Abstract
Amorpha-4,11-diene synthase (ADS) is the first key enzyme of artemisinin biosynthetic pathway in Artemisia annua L. In this study, the promoter region of the ADS gene has been cloned and used to demonstrate the expression of GUS reporter gene in both glandular trichomes of A. annua and non-glandular trichomes of Arabidopsis thaliana following homologous and heterologous expression of ADS promoter–GUS fusion. Subsequently, 5′ sequential deletion analysis of the ADS promoter revealed that a short sequence, −350 upstream of the transcription start site, was sufficient for trichome-specific expression in A. thaliana and that the region from −350 to −300 contained essential elements for this observed specificity. However, frequencies of transgenic A. thaliana plants displaying trichome-specific expressions varied between different lines, and all the lines with deleted fragments of the ADS promoter showed lower frequencies than the line with full-length ADS promoter. Most lines with deleted ADS promoter–GUS fusions showed GUS expressions in the guard cells of stomata as well, which was not observed in A. thaliana plants transformed with the full-length ADS promoter. GUS activities varied among different transgenic lines as well, both in transiently transformed Nicotiana benthamiana and stably transformed A. thaliana, with promoter–deletion lines exhibiting higher GUS activities than the full-length ADS promoter line.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
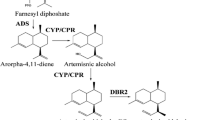
Artemisinin, a sesquiterpene lactone endoperoxide isolated from Artemisia annua L. (A. annua) of Asteraceae family, is a widely used anti-malarial drug. Malaria is a pandemic that threatens people in most tropical and subtropical regions of the world for decades. According to World Malaria Report 2010 (WHO 2011), there were an estimated 225 million cases of malaria worldwide in 2009, and among them 781,000 people died, accounting for 2.23 % of deaths across the world. Since resistance to traditional anti-malarial drugs has developed (Wellems 2002), such as chloroquine, and artemisinin as well as its derivatives are highly effective against the most severe and multidrug-resistant Plasmodium falciparum strains (Liu et al. 2006a), they now serve as primary components of the standard treatment worldwide for malaria. Nowadays, artemisinin is in great demand on international markets; however, A. annua plants, the main source of artemisinin, are of low content of this vital compound (0.1–0.8 %) (Abdin et al. 2003), resulting in the unaffordable cost of artemisinin-based treatment.
To enhance artemisinin yield, researchers have taken great efforts for the past two decades in its production through varied approaches, including total chemical synthesis (Xu et al. 1986; Avery et al. 1992), in vitro culture (Arsenault et al. 2008), microbial production (Ro et al. 2006; Zeng et al. 2008), and in vivo production in transgenic plants (Chen et al. 2000; Sa et al. 2001; Zhang et al. 2009; Westfall et al. 2012); however, to date, not a single sound industrial-scale production system has been established. To address such a problem, thorough insights into the mechanisms of artemisinin biosynthesis are required.
Artemisinin is specifically produced and stored in glandular secretory trichomes (Nguyen et al. 2011; Ferreira and Janick 1996), and quite a few studies indicate that trichomes are of great consequence to the production of artemisinin (Kapoor et al. 2007; Arsenault et al. 2010). Therefore, it is considerably attractive for researchers to find a way of enhancing artemisinin through regulation of trichomes or reactions occurring in them, such as employing trichome-specific promoters.
Amorpha-4,11-diene synthase (ADS) is the first key enzyme of artemisinin biosynthetic pathway in A. annua. It catalyzes farnesyl pyrophosphate (FPP) into amorpha-4,11-diene to provide the carbon skeleton for artemisinin (Nguyen et al. 2011). Kim et al. (2008) revealed that the promoter of the ADS gene specifically drove downstream gene expression in non-glandular trichomes of Arabidopsis thaliana. Later on, Wang et al. (2011) cloned a short version of ADS promoter (1,929 bp) and found that it was trichome-specific in A. annua L. From these studies, it seems that ADS promoter can be a promising tool for artemisinin phyto-genetic engineering. However, the expression of ADS gene in A. annua features considerable variations. It not only responds to diverse environmental signals (Nguyen et al. 2011), but also displays temporal expression profile, with remarkably decreased levels as A. annua plant matures (Olofsson et al. 2011). Therefore it is worthy of modifying the ADS promoter to a simplified one, which is stronger than the original one, much shorter in length to make it easily manipulated both in promoter reconstruction and genetic engineering, while retaining its trichome specificity as much as possible.
In this study, we cloned a 2935-bp DNA fragment upstream of the ADS coding sequence from A. annua L. We demonstrated that this ADS promoter is specific for gene expression in both glandular trichomes of A. annua and non-glandular trichomes of A. thaliana, and its truncated versions (no less than −350 upstream of the transcription start site) are capable of retaining trichome specificity in A. thaliana, while exhibiting stronger activities (around two to four times) than the full-length one. Therefore, compared to the full-length ADS promoter, the deleted ones are probably more promising for future genetic engineering use.
Materials and Methods
Extraction of Genomic DNA of A. annua
The genomic DNA was extracted from fresh young leaves of A. annua using CTAB method. Around 0.1 g fresh young leaves were collected and ground in liquid nitrogen; 700 μl preheated CTAB buffer containing both 2 % β-mercaptoethanol (v/v) and 1 % PVP (w/v) was then added to the ground leaves and then incubated in water bath at 65 °C for 1 h; 700 μl chloroform–isopentanol mixture (v/v=24:1) was then added to the CTAB–leaves mixture and shaken for 20 min (60 rev/min). The resulting mixture was then centrifuged at 10,000 rpm for 10 min, and after that the supernatant was separated with 600 μl chloroform–isopentanol mixture (v/v=24:1) added into it and shaken for 20 min (60 rev/min). The resulting mixture was then centrifuged at 10,000 rpm for 10 min, and then the supernatant was separated; 300 μl cold isopropanol was subsequently added to the supernatant, and the resulting mixture was then incubated at 4 °C for 1 h. After that, the mixture was centrifuged at 10,000 rpm for 10 min, and the resulting supernatant was discarded, with the deposit kept and washed with 70 % EtOH twice. The washed deposit was then dried and dissolved with 100 μl TE buffer, making the extracted A. annua genomic DNA solution.
Cloning of the Promoter Region of the ADS Gene in A. annua
According to the published ADS gene promoter sequence (GenBank No.: AY528931.1), a pair of specific primers was designed and the digestion sites of two restriction enzymes BamHI and NcoI (Fermentas FastDigest) were incorporated into these two primers as well. Using this pair of primers (pADS-BamHI-F and pADS-NcoI-R; see details in Table 1), high-fidelity PCR (KOD Plus, TOYOBO) was conducted as follows: 94 °C for 3 min; 30 cycles at 94 °C for 45 s, 55 °C for 45 s, 72 °C for 3 min; 72 °C for 10 min. The PCR products were then examined in gel electrophoresis and target fragments (~3,000 bp) were retrieved using JETQUICK Gel Extraction Spin Kit (GENOMED). The retrieved fragments were cloned into pMD18-T (Takara) and then transformed into competent cells of Escherichia coli strain DH5α through heat shock (42 °C for 90 s). Positive clones were picked out on LB/amp, and their plasmids were extracted using JETQUICK Plasmid Miniprep Spin Kit (GENOMED) for sequencing of the target DNA fragment in BGI.
Construction of the Expression Vector ADSpro::GUS and ADS Promoter Deletion Vectors
Both recombinant pMD18-T and another vector pCAMBIA1305.1 (Cambia) were double-digested with restriction enzymes BamHI and NcoI, and the target DNA fragment and the resulting linear vector pCAMBIA1305.1 were ligated with T4 DNA ligase (Thermo Scientific) (3 μl of the target DNA fragment, 1 μl of the linear vector, 1 μl of T4 DNA ligase buffer, 4.5 μl of ddH2O) under the following condition: 22 °C for 30 min. This recombinant expression vector was designated ADSpro::GUS.
To construct promoter deletion vectors, several primers were designed according to the ADS promoter sequence (see details in Table 1). With these specific primers, deleted fragments of the ADS promoter were isolated from the cloning vector ADSpro-MD18-T. All these fragments and pCAMBIA1391z (Cambia) were double-digested by two restriction enzymes, BamHI and NcoI, and the resulting digested ADS promoter fragments were constructed into pCAMBIA1391z, respectively, with T4 DNA ligase, making recombinant expression vectors designated as Δ-800, Δ-600, Δ-550, Δ-500, Δ-450, Δ-400, Δ-350, and Δ-300, respectively.
All of those expression vectors stated earlier as well as an intact pCAMBIA1305.1 vector (designated as 35Spro::GUS) were introduced into the competent cells of Agrobacterium tumefaciens strain EHA105 using freeze–thaw method (frozen in liquid nitrogen for 5 min and then thawed at 37 °C for 3 min). Positive clones were selected out on LB (with kanamycin, rifampicin, and streptomycin) and then further examined for target DNA fragments via PCR amplification. These transgenic A. tumefaciens were prepared for further plant transformation.
A. annua and A. thaliana Stable Transformation
Plant materials for transformations are A. annua wild type Youqing (collected in Youyang County, Chongqing City of China) and A. thaliana wild type Columbia, respectively.
Following the method of Zhang et al. (2009), A. annua was transformed with both of the two types of recombinant A. tumefaciens containing ADSpro::GUS or 35Spro::GUS fusions and selected with hygromycin resistance.
A. thaliana transformation was conducted following the floral dip protocol of Zhang et al. (2006) with several recombinant vectors, respectively: ADSpro::GUS, Δ-800, Δ-600, Δ-550, Δ-500, Δ-450, Δ-400, Δ-350, and Δ-300. Transgenic seeds were screened on MS (with hygromycin and cefalexin) medium, and the obtained seedlings were grown in soil under 16-h days and 8-h nights at 22 °C.
Transient Gene Expression in Nicotiana benthamiana
Transient expression in N. benthamiana was achieved through agroinfiltration as described (Kapila et al. 1997) with modifications. N. benthamiana of around 40 days after sowing were used for transformation. A. tumefaciens EHA105 stains transformed with ADSpro::GUS, 35Spro::GUS, Δ-800, Δ-600, Δ-550, Δ-500, Δ-450, Δ-400, Δ-350, or Δ-300 were prepared in LB (with kanamycin, rifampicin, and streptomycin). These bacteria liquid cultures were centrifuged, and the obtained deposit cells were re-suspended in transformation buffer (10 mM MES, 10 mM MgCl2, and 100 μM acetosyringone) with a final OD600 of around 1.0 and incubated for 3 h. The re-suspended cultures were then injected into the abaxial side of N. benthamiana leaves via a 1-ml syringe (needle removed). For each construct, at least three leaves were infiltrated to make sure of three replicates. For the purpose of quantitative expression comparisons between the full-length ADS promoter and other constructs, each leaf was treated with the left half injected with ADSpro::GUS A. tumefaciens stain while the right half was injected with stain of another construct. Infiltrated N. benthamiana plants were grown under 16-h day/8-h night at room temperature for 2–3 days before GUS assay.
Histochemical GUS Staining
After around 20 days following regeneration, two to three fresh young leaves of non-transgenic A. annua plants and transgenic A. annua plants with ADSpro::GUS and with 35Spro::GUS were sampled respectively for histochemical GUS staining using the method of Jefferson (1987).
For A. thaliana transgenic lines, lotus leaves of each regenerated plant of around 2–3 weeks old were sampled for GUS staining. The specificity percentage was thus calculated based on the staining results.
For A. annua and A. thaliana leaves bearing good GUS staining effects, optical microscope was employed to picture their staining patterns.
Quantitative Analysis of GUS Gene Expression
The fluorogenic assay was performed as described (Jefferson 1987).
For each agroinfiltrated N. benthamiana leaf, the left half and the right half were sampled separately in that they were transformed with different constructs (as stated earlier). The GUS activity of the right half of the leaf was then normalized to that of the left half so that results would be in relative forms (relative activity of a certain construct to that of ADSpro::GUS) and simultaneously the error between different leaves could be minimized.
For each Arabidopsis transgenic line (ADSpro::GUS, Δ-800, Δ-600, Δ-550, Δ-500, Δ-450, Δ-400, and Δ-350) of around 2–3 weeks old, no less than three lotus leaves were sampled and assayed separately to constitute at least three biological replicates. The detected GUS activities of all deleted ADS promoter constructs were then normalized to that of ADSpro::GUS, resulting in relative GUS activities.
Statistical analysis was carried out with the methods of one-way ANOVA and paired-samples t-test using SPSS Version 20 software (IBM Corp.).
Results
Isolation and Bioinformatic Analysis of the ADS Gene Promoter
We cloned a 2,935-bp DNA fragment upstream of the translation initial codon (ATG) of the ADS gene from the genomic DNA of A. annua L. through high-fidelity PCR amplification and followed by sequencing.
To achieve an in-depth understanding of this sequence, in silico analysis was conducted via several online tools. As TSSP (http://linux1.softberry.com/berry.phtml?topic=tssp&group=programs&subgroup=promoter) predicted, three promoter/enhancer(s) are located at 2868 LDF-, 1247 LDF-, and 774 LDF-, respectively; the 2868 LDF- site is then considered as the transcription start site (+1). Through PlantCARE (Rombauts et al. 1999; Lescot et al. 2002) (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) assay of this sequence, quite a number of putative cis-acting elements are present as shown in Fig. 1. For general promoter features, a well-conserved TATA-box (TATAAA) (a core promoter element) was found −12 bp upstream of the transcription start site (TSS); furthermore, several putative CAAT boxes, a common type of cis-acting elements in promoter and enhancer regions, are present at around −50 to −200 bp upstream of TSS.
Nucleotide sequence and putative cis-acting elements of the cloned ADS promoter. A brief introduction of cis-acting elements present in this figure is as follows: ARE, a cis-element related to anaerobic induction; CAAT box, a common type of cis-acting elements in promoter and enhancer regions; GARE motif, GA-responsive element; GCN4 motif, a cis-acting element required for endosperm expression; MBS, MYB binding site involved in drought inducibility; Py-rich stretch, a cis-element responsible for high transcription levels; Skn-1 motif, a cis-acting element required for endosperm expression; TATA box, a core promoter element; TCA element, a cis-element involved in salicylic acid responsiveness; TC-rich repeats, a cis-element involved in defense and stress responsiveness; TGA element, an auxin-responsive element; W box, the binding site of WRKY transcription factors; elements in relation to light response: AE box, ATC motif, ATCT motif, Box 4 site, Box I site, chs-CMA1a site, GAG motif, GA motif, Gap box, G box, GT1 motifs, I box, MNF1, MRE site, TCT motif
Previous studies showed that expression of the ADS gene is regulated by abiotic stress, biotic stress, and multiple hormone treatments. Our in silico analysis also reveals multiple binding sites of different transcription factor families.
First of all, three W boxes (TTGACC), known as the binding site of WRKY transcription factors, are predicted, and this consists well with the previous discovery that AaWRKY1 protein binds to W box in ADS promoter (Ma et al. 2009). In addition, two putative MBSs (CAACTG, TAACTG), the MYB binding site involved in drought inducibility (Yamaguchi-Shinozaki and Shinozaki 1993), are localized at −1485(+) and −2744(−) bp respectively. These are probably associated with the discovery (Yang et al. 2010) that expression of the ADS gene in shoots rises when A. annua roots were dried for 6 h. Moreover, six elements related to anaerobic induction (ARE, TGGTTT) and one involved in defense and stress responsiveness (TC-rich repeats, ATTTTCTTCA) are found as well, indicating that the ADS gene expression may be controlled by stress through those elements.
In addition to the stress (abiotic and biotic)-responsive elements, several cis-acting elements linked to phytohormone or signal molecule responsiveness were found as well, including one GARE motif (TCTGTTG, GA responsive), one TCA element (GAGAAGAAAA, involved in salicylic acid responsiveness), and one TGA element (AACGAC, auxin-responsive). Such prediction results are closely related to previous studies: An abrupt rise of ADS expression occurred at 6 h post GA3 application (Banyai et al. 2011); SA application gave rise to a temporary peak in ADS gene expression at 24 h after application (Pu et al. 2009).
Moreover, a 5′ UTR Py-rich stretch (TTTCTTCTCT), an element responsible for high transcription levels, has been predicted at −1190(−) bp, implying that ADS promoters are capable to drive the expression of downstream genes actively.
Moreover, various putative elements in relation to light response are abundant in ADS promoter sequence, signifying that this promoter is probably subject to light regulation. These elements include two AE boxes (AGAAACAA), one ATC motif (AGTAATCT), one ATCT motif (AATCTAATCT), four Box 4 sites (ATTAAT), four Box I sites (TTTCAAA), two chs-CMA1a sites (TTACTTAA ), five G boxes (CACGAC, CACGTC, CACGTT, CACATGG), one GA motif (ATAGATAA), one GAG motif (GGAGATG), one Gap box (AAATGGAGA), three GT1 motifs (GGTTAA or AATCCACA), one I box (ATGATATGA), one MNF1 (GTGCCC), one MRE site (AACCTAA), and one TCT motif (TCTTAC).
Interestingly, two types of cis-acting elements required for endosperm expression are found in the ADS promoter region: nine Skn-1 motifs (GTCAT) and three GCN4 motifs (TGTGTCA, TCAGTCA), suggesting that this promoter may also have activity during seed development. This may explain that, although A. annua seeds are trichome-absent, several artemisinin precursors are still found in them (Brown et al. 2003).
Those in silico analysis results reveal that the ADS promoter consists of multiple regulatory elements, which is comported well with previous study that ADS is under complex control.
The ADS Promoter is Specific for Gene Expression in Both Glandular Trichomes of A. annua L. and Non-glandular Trichomes of A. thaliana
To explore the expression pattern of the ADS promoter, we constructed the ADSpro::GUS fusion vector (along with 35Spro::GUS as a positive control) and transformed wild-type A. annua with them. As shown in Fig. 2, while most leaf veins but not trichomes of the transformants with 35Spro::GUS were stained blue (Fig. 2c), in contrast blue signals of the leaf transformed with ADSpro::GUS fusion were well confined to glandular trichomes (T-shaped trichomes not included) (Fig. 2b, d). No blue traces occurred in the non-transgenic A. annua leaf (Fig. 2a), which serves as a negative control for GUS assay. These results indicate that the ADS promoter specifically drives downstream gene expression in A. annua trichomes.
Histochemical GUS staining of transgenic A. annua plants. a Juvenile leaf of non-transgenic plants (control). b Juvenile leaf of plants stably transformed with ADSpro::GUS fusion. c Juvenile leaf of plants stably transformed with 35Spro::GUS fusion. d Trichomes on juvenile leaf epidermis of plants stably transformed with ADSpro::GUS fusion
A. thaliana trichomes have long been utilized as a model for plant trichome development researches. Kim et al. (2008) have previously isolated an ADS promoter sequence (2,574 bp, Genbank accession number DQ448294), created its corresponding transgenic A. thaliana plants, and demonstrated that sequence is specifically expressed in Arabidopsis non-glandular trichomes. The ADS promoter sequence we obtained is around 400 bp longer than DQ448294 at the 5′ end and features several base differentiations as well (data not shown). Therefore, similarly, we introduced our ADSpro::GUS fusion into wild-type A. thaliana Columbia, and GUS staining assay revealed that blue staining occurred specifically in Arabidopsis trichomes as well despite the existence of discrepancies between the two sequences, as is shown in Fig. 3a. Such results reveal that the trichome specificity of the ADS promoter in A. thaliana is not determined by those discrepancies.
Histochemical GUS staining of transgenic A. thaliana leaves. a A. thaliana plants transformed with full-length-ADSpro::GUS fusion. b A. thaliana plants transformed with Δ-800/Δ-600/Δ-550/Δ-500/Δ-450/Δ-400/Δ-350. c Closer view of A. thaliana trichomes with GUS staining. d A. thaliana plants transformed with Δ-300
Sequentially Deleted Fragments of the ADS Promoter Resulted in Unexpected Expression Patterns in A. thaliana
To thoroughly investigate the ADS promoter, especially what it is that endows it with trichome specificity, we conducted 5′ sequential deletion, obtained eight 5′ deleted fragments of the ADS promoter, used each of them into the expression vector pCAMBIA1391z containing GUS as the downstream gene of those promoter fragments, and numbered them as Δ-800, Δ-600, Δ-550, Δ-500, Δ-450, Δ-400, Δ-350, and Δ-300, respectively, as is shown in Fig. 4. Stably transformed A. thaliana wild type Columbia with those fusion vectors exhibited differentiated expression patterns after being stained with X-gluc, as Fig. 3b, d shows. Through 5′ sequential deletion, we found that, as short as merely −350 upstream the transcription start site (TSS), the promoter fragment still confers GUS expression specificity in A. thaliana trichomes, and when it was shortened to −300, we did not obtain any transformed lines displaying trichome-specific GUS staining pattern.
However, not all transgenic lines containing the deleted fragments of the ADS promoter equal to or longer than −350 exhibit trichome specificity; matter-of-factly, quite a small portion of them do so, and the rest feature constitutive GUS expression pattern, with blue staining observed in trichomes as well as in other parts of transformed Arabidopsis leaves. Figure 5 reveals the specificity percentages of all transgenic lines, from which we found that transgenic lines such as Δ-800 and Δ-550 featured specificity percentages as low as around one fifth, and from Δ-550 on, the percentage increased until the highest point of 45.83 %, which lay in Δ-400, and then plummeted to below 25 % when deleted to −350. However, the specificity percentage of the full-length ADS promoter transgenic line was around 70 %.
Arabidopsis trichomes are unicellular, protuberating from leaf or stem epidermis with a short stem and generally three branches (Folkers et al. 1997). Regarding the detailed trichome-specific GUS staining feature of both the full-length and all deleted ADS promoter fragments, blue staining was basically found in the lower region of the trichome stem close to the epidermis and could also be observed in branches and basal cells in some transgenic plants with relatively high GUS expressions. Generally, trichomes located at the leaf margin exhibited higher GUS expressions than those close to the central leaf vein, with the highest expression occurring in the ones at the very tip of the leaf.
Besides the trichome specificity of all deleted fragments of the ADS promoter, which is similar to that of the full-length ADS promoter, we have observed an unusual expression pattern: Apart from transgenic Arabidopsis lines with the full-length ADS promoter, in all the other lines with deleted fragments (except Δ-500), blue staining was commonly found in the guard cells of stomata, as shown in Fig. 6.
GUS staining in the guard cells of stomata exhibited on the leaf epidermis of A. thaliana plants transformed with constructs of the deleted fragments of the ADS promoter (Δ-800, Δ-600, Δ-500, Δ-450, Δ-400, or Δ-350). a Transgenic A. thaliana leaf surface with GUS staining in both trichomes and guard cells of stomata. b Closer view of GUS staining in guard cells of stomata (arrows)
Quantitative Analysis of GUS Gene Expression
To explore the driving capabilities of the full-length ADS promoter as well as all its deleted promoter fragments, we conducted GUS quantitative assays in both transiently and stably transformed lines of full-length/deleted ADS promoter (fragments).
We first injected transformed A. tumefaciens into N. benthamiana leaves to achieve transient expressions of all the deleted fragments of the ADS promoter and the full-length one along with CaMV 35S promoter as the positive control. Quantitative analysis of the transformed tobacco leaves revealed diversified GUS expression profiles. Compared with leaves carrying the full-length ADS promoter, those with 5′ sequentially deleted fragments expressed higher GUS activities, ranging from around 1.5 to three times that of the full-length one. Of the 5′ deleted fragments, from Δ-800 to Δ-300, GUS activities of the corresponding leaves fluctuated upward, with a minimum of 1.58 times (Δ-800) that of the original ADS promoter and a maximum of 2.89 times (Δ-300) (Fig. 7a). As expected, the positive control of 35S promoter led to considerably strong GUS expression, with 8.16 times of activity that of the original ADS promoter. Statistical analysis results showed that there were no significant differences between the activities of the eight deleted fragments; however, significant differences were found between the activity of 35S promoter and that of those deleted fragments. Paired-samples t-test results showed that activities of Δ-600, Δ-400, Δ-350, and 35S promoter were significantly higher than that of the full-length ADS promoter.
GUS quantitative analysis of the ADS promoter. a Relative GUS activities of N. benthamiana leaves transiently transformed with the deleted fragments of the ADS promoter compared to that of the full-length ADS promoter. b Relative GUS activities of A. thaliana leaves stably transformed with the deleted fragments of the ADS promoter compared to that of the full-length ADS promoter
In stably transformed Arabidopsis plants, as is mentioned earlier, all deleted ADS promoter fragments except Δ-300 rendered trichome specificity. We therefore examined the GUS activities of both the full-length ADS promoter and all the other deleted fragments (Δ-300 not included) in transgenic A. thaliana. As is displayed in Fig. 7b, all the examined 5′ deleted fragments exhibited higher expression activities than the original ADS promoter, with a minimum of 2.09 times (Δ-400) and a maximum of 3.27 times (Δ-350). From Δ-800 to Δ-450, activities change fairly slightly, followed by a drop of Δ-400 and a soaring rise of Δ-350. Statistical analysis results showed that activities of the seven deleted fragments of the ADS promoter were all significantly higher than that of the full-length one, and the activity of Δ-350 is significantly higher than that of Δ-400.
Discussion
The Cross-Species Feature of Trichome Specificity of the ADS Promoter
Trichomes differentiated from plant epidermis are generally categorized into two types: non-glandular trichomes and glandular trichomes (Werker 2000). From Arabidopsis trichomes which serve as a research model for cell differentiation to cotton fibers which dictate huge economic values, non-glandular trichomes have long been the research focus for decades. Glandular trichomes are economically important as well. However, they are far more complex than their non-glandular counterparts (Tissier 2012a). From multicellular trichomes on most composite plants to unicellular ones on tobacco, glandular trichomes exhibit diverse forms that we are striving to attain deeper comprehension of. In this study, the ADS promoter derived from Asteraceae species A. annua is specifically expressed not only in A. annua multicellular glandular trichomes but in A. thaliana single-cell non-glandular trichomes as well. This result indicates the conservation of gene regulations between non-glandular and glandular trichomes.
Complexity of the Trichome-Specific Gene Expression Directed by the ADS Promoter
Efforts have long been contributed to the elucidation of the mechanism that regulates trichome-specific expressions. For the past decade, sub-regions of trichome-specific promoters from various species (Gossypium hirsutum, Nicotiana tabacum, A. thaliana, Nicotiana sylvestris, etc.) that contribute to trichome specificity have been discovered, with lengths ranging from 50 to over 300 bp (Hsu et al. 1999; Wang et al. 2002; Gutierrez-Alcala et al. 2005; Liu et al. 2006b; Wu et al. 2007; Ni et al. 2008; Ennajdaoui et al. 2010). However, until now, few cis-acting elements that control such specificity are discovered. The only discovery lies in the promoter of a cotton fiber gene RDL1 which contains an L1 box and a MYB binding motif that synergistically determine the promoter’s trichome specificity in A. thaliana (Wang et al. 2004). Mutation of each cis-element alone did not deprive the promoter of trichome specificity, only reducing its activity greatly; double mutation of both elements completely inactivated the promoter. In this study, with the online tool PLACE (http://www.dna.affrc.go.jp/PLACE/index.html) (Prestridge 1991; Higo et al. 1999), no L1 boxes (TGCATTTA) were found in the full-length ADS promoter. Concerning the MYB binding motif, seven MYBCORE elements (CNGTT[A/G]) were discovered across the ADS promoter (−2758(−), −2745(−), −2713(−), −1491(+), −960(+), −680(−), and −162(−)); however, none of these was located in the region between −350 and −300. These findings apparently illustrated that the mechanisms controlling the trichome specificity of the ADS promoter are far distinguished from the “L1 box + MYB binding motif” mechanism of the RDL1 promoter.
To investigate the mechanism of the trichome specificity of the ADS promoter, at first, we considered the model plant A. thaliana an efficient shortcut with which to pin down a unique sub-region (even element) of the ADS promoter that explains its trichome specificity since fortunately it is specifically expressed in Arabidopsis trichomes as well. However, our study reveals that it is not that simple as we previously envisaged. With the sequential deletion progressing, remaining fragments as short as barely −350 still confers trichome specificity, however, at the cost of specificity ratio, namely, the ratio between transgenic lines with trichome-specific GUS expression and the ones with constitutive GUS expression dropped compared with that of the full-length ADS promoter transgenic lines. Such a result indicates that regions upstream of −800 probably contribute to reflect partial trichome specificity, not qualitatively but quantitatively, through a mechanism we have not yet known. Within the progressive deleted region, the specificity percentages of transformed Arabidopsis lines with different deleted fragments varied considerably as well. The trend did not display as total decrease or total increase but rather a peak shape with a long tail. It seems that within the deeply investigated region from −800 to −300, all the small sub-regions of 50–200 bp all contribute to trichome specificity. Some sub-regions do it negatively, such as the sub-region of −550 to −500, deletion of which increases trichome specificity, while others positively, such as the sub-region −400 to −350, deletion of which decreases trichome specificity sharply. Therefore, from this study, the trichome specificity of ADS promoter is more like a quantitative trait than a qualitative one. These unexpected results inevitably suggest that the mechanism that lies behind trichome-specific expression is far more complicated then we previously perceived it, at least in the case of the ADS promoter. Further studies based on alternative methods are probably needed to eventually elucidate the mechanism.
Deleted Fragments of the ADS Promoter Yielded Unexpected GUS Gene Expression in SGuard Cells of Leaf Tissues
It is intriguing that besides GUS expression in Arabidopsis trichomes, guard cells of stomata of most transformed A. thaliana lines with deleted fragments of the ADS promoters were found to be with GUS staining as well. Apart from the full-length ADS promoter and Δ-500, all the other deleted ADS promoters are able to confer guard cell expressions in their corresponding transgenic Arabidopsis plants. We therefore scrutinized the ADS promoter sequence for particular cis-acting elements that are putatively responsible for the specific expression in the guard cells of stomata and eventually found 37 Dof binding elements across the whole ADS promoter sequence with the online tool PLACE (http://www.dna.affrc.go.jp/PLACE/index.html) (Prestridge 1991; Higo et al. 1999). Dof proteins are transcription factors that constitute the Dof family, and they play diverse roles in plants, ranging from light and defense responses, seed development, and germination, etc. (Yanagisawa 2002). Recent studies reveal that they regulate guard cell-specific gene expression as well through binding a short cis-acting DNA sequence [A/T]AAAG, namely, Dof binding element (Plesch et al. 2001; Galbiati et al. 2008; Yang et al. 2008; Gardner et al. 2009). Among the 37 Dof binding elements found in ADS promoter sequence, six of them (16.2 %) are located in the region from −800 to the transcription start site, and the location sites are −543(−),−473(−),−415(−), 198(+),−17(−), and +52(+), respectively. However, the full-length ADS promoter containing a staggering 37 Dof binding elements fails to direct guard cell expression, while owners of no more than six Dof binding elements (Δ-800, Δ-600, Δ-550, Δ-450, Δ-400, Δ-350) succeed in it. Besides, among all of the deleted ADS promoters, only Δ-500 lines did not display any staining in guard cells. From these findings, we may easily conclude that only Dof binding elements themselves are not sufficient for guard cell expressions. There ought to be hidden mechanisms that regulate these expressions of ADS promoter in a more complicated yet unknown way.
Prospects for Artemisinin Genetic Engineering via Deleted Fragments of the ADS Promoters
Serving as the currently most effective anti-malarial compound, artemisinin dictates almost all economic values of A. annua L. With no other industrial-scale production systems available, land-produced A. annua plants serve as the paramount source for artemisinin-based anti-malarial medication. To improve artemisinin yield in A. annua is thus of great value.
CaMV 35S promoter is a constitutive promoter that is extensively used in both basic and applied research. Recognized bottlenecks occur in the application of constitutive promoters in that the resulting overexpression of target genes is non-economical, bringing about waste of energy, and over-expressed products in non-target areas can be harmful to the plant itself. Artemisinin is specifically biosynthesized and sequestered in A. annua glandular trichomes (Nguyen et al. 2011) partly due to its phytotoxic properties (Duke et al. 1987). From this study, an attractive phenomenon occurred in that CaMV 35S promoter did not guide the GUS expression in A. annua trichomes. Although the causes of the occurrence are to be further elucidated, together with those bottlenecks of constitutive promoters, it does not appear promising to employ constitutive promoters such as 35S promoter to genetically modify A. annua for high artemisinin yield when target genes are specially expressed in glandular trichomes.
Therefore, highly effective glandular trichome-specific promoters play a critical role in genetic engineering-based A. annua breeding. A fairly large number of trichome-specific promoters from various other species have been discovered (Tissier 2012b), but none of them have been demonstrated to be able to retain their trichome specificity and function effectively in gene engineering in A. annua. The ADS promoter is one of only a few trichome-specific promoters that are isolated from A. annua. Whether it is a good choice for engineering is still an open question in that the ADS gene is expressed in not only spatial but also highly temporal way—with extraordinarily high expression in juvenile stage of A. annua but extremely low level in the senescent stage (Olofsson et al. 2011). The ADS promoter corresponds well to its gene expression profile just as expected, with high GUS activity in A. annua young leaves while nearly not detectable in old leaves (data not shown). It is hence fairly worthwhile to demobilize ADS promoter, i.e., to achieve a compact trichome-specific core promoter through removing all the unnecessary negative-regulatory elements. Beyond the core promoter, enhancers can probably be added to create a “super promoter” that embraces both trichome-specific and high-level expression. In this study, we shortened the ADS promoter to various extents and investigated the resulting promoter fragments’ capabilities of driving downstream gene expression. It is worth noting that all of the deleted fragments of the ADS promoter are more powerful in directing gene expression than the full-length one, among which Δ-350 conferred the highest GUS expression (over three times that of the full-length promoter). Therefore, these deleted fragments can be candidates for future studies on “super promoters” and engineering application. For example, although the specificity percentage of the Δ-400 transgenic line is lower (around 45 %) than that of the full-length ADS promoter, Δ-400 can still probably serve as the backbone for the construction of a super promoter or at least be employed for the trial of artemisinin genetic engineering in that it is far easier to be manipulated (significantly shorter than the original ADS promoter), boasting trichome specificity and higher activity.
References
Abdin MZ, Israr M, Rehman RU, Jain SK (2003) Artemisinin, a novel antimalarial drug: biochemical and molecular approaches for enhanced production. Planta Med 69(4):289–299. doi:10.1055/s-2003-38871
Arsenault PR, Vail D, Wobbe KK, Erickson K, Weathers PJ (2010) Reproductive development modulates gene expression and metabolite levels with possible feedback inhibition of artemisinin in Artemisia annua. Plant Physiol 154(2):958–968. doi:10.1104/pp. 110.162552
Arsenault PR, Wobbe KK, Weathers PJ (2008) Recent advances in artemisinin production through heterologous expression. Curr Med Chem 15(27):2886–2896. doi:10.2174/092986708786242813
Avery MA, Chong WKM, Jenningswhite C (1992) Stereoselective total synthesis of (+)-artemisinin, the antimalarial constituent of Artemisia annua L. J Am Chem Soc 114(3):974–979. doi:10.1021/ja00029a028
Banyai W, Mii M, Supaibulwatana K (2011) Enhancement of artemisinin content and biomass in Artemisia annua by exogenous GA(3) treatment. Plant Growth Regul 63(1):45–54. doi:10.1007/s10725-010-9510-9
Brown GD, Liang G-Y, Sy L-K (2003) Terpenoids from the seeds of Artemisia annua. Phytochemistry 64(1):303–323. doi:10.1016/s0031-9422(03)00294-2
Chen D-H, Ye H-C, Li G-F (2000) Expression of a chimeric farnesyl diphosphate synthase gene in Artemisia annua L. transgenic plants via Agrobacterium tumefaciens-mediated transformation. Plant Sci 155(2):179–185. doi:10.1016/s0168-9452(00)00217-x
Duke SO, Vaughn KC, Croom EM Jr, Elsohly HN (1987) Artemisinin, a constituent of annual wormwood (Artemisia annua), is a selective phytotoxin. Weed Sci 35:499–505
Ennajdaoui H, Vachon G, Giacalone C, Besse I, Sallaud C, Herzog M, Tissier A (2010) Trichome specific expression of the tobacco (Nicotiana sylvestris) cembratrien-ol synthase genes is controlled by both activating and repressing cis-regions. Plant Mol Biol 73(6):673–685. doi:10.1007/s11103-010-9648-x
Ferreira JFS, Janick J (1996) Distribution of artemisinin in Artemisia annua. In: Progress in new crops. ASHS, Arlington, pp 579–584
Folkers U, Berger J, Hulskamp M (1997) Cell morphogenesis of trichomes in Arabidopsis: differential control of primary and secondary branching by branch initiation regulators and cell growth. Development 124(19):3779–3786
Galbiati M, Simoni L, Pavesi G, Cominelli E, Francia P, Vavasseur A, Nelson T, Bevan M, Tonelli C (2008) Gene trap lines identify Arabidopsis genes expressed in stomatal guard cells. Plant J 53(5):750–762. doi:10.1111/j.1365-313X.2007.03371.x
Gardner MJ, Baker AJ, Assie JM, Poethig RS, Haseloff JP, Webb AAR (2009) GAL4 GFP enhancer trap lines for analysis of stomatal guard cell development and gene expression. J Exp Bot 60(1):213–226. doi:10.1093/jxb/ern292
Gutierrez-Alcala G, Calo L, Gros F, Caissard JC, Gotor C, Romero LC (2005) A versatile promoter for the expression of proteins in glandular and non-glandular trichomes from a variety of plants. J Exp Bot 56(419):2487–2494. doi:10.1093/jxb/eri241
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27(1):297–300
Hsu CY, Creech RG, Jenkins JN, Ma DP (1999) Analysis of promoter activity of cotton lipid transfer protein gene LTP6 in transgenic tobacco plants. Plant Sci 143(1):63–70. doi:10.1016/s0168-9452(99)00026-6
Jefferson R (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5(4):387–405. doi:10.1007/BF02667740
Kapila J, DeRycke R, VanMontagu M, Angenon G (1997) An Agrobacterium-mediated transient gene expression system for intact leaves. Plant Sci 122(1):101–108. doi:10.1016/s0168-9452(96)04541-4
Kapoor R, Chaudhary V, Bhatnagar AK (2007) Effects of arbuscular mycorrhiza and phosphorus application on artemisinin concentration in Artemisia annua L. Mycorrhiza 17(7):581–587. doi:10.1007/s00572-007-0135-4
Kim SH, Chang YJ, Kim SU (2008) Tissue specificity and developmental pattern of amorpha-4,11-diene synthase (ADS) proved by ADS promoter-driven GUS expression in the heterologous plant, Arabidopsis thaliana. Planta Med 74(2):188–193. doi:10.1055/s-2008-1034276
Lescot M, Dehais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouze P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327. doi:10.1093/nar/30.1.325
Liu CZ, Zhao Y, Wang YC (2006a) Artemisinin: current state and perspectives for biotechnological production of an antimalarial drug. Appl Microbiol Biotechnol 72(1):11–20. doi:10.1007/s00253-006-0452-0
Liu J, Xia KF, Zhu JC, Deng YG, Huang XL, Hu BL, Xu XP, Xu ZF (2006b) The nightshade proteinase inhibitor IIb gene is constitutively expressed in glandular trichomes. Plant Cell Physiol 47(9):1274–1284. doi:10.1093/pcp/pcj097
Ma DM, Pu GB, Lei CY, Ma LQ, Wang HH, Guo YW, Chen JL, Du ZG, Wang H, Li GF, Ye HC, Liu BY (2009) Isolation and characterization of AaWRKY1, an Artemisia annua transcription factor that regulates the amorpha-4,11-diene synthase gene, a key gene of artemisinin biosynthesis. Plant Cell Physiol 50(12):2146–2161. doi:10.1093/pcp/pcp149
Nguyen KT, Arsenault PR, Weathers PJ (2011) Trichomes plus roots plus ROS = artemisinin: regulating artemisinin biosynthesis in Artemisia annua L. Vitro Cell Dev Biol-Plant 47(3):329–338. doi:10.1007/s11627-011-9343-x
Ni SM, Meng LJ, Zhao J, Wang XC, Chen J (2008) Isolation and characterization of the trichome-specific AtTSG1 promoter from Arabidopsis thaliana. Plant Mol Biol Rep 26(4):263–276. doi:10.1007/s11105-008-0036-5
Olofsson L, Engstrom A, Lundgren A, Brodelius PE (2011) Relative expression of genes of terpene metabolism in different tissues of Artemisia annua L. BMC Plant Biol 11. doi:10.1186/1471-2229-11-45
Plesch G, Ehrhardt T, Mueller-Roeber B (2001) Involvement of TAAAG elements suggests a role for Dof transcription factors in guard cell-specific gene expression. Plant J 28(4):455–464. doi:10.1046/j.1365-313X.2001.01166.x
Prestridge DS (1991) SIGNAL SCAN: a computer program that scans DNA sequences for eukaryotic transcriptional elements. Computer Applications in the Biosciences : CABIOS 7(2):203–206
Pu GB, Ma DM, Chen JL, Ma LQ, Wang H, Li GF, Ye HC, Liu BY (2009) Salicylic acid activates artemisinin biosynthesis in Artemisia annua L. Plant Cell Reports 28(7):1127–1135. doi:10.1007/s00299-009-0713-3
Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM, Ho KA, Eachus RA, Ham TS, Kirby J, Chang MCY, Withers ST, Shiba Y, Sarpong R, Keasling JD (2006) Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440(7086):940–943. doi:10.1038/nature04640
Rombauts S, Dehais P, Van Montagu M, Rouze P (1999) PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res 27(1):295–296. doi:10.1093/nar/27.1.295
Sa G, Mi M, He-chun Y, Ben-ye L, Guo-feng L, Kang C (2001) Effects of ipt gene expression on the physiological and chemical characteristics of Artemisia annua L. Plant Sci 160(4):691–698. doi:10.1016/s0168-9452(00)00453-2
Tissier A (2012a) Glandular trichomes: what comes after expressed sequence tags? Plant J 70(1):51–68. doi:10.1111/j.1365-313X.2012.04913.x
Tissier A (2012b) Trichome specific expression: promoters and their applications. Transgenic Plants - Advances and Limitations. In Tech. doi:10.5772/32101
Wang EM, Gan SS, Wagner GJ (2002) Isolation and characterization of the CYP71D16 trichome-specific promoter from Nicotiana tabacum L. J Exp Bot 53(376):1891–1897. doi:10.1093/jxb/erf054
Wang H, Olofsson L, Lundgren A, Brodelius PE (2011) Trichome-specific expression of amorpha-4,11-diene synthase, a key enzyme of artemisinin biosynthesis in Artemisia annua L., as reported by a promoter–GUS fusion. Am J Plant Sci 2(4):619–628
Wang S, Wang JW, Yu N, Li CH, Luo B, Gou JY, Wang LJ, Chen XY (2004) Control of plant trichome development by a cotton fiber MYB gene. Plant Cell 16(9):2323–2334. doi:10.1105/tpc.104.024844
Wellems TE (2002) Plasmodium chloroquine resistance and the search for a replacement antimalarial drug. Science 298(5591):124–126. doi:10.1126/science.1078167
Werker E (2000) Trichome diversity and development. Advances in Botanical Research Incorporating Advances in Plant Pathology 31:1–35. doi:10.1016/s0065-2296(00)31005-9
Westfall PJ, Pitera DJ, Lenihan JR, Eng D, Woolard FX, Regentin R, Horning T, Tsuruta H, Melis DJ, Owens A, Fickes S, Diola D, Benjamin KR, Keasling JD, Leavell MD, McPhee DJ, Renninger NS, Newman JD, Paddon CJ (2012) Production of amorphadiene in yeast, and its conversion to dihydroartemisinic acid, precursor to the antimalarial agent artemisinin. Proc Natl Acad Sci USA 109(3):111–118. doi:10.1073/pnas.1110740109
World Health Organization (2011) World malaria report 2010. WHO, Geneva
Wu AM, Lv SY, Liu JY (2007) Functional analysis of a cotton glucuronosyltransferase promoter in transgenic tobaccos. Cell Res 17(2):174–183. doi:10.1038/sj.cr.7310119
Xu XX, Zhu J, Huang DZ, Zhou WS (1986) Total synthesis of arteannuin and deoxyarteannuin. Tetrahedron 42(3):819–828
Yamaguchi-Shinozaki K, Shinozaki K (1993) Arabidopsis DNA encoding two desiccation-responsive rd29 genes. Plant Physiol 101(3):1119–1120. doi:10.1104/pp. 101.3.1119
Yanagisawa S (2002) The Dof family of plant transcription factors. Trends Plant Sci 7(12):555–560. doi:10.1016/s1360-1385(02)02362-2
Yang RY, Zeng XM, Lu YY, Lu WJ, Feng LL, Yang XQ, Zeng QP (2010) Senescent leaves of Artemisia annua are one of the most active organs for overexpression of artemisinin biosynthesis responsible genes upon burst of singlet oxygen. Planta Med 76(7):734–742. doi:10.1055/s-0029-1240620
Yang YZ, Costa A, Leonhardt N, Siegel RS, Schroeder JI (2008) Isolation of a strong Arabidopsis guard cell promoter and its potential as a research tool. Plant Methods 4. doi:10.1186/1746-4811-4-6
Zeng QP, Qiu F, Yuan L (2008) Production of artemisinin by genetically-modified microbes. Biotechnol Lett 30(4):581–592. doi:10.1007/s10529-007-9596-y
Zhang L, Jing F, Li F, Li M, Wang Y, Wang G, Sun X, Tang K (2009) Development of transgenic Artemisia annua (Chinese wormwood) plants with an enhanced content of artemisinin, an effective anti-malarial drug, by hairpin-RNA-mediated gene silencing. Biotechnol Appl Biochem 52(3):199–207. doi:10.1042/ba20080068
Zhang XR, Henriques R, Lin SS, Niu QW, Chua NH (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1(2):641–646. doi:10.1038/nprot.2006.97
Acknowledgments
This work was funded by China “863” Program (grant no. 2011AA100605), China Transgenic Research Program (grant no. 2013ZX08002001-006), and Shanghai Leading Academic Discipline Project (Horticulture).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhu, M., Zhang, F., Lv, Z. et al. Characterization of the Promoter of Artemisia annua Amorpha-4,11-diene Synthase (ADS) Gene Using Homologous and Heterologous Expression as well as Deletion Analysis. Plant Mol Biol Rep 32, 406–418 (2014). https://doi.org/10.1007/s11105-013-0656-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-013-0656-2