Abstract
Aims
Tomato is an important vegetable plant worldwide. Previously, we isolated a soilborne Actinobacteria species, Streptomyces lydicus A01, which promotes the growth of tomato seedlings. The related mechanisms are needed to study.
Methods
RNA sequencing and gas chromatography–mass spectrometry were used reveal the global effect of S. lydicus A01 on tomato seedlings. Liquid chromatography–mass spectrometry was used to detect plant hormones in tomato seedlings during the growth-promotion procedure.
Results
After administration of S. lydicus A01, tomato seedlings exhibit more vigorous growth with more leaflets and improved photosynthesis. It was revealed that S. lydicus A01 has some effects on the basic metabolism of tomato seedlings. Meanwhile, the biosynthesis of secondary metabolites related to plant growth was regulated, including cutin, suberine, wax and flavonoid. Additionally, plant hormones including abscisic acid, salicylic acid and jasmonic acid were discovered to involve in the plant-growth-promoting activity of S. lydicus A01.
Conclusions
This study revealed some growth-promoting mechanisms of tomato seedlings by S. lydicus A01, laying the groundwork for further research.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
China is the largest tomato cultivator in the world, and tomatoes have become one of the most important vegetable crops in China. In 2015, tomato production was 55.94 million tons, accounting for 7.1% of the total output of vegetables in China (China 2017). Tomatoes are perennial herbs in the tropics, whereas they grow as annuals in temperate areas. According to previous research, 30 diseases threaten the tomato yield in China, including at least 10 diseases that are known to cause an obvious yield reduction (Li 2011). Currently, disease control in tomato production or the post-harvest process mainly relies on using chemical pesticides; however, pollution and pesticide residues are a public concern. Therefore, novel, environmentally friendly disease control strategies are urgently needed.
The existence of plant-growth-promoting rhizobacteria (PGPRs) was first reported in 1978 (Buff et al. 1978). Bacillus, Pseudomonas and Streptomyces spp. are the most common and widely used biocontrol and PGPR organisms (Adesemoye et al. 2008; Qin et al. 2015; Salla et al. 2014). As beneficial soil microorganisms, PGPRs facilitate plant growth both directly and indirectly (Chaiharn and Lumyong 2009; Mazhar et al. 2016; Saleem et al. 2007). For example, they are involved in biological nitrogen fixation (Mia and Shamsuddin 2010); pollutant degradation (Hou et al. 2015; Myresiotis et al. 2012); and induction of plant systemic resistance (Choudhary et al. 2007). Given these advantages, several PGPRs have been developed into potential biocontrol agents (Lewis and Papavizas 1991). Promotion of plant growth and resistance by Streptomyces spp. (Dias et al. 2017), as well as by the strain Streptomyces lydicus A01 used in the present study (Lu et al. 2008, and our unpublished data) have also been reported.
The mechanisms underlying the biocontrol activity of PGPRs are unclear. Some reports have focused on single or a specific category of metabolites related to plant growth promotion and inhibition (Ding et al. 2010; Mei et al. 2015; Sobolev et al. 2006). Other studies have investigated gene pathways that play an important role in plant growth and response to abiotic/biotic stressors (Stevenson et al. 2000; Tang et al. 2011; Vasconcelos et al. 2016; Xu and Zhang 2015). However, global changes in primary and secondary metabolism in plants during the growth-promotion process of rhizobacteria, and the corresponding gene expression, lack sufficient systematic research.
In the present study, omics research strategies, including high-throughput sequencing and large scale gas chromatography–mass spectrometry (GC-MS), were used to explore the plant growth-promoting activity of S. lydicus strain A01.
Materials and methods
Growth promotion experiments
S. lydicus A01 was isolated from the soil of a vegetable field in a suburb of Beijing by the Institute of Plant and Environment Protection, Beijing Academy of Agriculture and Forestry Sciences (IPEP-BAAFS), and has been deposited in the Chinese General Microbiological Collection Centre (Beijing, China) as strain CGMCC No.1653. Tomato seed Jiahong No. 4 was purchased from the Beijing Jing Yan Yi Nong Sci-Tech Development Center.
S. lydicus A01 was cultured on GAOs No. 1 agar plates (Ruan 1977) at 28 °C for 2 weeks. The mature spores were washed from the plate with sterilized distilled water and diluted into the suspension at 106 spore mL−1. Two milliliters of this suspension was inoculated into 100 mL fermentation broth (glucose 2%, peptone 0.6%, yeast powder 0.6% and NaCl 1%) on a rotary shaker (180 rpm) for 3 days at 28 °C (Lu et al. 2008). The broth was centrifuged at 6000 g for 5 min at 25 °C, after which the pseudomycelia pellet was collected.
Peat soil, vermiculite and perlite were mixed at a ratio of 1:1:0.5 (v:v:v). S. lydicus A01 pseudomycelia collected by centrifugation were also mixed with the soil at a ratio of 1:9 (w:w). Soil without S. lydicus A01 was designated as a control. Tomato seeds were germinated at 28 °C on Petri dishes containing moistened filter paper for 5 days. The plants were grown in soil (with or without S. lydicus A01 treatment) under a 10-h light/14-h dark cycle with a light intensity of 150–200 μmol m−2 s−1 and 60–70% humidity in a 28 °C phytotron for 21 days. Then, we measured the height, fresh weight, number of leaves and fresh weight of tomato seedlings, and each experiment was repeated three times.
Analysis of the effect of S. lydicus A01 on photosynthesis in tomato leaves
The net photosynthetic rate was measured on attached tomato leaves with a portable open-flow gas exchange system (LI-6400XT; LI-COR, Lincoln, NE, USA). Measurements were conducted from 10:00 to 13:00 h (solar time) under artificial PPFD at 1200 μmol photons m−2 s−1 at the leaf level and 400 μmol CO2 mol−1 air. During measurements, the leaf-to-air vapor pressure deficit was 1.0 kPa (Kang et al. 2012).
RNA sequencing detection of global effect of S. lydicus A01 on transcription in tomato leaves
On the 21st day after inoculation (DAI), the tomato leaves with or without S. lydicus A01 treatment were collected. Total RNA was extracted using RNAqueous™ phenol-free total RNA (Ambion, Austin, TX, USA), and the RIN number from an Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA) was used to assess RNA integrity. Qualified total RNA was further purified using an RNAClean XP Kit (Beckman Coulter, Kraemer Boulevard Brea, CA, USA) and an RNase-Free DNase Set (Qiagen, Hilden, Germany). Then, a cDNA library was built. Solexa sequencing of paired-ends (2 × 100) was performed, which produced at least 100 million reads per sample.
Conventional analysis was performed on the sequencing data obtained from the transcriptome samples, including data preprocessing, genomic mapping, gene expression analysis, transcript expression analysis, alternative splicing analysis, analysis of differential genes and GO/KEGG enrichment analysis between differentially and non-differentially expressed genes (Trapnell et al. 2009, 2010). The reference genome for transcriptome analysis was derived from Ensembl Plants (http://plants.ensembl.org/Solanum_lycopersicum/Info/Index). Changes were considered significantly down-regulated when the log2 fold change was <−1, and significantly up-regulated when log2 fold change was >1.
Specific analyses of the tricarboxylic acid (TCA) cycle, and cutin, suberine, wax and flavonoid biosynthesis pathways were conducted via Sol Genomics Network online blast searches with related reference genes, and the genes with the highest homology were selected for the analysis on the basis of RNA sequencing (RNA-seq) data and drawing diagrams (Bordag et al. 2015; Cheah et al. 2014; Omann and Zeilinger 2010; Shibayama et al. 2015; Zeilinger and Omann 2007).
GC-MS detection of global effect of S. lydicus A01 on primary metabolome of tomato leaves
At 21st DAI, the tomato leaves with or without S. lydicus A01 treatment were collected. Four volumes of pre-cooled 60% methanol at −50 °C were added to quench for 20 min, and the mixture was centrifuged at 4 °C at 6000 rpm for 5 min. The precipitate was collected, washed twice with sterile water at 4 °C and freeze-dried under a vacuum until all water was removed. After extraction and derivation, intracellular substances were analyzed by GC-MS (Agilent 7890A/5975C). The chromatography conditions were as follows: HP-5MS capillary column (5% phenyl methyl silox: 30 mx 250 μm id, 0.25 μm; Agilent J & W Scientific, Folsom, CA, USA); split injection, injection volume of 1 μL, split ratio of 20:1. The inlet temperature was 280 °C, the ion source temperature was 250 °C, and the interface temperature was 150 °C. The temperature program was started at 40 °C for 5 min, increased to 300 °C at 10 °C min−1 stepwise intervals, and held for 5 min. The carrier gas was helium, with a carrier gas flow rate of 1 mL min−1. The MS conditions were set as follows: electrospray ionization source, full scan mode, electron energy 70 eV and quadrupole scan range m/z 35–780.
Data obtained by comparison and analysis of the information derived from the samples were further analyzed using bioinformatics, which included pre-processing of data, identification of compounds and differential screening of compounds (Smith et al. 2006; Vanholme et al. 2012). Metabolites were annotated on the basis of metabolome databases from the NIST commercial database (Babushok et al. 2007) and Wiley 9 (Kanehisa and Goto 2000; Kopka et al. 2005). Changes were considered significant at P < 0.05 and VIP > 1.
Liquid chromatography (LC)-MS detection of the effect of S. lydicus A01 on plant hormones in tomato leaves
Leaves were collected from plants grown with or without S. lydicus A01, and ground with liquid nitrogen; 100 mg of sample was then added to a 2-mL centrifuge tube with 500 μL solution of normal propyl alcohol–water–concentrated hydrochloric acid (2:1:0.002, v:v:v). The mixture was placed at 4 °C for 30 min and was extracted with 1 mL methylene chloride at 4 °C for 30 min. After centrifugation at 4 °C at 5000 rpm for 10 min, 1 mL of the lower layer was collected. The sample was centrifuged and concentrated, re-dissolved with 200 μL 80% methanol, filtered with a 0.22-μm microporous filter and injected into the LC-MS. The chromatographic conditions were as follows: an ACQU ITY UPLC® BEH C18 column was used (2.1 × 100 mm, 1.7 μm; Waters, USA), the injection volume was 5 μL, the column temperature was 40 °C, mobile phase A was acetonitrile, and mobile phase B was water with 10 mM ammonium acetate. The gradient elution conditions were 0–2 min, 10–25% A; 2–3 min, 25% A; 3–4 min, 25–90% A; 4–5 min, 90% A; 5–6 min, 90–10% A; 6–8 min, 10% A. The MS conditions were as follows: electrospray ionization source, positive and negative ion ionization mode, ion source temperature of 500 °C, ion source voltage of 5000 V/−4500 V, collision gas at 6 psi, air curtain gas at 30 psi, and atomization and auxiliary gases at 50 psi. Multipath monitoring was used for scanning (Bajad et al. 2006). The linearity of measurement was evaluated by analyzing different concentrations of the standard solutions of the abscisic acid (ABA), indoleacetic acid, gibberellic acid, salicylic acid (SA) and jasmonic acid (JA). Calibration standards were prepared by diluting the stock solutions to obtain the concentrations indicated in Supplementary Table 5.
Identification of important node genes by qRT-PCR
Representative genes were validated by qRT-PCR in a Roche LightCycler 480. The primers used are listed in Supplementary Table S6 (Du et al. 2014). The qRT-PCR system with a final volume of 20 μL was set up as follows: 10 μL 2 × SYBR Green PCR buffer, 0.5 μL 10 μM forward primer, 0.5 μL 10 μM reverse primer, 5 ng cDNA and deionized distilled water to a volume of 20 μL. The qRT-PCR amplification conditions were as follows: 50 °C for 2 min; 95 °C for 10 min; and 95 °C for 15 s, and 60 °C for 1 min, for a total of 40 cycles. Data analysis was conducted following the method outlined in the Bio-Rad quantitative PCR Application Guide 2−ΔΔCT (Livak and Schmittgen 2001).
Statistical analysis
Data are presented as mean ± SD. We used Student’s t test to examine the statistical significance between the two comparison groups, which was defined as P < 0.05 or P < 0.01, as indicated.
Results
Effect of S. lydicus A01 on the growth and photosynthetic rate of tomato seedlings
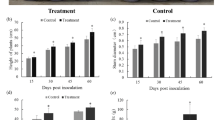
At 21st DAI, the height of the plants treated with S. lydicus A01 was 1.52 times higher than that of the controls (22.33 cm vs. 14.67 cm), and the fresh weight was 2.68 times higher than that of the controls (1.34 g vs. 0.50 g) (Fig. 1a–c). The leaves from treated plants were greener and had more vigor. The leaf number of the treated plants was twice that of the controls (12 vs. 6); the fresh weight was 2.96 times that of the controls (0.54 g vs. 0.18 g); and the total photosynthetic rate was 2.28 times that of the controls (114.62 μmol CO2 m−2 s−1 vs. 50.22 μmol CO2 m−2 s−1) (Fig. 1d–f).
Effect of S. lydicus A01 on tomato seedlings. a Photograph of 21-day-old plants. Left, A01-treated plants; right, controls; seedling height (b), fresh weight (c), leaf numbers (d) measured in control and A01-treated plants. Mean ± SD (n = ≥20 seedlings per group). e Fresh weight of tomato leaves. f Photosynthetic rate of tomato leaves; all leaves from each plant were measured. Mean ± SD (n = ≥20 seedlings per group). ** P < 0.01 by Student’s t test
Global effects of S. lydicus A01 on tomato leaves, determined by RNA-seq and GC-MS analysis
A total of 24,056 different transcripts, corresponding to ~71% of the reference transcripts (n = 33,810), were detected in tomato leaves. Compared with the tomatoes grown in control soil, after tomatoes were grown in soil mixed with S. lydicus A01 for 21 days, 986 genes were up-regulated, and 1239 were down-regulated. The proportion was 9.25% (Fig. 2a). The transcriptome data have been deposited in the Sequence Read Archive under accession numbers SRR6392273 and SRR6392274. Seventy-five compounds were identified from tomato leaves by GC-MS, including 19 amino acids, three amines, seven sugars, nine fatty acids, 26 organic acids, two phosphoric acids, six polyols and three compounds from other categories. Comprehensive analysis showed that, compared with the controls, the tomato seedlings grown with S. lydicus A01 for 21 days had 23 compounds that were significantly changed. Their proportion was 30.67% (Fig. 2b).
Effect of S. lydicus A01 on tomato transcriptome and metabolome. a Volcano plot representing the comparison of significantly up-regulated and down-regulated genes in leaves of 21-day-old tomatoes, determined by RNA-seq. Red points represent up-regulation, blue points represent down-regulation, and gray points represent no difference. b Metabolomic profiles and hierarchical clustering of metabolites in A01-treated and control plants (21 days old). Six biological replicates per group are shown. Red and blue indicate high and low levels, respectively
Effect of S. lydicus A01 on basic metabolism in tomato leaves, determined by RNA-seq and GC-MS analysis
The TCA cycle is the basis for biological metabolism and is the hub of carbohydrate, fatty acid and amino acid metabolic pathways; therefore, the corresponding RNA-seq data of tomato leaves were analyzed. In the S. lydicus A01-treated plants compared with the controls, only the malate dehydrogenase gene, but not the rate-limiting enzyme gene, changed significantly. GC-MS showed that the yields of citric acid, succinate, fumaric acid and malic acid did not significantly change between the S. lydicus A01-treated and control groups. Comprehensive analysis of RNA-seq and GC-MS data showed that S. lydicus A01 did not significantly affect the TCA cycle in tomato leaves (Fig. 3a and Supplementary Tables S1 and S2).
Effect of S. lydicus A01 on TCA cycle and related primary metabolic pathways in tomato leaves, determined by RNA-seq and GC-MS. a Effect of S. lydicus A01 on the genes and compounds in the TCA cycle. b Effect of S. lydicus A01 on sugar. c Effect of S. lydicus A01 on fatty acids. d Effect of S. lydicus A01 on amino acids. Red represents significantly up-regulated genes, blue represents significantly down-regulated genes, and gray represents genes with no significant change. Purple represents a rate-limiting step
RNA-seq analysis showed that 16 genes were significantly up-regulated in starch, sucrose and carbon metabolism, and 14 genes were significantly down-regulated (Table 1). Five genes were significantly up-regulated in fatty acid biosynthesis and degradation pathways (Table 1). In leaves from tomato seedlings treated with S. lydicus A01, among the amino acid metabolic pathways, 24 genes were significantly up-regulated and 16 were significantly down-regulated (Table 1).
GC-MS showed that one sugar (maltose) was significantly down-regulated (Fig. 3b and Supplementary Table S2). Three fatty acids (eicosanoic acid, docosanoic acid and tetracosanoic acid) were up-regulated (Fig. 3c and Supplementary Table S2). One amino acid (glutamate) was significantly up-regulated, and three (leucine, tyrosine and methionine) were significantly down-regulated (Fig. 3d and Supplementary Table S2).
Comprehensive analysis of the above RNA-seq and GC-MS data showed that S. lydicus A01 did not significantly affect the TCA cycle, but did affect related carbohydrate, lipid and amino acid metabolism in tomato leaves.
Effect of S. lydicus A01 on cutin, suberine and wax biosynthesis pathways, determined by RNA-seq and qRT-PCR analysis
RNA-seq analysis showed that in the cutin, suberine and wax biosynthesis pathways, all the significantly changed genes were up-regulated, including three genes related to the biosynthesis of unsaturated fatty acids, two genes encoding fatty acid-ω-hydroxylase (FAH), which is responsible for the metabolism of long-chain fatty acids to cutin and suberine; three genes encoding alcohol-forming fatty acyl-CoA reductase (FAR), which is responsible for the metabolism of long-chain acyl-CoA to wax; and one ω-hydroxypalmitate O-feruloyl transferase gene (Fig. 4a and Supplementary Table S3). FAR-c and FAH-a genes were detected by qRT-PCR, and the results were consistent with the RNA-seq results (Fig. 4b).
Effect of S. lydicus A01 on the flavonoid biosynthesis pathway, determined by RNA-seq and qRT-PCR analysis
RNA-seq data showed that in the flavonoid biosynthesis pathway, all the significantly changed genes were down-regulated or not expressed, including two chalcone synthase (CHS) genes, one chalcone isomerase (CHI) gene, one flavonoid 3′-monooxygenase (F3’M), two flavanone 4-reductases (F4R-a and F4R-b), one leucoanthocyanidin dioxygenase (LDOX), naringenin 3-dioxygenase (N3D), one flavonoid 3′,5′-hydroxylase gene (F3’5’H), one flavonoid 3′-hydroxylase gene (F3’H), one flavonoid 3-hydroxylase gene (F3H) and one flavonol synthase gene (FLS; Fig. 5a and Supplementary Table S4). CHS-b, LDOX and N3D genes were detected by qRT-PCR, and the results were consistent with the results of RNA-seq (Fig. 5b).
Effect of S. lydicus A01 and other metabolic pathways related to growth in tomato leaves
As well as reprogramming the primary and secondary metabolism pathways mentioned above, S. lydicus A01 also significantly affected many other pathways, such as phenylpropanoid, stilbenoid, diarylheptanoid, gingerol, carotenoid and brassinosteroid biosynthesis; aminobenzoate, limonene, pinene and bisphenol degradation; and linoleic acid, taurine and hypotaurine metabolism. These pathways are all related to tomato growth (Table 2).
Effect of S. lydicus A01 on plant hormones in tomato leaves
The content of ABA in leaves from S. lydicus A01-treated tomato plants (108.80 ng g−1 fresh weight) was 2.35 times that of the controls (46.20 ng g−1 fresh weight). The content of SA and JA in leaves of tomato plants treated with S. lydicus A01 was 2.45-fold (480 ng g−1 fresh weight) and 5.24-fold (1.19 ng g−1 fresh weight) lower than that in the controls (1180.0 ng g−1 and 6.22 ng g−1 fresh weight), respectively. Indoleacetic acid and gibberellic acid were not detected in either the S. lydicus A01-treated tomato leaves or the controls (Fig. 6).
Discussion
Treatment with S. lydicus A01 markedly affected the growth variables in tomato seedlings, including plant height, fresh weight, leaf number and photosynthetic rate. Numerous reports have shown the growth promotion effects of biocontrol agents (Adesemoye et al. 2008; Qin et al. 2015; Salla et al. 2014), including Streptomyces spp. (Dias et al. 2017); however, limited studies have investigated global variations in plants during their growth. To our knowledge, this is the first study to reveal the effect of S. lydicus on tomato leaves during the growth promotion process, including many pathways related to primary and secondary metabolism.
The TCA cycle is the basis for biological metabolism and is the hub of sugar, fatty acid and amino acid metabolism pathways (Bordag et al. 2015; Cheah et al. 2014). Comprehensive analysis of the RNA-seq and GC-MS results showed that S. lydicus A01 had scarcely affected on the TCA cycle, but led to significant down-regulation of maltose, leucine, tyrosine and methionine, and significant up-regulation of glutamate, eicosanoic acid, docosanoic acid and tetracosanoic acid in tomato leaves. We hypothesize that treatment with S. lydicus A01 plays roles in the cycling of maltose and results in a significant decrease in maltose content. The increase/decrease of these amino acids also seems to be modulated by S. lydicus A01 treatment.
Biosynthesis of cutin, suberine and wax is related to plant growth promotion (Yeats and Rose 2013). Unsaturated fatty acids are related to cutin, suberine and wax biosynthesis. RNA-seq and qRT-PCR analyses showed that soil mixed with S. lydicus A01 significantly increased expression of genes related to the synthesis of unsaturated fatty acids, as well as genes involved in the synthesis of cutin, suberine and wax in tomato leaves. The plant cuticle, including cutin, suberine and wax, is an extracellular hydrophobic layer that covers the aerial epidermis of all land plants, providing protection against desiccation and external environmental stressors. The physiological functions of the cuticle also extend beyond its primary function as a transpiration barrier, including important roles in processes including development and interaction with microbes (Yeats and Rose 2013).
The phenylpropanoid pathway is responsible for the biosynthesis of a variety of metabolites including flavonoids and lignin. Flavonoid accumulation in plants affects photosynthesis, auxin transport and plant growth (Besseau et al. 2007; Morales-Flores et al. 2015). Our data revealed that S. lydicus A01 significantly decreased the mRNA levels of flavonoid-synthesis-related genes (RNA-seq and qRT-PCR analysis), thus indicating that the flavonoid levels were decreased and confirming the negative effects of flavonoids on normal plant growth. The plasticity of flavonoid pathways and their ability depend on the demands of the local ecosystem to redirect the flows of intermediate molecules toward biosynthesis of the complex scaffolds of different chemicals (Mouradov and Spangenberg 2014). The decrease in flavonoids in this study was related to plant growth inhibition but not the bio-stress response. Moreover, genes involved in lignin catabolic and metabolic processes were also induced, thus increasing the mechanical strength of tomato seedlings (data not shown). Similar results have shown that silencing of CHS in strawberries leads to the loss of pigmentation, which is accompanied by a significant increase in the content of lignin (Ring et al. 2013).
S. lydicus A01 affects many other secondary metabolic pathways associated with plant growth, for example, increased biosynthesis of stilbenoids, which are usually produced by peanut plants as a defense response to fungal invasion and other exogenous stimuli (Sobolev et al. 2006), as well as the defense response to fungal pathogens in grapes (Chitarrini et al. 2017). p-Aminobenzoic acid (pABA) plays important roles in a wide variety of metabolic processes, including plant growth regulatory properties (Crisan et al. 2014). Here, we demonstrated specific inhibition of ABA biosynthesis. A similar result (controlled by rubreserine) has been reported to induce growth limitation in plants (Camara et al. 2012).
With both the RNA-seq results and plant phenotype measurements, we demonstrated that the control plants were under stress conditions, especially nutritional deficiencies. Therefore, we used LC-MS to detect the plant hormone ABA, which is important for mediating abiotic stress responses and plays a multifaceted and essential role in plant immunity (Cao et al. 2011). S. lydicus A01 significantly enhanced the level of ABA, thus indicating that the ABA-mediated stress response was induced. In contrast, we observed a decrease in JA and SA levels in plants, which is related to plant resistance to disease. The co-expression pattern of ABA, JA and SA may indicate the crosstalk among them. Specifically, SA- and JA-dependent signaling antagonizes ABA biosynthesis and signaling pathways during the early stages of stress (Gupta et al. 2017). In the present study, although the seedlings were grown in the same soil environment, S. lydicus A01 conferred upon the seedlings some adaptability to counter the stress.
Conclusions
In this study, it was found that S. lydicus A01 could promote the growth and photosynthetic rate of tomato seedling and led to secondary metabolite changes related to plant growth, including cutin, suberine, wax and flavonoid. Additionally, plant hormones including abscisic acid, salicylic acid and jasmonic acid were also discovered to involve in the plant-growth-promoting activity of S. lydicus A01.
References
Adesemoye AO, Obini M, Ugoji EO (2008) Comparison of plant growth-promotion with Pseudomonas aeruginosa and Bacillus subtilis in three vegetables. Braz J Microbiol 39:423–426
Babushok VI, Linstrom PJ, Reed JJ, Zenkevich IG, Brown RL, Mallard WG, Stein SE (2007) Development of a database of gas chromatographic retention properties of organic compounds. J Chromatogr A 1157:414–421
Bajad SU, Lu W, Kimball E, Yuan J, Peterson CN, Rabinowitz JD (2006) Separation and quantitation of water soluble cellular metabolites by hydrophilic interaction chromatography-tandem mass spectrometry. J Chromatogr A 1125:76–88
Besseau S, Hoffmann L, Geoffroy P, Lapierre C, Pollet B, Legrand M (2007) Flavonoid accumulation in Arabidopsis repressed in lignin synthesis affects auxin transport and plant growth. Plant Cell 19:148–162
Bordag N, Klie S, Jurchott K, Vierheller J, Schiewe H, Albrecht V, Tonn JC, Schwartz C, Schichor C, Selbig J (2015) Glucocorticoid (dexamethasone)-induced metabolome changes in healthy males suggest prediction of response and side effects. Sci Rep 5:15954
Buff TJ, Schroth MN, Suslow T (1978) Increased potato yields by treatment of seed pieces with specific strains of Pseudomonas fluorescens and P. putida. Phytopathology 68:1377–1383
Camara D, Bisanz C, Barette C, Van Daele J, Human E, Barnard B, Van der Straeten D, Stove CP, Lambert WE, Douce R, Marechal E, Birkholtz LM, Cesbron-Delauw MF, Dumas R, Rebeille F (2012) Inhibition of p-aminobenzoate and folate syntheses in plants and apicomplexan parasites by natural product rubreserine. J Biol Chem 287:22367–22376
Cao FY, Yoshioka K, Desveaux D (2011) The roles of ABA in plant-pathogen interactions. J Plant Res 124:489–499
Chaiharn M, Lumyong S (2009) Phosphate solubilization potential and stress tolerance of rhizobacteria from rice soil in Northern Thailand. World J Microbiol Biotechnol 25:305–314
Cheah HL, Lim V, Sandai D (2014) Inhibitors of the glyoxylate cycle enzyme ICL1 in Candida albicans for potential use as antifungal agents. PLoS One 9:e95951
China MOA (2017) Tomato 2016 market analysis and 2017 market forecast
Chitarrini G, Soini E, Riccadonna S, Franceschi P, Zulini L, Masuero D, Vecchione A, Stefanini M, Di Gaspero G, Mattivi F, Vrhovsek U (2017) Identification of biomarkers for defense response to Plasmopara viticola in a resistant grape variety. Front Plant Sci 5:1524
Choudhary DK, Prakash A, Johri BN (2007) Induced systemic resistance (ISR) in plants: mechanism of action. Indian J Microbiol 47(4):289–297
Crisan ME, Bourosh P, Maffei ME, Forni A, Pieraccini S, Sironi M, Chumakov YM (2014) Synthesis, crystal structure and biological activity of 2-hydro-xyethylammonium salt of p-Aminobenzoic Acid. PLoS One 9(7):e101892
Dias MP, Bastos MS, Xavier VB, Cassel E, Astarita LV, Santarém ER (2017) Plant growth and resistance promoted by Streptomyces spp. in tomato. Plant Physiol Biochem 118:479–493
Ding L, Jing H, Qin B, Qi L, Li J, Wang T, Liu G (2010) Regulation of cell division and growth in roots of Lactuca sativa L. seedlings by the Ent-Kaurene diterpenoid rabdosin B. J Chem Ecol 36:553–563
Du M, Zhai Q, Deng L, Li S, Li H, Yan L, Huang Z, Wang B, Jiang H, Huang T, Li C, Wei J, Kang L, Li J, Li C (2014) Closely related NAC transcription factors of tomato differentially regulate stomatal closure and reopening during pathogen attack. Plant Cell 26:3167–3184
Gupta A, Hisano H, Hojo Y, Matsuura T, Ikeda Y, Mori IC, Senthil-Kumar M (2017) Global profiling of phytohormone dynamics during combined drought and pathogen stress in Arabidopsis thaliana reveals ABA and JA as major regulators. Sci Rep 7:4017
Hou J, Liu W, Wang B, Wang Q, Luo Y, Franks AE (2015) PGPR enhanced phytoremediation of petroleum contaminated soil and rhizosphere microbial community response. Chemosphere 138:592–598
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kang Z, Li G, Huang J, Niu X, Zou H, Zang G, Wenwen Y, Wang G (2012) Photosynthetic and physiological analysis of the rice high-chlorophyll mutant (Gc). Plant Physiol Biochem 60:81–87
Kopka J, Schauer N, Krueger S, Birkemeyer C, Usadel B, Bergmuller E, Dormann P, Weckwerth W, Gibon Y, Stitt M, Willmitzer L, Fernie AR, Steinhauser D (2005) GMD@CSB.DB: the Golm metabolome database. Bioinformatics 21:1635–1638
Lewis JA, Papavizas GC (1991) Biocontrol of plant diseases: the approach for tomorrow. Crop Prot 10:95–105
Li J (2011) Chinese tomato breeding. China Ariculture Press, Beijing
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25:402–408
Lu CG, Liu WC, Qiu JY, Wang HM, Liu T, De Liu W (2008) Identification of an antifungal metabolite produced by a potential biocontrol Actinomyces strain A01. Braz J Microbiol 39:701–707
Mazhar R, Ilyas N, Raja NI, Saeed M, Hussain M, Seerat W, Qureshi H, Shabir S (2016) Plant growth-promoting rhizobacteria: biocontrol potential for pathogens. Pure Appl Biol 5(4):1288–1295
Mei C, Michaud M, Cussac M, Albrieux C, Gros V, Marechal E, Block MA, Jouhet J, Rebeille F (2015) Levels of polyunsaturated fatty acids correlate with growth rate in plant cell cultures. Sci Rep 5:15207
Mia MAB, Shamsuddin ZH (2010) Nitrogen fixation and transportation by rhizobacteria: a scenario of rice and banana. Int J Bot 6:235–242
Morales-Flores F, Olivares-Palomares KS, Aguilar-Laurents MI, Rivero-Cruz JF, Lotina-Hennsen B, King-Díaz B (2015) Flavonoids affect the light reaction of photosynthesis in vitro and in vivo as well as the growth of plants. J Agric Food Chem 63:8106–8115
Mouradov A, Spangenberg G (2014) Flavonoids: a metabolic network mediating plants adaptation to their real estate. Front Plant Sci 5:620
Myresiotis CK, Vryzas Z, Papadopoulou-Mourkidou E (2012) Biodegradation of soil-applied pesticides by selected strains of plant growth-promoting rhizobacteria (PGPR) and their effects on bacterial growth. Biodegradation 23(2):297–310
Omann M, Zeilinger S (2010) How a mycoparasite employs g-protein signaling: using the example of trichoderma. J Signal Transduct 2010:123–126
Qin Y, Han Y, Shang Q, Li P (2015) Complete genome sequence of Bacillus amyloliquefaciens L-H15, a plant growth promoting rhizobacteria isolated from cucumber seedling substrate. J Biotechnol 200:59–60
Ring L, Yeh SY, Huecherig S, Hoffmann T, Blanco-Portales R, Fouche M, Villatoro C, Denoyes B, Monfort A, Caballero JL, Muñoz-Blanco J, Gershenson J, Schwab W (2013) Metabolic interaction between anthocyanin and lignin biosynthesis is associated with peroxidase FaPRX27 in strawberry fruit. Plant Physiol 163:43–60
Ruan JS (1977) The basis of taxonomy of actinomycetes. The Chinese Academic Press, Beijing, pp 139–146
Saleem M, Arshad M, Hussain S, Bhatti AS (2007) Perspective of plant growth promoting rhizobacteria (PGPR) containing ACC deaminase in stress agriculture. J Ind Microbiol Biotechnol 34:635–648
Salla T D, da Silva R, Astarita L V and Santarém E R (2014) Streptomyces rhizobacteria modulate the secondary metabolism of Eucalyptus plants. Plant Physiol Biochem 85: 14–20
Shibayama J, Yuzyuk TN, Cox J, Makaju A, Miller M, Lichter J, Li H, Leavy JD, Franklin S, Zaitsev AV (2015) Metabolic remodeling in moderate synchronous versus dyssynchronous pacing-induced heart failure: integrated metabolomics and proteomics study. PLoS One 10:e118974
Smith CA, Want EJ, O'Maille G, Abagyan R, Siuzdak G (2006) XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem 78:779–787
Sobolev VS, Horn BW, Potter TL, Deyrup ST, Gloer JB (2006) Production of stilbenoids and phenolic acids by the peanut plant at early stages of growth. J Agric Food Chem 54:3505–3511
Stevenson JM, Perera IY, Heilmann I, Persson S, Boss WF (2000) Inositol signaling and plant growth. Trends Plant Sci 5:252–258
Tang W, Yuan M, Wang R, Yang Y, Wang C, Osesprieto JA, Kim T, Zhou H, Deng Z, Gampala SS (2011) PP2A activates brassinosteroid-responsive gene expression and plant growth by dephosphorylating BZR1. Nat Cell Biol 13:124–131
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25:1105–1111
Trapnell C, Williams B A, Pertea G, Mortazavi A, Kwan G, van Baren M J, Salzberg S L, Wold B J and Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28: 511–515
Vanholme R, Storme V, Vanholme B, Sundin L, Christensen JH, Goeminne G, Halpin C, Rohde A, Morreel K, Boerjan W (2012) A systems biology view of responses to lignin biosynthesis perturbations in Arabidopsis. Plant Cell 24:3506–3529
Vasconcelos MW, Menguer PK, Hu Y, Revers LF, Sperotto RA (2016) Stress signaling responses in plants. Biomed Res Int 2016:4720280
Xu J, Zhang S (2015) Mitogen-activated protein kinase cascades in signaling plant growth and development. Trends Plant Sci 20:56–64
Yeats TH, Rose JK (2013) The formation and function of plant cuticles. Plant Physiol 163:5–20
Zeilinger S, Omann M (2007) Trichoderma biocontrol: signal transduction pathways involved in host sensing and mycoparasitism. Gene Regul Syst Bio 1:227–234
Acknowledgements
This work was supported by Beijing Municipal Science and Technology Plan Projects (No. D151100003915002), Beijing Leafy Vegetables Innovation Team of Modern Agro-industry Technology Research System (BAIC07-2018), Science and Technology Innovation Ability Construction of BAAFS (No. KJCX20170415, No. KJCX20170107), Beijing Key Laboratory of Environment Friendly Management on Fruit Diseases and Pests in North China (No. BZ0432), National Key Research and Development Program of China (No. 2017YFD0200403, No. 2017YFD0201108), National Natural Science Foundation of China (No. 31270155), Science and Technology Innovation Team of Beijing Academy of Agriculture and Forestry Sciences (No. JNKST201607), and the National Special Project of Basic Work Project for Science and Technology (No. 2014FY120900). We sincerely thank Suzhou BioNovoGene Metabolomics Platform for the assistance in plant hormone detection with LC-MS and correlation analysis of GC-MS and RNA-seq data.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interests
The authors declare no competing financial interests.
Additional information
Responsible Editor: Hans Lambers.
Qiong Wu and Mi Ni are co-first authors
Rights and permissions
About this article
Cite this article
Wu, Q., Ni, M., Liu, WC. et al. Omics for understanding the mechanisms of Streptomyces lydicus A01 promoting the growth of tomato seedlings. Plant Soil 431, 129–141 (2018). https://doi.org/10.1007/s11104-018-3750-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-018-3750-2