Abstract
An improved bacterial preculture protocol for Agrobacterium-mediated genetic transformation was developed for an economic tomato cultivar (Solanum lycopersicum L. cv. Zhongshu No. 4). Frequencies of transient gene expression and stable transformation were influenced by the density of Agrobacterium preculture and not the density of Agrobacterium used for infection. The improved protocol presented in this study depends on the use of an overnight-grown Agrobacterium preculture density of OD600 nm = 1.0, diluted 1/10th with Luria-Bertani (LB) liquid medium, and grown for an additional 4 h. Cultures are collected and resuspended in a liquid cocultivation medium-I, adjusted to OD600 nm = 0.1. Using this modified Agrobacterium preparation, transient β-glucuronidase expression was higher than 90%, and transformation efficiency reached 44.7%. This improved transformation is simple, repeatable, does not require a feeder layer, and most notably, the transformation frequency is stable and highly efficient.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tomato (Solanum lycopersicum L.) is an economically important crop. However, abiotic stresses, including salinity, heat, drought, and nutrient deficiencies (such as phosphate and nitrate) often constrain fruit productivity (Raghothama 2000; Abel et al. 2002; Zhu 2002; Bhatnagar-Mathur et al. 2008). Developing transgenic plants is an effective approach for improving abiotic stress tolerance (Park et al. 2005; Bhatnagar-Mathur et al. 2008). Establishment of an efficient transformation system is essential. The first successful transformation of tomato was reported by Horsch et al. (1985) using Agrobacterium-mediated transformation. To date, various factors affecting the efficiency of Agrobacterium-mediated transformation have been investigated. These include cocultivation temperature (Dillen et al. 1997), type (Frary and Earle 1996; Ellul et al. 2003; Park et al. 2003; Wu et al. 2006) and developmental status of explants (Chyi and Phillips 1987; Hamza and Chupeau 1993; Tabaeizadeh et al. 1999; Dan et al. 2006; Sun et al. 2006), use of feeder-layer cells (van Roekel et al. 1993; Qiu et al. 2007), addition of phenolic compounds (Lipp João and Brown 1993; Cortina and Culiáňez-Macià 2004; Sun et al. 2006; Wu et al. 2006), vector constructs (van Roekel et al. 1993; Qiu et al. 2007), Agrobacterium strains (van Roekel et al. 1993; Wroblewski et al. 2005), and concentration (Ellul et al. 2003; Dan et al. 2006; Qiu et al. 2007; Wu et al. 2006) and composition of the medium (Hamza and Chupeau 1993; Frary and Earle 1996; Ling et al. 1998; Vidya et al. 2000; Krasnyanski et al. 2001; Pozueta-Romero et al. 2001; Park et al. 2003; Cortina and Culiáňez-Macià 2004; Wu et al. 2006). Current transformation efficiencies ranges from 6 to 40% (Hamza and Chupeau 1993; Frary and Earle 1996; Ling et al. 1998; Vidya et al. 2000; Krasnyanski et al. 2001; Park et al. 2003; Sun et al. 2006). In spite of these successes, most protocols rely on the use of either cumbersome feeder layers from petunia, tobacco, or tomato, complex media formulations, or time-consuming successive subcultures. Hence, optimizing the frequency of transformation of tomato is highly desirable.
Agrobacterium tumefaciens has been widely used to transform numerous plant species, including tomato. The ability of Agrobacterium to transform plants is under highly regulated genetic control, especially for the vir gene (Hansen et al. 1994; Gelvin 2003; Yuan et al. 2007). Over the years, optimal conditions for Agrobacterium vir gene induction have been extensively investigated (Joubert et al. 1995; Hwang and Gelvin 2004; Yuan et al. 2007). In addition, bacterial concentrations used for infiltration have also been optimized (Pozueta-Romero et al. 2001; Wu et al. 2006; Qiu et al. 2007). For tomato transformation, Agrobacterium concentrations usually range between 0.01 and 1.0 at OD600 nm (Lipp João and Brown 1993; Frary and Earle 1996; Ling et al. 1998; Krasnyanski et al. 2001; Pozueta-Romero et al. 2001; Park et al. 2003; Dan et al. 2006; Wu et al. 2006; Qiu et al. 2007), and overnight-grown Agrobacterium cultures are usually diluted prior to cocultivation (Krasnyanski et al. 2001; Dan et al. 2006; Wu et al. 2006). McCormick et al. (1986) incubated explants in a 1:20 dilution of Agrobacterium C58C1 (Rifr) culture; while, Lipp João and Brown (1993) incubated explants in a 1:10 dilution, grown for about 4 h with Agrobacterium C58C1 (Rifr) culture. On the other hand, Frary and Earle (1996) diluted an overnight-grown Agrobacterium strain LB4404 culture one to twofold to reach OD600 nm = 0.4 to obtain an optimal infection concentration; whereas, Raj et al. (2005) floated precultured cotyledons in a 1:15 dilution of Agrobacterium strain LB4404. Sun et al. (2006) used a 1:20–40 dilution of Agrobacterium C58C1 (Rifr) culture; while, Qiu et al. (2007) diluted an overnight-grown culture of Agrobacterium strain EHA101 to OD600 nm = 0.2. In our pilot experiment on tomato transformation, we found that when using cotyledons as explants, transformation frequencies varied greatly, even with the same Agrobacterium strain at the same inoculation concentration and length of inoculation followed by the same cocultivation conditions. These results indicated that there might be other factors that influenced transformation frequency of tomato.
This study reports on the effects of density of Agrobacterium preculture of tomato cotyledons under different compositions of cocultivation media on frequency of transformation of tomato.
Materials and methods
Plant material
Seeds of tomato (Solanum lycopersicum L.) cultivar Zhongshu No. 4 were surface-sterilized for 30 s with 75% ethanol, 15 min in a 2% NaClO, followed by eight washes with sterilized-distilled water. Sterile seeds were grown on a germination medium (Table 1 unless otherwise stated). Cultures were kept at 22–24°C under a 16 h photoperiod with a light intensity of 72 μmol m−2 s−1.
Cotyledons were excised from 6 to 8-day-old seedlings, and cotyledonary explants, ~25 mm2, were incubated, placed abaxially, on a preculture medium as described in Table 1. Preculture was conducted for a period of 2 days at 22–24°C under a 16 h photoperiod and a light intensity of 72 μmol m−2 s−1.
Agrobacterium strain and cloning vectors
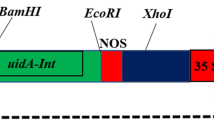
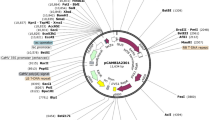
Agrobacterium strain LB4404 carrying the pBI121 binary vector (Jefferson et al. 1987) was used. The binary vector contains the selectable marker gene neomycin phosphotransferase II (nptII), driven by the nopaline synthase (nos) promoter, and a β-glucuronidase (GUS) gene under the control of the cauliflower mosaic virus 35S promoter (CaMV35S). Besides pBI121, three pBI121-derived vectors, designated B, C, and D (Fig. 1), were also constructed to. Vector B contained a phosphorus-related gene 172, a small RNA DBE#172 from Arabidopsis thaliana ecotype Columbia, under the control of CaMV35S, used as a control; while, vectors C and D carried the phosphorus-related gene ath-microRNA399d, a microRNA from A. thaliana ecotype Columbia under the control of CaMV35S and rd29A promoter from A. thaliana ecotype Columbia, respectively, for transformation experiments.
Schematic illustration of the binary vector plasmid constructs. LB left border, P nos nopaline synthase promoter, nptII neomycin phosphotransferase, T nos nopaline synthase terminator, RD29A RD29A promoter, MIR399 microRNA 399d (a phosphorus-related gene), RB right border, CaMV35S cauliflower mosaic virus 35S promoter, 172 small RNA DBE#172 (a reference gene)
Agrobacterium culture and cocultivation
A single colony of A. tumefaciens carrying the binary vector was inoculated in 10 ml Luria-Bertani (LB) broth containing 100 mg l−1 kanamycin, 25 mg l−1 rifampicin, and 50 mg l−1 streptomycin, and grown overnight on a rotary shaker at 28°C. Overnight-grown bacterial precultures were incubated to OD600 nm = 1.0, unless otherwise stated, diluted tenfold with LB liquid medium, and grown for another 4 h. Cultures were collected by centrifugation at 4,000 × g for 10 min at room temperature, and resuspended in a liquid cocultivation medium-I (Table 1), and adjusted to OD600 nm = 0.1. Precultured cotyledons were submerged in the bacterial solution for 15 min. Agrobacterium-infected explants were blotted onto a sterile paper towel, and cultured on cocultivation medium-II (Table 1, unless otherwise stated) for 3 days at 22–24°C in the dark.
To investigate the influence of Agrobacterium preculture density on transformation efficiency (TE) and transient GUS expression, the overnight Agrobacterium preculture was cultured to OD600 nm = 0.1, 0.5, 1.0, 1.5, and 2.0, and then diluted with LB liquid medium to OD600 nm = 0.1 and grown for another 4 h. Apart from Agrobacterium preculture densities, all other parameters remained the same as described above. To investigate the effects of Agrobacterium preculture density in combination with acetosyringone (AS), zeatin (ZT), and pH of cocultivation medium-II on transient GUS expression, an overnight Agrobacterium preculture was cultured to OD600 nm = 0.1, 1.0, and 2.0, and then diluted with LB liquid medium to OD600 nm = 0.1 and grown for another 4 h. Apart from Agrobacterium culture densities, all other parameters remained the same as described above.
To investigate whether Agrobacterium density effects on transient GUS expression levels were influenced by the concentration of AS in cocultivation medium-II, 0, 100, and 200 μM AS were added to cocultivation medium-II (apart from AS concentration, other parameters used in this study are listed in Table 1). To investigate whether Agrobacterium density effects on transient GUS expression level were influenced by ZT in cocultivation medium, 0 and 2 mg l−1 ZT were added to medium-II (apart from the ZT concentration, other parameters used in this study are listed in Table 1). To investigate whether Agrobacterium density effects on transient GUS expression levels were influenced by the pH of the cocultivation medium, medium-II was adjusted to pH 5.2, 5.5 or 5.8 using diluted NaOH solution (apart from pH, other parameters used in this study are listed in Table 1).
TE was calculated as: number of cotyledons with putative transformants/the total number of cotyledons per treatment, and the transient GUS levels were evaluated as: number of cotyledons with blue-stained/the total number of cotyledons per treatment.
Selection of transformants
Agrobacterium-infected explants cocultivated for 3 days were then transferred to selection medium (Table 1), and subcultured onto fresh medium once every 3 weeks until shoot buds were observed. When shoots where 5 mm in length, they were excised from callus and transferred to rooting medium-I (Table 1) for 2 weeks. Rooted shoots were then transferred to rooting medium-II (Table 1) for an additional week. Finally, putative transformed plants were transferred to 100% sterilized vermiculite and watered with 1/50 MS basal medium (Murashige and Skoog 1962). Pots with plants were covered with a plastic bag to maintain high relative humidity. The plastic bag was opened for 1 h daily to add fresh air, and the plants were lightly watered. About 7 days later, the plastic bags were removed and plants were grown under greenhouse conditions. Plants later were transferred to an agriculture soil into larger pots for maturation and seed collection of self-pollinated T1 generation.
GUS assays
For transient histochemical GUS assay, 3-day-old co-cultured cotyledons were immersed in the staining solution (50 mM sodium phosphate buffer, pH 7.0, 0.1% Triton X-100, 1 mM 5-bromo-4-chloro-3-indole-β-d-glucuronide) overnight (16 h) at 37°C as described by Jefferson et al. (1987). These were rinsed successively with 70% ethanol and 30% acetic acid for 24 h, and numbers of blue-stained explants were counted. For stable histochemical GUS assay, plantlets were incubated in the staining solution, as described above. After 16 h at 37°C, plantlets were rinsed successively with 70% ethanol and 30% acetic acid for 24 h.
Identification and analysis of putative transformants with polymerase chain reaction (PCR), Southern blot, and reverse transcription PCR (RT-PCR) analysis
Genomic DNA was extracted from leaves of greenhouse-grown tomato plants, and the presence of a 492-bp nptII gene-specific fragment was used to identify transformants. Approximately 10 ng of genomic DNA was used as template for PCR analysis. The nptII gene-specific primers used in this analysis were NptII F1-252 (5′-CACTGAAGCGGGAAGGGACT-3′) and NptII R1-743 (5′-GCGGCGATACCGTAAAGCAC-3′).
To confirm integration of the transgene, Southern blotting was conducted. Genomic DNA from PCR-positive transformants was subjected to enzymatic digestion. About 30 μg of genomic DNA was completely digested with EcoRI or BamHI, separated on a 1.0% agarose gel, and transferred to a Hybond H+ membrane (Amersham Biosciences, Buckinghamshire, UK). The membrane was hybridized with a nptII Dig-labeled probe (Roche Applied Science, Mannheim, Germany), following the manufacturer’s protocol. For the nptII probe, an internal fragment of the gene was obtained by PCR amplification.
For expression analysis, total RNA was extracted from leaves of greenhouse-grown tomato seedlings using RNAsio Reagent (TaKaRa, Tokyo, Japan) according to the manufacturer’s protocol. To ensure the absence of genomic DNA contamination, RNA preparations were tested for the amplification of an α-tubulin gene fragment using the following primer pairs: tub F (5′-TGAACAACTCATAAGTGGCAAAG-3′) and tub R (5′-TCCAGCAGAAGTGACCCAAGAC-3′). Only RNAs without DNA contamination (negative α-tubulin gene fragment amplification) were used for subsequent preparation in cDNA synthesis. One microgram of total RNA was used to synthesize cDNA using reverse transcriptase XL (AMV) (TaKaRa) following the manufacturer’s protocol. The cDNA samples were used as templates for RT-PCR analysis. Primers NptII F2-61 (5′-GGCTATGACTGGGCACAACA-3′) and NptII R2-329 (5′-GCAGGAGCAAGGTGAGATGAC-3′) were used to amplify the 269-bp nptII gene fragment. PCR and RT-PCR reactions were performed in a 10-μl total volume consisting of 1× reaction buffer, 1.5 mM MgCl2, 0.2 mM dNTPs, 0.2 μM of each primer, and 0.25 U Taq DNA polymerase (TaKaRa). The PCR cycles included an initial 5 min denaturation at 94°C, followed by 30 cycles each of 30 s at 94°C, 30 s at 55°C, 30 s at 72°C, and a final 7 min extension at 72°C.
Results
Influence of Agrobacterium preculture density on TE
As it was important to optimize the recovery of putative transformants of tomato to increase the likelihood of stable transformation, we evaluated several Agrobacterium preculture densities for their ability to recovery of putative transformants in cotyledon. TE was calculated as: number of cotyledons with putative transformants/the total number of cotyledons per treatment. An Agrobacterium preculture absorption OD600 nm of 1.0 produced the highest TE of 44.7% of tomato cotyledonary explants. TE decreased dramatically when the Agrobacterium preculture OD600 nm was either 0.1 or 2.0 (Fig. 2). As the concentration of Agrobacterium solution used for infection was the same, this indicated that the density of Agrobacterium preculture significantly affected TE values.
Transformation efficiency (TE) with different Agrobacterium preculture densities. OD600 nm = 0.1, 0.5, 1.0, 1.5, and 2.0 in Agrobacterium LB4404 preculture. TE was calculated as: number of cotyledons with putative transformants/the total number of cotyledons per treatment. TE is expressed as the mean ± SD. Differences at the 5% significance level were obtained using least significance tests (small letters). Media used in tomato tissue culture and Agrobacterium transformation are described in Table 1
Effect of Agrobacterium preculture density on transient GUS expression
When the preculture OD600 nm was 0.1, transient GUS expression in cotyledons was 78.8%; while, when the preculture OD600 nm was 2.0, transient GUS expression in cotyledons was 65.0%. The highest transient GUS expression in cotyledons was obtained at OD600 nm of 1.0 (Fig. 3). Using an optimized Agrobacterium preculture density (OD600 nm = 1.0), the transient GUS expression efficiency reached 90%. A strong correlation between transient GUS expression and TE (R 2 = 0.7073; Fig. 4) was observed, indicating that a transient GUS expression level was critical for achieving high rates of TE.
Transient GUS expression with different Agrobacterium preculture OD600 nm of 0.1, 0.5, 1.0, 1.5, and 2.0 in Agrobacterium LB4404 preculture. The transient GUS expression levels were evaluated as: number of cotyledons with blue-stained/the total number of cotyledons per treatment in a total of 80 explants pooled from two independent experiments. Differences in transient GUS expression levels were tested at the 5% significance level using least significance tests (small letters). Media used in Agrobacterium transformation are described in Table 1 unless otherwise stated
Correlation between transient GUS expression and transformation efficiencies (TE) in cotyledons. Data were collected from Figs. 2 and 3. TE was calculated as: number of cotyledons with putative transformants/the total number of cotyledons per treatment. The transient GUS expression levels were evaluated as: number of cotyledons with blue-stained/the total number of cotyledons per treatment in a total of 80 explants pooled from two independent experiments
Effects of Agrobacterium preculture density in combination with AS, ZT, and the pH of cocultivation media-II on transient GUS expression
To investigate whether bacterial density effects on transient GUS expression are influenced by the presence of a phenolic compound such as AS, cytokinin such as ZT, or pH of the cocultivation medium, the effects of Agrobacterium preculture density on TE was further investigated using a transient GUS expression system. An Agrobacterium preculture OD600 nm of 1.0 resulted in the highest transient GUS expression with various phenolic compound, pH, and composition of the cocultivation medium-II.
To investigate whether effects of bacterial density on transient GUS expression levels were influenced by the concentration of AS in the cocultivation medium-II, different levels of AS were added to cocultivation medium-II. As noted above, with the same AS concentration, highest transient GUS expression levels of cotyledons were obtained at bacterial density of OD600 nm = 1.0. At this optimal bacterial density, 100 μM AS in the cocultivation medium yielded the highest transient GUS expression in cotyledonary explants. At both optimal bacterial density and AS concentration in the cocultivation medium-II, the transient GUS expression efficiency was over 90% (Fig. 5a).
Effect of Agrobacterium LB4404 preculture density on the transient GUS expression levels at different cocultivation medium-II concentrations. Values of 0.1, 1.0, and 2.0 correspond to Agrobacterium LB4404 preculture absorption at OD600 nm = 0.1, 1.0, and 2.0, respectively; the transient GUS expression levels were evaluated as: number of cotyledons with blue-stained/the total number of cotyledons per treatment in a total of 80 explants pooled from two independent experiments; differences in transient GUS expression levels were tested at the 5% significance level using least significance tests (small letters). a Effect of Agrobacterium LB4404 preculture density on the transient GUS expression level with different acetosyringone (AS) concentrations in cocultivation medium-II; b Effect of Agrobacterium LB4404 preculture density on the transient GUS expression level at different zeatin (ZT) concentrations in cocultivation medium-II; c Effect of Agrobacterium LB4404 preculture density on the transient GUS expression level at different pH values of cocultivation medium-II
As the effect of bacterial density on transient GUS expression might be influenced by ZT in the cocultivation medium, ZT was either added to medium-II or left out. At the same ZT concentration, the highest transient GUS levels in cotyledonary explants were obtained at bacterial density of OD600 nm of 1.0 during preculture. In contrast to AS, the presence of ZT in cocultivation medium-II did not significantly improve transient GUS expression. Using optimal Agrobacterium densities (OD600 nm = 1.0), the transient GUS expression efficiency of cotyledon explants was over 90% (Fig. 5b).
To investigate whether Agrobacterium density effects on transient GUS expression levels were influenced by the pH of the cocultivation medium, medium-II was adjusted to different pH value before autoclaved. At the same pH, the highest transient GUS levels of cotyledonary explants were obtained with an Agrobacterium LB4404 preculture OD600 nm of 1.0. Under optimal tenfold bacterial cell dilution, the best pH for the cocultivation medium was 5.2 (Fig. 5c). Using an optimal bacterial density (OD600 nm = 1.0) and pH of cocultivation medium-II, transient GUS expression efficiency of cotyledonary explants was over 90%.
Effects of different constructs on TE
When three different constructs, carrying phosphorus-related genes, were used for transformation, in conjunction with Agrobacterium LB4404 preculture density of OD600 nm = 1.0, no significant differences in TEs were observed. TEs of constructs B, C, and D were 47.4 ± 11.9%, 44.9 ± 5.7%, and 35.0 ± 2.4%, respectively.
PCR, Southern blot, and RT-PCR analysis
All of putative transformed shoots were successfully rooted on Rooting medium-I. The putative transgenic plants were identified by PCR amplification for presence of the nptII transgene (Fig. 6a). The expected 490-bp band was detected in 61 out of 70 randomly selected plantlets, but it was absent from the negative control (non-transformed) plant (Fig. 6a). Putative transgenic plants were further confirmed by Southern blot analysis (Fig. 6b). Three randomly selected genomic samples were completely digested with EcoRI or BamHI, and hybridized with an nptII Dig-labeled probe (Roche Applied Science), following the manufacturer’s protocol. All candidate transformants were detected with the nptII gene fragment. Southern blot analysis of transformant plants confirmed the integration of the T-DNA into the tomato genome. Positive hybridization signals were observed in all transformants, whereas, the signal was absent in untransformed plant. Southern blot hybridization has revealed multiple insertion copy numbers in genomic DNA of tomato plants. Approximately one to three copies of nptII gene, ranging between 3.3 and 9.4 kb in size, are observed in these transgenic plants (Fig. 6b).
PCR, Southern blot, and RT-PCR analysis of putative transformants. a PCR amplification of the nptII fragment from putative transformants DNA (491 bp): lane M DL2000-bp marker (TaKaRa), lanes 1–12 putative transformants, lane P positive control (plasmid pBI121 DNA), lane N negative control (wild-type tomato); b Southern blot analysis of genomic DNA from three randomly selected tomato plants (T0). Thirty micrograms of DNA, following digestion with EcoRI (E) or BamHI (B), were loaded in each lane. Lane P positive control, pMD18 vector containing the nptII gene fragment (total length = 3,375 bp); Lane N negative control, untransformed tomato plants; Lanes 1–3 transformed tomato plants. For the nptII probe, an internal fragment of the gene was obtained by PCR amplification; c PCR amplification of nptII fragment from putative transformants RNA (269 bp): lane M DL2000-bp marker (TaKaRa), lanes 1–12 putative transformants, lane P positive control (plasmid pBI121 DNA), lane N negative control (wild-type tomato DNA)
Confirmed transformants were further analysis for levels of gene expression by RT-PCR using primers for the nptII gene. The expected 270-bp band was found in the positive control plasmid and in 44 out of 48 randomly selected plantlets, but it was absent from the negative control (Fig. 6c). We also found that the nptII gene expression level was not significantly changed (data not shown).
Genetic analysis
To investigate inheritance of the kanamycin-resistance nptII transgene, six independent transgenic lines were random selected. T0 seeds were harvested and germinated on germination medium containing 25 mg l−1 kanamycin. After 10 days, kanamycin-resistant seedlings with more than 5-cm long primary roots and green leaves were clearly distinguished from sensitive seedlings of less than 3-cm primary roots and yellow leaves. Segregation analysis of the T1 generation gave a Mendelian ratio of 3:1 (Table 2).
Comparison of previous and current protocols of tomato transformation
To compare earlier protocols with the current protocol of tomato transformation, we performed six transient GUS expression experiments according to the previous protocol (overnight Agrobacterium culture diluted with LB liquid medium to OD600 nm = 0.1, grown for another 4 h, with other procedures as described in “Materials and methods”) (McCormick et al. 1986; Lipp João and Brown 1993; Frary and Earle 1996; Raj et al. 2005; Sun et al. 2006; Qiu et al. 2007). We also conducted six transient GUS expression experiments according to the improved protocol (overnight Agrobacterium LB4404 preculture was cultured to OD600 nm = 1.0, then diluted with LB liquid medium to OD600 nm = 0.1, and grown for another 4 h, with other procedures as described in “Materials and methods”). The results (Table 3) showed that the current protocol yielded a higher efficiency of transformation and resulting in a higher frequency of transformation (Fig. 7).
Tomato transformation. a Transient GUS expression visualized by dark grey spots (in print version) or blue spots (in electronic version) in cotyledons; b rooting of transgenic shoots with 50 mg l−1 kanamycin; c stable GUS expression of a transgenic tomato plant; d transgenic tomato plant growing in the greenhouse
Discussion
Agrobacterium-mediated transformation is a common approach for introducing elite genes into plant genotypes. Several factors are taken into consideration to develop highly efficient transformation protocols, and among those, the concentration of Agrobacterium infiltration solution is an important factor in these protocols.
Levels of Agrobacterium infiltration solutions have been optimized, with OD600 nm ranging between 0.01 and 1.0 (Lipp João and Brown 1993; Frary and Earle 1996; Ling et al. 1998; Krasnyanski et al. 2001; Pozueta-Romero et al. 2001; Park et al. 2003; Dan et al. 2006; Qiu et al. 2007; Wu et al. 2006). Bacterial cells are usually cultured overnight and then diluted 2–50 times to achieve an appropriate infection concentration (McCormick et al. 1986; Lipp João and Brown 1993; Frary and Earle 1996; Raj et al. 2005; Sun et al. 2006; Qiu et al. 2007). In this study, bacterial cell density during preculture has a significantly effect on both transient GUS expression and TE (Figs. 2, 3). For strain LB4404, the optimal density is OD600 nm = 1.0.
In this study, the influence of the bacterial density during preculture under different AS and ZT concentrations, and different pH, was investigated revealing that regardless of all these factors, the level of transient GUS expression was highest when OD600 nm of bacterial cells was 1.0 during preculture. This further confirmed the importance of the density of bacterial cells in the preculture medium on successful transformation of tomato. Moreover, this is not unique to Agrobacterium strain LB4404. In this study, an agropine/succinamopine-type Agrobacterium strain EHA101 harboring pUbi, carrying the GUS gene, and a nopaline-type Agrobacterium strain C58 carrying pBI121, also carrying the GUS gene, were investigated. For Agrobacterium EHA101, bacterial density of OD600 nm = 1.5 or 3.0 during preculture resulted in highest frequencies of transient GUS expression, 71.3 ± 1.8% and 71.3 ± 5.3%, respectively. However, for the nopaline Agrobacterium strain C58, no bacterial density effects were detected as 100% transient GUS expression levels of cotyledons were obtained regardless of OD600 nm. As all infection experiments were performed at similar bacterial densities, it is likely that the effect of bacterial density during preculture is likely attributed to the growth stage of the bacterial preculture. Growth curves of bacterial cells showed that the concentrations of optimal dilutions for LB4404 and EHA101 bacterial cells fit the early-middle exponential (logarithmic) growth phase (data not shown). This indicated that in addition to the population effect; i.e., proper densities of the infection solution, of Agrobacterium, developmental stages of individual cells likely contributed to the vir gene-mediated gene transfer and incorporation into target plant tissues.
Cotyledons have been commonly used as explants in tomato transformation (Chyi and Phillips 1987; Lipp João and Brown 1993; Frary and Earle 1996; Krasnyanski et al. 2001; Park et al. 2003; Raj et al. 2005; Sun et al. 2006; Wu et al. 2006). Cotyledon explants were wounded and infected with Agrobacterium followed by cocultivation for 3 days in the dark. Longer exposure of explants to Agrobacterium resulted in 100% Agrobacterium regrowth after this cocultivation period and made it more difficult to eliminate Agrobacterium (data not shown). This phenomenon was also observed by Krasnyanski et al. (2001).
In this study, presence of AS, and ZT, as well as pH of cocultivation medium are critical factors in transformation and regeneration of transformants from explants. Addition of AS to cocultivation media improves transient GUS expression level (Figs. 3, 5) and TE (Fig. 2; Lipp João and Brown 1993; Uranbey et al. 2005; Wu et al. 2006). The infection capability of Agrobacterium is dependent on the vir gene located on the Ti plasmid. The vir gene has been reported to be highly expressed at an appropriate low pH (Lipp João and Brown 1993; Ogaki et al. 2008). This has been also confirmed in this study whereby a lower pH (pH 5.2) produced the highest transient GUS expression in cotyledonary explants (Fig. 5c).
Conclusions
Taken together, our results clearly show that efficiency of transformation in tomato is highly influenced by the density of preculture of bacterial cells. The improved protocol presented in this study depends on the use of an overnight-grown Agrobacterium preculture density of OD600 nm = 1.0, diluted 1/10th with LB liquid medium, and grown for an additional 4 h. Cultures are collected and resuspended in a liquid cocultivation medium-I, adjusted to OD600 nm = 0.1. With this improved protocol, a transient GUS expression is more than 90% and a TE of 44.7% is obtained. This transformation protocol is simple, repeatable, does not require a feeder layer, and most importantly, it yields a high frequency of transformants. It will be important to evaluate this protocol with other genotypes of tomato.
References
Abel S, Ticconi CA, Delatorre CA (2002) Phosphate sensing in higher plants. Physiol Plant 115:1–8. doi:10.1034/j.1399-3054.2002.1150101.x
Bhatnagar-Mathur P, Vadez V, Sharma KK (2008) Transgenic approaches for abiotic stress tolerance in plants: retrospect and prospects. Plant Cell Rep 27:411–424. doi:10.1007/s00299-007-0474-9
Chyi YS, Phillips GC (1987) High efficiency Agrobacterium-mediated transformation of Lycopersicon based on conditions favorable for regeneration. Plant Cell Rep 6:105–108. doi:10.1007/BF00276664
Cortina C, Culiáňez-Macià FA (2004) Tomato transformation and transgenic plant production. Plant Cell Tissue Organ Cult 76:269–275. doi:10.1023/B:TICU.0000009249.14051.77
Dan Y, Yan H, Munyikwa T, Dong J, Zhang Y, Armstrong CL (2006) MicroTom—a high-throughput model transformation system for functional genomics. Plant Cell Rep 25:432–441. doi:10.1007/s00299-005-0084-3
Dillen W, DeClercq J, Kapila J, Zambre M, VanMontagu M, Angenon G (1997) The effect of temperature on Agrobacterium tumefaciens-mediated gene transfer to plants. Plant J 12:1459–1463. doi:10.1046/j.1365-313x.1997.12061459.x
Ellul P, Garcia-Sogo B, Pineda B, Ríos G, Roig LA, Moreno V (2003) The ploidy level of transgenic plants in Agrobacterium-mediated transformation of tomato cotyledons (Lycopersicon esculentum L.) is genotype and procedure dependent. Theor Appl Genet 106:231–238. doi:10.1007/s00122-002-0928-y
Frary A, Earle ED (1996) An examination of factors affecting the efficiency of Agrobacterium-mediated transformation of tomato. Plant Cell Rep 16:235–240. doi:10.1007/BF01890875
Gelvin SB (2003) Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev 67:16–37. doi:10.1128/MMBR.67.1.16-37.2003
Hamza S, Chupeau Y (1993) Re-evaluation of conditions for plant regeneration and Agrobacterium-mediated transformation from tomato (Lycopersicon esculentum). J Exp Bot 44:1837–1845. doi:10.1093/jxb/44.12.1837
Hansen G, Das A, Chilton MD (1994) Constitutive expression of the virulence genes improves the efficiency of plant transformation by Agrobacterium. Proc Natl Acad Sci USA 91:7603–7607
Horsch RB, Fry JE, Hoffmann NL, Eichholtz D, Rogers SG, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231. doi:10.1126/science.227.4691.1229
Hwang HH, Gelvin SB (2004) Plant proteins that interact with VirB2, the Agrobacterium tumefaciens pilin protein, mediate plant transformation. Plant Cell 16:3148–3167. doi:10.1105/tpc.104.026476
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Joubert P, Sangwan RS, EI Arabi Aouad M, Beaupère D, Sangwan-Norreel BS (1995) Influence of phenolic compounds on Agrobacterium vir gene induction and onion gene transfer. Phytochemistry 40:1623–1628. doi:10.1016/0031-9422(95)00539-J
Krasnyanski SF, Sandhu J, Domier LL, Buetow DE, Korban SS (2001) Effect of an enhanced CaMV 35S promoter and a fruit-specific promoter on uida gene expression in transgenic tomato plants. In Vitro Cell Dev Biol Plant 37:427–433. doi:10.1007/s11627-001-0075-1
Ling HQ, Kriseleit D, Ganal MW (1998) Effect of ticarcillin/potassium clavulanate on callus growth and shoot regeneration in Agrobacterium-mediated transformation of tomato (Lycopersicon esculentum Mill.). Plant Cell Rep 17:843–847. doi:10.1007/s002990050495
Lipp João KH, Brown TA (1993) Enhanced transformation of tomato co-cultivated with Agrobacterium tumefaciens C58C1Rifr::pGSFRll61 in the presence of acetosyringone. Plant Cell Rep 12:422–425. doi:10.1007/BF00234705
McCormick S, Niedermeyer J, Fry J, Barnason A, Horsch R, Fraley R (1986) Leaf disc transformation of cultivated tomato (L. esculentum) using Agrobacterium tumefaciens. Plant Cell Rep 5:81–84. doi:10.1007/BF00269239
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Ogaki M, Furuichi Y, Kuroda K, Chin DP, Ogawa Y, Mii M (2008) Importance of co-cultivation medium pH for successful Agrobacterium-mediated transformation of Lilium × formolongi. Plant Cell Rep 27:699–705. doi:10.1007/s00299-007-0481-x
Park SH, Morris JL, Park JE, Hirschi KD, Smith RH (2003) Efficient and genotype-independent Agrobacterium-mediated tomato transformation. J Plant Physiol 160:1253–1257. doi:10.1078/0176-1617-01103
Park S, Li J, Pittman JK, Berkowitz GA, Yang H, Undurraga S, Morris J, Hirschi KD, Gaxiola RA (2005) Up-regulation of a H+-pyrophosphatase (H+-PPase) as a strategy to engineer drought-resistant crop plants. Proc Natl Acad Sci USA 102:18830–18835. doi:10.1073/pnas.0509512102
Pozueta-Romero J, Houlné G, Cañas L, Schantz R, Chamarro J (2001) Enhanced regeneration of tomato and pepper seedling explants for Agrobacterium-mediated transformation. Plant Cell Tissue Organ Cult 67:173–180. doi:10.1023/A:1011997926381
Qiu D, Diretto G, Tavarza R, Giuliano G (2007) Improved protocol for Agrobacterium mediated transformation of tomato and production of transgenic plants containing carotenoid biosynthetic gene CsZCD. Sci Hortic 112:172–175. doi:10.1016/j.scienta.2006.12.015
Raghothama KG (2000) Phosphate transport and signaling. Curr Opin Plant Biol 3:182–187. doi:10.1016/S1369-5266(00)80063-1
Raj SK, Singh R, Pandey SK, Singh BP (2005) Agrobacterium-mediated tomato transformation and regeneration of transgenic lines expressing tomato leaf curl virus coat protein gene for resistance against TLCV infection. Curr Sci 88:1674–1679
Sun HJ, Uchii S, Watanabe S, Ezura H (2006) A highly efficient transformation protocol for Micro-Tom, a model cultivar for tomato functional genomics. Plant Cell Physiol 47:426–431. doi:10.1093/pcp/pci251
Tabaeizadeh Z, Agharbaoui Z, Harrak H, Poysa V (1999) Transgenic tomato plants expressing a Lycopersicon chilense chitinase gene demonstrate improved resistance to Verticillium dahliae race 2. Plant Cell Rep 19:197–202. doi:10.1007/s002990050733
Uranbey S, Sevimay CS, Kaya MD, İpek A, Sancak C, Başalma D, Er C, Őzcan S (2005) Influence of different co-cultivation temperatures, periods and media on Agrobacterium tumefaciens-mediated gene transfer. Biol Plant 49:53–57. doi:10.1007/s10535-005-3057-z
van Roekel JSC, Damm B, Melchers LS, Hoekema A (1993) Factors influencing transformation frequency of tomato (Lycopersicon esculentum). Plant Cell Rep 12:644–647. doi:10.1007/BF00232816
Vidya CSS, Manoharan M, Kumar CTR, Savithri HS, Sita GL (2000) Agrobacterium-mediated transformation of tomato (Lycopersicon esculentum var. Pusa Ruby) with coat-protein gene of Physalis mottle tymovirus. J Plant Physiol 156:106–110
Wroblewski T, Tomczak A, Michelmore R (2005) Optimization of Agrobacterium-mediated transient assays of gene expression in lettuce, tomato and Arabidopsis. Plant Biotechnol J 3:259–273. doi:10.1111/j.1467-7652.2005.00123.x
Wu Y, Chen Y, Liang X, Wang X (2006) An experimental assessment of the factors influencing Agrobacterium-mediated transformation in tomato. Russ J Plant Physiol 53:252–256. doi:10.1134/S1021443706020166
Yuan ZC, Edlind MP, Liu P, Saenkham P, Banta LM, Wise AA, Ronzone E, Binns AN, Kerr K, Nester EW (2007) The plant signal salicylic acid shuts down expression of the vir regulon and activates quormone-quenching gene in Agrobacterium. Proc Natl Acad Sci USA 104:11790–11795. doi:10.1073/pnas.0704866104
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273. doi:10.1146/annurev.arplant.53.091401.143329
Acknowledgments
The authors thank Dr. Sumei Li from the Institute of Soil Science, Chinese Academy of Sciences, for valuable suggestions on transgenic techniques. The authors also thank the anonymous reviewers for their valuable comments and suggestions that had improved this manuscript greatly. This work was supported by the National Natural Science Foundation of China (40671100) and the National Basic Research Program of China (2007CB109303).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gao, N., Shen, W., Cao, Y. et al. Influence of bacterial density during preculture on Agrobacterium-mediated transformation of tomato. Plant Cell Tiss Organ Cult 98, 321–330 (2009). https://doi.org/10.1007/s11240-009-9566-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-009-9566-2