were excised 2 mm from the cotyledonary nodes (conjunction sites between cotyledons and hypocotyls). Each cotyledon was then cut into 6 equally-sized explants of approximately 3 mm × 2 mm.
Shoot and root regeneration
Six regeneration media, RM1 (Ficcadenti and Rotino 1995), RM2 (Guis et al. 2000), RM3 (Yadav et al. 1996), RM4 (Fang and Grumet 1990), RM5 (Souza et al. 2006), and RM6 (Bordas et al. 1997), were evaluated with our melon genotypes (Table 1). Each medium represented the optimal composition to give the highest regeneration frequency in each study reported. All media were based on MS salts supplemented with 30 g l−1 sucrose and various combinations of plant growth regulators (BA, ABA, IAA, 2iP and kinetin). Media were solidified using agar 7 g l−1 except in RM3 where agar was replaced by 2.6 g l−1 Phytagel™.
Cotyledonary explants were placed adaxially on a regeneration medium. Tissue cultures were placed in a room maintained at 25 ± 2°C under cool white fluorescent lights with 16 h light/8 h dark photoperiod and 60–80 μmol m−2 s−1 light intensity. After 4 weeks, the induction of calli and shoot primordia was recorded and scored from 0 to 100% (Table 2). Shoots or shoot primordia were transferred to shoot elongation medium, which consisted of MS basal medium supplemented with 30 g l−1 sucrose, 8 g l−1 agar and 0.01 mg l−1 BA. Shoots that failed to root on elongation medium were transferred to a rooting medium (MS basal medium supplemented with 30 g l−1 sucrose and 8 g l−1 agar).
Preparation of Agrobacterium tumefaciens
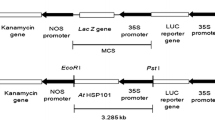
A.tumefaciens strains EHA105 and LBA4404 carrying pCNL56, harboring a CaMV 35S promoter for constitutive gene expression of neomycin phosphotransferase II (nptII) and gusA/intron genes (Li et al. 1992), were used in transformation studies. A single colony of A. tumefaciens was inoculated in 3 ml of YEP liquid medium supplemented with 50 mg l−1 kanamycin and 20 mg l−1 rifampicin and grown at 200 rpm at 28°C for 24 h. The 3 ml culture was added to 20 ml of YEP liquid medium supplemented with the same concentrations of antibiotics and cultured for 3–5 h until an OD600 reached between 0.7 and 1.0. Bacterial cells were collected using centrifugation at 4,000 rpm for 10 min at room temperature and then re-suspended in liquid RM4 medium supplemented with 100 μM acetosyringone.
Inoculation, co-cultivation and light test
Cotyledonary explants were pre-cultured on RM4 medium for 2 d and then inoculated in the A. tumefaciens suspension for 10 min. They were removed from the suspension, dried on sterile filter paper and then transferred to RM4 medium supplemented with 100 μM acetosyringone (pH 5.5). The effect of light during co-cultivation was compared by maintaining inoculated explants on co-cultivation medium for 3 d in dark (22°C) or for 3 d in light (24°C, fluorescent desk lamp). Seven days after inoculation, GUS transient expression in cotyledonary explants was assessed.
Antibiotic testing and plant regeneration
To determine sensitivity of cotyledonary explants to kanamycin, both cantaloupe ‘F39’ and honeydew ‘150’ genotypes were tested in RM4 medium with various kanamycin concentrations: 0, 100, 125, 150, 175 and 200 mg l−1. Appropriate concentration was determined for each genotype. To eliminate A. tumefaciens overgrowth during selection, the effects of Clavamox® (amoxicillin trihydrate/clavulanate potassium tablets, Pfizer Animal Health) and Timentin™ (ticarcillin disodium/potassium clavulanate powders, Duchefa Direct, St. Louis) on the regeneration of both genotypes were compared by supplementing media with 250 mg l−1 Clavamox® or 300 mg l−1 Timentin™ throughout the regeneration process (shoot regeneration, shoot elongation and rooting).
After co-cultivation at 22°C in the dark for 3 d, explants were transferred to medium RM4 containing kanamycin and 250 mg l−1 Clavamox®, and subcultured every 14 d. Three to four weeks later, regenerated shoot primordium aggregates were excised into small pieces (3 mm × 3 mm) and transferred onto shoot elongation medium supplemented with 50 mg l−1 kanamycin and 250 mg l−1 Clavamox®. Some elongated shoots produced roots in 14–28 d on this medium. Large shoots that failed to produce roots were transferred to rooting medium supplemented with 50 mg l−1 kanamycin as described by Compton et al. (2004). When root systems were well developed, plants were transferred to soil and acclimatized in the tissue culture room (24 ± 2°C, 16 h light/8 h dark photoperiod). Plants were then transferred to a greenhouse where they were allowed to grow to maturity.
Histochemical GUS assay, polymerase chain reaction (PCR) and Southern blot analyses
To examine the expression of the gusA gene in putative transformed regenerants, tissues were incubated in X-glucuronide staining solution at 37°C overnight followed by washing with 95% ethanol to remove chlorophyll (Jefferson et al. 1987).
Genomic DNA was isolated from leaf tissues of wild type and putative transformants by the method of Skroch and Nienhuis (1995). As an initial screen for presence of the transferred gene, PCR primers were designed to amplify fragments from nptII under the following conditions: forward and reverse primers were 5′-CCC GGT TCT TTT TGT CAA GAC CGA CCT-3′ and 5′-GTT TGC GCG CTA TAT TTT GTT TTC TAT CGC-3′, respectively. The PCR reaction mixture contained 50 ng of genomic DNA, 1 × PCR buffer, 200 μM dNTP, 0.2 μM forward and reverse primers and 1 μl of polymerase mix (Clontech Laboratories, Inc. Mountain View, CA, USA) in a total volume of 50 μl. The reaction consisted of an initial denaturation at 94°C for 3 min, followed by 30 cycles of denaturation at 94°C for 30 s, annealing at 68°C for 30 s, and elongation at 72°C for 2 min, and a final elongation at 72°C for 10 min. Amplified products were visualized on 1% (w/v) agarose gels.
Twenty micrograms of genomic DNA was digested with EcoRI at 37°C overnight and separated by electrophoresis on a 1% (w/v) agarose gel at 24 V overnight. DNA fragments were then denatured and transferred onto an N+ Hybond nylon membrane (Amersham Hybond™ -N, GE Healthcare Life Sciences). A gusA probe was labeled using the PCR DIG Probe Synthesis Kit (Roche Applied Science, Mannheim, Germany). The membrane was then hybridized with the DIG-labeled probe in DIG Easy Hyb solution (Roche Applied Science, Mannheim, Germany) at 45°C overnight. DIG-labeled nucleic acids were detected with CDP-Star (Roche Applied Science, Mannheim, Germany) by exposing the membrane under a LAS-4,000 Chemiluminescent Image System (Fuji Film Life Science, USA). Probe labeling, hybridization, washing and detection were conducted according to the manufacturer’s instruction.
Statistical analysis
Regeneration experiments were conducted with two factors (genotype × medium composition) in a randomized complete block design (12 explants/dish; 5 replicates/treatment) and analyzed by two-way analysis of variance. Differences between the means were performed using Duncan’s Multiple Range Test where the 5% probability level was considered significant. Kanamycin sensitivity was tested in a randomized complete block design (4 explants/dish; 8 replicates/treatment) and analyzed by one-way analysis of variance. Each dish was considered as a replicate in all the experimental designs.
Results and discussion
Plant regeneration
Six media, previously reported to be effective from other melon regeneration and/or transformation protocols, were screened for our three different genotypes (Fig. 1). Calli and shoot primordia were usually formed within 3–4 weeks on the initial regeneration media. Shoots developed from shoot primordia approximately 2–3 weeks later. Significant differences appeared in the frequency of explants producing calli and shoot primordia on the various media, and an interaction between genotype and medium composition was also observed (Table 3). Overall, media RM3, RM4 and RM1 were best for shoot regeneration in ‘141’, ‘F39’ and ‘TMS’, respectively (Fig. 2a–c). RM3 and RM4 induced the highest frequency of both callus and shoot primordium regeneration in ‘F39’; however, RM4 performed better in terms of producing fewer vitrified shoots (Fig. 2d, e).
Effect of six different media on the production of calli and shoot primordia from cotyledonary explants of three melon genotypes ‘141, ‘F39’ and ‘TMS’. a The frequency of explants producing calli, b the frequency of explants producing shoot primordia. Bars with same letters are not significantly different by Duncan’s Multiple Range Test at 5% probability level. Vertical bars show standard errors
Callus production and shoot regeneration on the optimal medium from cotyledonary explants of three different melon genotypes after 4 weeks of culture. a ‘141’ on RM3 [MS basal supplemented with 1 mg l−1 BA, 0.26 mg l−1 ABA, 0.8 mg l−1 IAA and 5.4 mg l−1 AgNO3], b ‘F39’ on RM4 [MS basal supplemented with 1.13 mg l−1 BA, 0.26 mg l−1 ABA and 0.88 mg l−1 IAA], c ‘TMS’ on RM1 [MS salt supplemented with 0.63 mg l−1 BA and 0.26 mg l−1 ABA], d ‘F39’ on RM4 (less vitrification), e ‘F39’ on RM3 (more vitrification) (bars = 1 cm)
Subsequently, we compared shoot primordium regeneration efficiency from each region of the cotyledonary explants: proximal, middle and distal. No significant differences were detected (data not shown), which was not in accord with the results reported by Gonsalves et al. (1994). Their research showed that regeneration frequency of the proximal side of the melon explant was significantly higher than that of the distal region.
Regenerated shoots proliferated but did not elongate on MS basal medium supplemented with BA levels higher than 0.1 mg l−1. A series of low BA concentrations, 0, 0.01, 0.025, 0.05 and 0.1 mg l−1, was examined on both ‘F39’ and ‘150’ for shoot elongation. Low levels of BA (0 and 0.01 mg l−1) treatments reduced the prevalence of multiple shoots and allowed shoots to elongate and produce roots (data not shown).
Selection for transformation
To determine the concentration of kanamycin for the selection of transformed shoots, explants were cultured on regeneration media containing a range of antibiotic concentrations (Fig. 3). In vitro shoot regeneration frequencies of both genotypes were approximately 90% in medium RM4 without kanamycin. Some tissues produced calli and shoot primordia on 100–150 mg l−1 kanamycin, but failed to develop normal shoots. In ‘F39’ and ‘150’, shoot regeneration was completely inhibited on 200 and 150 mg l−1 kanamycin, respectively. These two concentrations were chosen as the selection thresholds for those genotypes.
The beta-lactam antibiotics are used to eliminate A. tumefaciens from inoculated tissues. Timentin™ is one of the most expensive antibiotics used widely in transformation studies (Ieamkhang et al. 2005; Slater et al. 2011; Thiruvengadam et al. 2011). Clavamox® is relatively inexpensive and appears to work effectively at a low dose (250 mg l−1). Even a high one time dose of 10,000 mg l−1 was reported to be non-toxic to plant tissues (Gould and Magallens-Cedeno 1998). Clavamox® tablets are sterile and individually packaged. The antibiotic is stable in this form and convenient to use since there is no need to prepare a stock solution. Cheng et al. (1998) reported that Timentin™ stock solution was stable only for 4 weeks at −20 or −80°C, which is another disadvantage compared to Clavamox®. Our results indicated that Clavamox® was superior to Timentin™ with regard to the impact on tissue regeneration efficiency. Clavamox® has been compared with two other commonly used beta-lactam antibiotics, carbenicillin and cefotaxime, on the growth of A. tumefaciens strain LBA4404 and the shoot regeneration of tomato. No significant differences were found (Hussain et al. 2008).
Clavamox® and Timentin™ were tested on non-inoculated explants of ‘F39’ and ‘150’ to examine possible adverse effects on plant regeneration. In ‘150’, no significant differences were detected during the early stages of shoot regeneration (first 4 weeks); however, inclusion of Timentin™ resulted in vitrification and/or aberrant morphology in 70% of the shoots over time (Fig. 4 and Table 4). This vitrification severely reduced shoot elongation frequency and ability to produce roots. In contrast, only 5% of the shoots cultured on the medium containing Clavamox® became vitrified and/or abnormal. This result was similar to results seen in the non-antibiotic treatments. For ‘F39’, Clavamox® and Timentin™ treatments failed to show any significant differences in shoot regeneration and production of abnormal types, suggesting the effects of these antibiotics could be genotype-dependent.
Like many other melon transformation reports (Dong et al. 1991; Akasaka-Kennedy et al. 2004; Wu et al. 2009), the common problem of ‘escapes’ was also identified in our study. By reducing explant size (6 explants per cotyledon, 3 mm × 2 mm) and increasing the volume of selection medium per petri dish (45 ml), we were able to reduce the curling of cotyledonary explants and the incidence of ‘escapes’. A similar result was obtained by Wu et al. (2009) in oriental melon transformation.
Inoculation and co-cultivation
Initially, A. tumefaciens strains LBA4404 and EHA105 were tested for their ability to transform explants based on GUS histochemical staining. Both strains showed fairly good inoculation efficiencies demonstrated by GUS assays (Fig. 5). EHA105 was used in subsequent studies as it was reported to be more efficient in producing stable transformants compared to LBA4404 (Galperin et al. 2003).
Previous reports of melon transformation indicated optimal inoculation times of 20 s to 30 min (Ayub et al. 1996; Guis et al. 2000; Vallés and Lasa 1994) depending on the Agrobacterium strain used, concentration of the bacterial suspension, plant species, explant size and thickness. We evaluated 10, 15 and 20 min inoculation times. Three days after inoculation, the explants inoculated for 10 min remained healthy, but the longer inoculation treatments resulted in the explants becoming brown or necrotic in some cases (data not shown). Therefore, an inoculation period of 10 min was adopted.
Co-cultivation is one of the most important steps in a transformation procedure. Factors during co-cultivation can enhance transformation efficiency, i.e., temperature (Fullner and Nester 1996; Yasmin and Debener 2010; Sharma et al. 2011; Seo et al. 2011), lighting conditions (Zambre et al. 2003), co-cultivation period (Fang and Grumet 1990; Shilpa et al. 2010; Seo et al. 2011), addition of acetosyringone (Costa et al. 2006; Afroz et al. 2010; Sharma et al. 2011) and antioxidants (Dan et al. 2009; Olhoft and Somers 2001; Ostergaard and Yanofsky 2004; Toldi et al. 2002; Zheng et al. 2005; Kumar et al. 2011; Dutt et al. 2011). Co-cultivation periods of 2–6 d were reported to be optimal for melons (Galperin et al. 2003; Vallés and Lasa 1994; Akasaka-Kennedy et al. 2004; Dong et al. 1991). Under our co-cultivation conditions at 22°C, A. tumefaciens strain EHA105 began to overgrow the explant 3 d following the inoculation (data not shown).
Zambre et al. (2003) reported a positive effect of light on gene transfer from A. tumefaciens to callus explants of Phaseolus acutifolius (tepary bean) and root explants of Arabidopsis thaliana. Our results showed there were no significant differences in transformation rates between light and dark treatments (P ≤ 0.05) during co-cultivation (Table 5). Plausible reasons for this result may be: (1) the light/dark influence on Agrobacterium-mediated transformation may be genotype dependent; (2) the light/dark influence may depend on explant types (callus and/or root); (3) the co-cultivation temperatures of our light and dark treatments of 24 and 22°C, respectively, may have influenced our results. Temperatures of 19–22°C have been reported to be critical for high efficiency of Agrobacterium-mediated transformation while temperatures higher than 22°C dramatically decreased this efficiency (Fullner and Nester 1996).
GUS staining, PCR and Southern blot analysis
Cotyledonary explants were inoculated with A. tumefaciens strain EHA105. A total of 1,075 explants of ‘F39’ (from two experiments) and 1,205 explants of ‘150’ (from three experiments) were used. GUS histochemical analysis was conducted throughout the selection procedure to identify putative transformants, as well as non-transformed or chimeric tissues. Isolated pieces of leaf tissue from regenerating shoots were examined for GUS activity (Fig. 6a–d). All GUS positive tissues were then analyzed further for expression of the GUS gene in apical shoots (Fig. 6e) and roots (Fig. 6f). During large scale GUS assays, we found a large proportion of chimeras in both genotypes (9 out of 12 samples of ‘F39’ and 9 out of 13 samples of ‘150’). Chimeric shoots exhibited blue sectors of various sizes following GUS histochemical staining as reported in other crop species (Moore 1995; Mollel et al. 2004; Kathiravan et al. 2006). Most chimeric shoots did not survive kanamycin selection and died before or during rooting.
Callus and shoot development from explants and GUS histochemical assay. a and c A regenerated shoot of cantaloupe ‘F39’ before (a) and after (c) GUS staining, b and d a cotyledonary explant with callus producing shoot primordia of honeydew ‘150’ before (b) and after (d) GUS staining, e an apical shoot of a GUS-positive plant of ‘150’, f GUS expression in roots of a kanamycin-resistant plant of ‘150’
The binary plasmid pCNL56 harbors both nptII and gusA genes. All GUS-positive plants that rooted on media containing kanamycin were selected for PCR analysis to screen for the presence of nptII. Genomic DNA from all rooted GUS-positive plants yielded a 1.2 kb fragment identical to the expected fragment amplified from the nptII gene (Fig. 7a). Plants regenerated from non-inoculated explants failed to produce a fragment. Most non-transformed shoots were killed by kanamycin selection during regeneration, elongation and rooting. Although some shoots survived kanamycin (150 and 200 mg l−1), many failed to root in the rooting medium containing 50 mg l−1 kanamycin. Based on PCR analysis, we found only one ‘F39’ and two ‘150’ regenerated plants to be non-transgenic escapes.
PCR and southern blot analyses of putative transformants. a Preliminary screen for detecting the presence of the nptII gene in putative transformants using PCR. Lane 1 100 bp ladder, lane 2 pCNL56 (positive control), lane 3 wild type honeydew ‘150’ (negative control), lanes 4–8 GUS-positive plants of honeydew ‘150’, lane 9 wild type cantaloupe ‘F39’ (negative control), lanes 10–12 GUS-positive plants of cantaloupe ‘F39’, b integration of the gusA gene as detected by Southern blotting. Lane 1 pCNL56 (positive control), lane 2 wild type honeydew ‘150’ (negative control), lanes 3–7 PCR-positive plants of honeydew ‘150’, lane 8 wild type cantaloupe ‘F39’ (negative control), lanes 9–11 PCR-positive plants of cantaloupe ‘F39’
Southern blot analysis was performed on all PCR-positive plants to identify genomic integration of gusA (Fig. 7b). Transformation efficiency was estimated using the total number of plants exhibiting genomic incorporation, divided by the total number of inoculated explants. The transformation efficiency for ‘F39’ ranged between 0.2 and 0.4% (average efficiency was 0.3 ± 0.1%), and for ‘150’ the range was between 0.2 and 0.8% (average was 0.5 ± 0.3%).
Phenotypes of regenerated transformants
Abnormal morphological characteristics were observed in the transgenic melon plants (Fig. 8a–g). The most severe phenotype was a lack of apical dominance (Fig. 8a) resulting in abnormal growth. Other aberrant morphologies included shorter (Fig. 8d) and longer (Fig. 8g) internodal growth and irregular leaf shapes (Fig. 8f). Plants exhibiting lack of apical dominance and shorter/longer internodes died prematurely in the greenhouse. Morphologically normal transgenic plants produced both male (Fig. 8h) and perfect (Fig. 8i) flowers.
Morphological characteristics of transgenic melon plants. a Transgenic honeydew line ‘090129’ showing a lack of apical dominance, b wild type honeydew ‘150’, c wild type cantaloupe ‘F39’, d transgenic cantaloupe line ‘082417’, e a leaf of wild type cantaloupe ‘F39’, f a leaf of transgenic cantaloupe line ‘082417’, g transgenic honeydew line ‘081105’, h male flowers of wild type (left) and transgenic (right) honeydew ‘150’, i perfect flowers of wild type (left) and transgenic (right) honeydew ‘150’, j transgenic melon plants in the greenhouse
Similar abnormalities reported in western shipper melon ‘Topmark’ and other melon types were suggested to be caused by a high kanamycin concentration (150 mg l−1) in the selection medium (Gonsalves et al. 1994). In our study, kanamycin at 200 and 150 mg l−1 was used for in vitro selection of ‘F39’ and ‘150’, respectively, similar to the levels reported by Gonsalves et al. (1994). However, in our study, abnormal in vitro regenerated wild-type plants were also observed in the absence of kanamycin (data not shown). This observation suggests that the tissue culture process and plant growth regulator effects may induce aberrant morphology in regenerated melon shoots (Larkin and Scowcroft 1981).
In summary, we developed a genetic transformation protocol for two elite breeding lines of melon: western shipper cantaloupe ‘F39’ and honeydew ‘150’. The protocol developed here will serve to initiate successful transformation of these genotypes with novel genes to improve fruit quality or other desired traits, which cannot be obtained through conventional breeding. However, the protocol presented here cannot be considered routine. Further studies are needed to minimize the incidence of chimeras and abnormal plant development.
Abbreviations
- BA:
-
6-Benzyladenine
- IAA:
-
Indole-3-acetic acid
- ABA:
-
Abscisic acid
- nptII :
-
Neomycin phosphotransferase II
- GUS:
-
β-glucuronidase
- MS:
-
Murashige and Skoog
- PCR:
-
Polymerase chain reaction
References
Afroz A, Chaudhry Z, Rashid U, Ali GM, Nazir F, Iqbal J, Khan MR (2010) Enhanced resistance against bacterial wilt in transgenic tomato (Lycopersicon esculentum) lines expressing the Xa21 gene. Plant Cell Tissue Organ Cult 104:227–237
Akasaka-Kennedy Y, Tomita K, Ezura H (2004) Efficient plant regeneration and Agrobacterium-mediated transformation via somatic embryogenesis in melon (Cucumis melo L.). Plant Sci 166:763–769
Ayub R, Guis M, Amor MB, Gillot L, Roustan JP, Latché A, Bouzayen M, Pech JC (1996) Expression of ACC oxidase antisense gene inhibits ripening of cantaloupe melon fruits. Nat Biotechnol 14:862–866
Bordas M, Montesinos C, Dabauza M, Salvador A, Roig LA, Serrano R, Moreno V (1997) Transfer of the yeast salt tolerance gene HAL1 to Cucumis melo L. cultivars and in vitro evaluation of salt tolerance. Trans Res 6:41–50
Castelblanque L, Marfa V, Claveria E, Martinez I, Perez-Grau L, Dolcet-Sanjuan R (2008) Improving the genetic transformation efficiency of Cucumis melo subsp. melo ‘Piel de Sapo’ via Agrobacterium. Cucurbitaceae 2008. In: Pitrat M (ed) Proc IXth EUCARPIA Mtg on Gene and Breed of Cucurbitaceae, INRA, Avignon, France, 21–24 May 2008, pp 627–631
Cheng ZM, Schnurr JA, Kapaun JA (1998) Timentin as an alternative antibiotic for suppression of Agrobacterium tumefaciens in genetic transformation. Plant Cell Rep 17:646–649
Chovelon V, Restier V, Dogimont C, Aarrouf J (2008) Histological study of shoot organogenesis in melon (Cucumis melo L.) after genetic transformation. Cucurbitaceae 2008. In: Pitrat M (ed) Proc IXth EUCARPIA mtg on Gene and Breed of Cucurbitaceae, INRA, Avignon, France, 21–24 May 2008, pp 633–637
Clough GH, Hamm PB (1995) Coat protein transgenic resistance to watermelon mosaic and zucchini yellows mosaic-virus in squash and cantaloupe. Plant Dis 79:1107–1109
Compton ME, Gray DJ, Gaba VP (2004) Genetic transformation of watermelon. In: Curtis IS (ed) Transgenic crops of the world-essential protocols. Kluwer Academic Publishers, Dordrecht, pp 425–433
Costa MS, Miguel C, Oliveira MM (2006) An improved selection strategy and the use of acetosyringone in shoot induction medium increase almond transformation efficiency by 100-fold. Plant Cell Tissue Organ Cult 85:205–209
Dan Y, Charles L, Armstrong Dong J, Feng X, Fry JE, Keithly GE, Martinell BJ, Roberts GA, Smith LA, Tan LJ, Duncan DR (2009) Lipoic acid—a unique plant transformation enhancer. In Vitro Cell Dev Biol Plant 45:630–638
Dong JZ, Yang MZ, Jia SR, Chua NH (1991) Transformation of melon (Cucumis melo L.) and expression from the cauliflower mosaic virus-35 s promoter in transgenic melon plants. Biotechnology 9:858–863
Dutt M, Vasconcellos M, Grosser JW (2011) Effects of antioxidants on Agrobacterium-mediated transformation and accelerated production of transgenic plants of Mexican lime (Citrus aurantifolia Swingle). Plant Cell Tissue Organ Cult Online First™, 6 May 2011
Fan JD, He QW, Wang XF, Yu XY (2007) Antisense acid Invertase (anti-MAI1) gene alters soluble sugar composition and size in transgenic muskmelon fruits. Acta Hort Sinica 34:677–682
Fang G, Grumet R (1990) Agrobacterium tumefaciens mediated transformation and regeneration of muskmelon plants. Plant Cell Rep 9:160–164
Ficcadenti N, Rotino GL (1995) Genotype and medium affect shoot regeneration of melon. Plant Cell Tissue Organ Cult 40:293–295
Fuchs M, McFerson JR, Tricoli DM, McMaster JR, Deng RZ, Boeshore ML, Reynolds JF, Russell PF, Quemada HD, Gonsalves D (1997) Cantaloupe line CZW-30 containing coat protein genes of cucumber mosaic virus, zucchini yellow mosaic virus, and watermelon mosaic virus-2 is resistant to these three viruses in the field. Mol Breed 3:279–290
Fullner KJ, Nester EW (1996) Temperature affects the T-DNA transfer machinery of Agrobacterium tumefaciens. J Bacteriol 178:1498–1504
Gaba V, Elman C, Watad AA, Gray DJ (1996) Ancymidol hastens in vitro bud development in melon. HortScience 31:1223–1224
Galperin M, Patlis L, Ovadia A, Wolf D, Zelcer A, Kenigsbuch D (2003) A melon genotype with superior competence for regeneration and transformation. Plant Breed 122:66–69
Gonsalves C, Xue B, Yepes M, Fuchs M, Ling KS, Namba S, Chee P, Slightom JL, Gonsalves D (1994) Transferring cucumber mosaic virus-white leaf strain coat protein gene into Cucumis melo L. and evaluating transgenic plants for protection against infections. J Am Soc Hort Sci 119:345–355
Gould JH, Magallens-Cedeno M (1998) Adaptation of cotton shoot apex culture to Agrobacterium-mediated transformation. Plant Mol Biol Rep 16:1–10
Gray DJ, McColley DW, Compton ME (1993) High-frequency somatic embryogenesis from quiescent seed cotyledons of Cucumis melo cultivars. J Am Soc Hort Sci 118:425–432
Guis M, Latché A, Pech JC, Roustan JP (1997) An efficient method for production of diploid cantaloupe charentais melon (Cumcumis melo L. var cantalupensis) by somatic embryogenesis. Scientia Hort 69:199–206
Guis M, Amor MB, Latché A, Pech JC, Roustan JP (2000) A reliable system for the transformation of cantaloupe charentais melon (Cucumis melo L. var. cantalupensis) leading to a majority of diploid regenerants. Scientia Hort 84:91–99
Hao J, Niu Y, Yang B, Gao F, Zhang L, Wang J, Hasi A (2011) Transformation of a marker-free and vector-free antisense ACC oxidase gene cassette into melon via the pollen-tube pathway. Biotechnol Lett 33:55–61
Hoekema A, Hirsch PR, Hooykaas PJJ, Schilperoort RA (1983) A binary plant vector strategy based on separation of vir- and T-region of the Agrobacterium tumefaciens Ti-plasmid. Nature 303:179–180
Hood EE, Gelvin SB, Melchers LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plant. Transgenic Res 2:208–218
Hussain AF, Anfoka GH, Hassawi DS (2008) Transformation of tomato with TYLCV gene silencing construct using optimized Agrobacterium-mediated protocol. Biotechnology 7:537–543
Ieamkhang S, Chatchawankanphanich O (2005) Augmentin® as an alternative antibiotic for growth suppression of Agrobacterium for tomato (Lycopersicon esculentum) transformation. Plant Cell Tissue Organ Cult 82:213–220
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuonidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 13:3901–3907
Kathiravan K, Vengedesan G, Singer S, Steinitz B, Paris HS, Gaba V (2006) Adventitious regeneration in vitro occurs across a wide spectrum of squash (Cucurbita pepo) genotypes. Plant Cell Tissue Organ Cult 85:285–295
Keng CL, Hoong LK (2005) In vitro plantlets regeneration from nodal segments of muskmelon (Cucumis melo L.). Biotechnology 4:354–357
Kintzios SE, Taravira N (1997) Effect of genotype and light intensity on somatic embryogenesis and plant regeneration in melon (Cucumis melo L.). Plant Breed 116:359–362
Kintzios S, Stavropoulou Er, Skamneli S (2004) Accumulation of selected macronutrients and carbohydrates in melon tissue cultures: association with pathways of in vitro dedifferentiation and differentiation (organogenesis, somatic embryogenesis). Plant Sci 167:655–664
Kumar V, Campbell LM, Rathore KS (2011) Rapid recovery- and characterization of transformants following Agrobacterium-mediated T-DNA transfer to sorghum. Plant Cell Tissue Organ Cult 104:137–146
Larkin PJ, Scowcroft WR (1981) Somaclonal variation—a novel source of variability from cell cultures for plant improvement. Theor Appl Genet 60:197–214
Li X, Liu C, Ritchie SW, Peng J, Gelvin SB, Hodges TK (1992) Factors influencing Agrobacterium-mediated transient expression of gusA in rice. Plant Mol Biol 20:1037–1048
Mollel MHN, Goyvaerts EMA (2004) Preliminary examination of factors affecting Agrobacterium tumefaciens-mediated transformation of marula, Sclerocarya birrea subsp. caffra (Anacardiaceae). Plant Cell Tissue Organ Cult 79:321–328
Moore GA (1995) Phenotypic stability of transgenic citrus. HortScience 30:903–904 (Abstract)
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Nakagawa H, Saijyo T, Yamauchi N, Shigyo M, Kako S, Ito A (2001) Effects of sugars and abscisic acid on somatic embryogenesis from melon (Cucumis melo L.) expanded cotyledon. Scientia Horti 90:85–92
Nuňez-Palenius HG, Cantliffe DJ, Huber DJ, Ciardi J, Klee HJ (2006) Transformation of a muskmelon ‘Galia’ hybrid parental line (Cucumis melo L. var. reticulatus Ser.) with an antisense ACC oxidase gene. Plant Cell Rep 25:198–205
Nuňez-Palenius HG, Grumet R, Lester G, Cantliffe D (2008) Melon fruits: genetic diversity, physiology, and biotechnology features. Crit Rev Biotechnol 28:13–55
Olhoft PM, Somer DA (2001) l-Cysteine increases Agrobacterium-mediated T-DNA delivery into soybean cotyledonary-node cells. Plant Cell Rep 20:706–711
Oridate T, Oosawa K (1986) Somatic embryogenesis and plant regeneration from suspension callus culture in melon. Jpn J Breed 36:424–428
Oridate T, Atsumi H, Ito S, Araki H (1992) Genetic differences in somatic embryogenesis from seeds in melon (Cucumis melo L.). Plant Cell Tissue Organ Cult 29:27–30
Orts MC, Garciasogo B, Roche MV, Roig LA, Moreno V (1987) Morphogenetic response of calli derived from primary explants of diverse cultivars of melon. HortScience 22:666
Ostergaard L, Yanofsky M (2004) Establishing gene function by mutagenesis in Arabidopsis thaliana. Plant J 39:682–696
Seo MS, Takahashi S, Kadowaki K, Kawamukai M, Takahara M, Takamizo T (2011) Expression of CoQ10-producing ddsA transgene by efficient Agrobacterium-mediated transformation in Panicum meyerianum. Plant Cell Tissue Organ Cult Online First™ 10 June 2011
Sharma M, Kothari-Chajer A, Jagga-Chugh S, Kothari SL (2011) Factors influencing Agrobacterium tumefaciens-mediated genetic transformation of Eleusine coracana (L.) Gaertn. Plant Cell Tissue Organ Cult 105:93–104
Shilpa KS, Kumar VD, Sujatha M (2010) Agrobacterium-mediated genetic transformation of safflower (Carthamus tinctorius L.). Plant Cell Tissue Organ Cult 103:387–401
Skroch PW, Nienhuis J (1995) Qualitative and quantitative characterization of RAPD variation among snap bean genotypes. Theor Appl Genet 91:1078–1085
Slater SMH, KellerWA, Scoles G (2011) Agrobacterium-mediated transformation of Eruca sativa. Plant Cell Tissue Organ Cult Online First™ 5 January 2011
Souza FVD, Garcia-Sogo B, da Silva Souza A, San-Juán P, Moreno V (2006) Morphogenetic response of cotyledon and leaf explants of melon (Cucumis melo L.) cv. Amarillo Oro. Braz Arch Biol Technol 49:21–27
Thiruvengadam M, Hsu W, Yang C (2011) Phosphomannose-isomerase as a selectable marker to recover transgenic orchid plants (Oncidium Gower Ramsey). Plant Cell Tissue Organ Cult 104:239–246
Toldi O, Tóth S, Pónyi T, Scott P (2002) An effective and reproducible transformation protocol for the model resurrection plant Craterostigma plantagineum Hochst. Plant Cell Rep 211:63–69
Vallés MP, Lasa JM (1994) Agrobacterium-mediated transformation of commercial melon (Cucumis melo L. cv. Amarillo Oro). Plant Cell Rep 13:145–148
Wu HW, Yu TA, Raja JAJ, Wang HC, Yeh SD (2009) Generation of transgenic oriental melon resistant to Zucchini yellow mosaic virus by an improved cotyledon-cutting method. Plant Cell Rep 28:1053–1064
Yadav RC, Saleh MT, Grumet R (1996) High frequency shoot regeneration from leaf explants of muskmelon. Plant Cell Tissue Organ Cult 45:207–214
Yasmin A, Debener T (2010) Transient gene expression in rose petals via Agrobacterium infiltration. Plant Cell Tissue Organ Cult 102:245–250
Zambre M, Terryn N, De Clercq J, De Buck S, Dillen W, Van Montagu M, Van Der Straeten D, Angenon G (2003) Light strongly promotes gene transfer from Agrobacterium tumefaciens to plant cells. Planta 216:580–586
Zheng Q, Ju B, Liang L, Xiao X (2005) Effects of antioxidants on the plant regeneration and GUS expressive frequency of peanut (Arachis hypogaea) explants by Agrobacterium tumefaciens. Plant Cell Tissue Organ Cult 811:83–89
Acknowledgments
This research was supported by an USDA-CSREES Special Grant “Designing Foods for Health” (2006-34402-17121, 2008-34402-19195 and 2009-34402-19831) through the Vegetable and Fruit Improvement Center, Texas A&M University. We wish to thank Drs. Rebecca Grumet, Keerti Rathore, Sung Hun Park, Sylvain Marcel, Selvakumar Veluchamy, Mehdi Kabbage, Ms. Marianne Arnold, Sue Hammar, and Jungeun Kim for technical advice and help.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ren, Y., Bang, H., Curtis, I.S. et al. Agrobacterium-mediated transformation and shoot regeneration in elite breeding lines of western shipper cantaloupe and honeydew melons (Cucumis melo L.). Plant Cell Tiss Organ Cult 108, 147–158 (2012). https://doi.org/10.1007/s11240-011-0024-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-011-0024-6