Abstract
Artemisinin, isolated from Artemisia annua L., is widely known as a functional anti-malaria drug. Due to the low content of artemisinin in A. annua plants, great efforts have been made to determine the artemisinin biosynthetic pathway by genetic engineering. ADH1, encoding an alcohol dehydrogenase, was cloned from the glandular secretory trichomes (GSTs) in A. annua. The gene expression analysis showed that ADH1 was predominately expressed in buds and young leaves, and the expression of ADH1 was the highest in the youngest leaves. To further investigate the expression pattern of ADH1 in A. annua, a 1070-bp promoter region of ADH1 was cloned. We found 14 putative cis-elements were presented in the ADH1 promoter sequence, indicating that ADH1 is complexly regulated. The ADH1 promoter sequence was fused to the β-glucuronidase reporter gene (GUS) and introduced into A. annua plants. GUS signals were only found in the glandular secretory trichomes of young tissues in transgenic A. annua plants. Besides, the treatment of A. annua seedlings with 100 μM methyl jasmonate (MeJA) and 100 μM abscisic acid (ABA), respectively, increased the ADH1 transcript levels. The dual luciferase (dual-LUC) assay demonstrated that the reported transcription factors, MYC2 and ERF1, activated the expression of ADH1 in vivo. Our study shows that ADH1 gene is exclusively expressed in the glandular secretory trichomes of young tissues of A. annua, it implies that the promoter of ADH1 gene could be used in engineering of A. annua for increasing artemisinin content.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
More than 214 million people are suffering from a threat of malaria all over the world. The case of malaria has led to approximately 4,38,000 deaths in 2015 (WHO 2016). Artemisinin-combination therapies (ACTs), the most effective available antimalarial cure at present, is globally recommended by WHO (Mutabingwa 2005). Artemisinin is an anti-malarial endoperoxide with a peroxide bridge. Unfortunately, the limited artemisinin content in Artemisia annua plants (0.01–0.8% DW) results in a worldwide shortage of the antimalarial drug (Abdin et al. 2003). Therefore, it is urgent to adopt new strategies for a large improvement of artemisinin accumulation in A. annua to obtain a number of stable high-producing plants. Great efforts have been expended in metabolic engineering to enhance the artemisinin accumulation. Several transcription factors regulating the biosynthetic pathways of artemisinin have been reported. In A. annua, WRKY1 is isolated firstly and reported as a positive regulator in artemisinin biosynthesis. Overexpressing WRKY1 driven by the glandular trichome-specific CYP71AV1 promoter can lead the artemisinin content to almost 1.9 times higher than that of controls (Han et al. 2014). Previous study revealed that the expressions of ADS, CYP71AV1, and DBR2 were strongly activated in ORA-overexpressing transgenic lines, resulting in an increase of artemisinin content by 40–53% compared to control plants (Lu et al. 2013a).
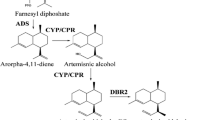
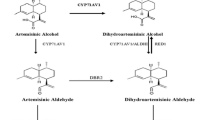
Numerous efforts have been made to clarify the artemisinin biosynthesis (Fig. 1). The reaction from farnesyl diphosphate (FPP) to amorpha-4,11-diene is catalyzed by amorpha-4,11-diene synthase (ADS), which is considered as the initial step specifically occurred in the glandular secretory trichomes (Banyai et al. 2010; Wang et al. 2011). Next, amorpha-4,11-diene is gradually catalyzed to artemisinic alcohol, and further yield artemisinic aldehyde under the catalysis of both cytochrome P450 monooxygenase (CYP71AV1) and cytochrome P450 reductase (CPR) (Ro et al. 2006; Teoh et al. 2006). Recently, alcohol dehydrogenase 1 (ADH1) has been proved to involve in the formation of artemisinic aldehyde in yeast (Paddon et al. 2013). Subsequently, the artemisinic aldehyde is deoxidized to dihydroartemisinic aldehyde by artemisinic aldehyde Δ11 (13) reductase (DBR2) (Zhang et al. 2008). Then dihydroartemisinic aldehyde is reduced to form dihydroartemisinic acid with the action of aldehyde dehydrogenase 1 (ALDH1) (Teoh et al. 2009). Artemisinin is formed from dihydroartemisinic acid in a photo-oxidative reaction rather than an enzymatic conversion (Sy and Brown 2002; Brown and Sy 2004). Alternatively, the artemisinic aldehyde is catalyzed to artemisinic acid, the precursor of arteannuin B, by the CYP71AV1 and ALDH1 (Ro et al. 2006; Teoh et al. 2009). The conversion of artemisinic acid to arteannuin B is also regarded as a photo-oxidative reaction without any enzyme involved in (Sy and Brown 2002; Brown and Sy 2004).
Artemisinin biosynthetic pathway in A. annua. HMGS 3-hydroxy-3-methyl-glutaryl coenzyme A synthase, HMGR 3-hydroxy-3-methyl-glutaryl coenzyme A reductase, DXS 1-deoxy-d-xylulose-5-phosphate synthase, DXR 1-deoxy-d-xylulose-5-phosphate reductoisomerase, FPS farnesyl diphosphate synthase, ADS amorpha-4,11-diene synthase, CYP71AV1 cytochrome P450 monooxygenase, CPR cytochrome P450 reductase, ADH1 alcohol dehydrogenase 1, DBR2 artemisinic aldehyde Δ11 (13) reductase, ALDH1 aldehyde dehydrogenase 1
Previous researches indicate that the glandular secretory trichomes can be considered as storages for artemisinin and infer the precursors of artemisinin biosynthesis are exclusively located in this type of trichomes (Ferreira and Janick 1996; Arsenault et al. 2010; Nguyen et al. 2011). Previous studies proved that ADS, CYP71AV1, DBR2 and ALDH1 are specifically expressed in the glandular secretory trichomes of young tissues (Wang et al. 2011, 2013; Jiang et al. 2014; Liu et al. 2016).
Recently, ADH1 was isolated from A. annua plants. The sequence analysis and in vitro characterization indicate that ADH1 protein, expressed and purified from Escherichia coli, encodes alcohol dehydrogenase, suggesting that ADH1 catalyzes artemisinic alcohol to form artemisinic aldehyde in artemisinin biosynthesis (Paddon et al. 2013). Moreover, there was 18% more artemisinic acid than the control when ADH1 together with ALDH1, CYP71AV1, CYB5 and CPR1 were expressed in yeast strain Y1368 (Paddon et al. 2013). However, the localization of ADH1 is not reported in A. annua.
An enormous amount of studies to improve the artemisinin content in A. annua have been made a great progress. When FPS gene was overexpressed in the A. annua plants, the transgenic plants exhibited the 2.5-fold artemisinin content compared to that in non-transgenic plants (Banyai et al. 2010). The artemisinin content was improved by 82% in the ADS over-expressing lines by GC–MS (Ma et al. 2009). Furthermore, co-overexpressing ADS, CPR and CYP71AV1 genes in A. annua led to the greater artemisinin accumulation than that in control plants (Lu et al. 2013b). In another example, HMGR and ADS genes were co-overexpressed in A. annua plants and one of the transgenic lines exhibited more than seven-fold higher artemisinin content than the control lines (Alam and Abdin 2011). Four artemisinin biosynthetic pathway genes (ADS, CYP71AV1, CPR and ALDH1) were overexpressed in A. annua and the transgenic lines showed higher artemisinin content, with 3.4-fold than that detected in wild-type plants (Shi et al. 2017). Therefore, the transgenic approach is vital to improve the production of artemisinin in A. annua.
In this study, we analyzed the transcript levels of ADH1 in different tissues and leaves at leaf development age by quantitative real-time PCR, which indicated that ADH1 was predominately expressed in buds and young leaves, and the expression of ADH1 gene was the highest in the youngest leaf and rapidly reduced during leaf development. To further investigate the expression pattern of ADH1, the promoter sequence of ADH1 was cloned. Then the ADH1 promoter sequence was fused to the GUS reporter gene and introduced into A. annua plants by Agrobacterium-mediated transformation system. GUS signals were only found in the glandular secretory trichomes of young tissues in transgenic A. annua plants. Besides, the treatment of seedlings with MeJA and ABA, respectively, increased the ADH1 transcript levels. Finally, a dual-LUC assay in tobacco leaves indicated that MYC2 and ERF1 activated the expression of ADH1, respectively.
Materials and methods
Relative expression analysis by RT-q-PCR
The different organs used in this study [roots, stems, young leaves (the two youngest leaves), old leaves (from the sixth to the ninth leaf), buds and flowers] and the leaves from different nodes were collected from 4-month-old A. annua for RNA extraction following the manufacturer’s protocol (Tiangen Biotech, Beijing, People’s Republic of China). First-strand cDNA was synthesized from the total RNA using the PrimeScript II first Strand cDNA Synthesis Kit (Takara Biotechnology, Dalian, China). We used the β-actin as a standard control (Liu et al. 2015, 2016). Primers involved in this expriment are listed in Table 1. Amplification was employed using the Roche LightcycleR® 96 (Roche, Mannheim, Germany) with the kit SuperReal PreMix Plus (Tiangen Biotech, Beijing, China) following the manufacturer’s instruction as followed: 94 °C for 8 min, followed by 40 cycles of 94 °C (20 s), 55 °C (20 s) and 72 °C (20 s), and a last extension at 72 °C for 8 min. Three technical replications were run for each cDNA sample. The transcription level of ADH1 was analyzed by RT-q-PCR using the 2−ΔΔCt method (Livak and Schmittgen 2001).
Cloning of the ADH1 promotor
We extracted genomic DNA from the youngest leaves of A. annua plants by the cetyltrimethylammonium bromide method (Fütterer et al. 1995). Based on our genomic databases (unpublished), the promoter region of ADH1 was amplified using high-fidelity PCR (Toyobo, Osaka, Japan) with ADH1-specific primers (Table 1). The amplified fragment of ADH1 promoter was purified and linked with pLB vector (Tiangen Biotech, Beijing, China) to sequence.
Analysis of promoter sequence
The TSSP software (http://linux1.soft-berry.com), the PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html), and the PLACE (http://www.dna.affrc.go.jp/PLACE/) databases were applied for the prediction of transcription start site (TSS) and the cis-acting elements of the cloned ADH1 promoter.
Vector construction
The ADH1 promoter was amplified with primers containing HindIII and NcoI restriction sites. The amplified fragment and pCAMBIA1391Z vector (CambiaLabs) carrying the GUS were digested with HindIII and NcoI (Fermentas FastDigest) respectively. The promoter region was inserted into pCAMBIA1391Z vector. Subsequently, the recombinant plasmid was introduced into Agrobacterium tumefaciens strain EHA105. The resulting A. tumefaciens EHA105 harboring the construct (pCAMBIA1391Z-proAHD1-GUS) were prepared for the plant transformation.
Plant materials and transformation of A. annua
Seeds called “Huhao 1”, originated from Chongqing, were cultivated by our group (Shen et al. 2016). Seeds were immersed in 75% ethanol for 60 s, and then soaked in 25% sodium hypochlorite solution for 8 min, followed by five rinses with sterile distilled water. All the germ-free seeds were plated on MS medium (Murashige and Skoog 1962) [MS powder 4.4 g L−1(Sigma–Aldrich, MO, USA), 30 g L−1 sucrose (Sangon Biotech, Shanghai, China), 2.6 g L−1 phytagel(Sigma–Aldrich, MO, USA), pH = 5.8] at 25 °C for 3 weeks. 20–25-day-old seedlings were used for transformation. Leaf discs with 0.5 cm diameter were cut and co-cultivated with A. tumefaciens strain EHA105 carrying the pCAMBIA1391Z-pADH1-GUS on co-cultivation medium in the dark at 28 °C for 72 h. Then the treated explants were transferred to selection medium MS1 and further to root-inducing medium MS2 as described previously (Liu et al. 2016). Finally, we got the plantlets and cultivated them for 2–3 months in the greenhouse at room temperature.
GUS assay
The leaf primordia, expanded leaves at different nodes, stems and roots were respectively collected from both transgenic A. annua plants and wild-type plants to carry out GUS analysis. GUS staining of these samples was performed according to previous protocol (Jefferson et al. 1987).
Hormonal treatments
MeJA (100 μM) (Sigma–Aldrich) and abscisic acid (100 μM) (Sigma–Aldrich) were prepared for hormone treatments, and 1% DMSO solution was used as a mock treatment. The cutting plants were prepared and sprayed with different hormone solutions. After treatment, they were cultivated in the greenhouse with the artificial light at a constant temperature of 25 °C. The youngest leaves (leaves at the first node) of the wild-type A. annua plants were respectively collected for RNA extraction at 0, 1, 3, 6, 9, 12 and 24 h after hormonal treatments. Leaves from three independent plants were collected for each determination. In addition, the leaves of transgenic A. annua plants were used for GUS staining after the hormonal treatments.
Dual-luciferase activity assay
The promoter fragment of ADH1 was inserted into pGreenII 0800-LUC vector. The recombinant plasmid was introduced into A. tumefaciens strain GV3101 with the plasmid pSoup19 as a reporter construct. For dual-luciferase activity assay, pHB-ERF1, pHB-bZIP1, pHB-WRKY1 and pHB-MYC2 (Ma et al. 2009; Lu et al. 2013c; Zhang et al. 2015; Shen et al. 2016) plasmids were respectively transformed into A. tumefaciens strain GV3101 to act as effectors (Luo et al. 2014). The A. tumefaciens strains were cultured overnight at 28 °C, collected by centrifugation and further resuspended using MS liquid medium to OD600 = 0.6. Then the mixture were infiltrated into tobacco leaves followed by the agroinfiltration procedure (Voinnet et al. 2003). The pHB construct was used as a negative control. The dual-LUC assay was performed using Dual-Luciferase® Reporter Assay System (Promega, Durham, USA). Three biological repeats were valued for each sample.
Statistical analysis
Data are presented as means ± SD unless otherwise stated. To compare group differences, paired or unpaired, two-tailed Student’s t tests were used. P < 0.05 was considered significant and P < 0.01 was considered extremely significant.
Results and discussion
Expression pattern of ADH1 in A. annua
To analyze the expression pattern of ADH1 in A. annua, the different organs were collected for RNA extraction from A. annua. The results showed that the ADH1 expression was the highest in buds and young leaves (Fig. 2a). The ADH1 expression was poorly detected in roots, old leaves and stems. We analyzed the expression of ADH1 in leaves at different developmental stages. The transcript level of ADH1 was the highest in the youngest leaf (leaf 0) and reduced gradually with age of the leaves (Fig. 2b). The ADH1 expression pattern is similar to that of the artemisinin biosynthetic pathway genes (Figs. 3, 4). Several researches indicate that the artemisinin-specific enzymes (ADS, CYP71AV1, DBR2 and ALDH1) are located to the glandular secretory trichomes, where artemisinin is biosynthesized and stored (Zhang et al. 2008; Wang et al. 2011, 2013; Liu et al. 2016). Our results suggest that ADH1 is also likely to be located to the glandular secretory trichomes.
Expression pattern of ADH1 in A. annua plants: a relative expression of ADH1 in different tissues (root, stem, old leaf, young leaf, flower, and bud). β-actin was used as the control gene. Error bars represent ± SD (n = 3). Statistical significance was determined by two-tailed Student’s t test with P values; **P < 0.01. Asterisks above the bar represent the significance of the difference between different tissues; b relative expression of ADH1 in expanded leaves at different nodes (leaf 0, leaf 1, leaf 2, leaf 3, leaf 4, leaf 5, leaf 9 and leaf 16). β-actin was used as the control gene. Error bars represent ± SD (n = 3). Statistical significance was determined by two-tailed Student’s t test with P values; **P < 0.01. Asterisks above the bar represent the significance of the difference between expanded leaves at different nodes
Expression patterns in different tissues of A. annua plants: a relative expression of ADS; b relative expression of CYP71AV1; c relative expression of DBR2; d relative expression of ALDH1. β-actin was used as the control gene. Error bars represent ± SD (n = 3). Statistical significance was determined by two-tailed Student’s t test with P values; **P < 0.01. Asterisks above the bar represent the significance of the difference between different tissues
Expression patterns of leaves at different nodes in A. annua: a relative expression of ADS; b relative expression of CYP71AV1; c relative expression of DBR2; d relative expression of ALDH1. β-actin was used as the control gene. Error bars represent ± SD (n = 3). Statistical significance was determined by two-tailed Student’s t test with P values; **P < 0.01. Asterisks above the bar represent the significance of the difference between expanded leaves at different nodes
Cloning and analysis of ADH1 promoter
To further investigate the expression of ADH1, we predicted the ADH1 promoter sequence from our genomic databases (unpublished) and cloned the 1070-bp ADH1 promoter.
The TSS of ADH1 promoter was located 53 bp upstream from the start codon (ATG) at position +1 (labeled +1 in Fig. 5). It was found that a putative TATA-box (TATAA), the well-conserved core promoter sequence, at position from −6 to −10 in the ADH1 promoter (Fig. 5). A CAAT-box generally accompanied by the consensus sequence (GGCCAATCT) is required for transcription. We found the CAAT-box (CAATT) was located 165–169 bp upstream from the putative TSS of ADH1 promoter (Fig. 5).
As shown in Table 2, 14 cis-acting elements in the ADH1 promoter were listed.
One G-box (GGTACAC; position −24 to −30) was found in the ADH1 promoter. The MYC2 transcription factor, one of the bHLH transcription factor family, is capable of binding G-box or G-box-like motifs in the promoter, exhibiting a strong JA-response ability (Shen et al. 2016). There is a strong possibility that the MYC2 binds the G-box of ADH1 promoter.
Five E-boxes were found at positions −24 to −29 (CATGTG), −151 to −156 (CATTTG), −237 to −242 (CATTTG), −370 to −375 (CAACTG), −642 to −647 (CATATG), respectively. It was reported earlier that bHLH TFs could induce gene expression through binding to the E-box sequence (5′-CANNTG-3′) (Toledoortiz et al. 2003). Biochemical analysis demonstrates that bHLH TFs of plants make contribution to the JA responsiveness (Miyamoto et al. 2012). Thus, the E-box in the ADH1 promoter region may act as the JA-responsive cis-element. Previous studies also showed that bHLH TFs play positive roles in ABA-mediated response of Arabidopsis (Kim and Kim 2006).
Two MBS (MYB binding sites) (CAACTG/TAACTG; position −370 to −375 and −980 to −985) were predicted in the ADH1 promoter. Earlier study demonstrated that a MYB-related transcription factor, MYB2, may regulate genes which are induced by water stress in Arabidopsis (Urao et al. 1993). It was also reported that Glabroos1 (GL1), another MYB-related gene, is involved in the formation of trichomes of leaves in Arabidopsis (Urao et al. 1993). All the above suggests that the ADH1 may have similar functions in A. annua.
In addition, the ADH1 promoter was predicted to carry other functional cis-elements. Two ARE (TGGTTT/TTTGGT) were found at the positions −868 to −863 and −28 to −23, which is the cis-element involved in the anaerobic induction. Further analysis of the promoter sequence showed that ADH1 may relate to the seed development since two Skn-1_motif (GTCAT; position −72 to −76 and 20–24) and one GCN4-motif (TGAGTCA; position −144 to −150), which are essential for endosperm expression, were all detected in the ADH1 promoter.
A number of light responsive elements are presented in ADH1 promoter sequence, including two ACE (ATTTGCTAAA; position −216 to −207 and 37–46), two Box 4 (ATTAAT; position −251 to −256 and −120 to −125), two GATA-box (TGATAA; position −283 to −288 and −3 to +3), one TCT-motif (CATTCT; position −502 to −507), one GA-motif (AAAGATGA; position −720 to −727), one GAG-motif (AGAGATG; position −624 to −630), one Box III (TCCCATTTAC; position −770 to −779) and two Sp1 (CCCCCCACTT/CCCTCCCTTG; position −752 to −761 and −852 to −861). This suggests that the ADH1 may be regulated by light.
Analysis of the activity of ADH1 promoter
To further analyze the expression pattern of ADH1 in A. annua, the ADH1 promoter sequence was inserted into the vector pCAMBIA1391Z carrying the GUS reporter gene. The recombination plasmid was introduced into A. annua plants as described above. The positive transgenic plants were selected by PCR. GUS staining was only observed in the glandular secretory trichomes of leaf primordia (Fig. 6a), young leaves (Fig. 6b–j), and stems of the transgenic A. annua plants (Fig. 6m, n). Besides, we did not find any GUS signal in old leaves and roots (Fig. 6k, l, o, p).
GUS staining of 1 month-old transgenic A. annua plants: a leaf primordia of 1 month-old transgenic A. annua plant; b expanded leaf at node 1 from a 1 month-old transgenic A. annua plant; c magnification (×4) of expanded leaf at node 1; d expanded leaf at node 2 from a 1 month-old transgenic A. annua plant; e magnification (×2) of expanded leaf at node 2; f magnification (×4) of GSTs in expanded leaf at node 2; g magnification (×8) of GSTs in expanded leaf at node 2; h expanded leaf at node 3 from a 1 month-old transgenic A. annua plant; i magnification (×2) of expanded leaf at node 3; j expanded leaf at node 4 from a 1 month-old transgenic A. annua plant; k old leaf of 1 month-old transgenic A. annua plant; l magnification (×2) of old leaf; m GSTs in stems; n magnification (×2) of GSTs in stems; o roots of 1 month-old transgenic A. annua plant; p magnification (×2) of roots. GST glandular secretory trichome and TST T-shaped trichome
Response of the ADH1 promoter to hormones
It is well established that the expression of ADS, CYP71AV1, DBR2 and ALDH1 are induced by MeJA in A. annua (Wu et al. 2011; Liu et al. 2016). Besides, the transcript levels of HMGR, FDS and CYP71AV1 were upregulated when A. annua plants were exposed to the ABA treatment (Jing et al. 2009).
We treated A. annua seedlings with MeJA and ABA separately as described above, and the transcript level of ADH1 gene was valued using RT-q-PCR. The relative expression levels at different time were normalized to the expression at 0 h. Compared to 0 h, the transcript level of ADH1 gene reached twofold expression at 9 h after spraying with MeJA (Fig. 7a). The relative expression of ADH1 gene was induced approximate 2.5-fold of the control at 6 h in the ABA responsiveness experiment (Fig. 7b). In addition, the result showed stronger GUS staining after hormonal treatments compared with the control (Fig. 8a–d). Observation obtained shows that the expression of ADH1 increased quickly to the peak at a point time and then slowly decreased after hormone treatment. Similarly, it was reported that the expression of MYC2 was upregulated rapidly and peaked at 6 h after MeJA treatment, and then declined gradually until 48 h (Shen et al. 2016). The transcript level of MYC2 after ABA treatment increased relatively slowly and declined rapidly (Shen et al. 2016). The ALDH1 response to MeJA treatment was also quick; it peaked at 3 h and then declined (Liu et al. 2016). The expression pattern of ADH1 after hormone treatment is similar to that of MYC2 and ALDH1.The results indicate that ADH1 gene shows hormone–dependent expression, which in agreement with the analysis that the ADH1 promoter carries a few motifs involved in hormonal induction.
Expression patterns of ADH1 in response to hormones: a relative expression of ADH1 in A. annua after treatment with MeJA; b relative expression of ADH1 in A. annua after treatment with ABA. β-actin was used as the control gene. Error bars represent ±SD (n = 3). Statistical significance was determined by two-tailed Student’s t test with P values; *P < 0.05; **P < 0.01. Asterisks above the bar represent the significance of the difference between hormone-treated and control plants
MYC2 and ERF1 activate the transcription of ADH1 in vivo
To investigate the regulation of ADH1 transcription, a dual-LUC assay was performed in tobacco leaves. MYC2 activated the expression of ADH1 promoter by showing a much higher value of relative LUC activity than the control (Fig. 9). ERF1 can also activate the expression of ADH1 promoter for the value of relative LUC activity reached 2.9 times compared with the control (Fig. 9).Together, these results suggest that MYC2 and ERF1 may recognize specific cis-regulatory sequences in the ADH1promoter, and then activate the transcription of ADH1.
Conclusion
Expression pattern of ADH1 indicates that ADH1 shows high expression in young leaves as the same as that of other crucial enzyme genes involved in biosynthetic pathways of artemisinin. Further analysis of ADH1 promoter sequence reveals that the expression of ADH1 may be complexly regulated. The GUS staining analysis shows that the ADH1 is located to the glandular secretory trichomes of young tissues, which are similar to the GUS staining pattern of ADS promoter. Not only does the expression of ADH1 in yeast increase the production of artemisinic acid observably, but also ADH1 is worth to be considered as an efficient participant in the biosynthesis of artemisinin. Besides, the GST-specific promoter (proADH1) may be a good substitute for CaMV 35S promoter to drive the expression of a known gene in the glandular secretory trichomes. Moreover, the expression of ADH1 can be obviously improved by various hormones, providing an evident for the enhancement of artemisinin content by treating with hormones. We may conclude that overexpression of ADH1 gene in A. annua plants will most likely has a positive effect on artemisinin biosynthesis.
References
Abdin MZ, Israr M, Rehman RU, Jain SK (2003) Artemisinin, a novel antimalarial drug: biochemical and molecular approaches for enhanced production. Planta Med 69:289–299
Alam P, Abdin MZ (2011) Over-expression of HMG-CoA reductase and amorpha-4,11-diene synthase genes in Artemisia annua L. and its influence on artemisinin content. Plant Cell Rep 30:1919–1928
Arsenault PR, Vail D, Wobbe KK, Erickson K, Weathers PJ (2010) Reproductive development modulates gene expression and metabolite levels with possible feedback inhibition of artemisinin in Artemisia annua. Plant Physiol 154:958–968
Banyai W, Kirdmanee C, Mii M, Supaibulwatana K (2010) Overexpression of farnesyl pyrophosphate synthase (FPS) gene affected artemisinin content and growth of Artemisia annua L. Plant Cell Tiss Org Cult 103:255–265
Brown GD, Sy LK (2004) In vivo transformations of dihydroartemisinic acid in Artemisia annua plants. Tetrahedron 60:1139–1159
Ferreira JFS, Janick J (1996) Distribution of artemisinin in Artemisia annua. In: Janick J (ed) Progress in new crops. ASHS, Arlington, pp 579–584
Fütterer J, Gisel A, Iglesias V, Klöti A, Kost B, Mittelsten Scheid O, Neuhaus G, Neuhaus-Url G, Schrott M, Shillito R, Spangenberg G, Wang ZY (1995) Standard molecular techniques for the analysis of transgenic plants. In: Potrykus I, Spangenberg G (eds) Gene transfer to plants. Springer, Berlin, pp 215–263
Han J, Wang H, Lundgren A, Brodelius PE (2014) Effects of overexpression of AaWRKY1 on artemisinin biosynthesis in transgenic Artemisia annua plants. Phytochemistry 102:89–96
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusion: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plant. EMBO J 6:3901–3907
Jiang WM, Lu X, Qiu B, Zhang FY, Shen QY, Lv ZY, Fu XQ, Yan TX, Gao ED, Zhu MM, Chen LX, Zhang L, Wang GF, Sun XF, Tang KX (2014) Molecular cloning and characterization of a trichome-specific promoter of artemisinic aldehyde Δ11 (13) reductase (DBR2) in Artemisia annua. Plant Mol Biol Rep 32:82–91
Jing FY, Zhang L, Li MY, Tang YL, Wang YL, Wang YY, Wang Q, Pan QF, Wang GF, Tang KX (2009) Abscisic acid (ABA) treatment increases artemisinin content in Artemisia annua by enhancing the expression of genes in artemisinin biosynthetic pathway. Biologia 64:319–323
Kim J, Kim HY (2006) Molecular characterization of a bHLH transcription factor involved in Arabidopsis abscisic acid-mediated response. BBA Gene Struct Exp 1759:191–194
Liu WH, Zhao TF, Wang HY, Zeng JL, Xiang LE, Zhu SQ, Chen M, Lan XZ, Liu XQ, Liao ZH (2015) Reference gene selection in Artemisia annua L., a plant species producing anti-malarial artemisinin. Plant Cell Tiss Org Cult 121:141–152
Liu M, Shi P, Fu XQ, Brodelius PE, Shen Q, Jiang WM, He Q (2016) Characterization of a trichome-specific promoter of the aldehyde dehydrogenase 1 (ALDH1) gene in Artemisia annua. Plant Cell Tiss Org Cult 126:469–480
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCt method. Methods 25:402–408
Lu X, Zhang L, Zhang FY, Jiang WM, Shen Q, Zhang LD, Lv ZY, Wang GF, Tang KX (2013a) AaORA, a trichome-specific AP2/ERF transcription factor of Artemisia annua, is a positive regulator in the artemisinin biosynthetic pathway and in disease resistance to Botrytis cinerea. New Phytol 198:1191–1202
Lu X, Shen Q, Zhang L, Zhang FY, Jiang WM, Lv ZY, Yan TX, Fu XQ, Wang GF, Tang KX (2013b) Promotion of artemisinin biosynthesis in transgenic Artemisia annua by overexpressing ADS, CYP71AV1 and CPR genes. Ind Crop Prod 49:380–385
Lu X, Jiang WM, Zhang L, Zhang F, Zhang FY, Shen Q, Wang GF, Tang KX (2013c) AaERF1 positively regulates the resistance to Botrytis cinerea in Artemisia annua. Plos One 8(2):e57657
Luo Q, Lian HL, He SB, Li L, Jia KP, Yang HQ (2014) COP1 and phyB physically interact with PIL1 to regulate its stability and photomorphogenic development in Arabidopsis. Plant Cell 26:2441–2456
Ma DM, Pu GB, Lei CY, Ma LQ, Wang HH, Guo YW, Chen JL, Du ZG, Wang H, Li GF, Ye HC, Liu BY (2009a) Isolation and characterization of AaWRKY1, an Artemisia annua transcription factor that regulates the amorpha-4,11-diene synthase gene, a key gene of artemisinin biosynthesis. Plant Cell Physiol 50(12):2146–2161
Ma C, Wang H, Lu X, Wang H, Xu G, Liu B (2009b) Terpenoid metabolic profiling analysis of transgenic Artemisia annua L. by comprehensive two-dimensional gas chromatography time-of-flight mass spectrometry. Metabolomics 5:497–506
Miyamoto K, Shimizu T, Lin FQ, Sainsbury F, Thuenemann E, Lomonossoff G, Nojiri H, Yamane H, Okada K (2012) Identification of an E-box motif responsible for the expression of jasmonic acid-induced chitinase gene OsChia4a in rice. J Plant Phsiol 169:621–627
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Mutabingwa TK (2005) Artemisinin-based combination therapies (ACTS): best hope for malaria treatment but inaccessible to the needy! Acta Trop 95:305–315
Nguyen KT, Arsenault PR, Weathers PJ (2011) Trichomes + roots + ROS = artemisinin: regulating artemisinin biosynthesis in Artemisia annua L.. In Vitro Cell Dev Biol Plant 47:329–338
Paddon CJ, Westfall PJ, Pitera DJ, Benjamin K, Fisher K, McPhee D, Leavell MD, Tai A, Main A, Eng D, Polichuk DR, Teoh KH, Reed DW, Treynor T, Lenihan J, Fleck M, Bajad S, Dang G, Dengrove D, Diola D, Dorin G, Ellens KW, Fickes S, Galazzo J, Gaucher SP, Geistlinger T, Henry R, Hepp M, Horning T, Iqba T, Jiang H, Kizer L, Lieu B, Melis D, Moss N, Regentin R, Secrest S, Tsuruta H, Vazquez R, Westblade LF, Xu L, Yu M, Zhang Y, Zhao L, Lievense J, Covello PS, Keasling JD, Reiling KK, Renninger NS, Newman JD (2013) High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496:528–532
Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM, Ho KA, Eachus RA, Ham TS, Kirby J, Chang MCY, Withers ST, Shiba Y, Sarpong R, Keasling JD (2006) Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440:940–943
Shen Q, Lu X, Yan TX, Fu XQ, Lv ZY, Zhang FY, Pan QF, Wang GF, Sun XF, Tang KX (2016) The jasmonate-responsive AaMYC2 transcription factor positively regulates artemisinin biosynthesis in Artemisia annua. New Phytol 210:1269–1281
Shi P, Fu XQ, Liu M, Shen Q, Jiang WM, Li L, Sun XF, Tang KX (2017) Promotion of artemisinin content in Artemisia annua by overexpression of multiple artemisinin biosynthetic pathway genes. Plant Cell Tiss Org Cult. doi:10.1007/s11240-017-1173-z
Sy LK, Brown GD (2002) The mechanism of the spontaneous autoxidation of dihydroartemisinic acid. Tetrahedron 58:897–908
Teoh KH, Polichuk DR, Reed DW, Nowak G, Covello PS (2006) Artemisia annua L. (Asteraceae) trichome-specific cDNAs reveal CYP71AV1, a cytochrome P450 with a key role in the biosynthesis of the antimalarial sesquiterpene lactone artemisinin. FEBS Lett 580:1411–1416
Teoh KH, Polichuk DR, Reed DW, Covello PS (2009) Molecular cloning of an aldehyde dehydrogenase implicated in artemisinin biosynthesis in Artemisia annua. Botany 87:635–642
Toledoortiz G, Hqu E, Quail PH (2003) The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15:1749–1770
Urao T, Yamaguchi-Shinozaki K, Urao S, Shinozaki K (1993) An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5:1529–1539
Voinnet O, Rivas S, Mestre P, Baulcombe DC (2003) An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J 33:949–956
Wang H, Olofsson L, Lundgren A, Brodelius PE (2011) Trichome-specific expression of amorpha-4,11-Diene synthase, a key enzyme of artemisinin biosynthesis in Artemisia annua L., as reported by a promoter-GUS fusion. Am J Plant Sci 2:619–628
Wang H, Han J, Kanagarajan S, Lundgren A, Brodelius PE (2013) Trichome-specific expression of the amorpha-4,11-diene 12-hydroxylase (cyp71av1) gene, encoding a key enzyme of artemisinin biosynthesis in Artemisia annua, as reported by a promoter-GUS fusion. Plant Mol Biol 81(1–2):119–138
World Health Organization (2016) World malaria report 2015. WHO, Brussels
Wu W, Yuan M, Zhang Q, Zhu YM, Yong L, Wang W, Qi Y, Guo DJ (2011) Chemotype-dependent metabolic response to methyl jasmonate elicitation in Artemisia annua. Planta Med 77:1048–1053
Zhang YS, Teoh KH, Reed DW, Maes L, Goossens A, Olson DJH, Ross ARS, Covello PS (2008) The molecular cloning of artemisinic aldehyde ∆11 (13) reductase and its role in glandular trichome-dependent biosynthesis of artemisinin in Artemisia annua. J Biol Chem 283:21501–21508
Zhang FY, Fu XQ, Lv ZY, Lu X, Shen Q, Zhang L, Zhu MM, Wang GF, Sun XF, Liao ZH, Tang KX (2015) A basic leucine zipper transcription factor, AabZIP1, connects abscisic acid signaling with artemisinin biosynthesis in Artemisia annua. Mol Plant 8:163–175
Acknowledgements
This work was funded by the China National Transgenic Plant Research and Commercialization Project (Grant No. 2016ZX08002-001), China National High-Tech ‘‘863’’ Program (Grant No. 2011AA100605), the Shanghai Key Discipline Cultivation and Construction Project (Horticulture), and the Shanghai Jiao Tong University Agri-Engineering Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
He, Q., Fu, X., Shi, P. et al. Glandular trichome-specific expression of alcohol dehydrogenase 1 (ADH1) using a promoter-GUS fusion in Artemisia annua L.. Plant Cell Tiss Organ Cult 130, 61–72 (2017). https://doi.org/10.1007/s11240-017-1204-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-017-1204-9