Abstract
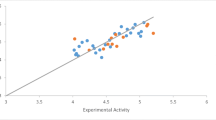
In this study, we investigated the structure-activity relationships of a series of β-carboline alkaloid derivatives using the 2D-QSAR and molecular docking, in order to identify the mode of interaction between β-carboline derivatives and the PLK1 kinase, and determine their key substituents responsible for the cytotoxic activity. The obtained QSAR models using multiple linear regression (MLR) and partial least squares (PLS) methods showed a high correlation between the experimental activity and the predicted one by PLS (R2PLS = 0.82, q2 = 0.72) and MLR (R2MLR = 0.82, q2 = 0.72). An external dataset was used to test the extrapolation power of the models which resulted in an R2PLS (EV) = 0.76; RMSE = 0.39. The 2D-QSAR analysis reveals that lipophilicity plays an important role in the cytotoxic activity of this group of β-carboline derivatives. Indeed, the molecular docking study into the active site of the polo-like kinase (PLK1) revealed that the most active ligand 57 shows higher binding energy and interacts, especially by H-bonds and hydrophobic interactions, with the active site of the PLK1 kinase. Consequently, the results obtained from the 2D-QSAR and docking studies provided a useful tool to design new and potent β-carboline derivatives as cytotoxic agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alkaloids constitute the largest group of plant secondary metabolites containing nitrogen and exhibit extensive pharmacological actions, such as cytotoxic and antitumor activities [1, 2]. Furthermore, β-carbolines are a group of natural alkaloids that possess a common tricyclic skeleton [3, 4]. These compounds are produced and stored by plants as products of different biosynthesis pathways from amino acids such as lysine, ornithine, tyrosine, and tryptophan [5]. They are also encountered in some animals such as insects and mammalians as well as human tissues and body fluids [6]. In recent decades, these compounds have attracted a great interest due to their pharmacological properties such as anxiolytic, hypnotic, anticonvulsant, sedative, antimicrobial, antiviral, parasitical, antidiabetic, and anti-inflammatory, as well as their potent antitumor activities [7,8,9,10,11,12,13,14]. Indeed, in vitro studies have demonstrated the decrease in cell viability of cancer cells from various tissues [15,16,17,18,19,20,21,22,23]. This anticancer activity is also confirmed in vivo since β-carbolines inhibit the tumor growth of various murine models [13, 15, 23,24,25]. In addition, the study by Zhiyong Chen and al., which examined the synthesis of a series of β-carboline derivatives, the evaluation of their antitumor activities, and the analysis of structure-activity relationships, showed that these compounds have potent antitumor activities, and that the antitumor potential is correlated to both the planarity of the molecule and the presence of the substituents on the central ring [26].
This activity is explained by the action of β-carbolines on different characteristics acquired by cancer cells, through multiple mechanisms of action, namely:
-
Inhibition of cyclin-dependent kinases (CDK), inducing a cell cycle arrest of different cancer cell lines. Thus, a state of senescence in breast cancer cells (MCF-7) by inhibiting the expression of telomerase [21].
-
Increased expression of tumor suppressor factor (p53), inducing apoptosis, cell cycle arrest, and inhibition of angiogenesis [18, 21, 24].
-
Inhibition of the regulated phosphorylated tyrosine kinase double-specified protein (DYRK1A) by degradation of the amplified and/or mutated epidermal growth factor receptor (EGFR) in glioblastomas, which will inhibit the growth of Tumor cells [27,28,29].
-
Induction of cell death by decreasing expression of several anti-apoptotic proteins (Bcl-xl, Bcl-2, and Mcl-1) and increasing expression of pro-apoptotic proteins (Bid, Bax) [13, 18, 20, 22, 23]. Thus, it has been shown that β-carbolines inhibit topoisomerase 1, which causes transcriptional changes, inhibition of replication, and DNA damage-inducing cell death [29, 30].
-
Inhibition of invasion, angiogenesis, and formation of metastases by the action of β-carbolines on the expression of various pro- and anti-metastatic and angiogenic factors [14, 22, 24].
Furthermore, it has been found that this class of molecule exhibits remarkable acute neurotoxicity in mice characterized by tremors, agitation, and convulsion movements [13, 31, 32].
As for targeted receptor of this chemical series, it has been shown that β-carbolines are potent inhibitors of polo-like kinases (PLK) which plays an essential role in the ordered execution of mitotic events [33]. PLK1 kinase, a member of the PLK family, is an attractive target for anticancer drugs.
In continuing of our work to develop novel more effective cytotoxic, antitumor, and less toxic agents from the β-carboline derivatives using experimental and theoretical studies [34,35,36,37], we studied a series of β-carboline derivatives exhibiting cytotoxic activity against the hepatocellular line (HepG2) using 2D-QSAR and molecular docking analysis. The aim of this study was to identify the mode of interaction between β-carboline derivatives and the PLK1kinase and determine their key substituents responsible for the cytotoxic activity, in order to guide the design of new β-carboline alkaloids with improved pharmacological activities and less neurotoxicity.
Materials and methods
Data collection
In this study, all β-carboline derivatives (40 molecules) have been taken from the work of Cao et al. [32]. The reported IC50 values for cytotoxic activity against the hepatocellular line (HepG2) were converted into the corresponding pIC50 values (pIC50 = -logIC50) and selected for this study (Table 1). Dataset was split into two sets; 34 molecules were chosen randomly to develop the QSAR model (training set) and the rest (7 molecules) were used to test the prediction performance of the proposed model (test set).
Molecular modeling
2D QSAR modeling
All compounds were sketched using the MarvinSketch program (version 16.2.1.0) [38], and various 2D descriptors were calculated using the MOE software (Molecular Operating Environment version 2008.10) [39]. After the calculation of descriptors provided by MOE software, a correlation matrix for variable selection was applied on the molecular descriptors to select only the appropriate ones. Therefore, the number was reduced to six descriptors which are used as input to perform multiple linear regression (MLR) and partial least squares (PLS) methods using MOE and XLSTAT 2014 software package [40], respectively. Subsequently, the quality of the models developed was examined by different statistical parameters [41], for instance, the square of correlation coefficient (R2), adjusted coefficient of determination (R2adj), root mean square error (RMSE), and variance ratio (F) at specified degrees of freedom (df). In addition, the models were validated using internal cross-validation (q2) and external validation (R2pred). [42,43,44]
Docking study
To explore the interaction and illustrate the accurate binding model for the active site of the PLK1 with ligands, a molecular docking study was performed using AutodockVina and Autodock tools 1.5.4 [45]. The X-ray crystal structure of the PLK1 (PDB code 2OWB) was used for the docking study. All small molecules were removed from the protein, and the receptor was used for the docking study by adding the polar hydrogen to assign appropriate ionization states to both acidic and basic amino acid residues. For ligands, they were sketched using the MarvinSketch program and minimized by MOE software, using MMFF94 (Merck Molecular Force Field) force field with the gradient convergence criterion set to 0.01 kcal/mol, and saved in pdb format. Then, four compounds (57, 58, 59, and 60) with better activity were docked within the prepared protein. The mode of interaction of the co-crystalized ligand against 2OWB was used a standard docked model.
AutoGrid was carried out for the preparation of the grid map using a grid box to enclose the binding site with dimensions of x = 20, y = 20, and z = 20 at 1 Å spacing. The Lamarckian genetic algorithm was used for the calculation of the docking possibilities. Then, the results were analyzed using Discovery studio 2016 and PyMol software’s [46, 47].
Results and discussion
2D QSAR modeling
In this work, we used principal component analysis by XLSTAT 2014 software to select six descriptors that show a high correlation with the response activity and no correlation between them owing to the fact that the greatest value of the correlation coefficient is 0.75; this one gives extra weight because they will be more effective at prediction [48]. Figure 1 presents the circles of correlations corresponding to a projection of the six variables on a two-dimensional plane constituted by the two factors (F1 and F2). Besides, the principal component analysis also gives a representation of the molecules of the database in a plan composed by the first two principal components (F1 and F2), which can show the presence of three subgroups of molecules. As shown in Fig. 2, molecules without hydrogen bond donor or acceptor are identified at the bottom left of the graph. At the top of the graph center, we can distinguish the group of molecules with one or two hydrogen bond acceptors and no hydrogen bond donor. Finally, molecules that have more than two hydrogen bond acceptors and one hydrogen bond donor are found at the bottom right of the graph.
From this first analysis, we can conclude that the number of hydrogen bond donor and acceptor is the main features for discriminating between the molecules within the overall database.
Thereafter, the QSAR model was performed using PLS and RLM methods correlating the cytotoxic activity with six descriptors. The generated models were thoroughly scrutinized for statistical validity and predictive potential, according to the criteria described in the experimental section.
The best QSAR model derived from modeling the cytotoxic activity of the 34 heterocyclic β-carboline derivatives is as follows.
-
RLM model:
-
PLS model:
where N is the number of training set, R2 is the squared correlation coefficient, R2adj is the adjusted coefficient of determination, RMSE is the root mean square error, and F represents the Fisher ratio between the variances of calculated and observed activities.
QSAR models were established using two statistical analysis methods (RLM and PLS), using experimental data (pIC50) for a series of β-carboline derivatives and a set of six descriptors. The model selection criteria were based on the correlation coefficient values derived from the correlation between the experimental and predicted activities, as well as the inter-correlations between the descriptors.
We found that for both methods used (PLS, RLM), the QSAR model obtained can predict about 82% of the experimental activity (pIC50) of the molecules. In addition, it has a high Fischer factor (F = 20.848) and a low error (RMSERLM = 0.254, RMSEPLS = 0.227), which means that the model explains the activity (dependent variable) in a statistically significant and satisfactory manner. The value of the test (t) is used to evaluate the importance of the descriptors involved in the model, which is in the following order:
A comparison of the quality of two models from MLR and PLS models (Table 2) shows that the PLS model outperforms slightly the MLR one when judged by the obtained RMSE values [49]. As PLS is a more robust multivariate statistical technique, we choose to use the PLS model as an in silico tool to predict the activity of β-carboline derivatives and to draw a SAR map for further design and chemical synthesis studies.
Taking into consideration the selected descriptors and their impacts on the PLS model, we were able to propose a structure-activity relationship map that highlights the structural motifs responsible for the cytotoxic activity of the β-carboline derivatives (Fig. 3):
Structure-activity relationship (SAR) analysis shows that the activity of the β-carboline derivatives can be influenced by:
-
The planarity of the molecule that was captured in part by the radius descriptor in agreement with the published work by Zhiyong Chen and al [26];
-
The electropositive character, captured by the descriptor vsa_base, of the nitrogen atom in position 2; and
-
The presence of various substituents on the structure of the β-carbolines essentially represented by the SMR_VSA3 descriptor.
The QSAR model revealed that the activity of β-carboline derivatives was represented by some of selected descriptors:
-
The number of hydrogen bond acceptor (a_acc) has a negative coefficient in the model equation, suggesting that increased activity can be achieved by decreasing the number of heteroatoms (nitrogen or oxygen atoms);
-
The number of hydrogen bond donor (a_don) has a positive coefficient in the model equation, suggesting that an increase in activity can be achieved by increasing heteroatoms with one or more hydrogen atoms;
-
The relative partial positive charge (PEOE_RPC+) obtained using the partial equalization of orbital electronegativities (PEOE) method is defined as the ratio of the largest positive partial charge to the total positive partial charge on the molecule [50]. This descriptor has a negative coefficient in the model equation, suggesting that increased activity can be obtained by decreasing the relative partial positive charge.
-
The basic character (vsa_base) of the molecules has a negative coefficient in the model equation, suggesting that increased activity can be obtained by increasing the polarity of the β-carboline derivatives;
-
Radius (radius) has a negative contribution to activity, which means that activity is improved by decreasing the radius. Thus, molecules that have an ionic bond between nitrogen and bromine (no path between the two atoms) have the smallest radius (radius = 0), suggesting that the positive charge of pyridinium is favorable to cytotoxic activity;
-
Molar refractivity based on approximate Van Der Waals surface area calculations (SMR_VSA3) is usually used to reflect the polarizability of molecule [51]. The positive sign of this descriptor suggests that the polarizability of the molecule is favorable for the cytotoxic activity.
Molecular docking
Docking study was carried out on the four most active compounds (57, 58, 59, 60) to identify the nature of the interactions between ligand and biological target taking into account the orientation and conformation of the ligand in the active site of the protein. In this study, docking protocol was validated by redocking of the co-crystallized ligand at the PLK1kinase (Fig. 4). The RMSD (root mean square distance) of the docked ligand was within the reliable range of 2 Å, suggesting that the docking procedure could be used to predict the binding mode of our compounds.
The interaction analysis of the ligand-protein indicates that important residues present at the binding site were polar (Lys66, Asp194, Arg57, Arg134, Lys61, Lys82, Arg136, Cys67, Glu131, Cys133) and non-polar (Leu132, Val114, Leu130, Gly62, Ala65, Leu59, Phe58, Phe183, Ala80). As shown in Fig. 4, we can see that the co-crystallized compound sits in the hydrophobic cavity and interacts by several types of interactions and more precisely by the conventional hydrogen bond with Glu131 and Cys133.
The ligands studied show a strong binding to the active sites of the target. As reported in Table 3, the biological activity values correlate with the estimated binding energy and the latter decrease with the increase of biological activity.
The analysis of interactions between the compound 57 and the binding site (Fig. 5) reveals that is surrounded by the hydrophobic region formed by Arg135, Arg134, Arg57, Glu69, Cys133, Gly193, and His105. Also, compound 57 exhibits a hydrogen binding interactions between three fluorines and oxygen of the pentafluorobenzyl moiety with Cys67 and Asp194 of PLK1. In another hand, compound 57 interacts with PLK1 by other binds such as Pi-sigma, Alkyl, Pi-Alkyl, and Pi-lone Pair with Leu130, Lys82, Val114, Leu132, Arg136 Ala80, and Phe183.
The results obtained during this work correlate well with each other and are in good agreement with those of the 3D-QSAR obtained by Cao et al. [32]. In fact, the electropositive parameter captured by 3D-QSAR in position 2 can be represented by the descriptor vsa_base obtained by the 2D-QSAR model; then, the presence of the electron-rich groups in positions 2 and 7 such as benzyl and pentafluorobenzyl, respectively, is favorable for cytotoxic activity since they make it possible to form pi-alkyl and alkyl bonds with PLK1. In addition, docking results show that the pentafluorobenzyl moiety plays a crucial role in improving binding energy by forming hydrogen bonds with PLK1, which can be captured by the selection of hydrogen bond donor and polarizability (a_don, SMR_VSA3) descriptors in 2D-QSAR which have a positive sign in the model. On the other hand, electrostatic repulsive interaction in position 3 due to the electronegative groups demonstrated by 3D-QSAR is unfavorable for cytotoxic activity; this may be explained by the negative sign obtained by 2D-QSAR for PEOE_RPC+ descriptor.
According to the above findings, three compounds were designed based on the structure of the compounds with the highest pIC50 values (57). The structures of the compounds designed, the pIC50 values theoretically predicted by the 2D QSAR, and their binding energy are listed in Table 4. According to the results of the predicted activities, it was observed that the compounds designed had higher predicted pIC50 values than the compounds studied in this work. Furthermore, newly designed compounds were docked at the PLK1 kinase. All compounds have better binding energy than the compounds studied in this work since they were well placed in the active site and indicate that they could be potential inhibitors of PLK1 kinase. The interaction analysis between the compound 3, which showed the highest activity and the low binding energy, and the PLK1 kinase reveals that the introduction of the methyl group in the meta-benzyl positions at the position 7 and the substitution of the ethyl group in position 9 with the amino group allow forming hydrophobic binding and establishing three hydrogen bonds (two bonds between fluorine and Lys82 and Asp194 and one bond between the amino group and Arg136) as well as other bonds: alkyl, Pi-Alkyl, Pi-sigma, and Pi-Pi Stacked with Val114, Cys67, Ala80, Leu130, His105, Leu59, and Phe183, respectively, as shown in Fig. 6.
Conclusions
In this paper, a quantitative analysis of structure-activity relationships and molecular docking studies was performed on a series of 40 β-carboline derivatives. The 2D-QSAR was performed using partial least squares regression (PLS) and multiple linear regression (RLM). The results found showed that the model proposed via the PLS method is able to accurately predict the cytotoxic activity and that the selected descriptors are relevant to explain the increase or decrease in cytotoxic activity against the hepatocellular (HepG2) line. In addition, the docking study showed that compound 57 has low binding energy and seems to have more binding affinities towards PLK1. Analysis of the structural interactions shows that this compound is in a hydrophobic pocket and interacts with the active site, especially through hydrogen bonds. Accordingly, obtained results were used to design three newly compounds with predicted low binding affinity and improved cytotoxic activity. Overall, these results can be used to perform virtual screening for new β-carboline and can also help to design new compounds to obtain potent and novel β-carboline with improved biological activities.
References
Mohan K, Jeyachandran R, Deepa (2012) Alkaloids as anticancer agents. Ann Phytomed 1:46–53
Byler KG, Wang C, Setzer WN (2009) Quinoline alkaloids as intercalative topoisomerase inhibitors. J Mol Model 15:1417–1426. https://doi.org/10.1007/s00894-009-0501-6
Abramovitch RA, Spenser ID (2008) The carbolines. Adv Heterocycl Chem 3:79–207. https://doi.org/10.1016/S0065-2725(08)60542-5
Gilbert J, Senyuva HZ (2008) Bioactive compounds in foods. Blackwell Pub, Oxford
Aniszewski T (2015) Alkaloids: chemistry, biology, ecology, and applications2nd edn. Elsevier, Waltham, MA
Zheng W, Wang S, Barnes LF et al (2000) Determination of harmane and harmine in human blood using reversed-phased high-performance liquid chromatography and fluorescence detection. Anal Biochem 279:125–129. https://doi.org/10.1006/abio.1999.4456
Braestrup C, Nielsen M, Olsen CE (1980) Urinary and brain beta-carboline-3-carboxylates as potent inhibitors of brain benzodiazepine receptors. Proc Natl AcadSci 77:2288–2292
Schupp P, Poehner T, Edrada R et al (2003) Eudistomins W and X, two new β-carbolines from the micronesian tunicate Eudistoma sp. J Nat Prod 66:272–275. https://doi.org/10.1021/np020315n
Nazari Formagio AS, Santos PR, Zanoli K et al (2009) Synthesis and antiviral activity of β-carboline derivatives bearing a substituted carbohydrazide at C-3 against poliovirus and herpes simplex virus (HSV-1). Eur J Med Chem 44:4695–4701. https://doi.org/10.1016/j.ejmech.2009.07.005
Di Giorgio C, Delmas F, Ollivier E et al (2004) In vitro activity of the β-carboline alkaloids harmane, harmine, and harmaline toward parasites of the species Leishmaniainfantum. ExpParasitol 106:67–74. https://doi.org/10.1016/j.exppara.2004.04.002
Waki H, Park KW, Mitro N et al (2007) The small molecule harmine is an antidiabetic cell-type-specific regulator of PPARγ expression. Cell Metab 5:357–370. https://doi.org/10.1016/j.cmet.2007.03.010
Yamazaki Y, Kawano Y (2011) Inhibitory effects of herbal alkaloids on the tumor necrosis factor-α and nitric oxide production in lipopolysaccharide-stimulated RAW264 macrophages. Chem Pharm Bull (Tokyo) 59:388–391
Chen Q, Chao R, Chen H et al (2005) Antitumor and neurotoxic effects of novel harmine derivatives and structure-activity relationship analysis. Int J Cancer 114:675–682. https://doi.org/10.1002/ijc.20703
Cao R, Peng W, Chen H et al (2005) DNA binding properties of 9-substituted harmine derivatives. BiochemBiophys Res Commun 338:1557–1563. https://doi.org/10.1016/j.bbrc.2005.10.121
Cao R, Chen Q, Hou X et al (2004) Synthesis, acute toxicities, and antitumor effects of novel 9-substituted β-carboline derivatives. Bioorg Med Chem 12:4613–4623. https://doi.org/10.1016/j.bmc.2004.06.038
Pérez Martín JM, Labrador V, Fernández Freire P et al (2004) Ultrastructural changes induced in HeLa cells after phototoxic treatment with harmine. J ApplToxicol 24:197–201. https://doi.org/10.1002/jat.972
Song Y, Kesuma D, Wang J et al (2004) Specific inhibition of cyclin-dependent kinases and cell proliferation by harmine. BiochemBiophys Res Commun 317:128–132. https://doi.org/10.1016/j.bbrc.2004.03.019
Hamsa TP, Kuttan G (2011) Harmine activates intrinsic and extrinsic pathways of apoptosis in B16F-10 melanoma. Chin Med 6:11
Frédérick R, Bruyère C, Vancraeynest C et al (2012) Novel trisubstituted harmine derivatives with original in vitro anticancer activity. J Med Chem 55:6489–6501. https://doi.org/10.1021/jm300542e
Liu H, Han D, Liu Y et al (2013) Harmine hydrochloride inhibits Akt phosphorylation and depletes the pool of cancer stem-like cells of glioblastoma. J Neuro-Oncol 112:39–48. https://doi.org/10.1007/s11060-012-1034-x
Zhao L, Wink M (2013) Theβ-carboline alkaloid harmine inhibits telomerase activity of MCF-7 cells by down-regulating hTERT mRNA expression accompanied by an accelerated senescent phenotype. PeerJ 1:e174. https://doi.org/10.7717/peerj.174
Zhang H, Sun K, Ding J et al (2014) Harmine induces apoptosis and inhibits tumor cell proliferation, migration and invasion through down-regulation of cyclooxygenase-2 expression in gastric cancer. Phytomedicine 21:348–355. https://doi.org/10.1016/j.phymed.2013.09.007
Cao M-R, Li Q, Liu Z-L et al (2011) Harmine induces apoptosis in HepG2 cells via mitochondrial signaling pathway. Hepatobiliary Pancreat Dis Int 10:599–604. https://doi.org/10.1016/S1499-3872(11)60102-1
Dai F, Chen Y, Song Y et al (2012) A natural small molecule harmine inhibits angiogenesis and suppresses tumour growth through activation of p53 in endothelial cells. PLoS One 7:e52162. https://doi.org/10.1371/journal.pone.0052162
Hamsa TP, Kuttan G (2011) Studies on anti-metastatic and anti-invasive effects of harmine using highly metastatic murine B16F-10 melanoma cells. J Environ Pathol Toxicol Oncol 30:123–137. https://doi.org/10.1615/JEnvironPatholToxicolOncol.v30.i2.40
Chen Z, Cao R, Shi B et al (2011) Synthesis and biological evaluation of 1,9-disubstituted β-carbolines as potent DNA intercalating and cytotoxic agents. Eur J Med Chem 46:5127–5137. https://doi.org/10.1016/j.ejmech.2011.08.027
Bain J, Plater L, Elliott M et al (2007) The selectivity of protein kinase inhibitors: a further update. Biochem J 408:297–315. https://doi.org/10.1042/BJ20070797
Göckler N, Jofre G, Papadopoulos C et al (2009) Harmine specifically inhibits protein kinase DYRK1A and interferes with neurite formation: harmine, a specific inhibitor of DYRK1A. FEBS J 276:6324–6337. https://doi.org/10.1111/j.1742-4658.2009.07346.x
Adayev T, Wegiel J, Hwang Y-W (2011) Harmine is an ATP-competitive inhibitor for dual-specificity tyrosine phosphorylation-regulated kinase 1A (Dyrk1A). Arch BiochemBiophys 507:212–218. https://doi.org/10.1016/j.abb.2010.12.024
Sobhani AM, Ebrahimi S-A, Mahmoudian M (2002) An in vitro evaluation of human DNA topoisomerase I inhibition by Peganumharmala L. seeds extract and its beta-carboline alkaloids. J Pharm PharmSci 5:19–23
Cao R, Chen H, Peng W et al (2005) Design, synthesis and in vitro and in vivo antitumor activities of novel β-carboline derivatives. Eur J Med Chem 40:991–1001. https://doi.org/10.1016/j.ejmech.2005.04.008
Cao R, Guan X, Shi B et al (2010) Design, synthesis and 3D-QSAR of β-carboline derivatives as potent antitumor agents. Eur J Med Chem 45:2503–2515. https://doi.org/10.1016/j.ejmech.2010.02.036
Han X, Zhang J, Guo L et al (2012) A series of beta-carboline derivatives inhibit the kinase activity of PLKs. PLoS One 7:e46546. https://doi.org/10.1371/journal.pone.0046546
Lamchouri F, Settaf A, Cherrah Y et al (2000) In vitro cell-toxicity of Peganumharmala alkaloids on cancerous cell-lines. Fitoterapia 71:50–54
Lamchouri F, Toufik H, Elmalki Z et al (2013) Quantitative structure–activity relationship of antitumor and neurotoxic β-carbolines alkaloids: nine harmine derivatives. Res ChemIntermed 39:2219–2236. https://doi.org/10.1007/s11164-012-0752-1
Lamchouri F, Zemzami M, Jossang A et al (2013) Cytotoxicity of alkaloids isolated from Peganumharmala seeds. Pak J Pharm Sci 26:699–706
Lamchouri F, Toufik H, Bouzzine SM et al (2010) Experimental and computational study of biological activities of alkaloids isolated from Peganum harmala seeds. J Mater EnvSci 1:343–352
ChemAxon, MarvinSketch 2.1.0 (2016). http://www.chemaxon.com. Accessed 01 Dec 2017
Molecular Operating Environment software, Chemical Computing Group Inc., Montreal, Quebec, Canada. http://www.chemcomp.com/
Addinsoft (2014) XLSTAT: data analysis and statistical solution for microsoft excel. Paris, France. http://www.xlstat.com. Accessed 25 Feb 2017
Tropsha A (2010) Best practices for QSAR model development, validation, and exploitation. Mol Inform 29:476–488. https://doi.org/10.1002/minf.201000061
Dehmer M, Varmuza K, Bonchev D (2012) Statistical modelling of molecular descriptors in QSAR/QSPR: DEHMER:MOL. DESCRIPTOR O-BK. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim
Roy K (2007) On some aspects of validation of predictive quantitative structure–activity relationship models. Expert Opin Drug Discov 2:1567–1577. https://doi.org/10.1517/17460441.2.12.1567
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26:694–701. https://doi.org/10.1002/qsar.200610151
Trott O, Olson AJ (2010) AutoDockVina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J ComputChem 31:455–461. https://doi.org/10.1002/jcc.21334
(2016) Dassault Systèmes BIOVIA, Discovery Studio Modeling Environment, Release 2017, San Diego: Dassault Systèmes. http://accelrys.com/products/collaborative-science/bioviadiscoverystudio/. accessed 01 Feb 2018
DeLano WL (2008) The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC. http://www.pymol.org. Accessed 01 Feb 2018
Consonni V, Ballabio D, Todeschini R (2010) Evaluation of model predictive ability by external validation techniques. J Chemom 24:194–201. https://doi.org/10.1002/cem.1290
Aptula AO, Jeliazkova NG, Schultz TW, Cronin MTD (2005) The better predictive model: high q2 for the training set or low root mean square error of prediction for the test set? QSAR Comb Sci 24:385–396. https://doi.org/10.1002/qsar.200430909
Gasteiger J, Marsili M (1980) Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36:3219–3288. https://doi.org/10.1016/0040-4020(80)80168-2
Labute P (2000) A widely applicable set of descriptors. J Mol Graph Model 18:464–477. https://doi.org/10.1016/S1093-3263(00)00068-1
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interest.
Ethical approval
This chapter does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Akabli, T., Toufik, H., Yasri, A. et al. Combining ligand-based and structure-based drug design approaches to study the structure-activity relationships of a β-carboline derivative series. Struct Chem 29, 1637–1645 (2018). https://doi.org/10.1007/s11224-018-1141-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-018-1141-1