Abstract
Aims
The goal of this study was to investigate the structure and functional potential of microbial communities associated with healthy and diseased tomato rhizospheres.
Methods
Composition changes in the bacterial communities inhabiting the rhizospheric soil and roots of tomato plants were detected using 454 pyrosequencing. Microbial functional diversity was investigated with BIOLOG technology.
Results
There were significant shifts in the microbial composition of diseased samples compared with healthy samples, which had the highest bacterial diversity. The predominant phylum in both diseased and healthy samples was Proteobacteria, which accounted for 35.7–97.4 % of species. The class Gammaproteobacteria was more abundant in healthy than in diseased samples, while the Alphaproteobacteria and Betaproteobacteria were more abundant in diseased samples. The proportions of pathogenic Ralstonia solanacearum and Actinobacteria species were also elevated in diseased samples. The proportions of the various bacterial populations showed a similar trend both in rhizosphere soil and plant roots in diseased versus disease-free samples, indicating that pathogen infection altered the composition of bacterial communities in both plant and soil samples. In terms of microbial activity, functional diversity was suppressed in diseased soil samples. Soil enzyme activity, including urease, alkaline phosphatase and catalase activity, also declined.
Conclusions
This is the first report that provides evidence that R. solanacearum infection elicits shifts in the composition and functional potential of microbial communities in a continuous-cropping tomato operation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Various studies have proven that microbial communities sustain soil ecosystems, and a wide range of microorganisms is involved in important soil functions. Some rhizosphere-inhabiting microbes are known to participate in detrimental interactions with plants that lead to the development of root diseases, while others are clearly beneficial to the plants by stimulating their growth (Bulgarelli et al. 2012), enhancing nutrient uptake, or playing a significant role in controlling soil-borne pathogens and the diseases they cause (Bais et al. 2006; Lugtenberg and Kamilova 2009; Trivedi et al. 2012). Plant and soil conditions play important roles in regulating microbial communities as well (Lundberg et al. 2012). Plants exude up to 21 % of their photosynthetically fixed carbon into the rhizosphere, thereby leading to significant changes in microbial diversity and function in the soil (Marschner et al. 2001; Baudoin et al. 2003). It has been postulated that plants actively recruit beneficial soil microorganisms in their rhizospheres to counteract pathogen assault (Cook et al. 1995). It has frequently been reported that plants limit the diversity of microbial communities in the rhizosphere (Rosario et al. 2007). The interplay between plant and microbial communities has long been thought to be one of key mechanisms that suppress soil-borne pathogens. In-spite of the tremendous progress made in the field of microbial ecology in recent years, our knowledge on the community structure of the rhizospheric microbial community and their principal functions is still its infancy (Trivedi et al. 2012).
Soil-borne pathogens of plants cause various diseases in crops, including root and crown rots, vascular wilting, take-all and damping-off. Pathogens thus pose a substantial threat to food production. Numerous studies have focused on the crop-defense systems against plant pathogens, the role of plant-beneficial bacteria in the control of plant pathogens (Harvell et al. 1999; Montesinos et al. 2002; Lugtenberg and Kamilova 2009), and the effects of plant pathogens on the structure of plant-associated bacterial communities (McSpadden Gardener and Weller 2001; Yang et al. 2001; Reiter et al. 2002; Trivedi et al. 2010, 2011). Although it has been postulated that the disruption of multitrophic interactions in a stable ecosystem under the influence of invading phytopathogens could cause community reorganization leading to massive collapse and severe degradation of soil ecosystems (van der Putten et al. 2007), the implications of shifts in diversity for ecosystem function are not well understood. The long-term sustainability of ecosystem productivity requires detailed knowledge on its biodiversity coupled with profound understanding of its function. Determining the functional role of complex and dynamic microbial communities is challenging because of our inability to culture the vast majority of microorganisms. For example, it is widely accepted that up to 99 % of soil microorganisms remains unculturable, and it a great challenge to elucidate the functional changes of microbial communities in the crop rhizosphere when infected by soil-borne pathogens (Nannipieri et al. 2003; Fitter et al. 2005). Recently developed metagenomic technologies, such as high-throughput 454 pyrosequencing, have revolutionized microbial research and provided a powerful fingerprinting strategy with unprecedented resolution. Massively parallel pyrosequencing allows for the detection of microbial communities as a whole, overcoming the inherent bias of culture-dependent and traditional molecular techniques (Glenn 2011; Diaz et al. 2012; Ye and Zhang 2013).
Bacterial wilt caused by Ralstonia solanacearum is one of the most devastating plant diseases worldwide. R. solanacearum affects a wide range of plants in more than 50 families (Hayward 1995). In China, its hosts include economically important crops such as tomato (Lycopersicon esculentum Miller), potato (Solanum tuberosum L.), tobacco (Nicotiana tabacum L.), eggplant (Solanum melongena L.), pepper (Capsicum annuum L.), peanut (Arachis hypogaea L.) and ginger (Zingiber officinale Roscoe). This disease has caused great losses in agriculture and horticulture. Bacterial wilt control approaches, including field sanitation, crop rotation and application of resistant varieties, have been tried with limited success (Ciampi-Panno et al. 1989). The application of chemical pesticides is generally considered to be the most effective and fastest strategy for plant disease management. However, no effective chemical product is available for bacterial wilt. Although streptomycin is a suitable bactericide for the control of this disease, Chinese farmers are reluctant to use it because large dosages are required for the expected effectiveness, probably due to bacterial resistance that has developed as a result of the repeated and abusive use of streptomycin for the last several decades in this country. Therefore, more effort must be devoted to biological control with living microbes.
Vegetables grown in greenhouses are more susceptible to diseases due to optimal growth conditions, including temperature, humidity, salinity and tillage (Abawi and Widmer 2000). We have established a greenhouse in which tomato plants have been grown continuously for 3 years. Some of the plants in our greenhouse exhibited diseased symptoms characteristic of Ralstonia solanacearum infection. The goal of this study was to understand the consequences on microbial community composition and their associated functions resulting from stress caused by the invading pathogen using the R. solanacearum-tomato model, and to elucidate microbial mechanisms of continuous cropping by deciphering the changes in function and diversity of rhizospheric microbial communities in healthy and diseased soil ecosystems.
Materials and methods
Site description and sample collection
Tomato plants (cv. Shanghai 903) were grown in a vegetable greenhouse located in the Suzhou city, Jiangsu province, China (36°26′N, 120°27′E). Plants were continuously cultivated for 3 years in the same field. Some plants in certain sections showed wilt symptoms associated with infection by R. solanacearum, while others were healthy, although all plants received the same management. The rhizospheric soil and plant root samples were collected separately from three diseased and three disease-free sites based on visual symptoms, including diseased roots, healthy roots, rhizospheric soil from diseased tomatoes (disease RS), and rhizospheric soil from healthy tomatoes (healthy RS) at the flowering stage in June 2011. Each rhizosphere sample consisted of the total root system with tightly adhering sediment from each individual plant. Roots were shaken gently to remove the loosely adhering soil. Rhizosphere soil was carefully removed from fine roots by gently scraping adhering soil using fine forceps. Bulk soil samples around the infected and healthy roots were also collected in triplicate for BIOLOG analysis and the detection of soil enzyme activity. Each soil sample was stored at 4 °C until it was brought to the lab for immediate processing. Soil samples were homogenized by passing them through a <2 mm sieve to remove aboveground plant materials, roots, and stones for downstream processing, including microbial genomic DNA extraction, pyrosequencing, isolation of bacterial species, determination of soil enzymatic activity, and microbial function. Healthy and diseased roots were also collected for DNA extraction from endophytic microorganisms.
Determination of R. solanacearum populations in soil and plant
Samples with three replicates were carefully collected to harvest R. solanacearum cells from rhizospheric soil and root samples of both healthy and diseased tomato using the semi-selection medium, SMSA, according to a previously described procedure (French et al. 1995; Kang et al. 2004). For plants samples, roots were washed under running water, surface-sterilized by immersion in 70 % ethanol for a few seconds followed by treatment in 0.1 % (w/v) mercuric chloride for 10 min, and then cut into fragments 0.5 cm long with a sterile knife. Fragments were placed in a sterilized mortar containing 0.5 ml of sterile distilled water and ground with a sterilized pestle. The resulting suspension was plated onto SMSA medium. For soil samples, rhizosphere soil was transferred to a sterilized Erlenmeyer flask and suspended in a sterilized 0.85 % NaCl solution at a ratio of 1:9 (v/v). All soil suspensions were incubated at 28 °C on a rotary shaker at 150 rpm for 30 min and then allowed to settle for 5 min. The resulting supernatant from each soil suspension was serially diluted with a sterilized 0.85 % NaCl solution, and 2 ml of each dilution was subsequently plated onto SMSA medium. Three plates were used for each sample. The plates were incubated at 28 °C for 2 days, and the number of colonies was then counted.
Soil and plant DNA extraction
Genomic DNA was extracted from 0.5 g of each soil sample using a FastDNA SPIN Kit for soil (MP Biomedicals, Santa Ana, CA) according to the manufacturer’s instructions. The extracted soil DNA was dissolved in 70 μL of TE buffer, quantified by spectrophotometry and stored at −20 °C for further analysis. To isolate genomic DNA from endophytic microbial communities, tomato roots were washed under running water, and surface-sterilized by immersion in 70 % ethanol (60 s) followed by treatment in 0.1 % (w/v) mercuric chloride for 10 min. Surface sterilization of the plant material was checked by rolling the sterilized plant material on nutrient agar plates, which were then incubated for up to 7 days at 28 °C. Root Genomic DNA was isolated using a modified CTAB method as previously described in greater detail (Kidwell and Osborn 1992).
High-throughput pyrosequencing
Bidirectional pyrosequencing was performed with a Roche 454 GS FLX Titanium sequencer by analyzing the V4 region of the 16S rRNA gene at the whole microbial community level as previously described Xia et al. (2011) with slight modifications. PCR-amplifying the 16S rRNA genes with the universal primers 515F and 907R were extended as amplicon fusion primers using the respective A and B primer adapters, a key sequence, and a barcode sequence. PCR reactions were performed in a 25-μL reaction mixture containing 0.5 μM of both forward and reverse primers, 1.0 μL of template DNA, and 12.5 μL of 2X Premix (TaKaRa Biotech, Dalian, China) as previously described (Xia et al. 2011). The triplicate PCR amplicons were pooled, gel-purified, and visualized on 2.0 % agarose gels. The concentration of purified PCR amplicons was determined with a Nanodrop® ND-1000 UV–vis Spectrophotometer, and the purified PCR amplicons were then combined in equimolar ratios into a single tube in preparation for pyrosequencing analysis.
Pyrosequencing data were processed and analyzed following a previously described procedure (Chu et al. 2010; Fierer et al. 2008; Lauber et al. 2009) using the Quantitative Insights Into Microbial Ecology (QIIME) pipeline (http://qiime.sourceforge.net). In brief, low quality sequence reads were filtered such that read lengths <200 bp, reads with an average quality score <25, reads with ambiguous nucleotides of >6 homopolymers and reads with mismatches in primers were excluded. The 6 bp barcode was examined to assign reads to individual samples. Phylotypes were identified using UCLUST (Edgar 2010), with a taxonomic resolution at the 97 % sequence similarity level. A representative sequence from each phylotype was aligned using PyNAST (Caporaso et al. 2010) with a relaxed neighbor-joining tree built using FastTree (Price et al. 2009). The taxonomic identity of each phylotype was determined using the RDP Classifier (Wang et al. 2007).
BIOLOG assay
For the BIOLOG assay, EcoPlate™ plates (BIOLOG Inc., Hayward, CA, USA) were used to trace the succession of the bacterial carbon source utilization profile. Approximately 1.0 g of fresh bulk soil from around either diseased or healthy roots was mixed with 99 ml of 1 % phosphate buffer solution for 20 min, and 100 μL of the solution was transferred into each well of the EcoPlate with 31 carbon sources. The plates were incubated at 25 °C in the dark in triplicate. Color development of each well was determined using a plate reader (Micro Station; BIOLOG Inc) at 24 h intervals over the course of 7 days of incubation. The value of the average well color development (AWCD) for each sample was then calculated to assess the functional diversity of microbial communities in the rhizospheric soils of either healthy or diseased tomato roots.
Soil enzyme activity assay
The activity of urease (EC 3.5.1.5), catalase (EC 1.11.1.6), and alkaline phosphatase (EC 3.1.3.1) in bulk soil was detected according to Guan (1986).
Data analysis
Duncan’s Test (P < 0.05) of one-way ANOVA analysis was performed to analyze soil enzyme activity, the activity of cell-well degradative enzymes, and the phosphate-solubilizing and nitrogen-fixing capacity of rhizobacterial isolates using the SPSS 13.0 (SPSS Inc, Chicago, IL) general linear model (GLM) procedure. UniFrac significance and principal coordinate analyses (PCoA) were calculated using Fast UniFrac analysis to evaluate the degree of similarity between microbial communities associated with different samples (diseased root, healthy root, disease RS, and healthy RS). UniFrac distances were calculated from phylogenetic distances, particularly the fraction of branch lengths shared between all samples (Ishak et al. 2011).
Results
Detection of R. solanacearum
R. solanacearum was separately isolated and quantified for diseased root, healthy root, diseased RS, and healthy RS by a culture-dependent technique using modified SMSA medium specific for pathogen detection (French et al. 1995). Mucous, opaque, pleomorphic and convex colonies with a red center and whitish periphery were isolated from infected samples on nutrient media. The population size of R. solanacearum in three diseased rhizospheric soil samples was 5.7, 7.3 and 7.6 × 107 CFU (colony forming unit)/g of soil on average. The pathogen was also detected in infected plants, where concentrations ranged from 2.3 to 3.6 × 108 CFU/g of root. As expected, R. solanacearum was not detected in the healthy rhizosphere soil and root samples. These results indicated that diseased tomato plants were infected by R. solanacearum.
Overall diversity of bacterial communities
Total microbial communities were characterized by high-throughput pyrosequencing of rhizosphere soil and tomato roots from diseased and healthy samples in triplicate. Across all 12 samples (6 soil and 6 plant samples), a total of 73,026 high-quality sequences (45,368 sequences for roots and 27,658 for rhizospheric soil) were obtained after the low quality reads were removed. There were an average of 4,510, 4,710, 7,177 and 7,945 effective sequence reads out of 6,045, 6,065 10,415 and 10,524 raw reads for healthy RS, diseased RS, healthy root, and diseased root samples, respectively (Table 1).
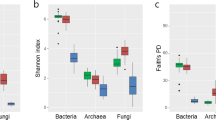
Rhizosphere soil had higher 16S rRNA gene diversity than did root. The highest OTU number was observed in healthy RS, i.e., 1,266 at a distance cutoff level of 3 % (Table 1), while the lowest number of OTUs was obtained for healthy root. Healthy RS had the highest bacterial diversity among the four samples, as shown by the Chao 1 index representative of bacterial phylotype richness levels (Table 1). Diseased RS and root had moderate richness and healthy root had the lowest richness. The rarefaction curves of the four samples at a distance cutoff level of 3 % are shown in Fig. 1. All amplified rarefaction curves increased rapidly from 0 to 1,000 sequences, indicating that sequence derived diversity and richness in this study were sufficient to characterize the species in each sample.
Bacterial community composition in soil and roots
The effective bacterial sequences in the 12 samples were all assigned to corresponding taxonomies using BLAST combined with MEGAN. From the phylum assignment results, we found that high-throughput pyrosequencing produced few reads classified as archaea, and 99.99 % of the 16S rRNA sequence reads were identified as bacteria belonging to 16 phyla: Acidobacteria, Actinobacteria, Bacteria_incertae_sedis, Bacteroidetes, Chloroflexi, Cyanobacteria, Deinococcus-Thermus, Firmicutes, Gemmatimonadetes, Nitrospira, OP10, Planctomycetes, Proteobacteria, Spirochaetes, TM7 and Verrucomicrobia. The bacterial phyla ranged from 0.11 % for Firmicutes to 97.49 % for Proteobacteria, on average. Proteobacteria was the dominant phylum across all samples, accounting for 35.70–97.49 % of total 16S rRNA gene reads sequence (Fig. 2) and higher relative proportions in plant than in soil samples. The relative abundance of the Actinobacteria was second to the Proteobacteria, with an average abundance of 2.43, 12.02, 14.02 and 21.65 % in healthy root, diseased root, healthy RS, and diseased RS, respectively. In addition to the phylum, bacterial diversity and abundance were also analyzed at the class level, and results indicated that the most dominant class was Gammaproteobacteria, with relative abundances of 94.01, 47.97, 44.92, and 32.10 % on average in healthy root, diseased root, healthy RS, and diseased RS, respectively (Fig. 2). A detailed comparison of these phyla is shown in Table S1.
Differences in bacterial communities from diseased and healthy samples
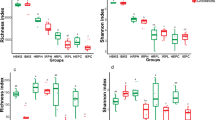
To verify the differences observed in bacterial communities from diseased and healthy samples, the relative abundances of different phyla (different classes for the Proteobacteria) from roots and rhizospheric soil from both healthy and diseased samples were compared (Fig. 3). The Gammaproteobacteria were significantly more abundant in healthy rhizospheric soil and root samples than in diseased samples (P < 0.05), but the abundance of Actinobacteria, Alphaproteobacteria, and Betaproteobacteria was higher in diseased samples (Fig. 3). These results indicated that the structure of bacterial communities showed a similar trend in both soil and plants between diseased and healthy samples.
Percent change of phylotype abundance in diseased relative to healthy samples in both soil (a) and plants (b). Percent change is the difference in the relative abundance of the main phylotype between diseased and healthy samples. A minus sign indicates that the phylotype abundance was greater in healthy samples than in diseased samples. Shown at left is the phylotype from top to bottom with decreasing abundance in the healthy samples
To further compare the variations in bacterial communities between healthy and diseased samples, lineage-specific, weighted UniFrac and principal coordinate analysis were employed. The first principal component of weighted algorithms separated bacterial communities in healthy root and diseased root from those of healthy RS and diseased RS, suggesting strong variations in bacterial community structures between root and soil samples (63.07 % of contribution rate). Along the second principal component axis (18.96 % of contribution rate), shifts in bacterial community composition between diseased and healthy samples, especially in the two root samples, were detected (Fig. 4).
Functional diversity analysis of microbial soil communities
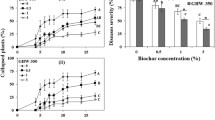
Average well color development (AWCD) was used as an indicator of microbial activity in healthy and diseased soils. The AWCD for both samples increased rapidly after incubation for 24 h and generally followed the same ‘S’ pattern with incubation time, but reached a maximum at different incubation times (Fig. 5). Diseased soil showed a delayed onset of AWCD increase, a reduced rate of AWCD increase and a reduced AWCD maximum when compared with healthy soils at each time point. Pairwise comparisons between diseased and healthy soils indicated a significant reduction in AWCD, ranging from 11.00 to 91.11 % at different time points during the incubation period (Fig. 5).
Soil enzyme activity for diseased and healthy rhizospheric soil
Soil urease, alkaline phosphatase, and catalase activity were determined to assess the potential turnover rates of carbon and nitrogen in rhizospheric soils. Enzymatic activities in diseased soils were inhibited by 12.12 to 25.00 %. For nitrogen cycle enzymes, the mean value of urease activity in healthy RS and diseased RS was 0.30 and 0.24 mg NH3–N·g−1·24 h−1, respectively. The activity of alkaline phosphatase was 0.42 mg hydroxybenzene·g−1·24 h−1 for Healthy RS compared with 0.36 mg·hydroxybenzene g−1·24 h−1 for diseased RS. Catalase activity in healthy RS and diseased RS soils was 1.11 and 0.99 mL·g−1·20 min−1, respectively.
Discussion
The diversity and stability of bacterial communities present in the rhizosphere significantly affect soil and plant quality and ecosystem sustainability (Trivedi et al. 2012). Bacterial community structure and function in the rhizosphere is determined by many selection factors that influence the growth and size of different bacterial populations. Introduction of phytopathogens and nonnative bacteria has been shown to change the amount and composition of organic acids, sugars and other essential nutrients in root exudates (Phillips et al. 2004; Kamilova et al. 2006). Researchers have postulated that changes in host physiology due to pathogen infection can also have differential influence on rhizosphere community composition and associated functions (van der Putten et al. 2007). At present, ecosystem simulation models do not include microbial composition and often do not explicitly consider the effects of phytopathogen infection on microbial activities or the interactions between diverse microbial processes (Bardgett et al. 2008). In this study, we evaluated whether changes in plant physiology due to infection with R. solanacearum might lead to variations in the bacterial community of the rhizosphere and roots of tomato plants. Our results clearly demonstrated the impact of soil-borne pathogens on the total microbial community using high-throughput pyrosequencing. Our results further indicated that pathogen infection could lead to significant changes in the function and composition of microbial communities in healthy and diseased samples.
The results of this study also showed that bacterial wilt influenced the composition of microbial communities inhabiting tomato root and rhizospheric soils (Fig. 2), in agreement with previous findings that ‘Ca. L. asiaticus’ could drastically influence the composition of a rhizobacterial community (Trivedi et al. 2012). Rarefaction curves clearly indicated that healthy RS harbored the highest diversity of bacteria (Fig. 1), suggesting a strong correlation between species richness and ecosystem productivity in healthy RS. Within the bacterial community, the phylum Proteobacteria was the predominant rhizosphere colonizer, presumably because of relatively rapid growth rates (Fierer et al. 2007; DeAngelis et al. 2009). As for the rhizospheric soil and tomato roots infected by the soil-borne pathogen, we observed a decline in the relative abundance of Proteobacteria, consistent with previous observations (Mendes et al. 2011). In contrast, pathogen exposure significantly promoted the proportions of Bacteroidetes and Actinobacteria in both soil and plants (Figs. 2, 3; table S1), suggesting the rapid propagation of these microorganisms in the presence of soil-borne pathogens. This is interesting because Bacteroidetes and Actinobacteria are typical bulk-soil inhabitants and represent stable components of the microbial ecosystem. It is generally accepted that these microorganisms are less affected by nutrient changes (Crawford 1978; Kielak et al. 2008; Yergeau et al. 2009). This study provided strong evidence for the response of microbial communities to soil-borne pathogens. Actinobacteria are known antibiotic producers, which might play a fundamental role in the maintenance of soil ecosystems through the production of antibiotics to counteract R. solanacearum infection, although mechanisms behind this process remain unknown (Wiedenbeck and Cohan 2011). Other reports have shown that the microbiota inhabiting the soil and roots of Arabidopsis thaliana changes when grown in different natural soils under controlled environmental conditions (Bulgarelli et al. 2012; Lundberg et al. 2012). In the present study, clear variations in bacterial community structure between root and soil samples have been identified (Fig. 4), indicating the plant is an important factor. These differences may relate to the plant immune system, which may preferentially promote the growth of beneficial soil microorganisms in the rhizosphere to counteract pathogen assault, leading to the reconstruction of microbial communities (Lundberg et al. 2012). The inhibition of Gammaproteobacteria (Pseudomonadaceae, Xanthomonadales) in the diseased samples may result from soil-borne pathogen exposure (Fig. 2). It has been shown that phytopathogens have profound impacts on the amounts and diversity of nutrients and plant secondary metabolites present in root exudates, which in turn suppresses microorganisms sensitive to alterations in these niches (Cook et al. 1995). Mendes et al. (2011) have also found that these bacteria were more abundant in suppressive rather than favorable soil when sugar beet seedlings were infected by Rhizoctonia solani. In the present study, the Betaproteobacteria were more abundant in the diseased rhizosphere, a phenomena partly attributed to pathogenic infection by R. solanacearum. Yang et al. (2001) detected significant changes in bacterial communities associated with healthy and Phytophthora-infected avocado roots. Similar results were obtained in microbial communities of rhizospheric soil when exposed to take-all disease of wheat caused by Gaeumannomyces graminis var. tritici (McSpadden Gardener and Weller 2001). The rapid advancement of molecular techniques will be of great help for deciphering the microbial mechanisms underlying the various difficulties encountered with continuous cropping.
Microbial communities of the rhizosphere control many belowground processes critical to ecosystem functioning through their influence on the decomposition of organic matter, nutrient cycling and the creation of soil structure (Nannipieri et al. 2003; Cardon and Whitbeck 2007). Shifts in the abundance and composition of the microbial community in soil can, in turn, significantly influence the dynamics of these processes. So we employed the BIOLOG and nutrient cycling-related soil enzymatic activity analysis to investigate changes in microbial function induced by R. solanacearum infection. BIOLOG analysis was used to assess the functional diversity of the soil microbial community, since the degradation of the soil ecosystem may not necessarily be associated with composition changes (Konopka 2009). Our results clearly indicated that the microbial communities of healthy soils have a greater capacity for carbon utilization (Fig. 5). Although AWCD values do not represent the activity of individual microorganisms (Garland and Mills 1991; Haack et al. 1995), these results provide direct evidence for the functional activities of microbes at the community level colonizing soils and could thus serve as a useful tool to monitor the succession patterns of soil microbial communities under disturbance. In addition, increases in soil enzyme activity are also thought to represent improvement in organic-matter content, microbial activity, and carbon and nutrient cycling (Tabatabai 1994). In fact, soil enzyme activity has been shown to change in response to various management practices, such as tillage, rotation, and fertilization, and therefore is considered to be an important indicator of soil quality and ecological stability (Dick 1994; Ajwa et al. 1999; Lagomarsino et al. 2009; Tian et al. 2010). Urease, phosphatase, and catalase are involved in nitrogen and phosphorus cycling in the soil. There was a positive correlation between plant health and soil enzyme activity. In general, enzyme activity in the soil has been shown to decrease in response to tomato bacterial wilt, which might be related to the dynamics of microbial populations in the soil. It has been suggested that the composition of the microbial community determines the potential for enzyme synthesis, and thus any modification in the microbial community due to environmental factors could be reflected on the level of soil enzymatic activity (Kandeler et al. 1996). Our results provide strong evidence that changes in the microbial community are linked to functional activity, although the microbial response to soil-borne pathogens require further investigation at the level of the individual microorganism.
Conclusion
Interactions between microorganisms and the soil are very complex, and have been shown to strongly influence both plant and microbial community composition and ecosystem processes. Our results revealed that there were significant shifts in the composition and functional potential of soil microbial communities in rhizospheric soil and tomato roots infected by soil-borne pathogens. These results provide guidance for a reduction in the amount of pathogens in continuous cropping by maintaining harmonious soil composition, regulating plant growth conditions, and preserving ecosystem equilibrium. However, further study is required to understand the relative importance of perturbations caused by phytopathogens on microbial biodiversity and activity and searching for appropriated methods to control plant diseases.
References
Abawi GS, Widmer TL (2000) Impact of soil health management practices on soilborne pathogens, nematodes and root diseases of vegetable crops. Appl Soil Ecol 15:37–47
Ajwa HA, Dell CJ, Rice CW (1999) Changes in enzyme activities and microbial biomass of tall grass prairie soil as related to burning and nitrogen fertilization. Soil Biol Biochem 31:769–777
Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266
Bardgett RD, Freeman C, Ostle NJ (2008) Microbial contributions to climate change through carbon cycle feedbacks. ISME J 2:805–814
Baudoin E, Benizri E, Guckert A (2003) Impact of artificial foot exudates on the bacterial community structure in bulk soil and maize rhizosphere. Soil Biol Biochem 35:1183–1192
Bulgarelli D, Rott M, Schlaeppi K, Loren V, van Themaat E, Ahmadinejad N, Assenza F, Rauf P, Huettel P, Reinhardt R, Schmelzer E, Peplies J, Gloeckner F-O, Amann R, Eickhorst T, Schulze-Lefert P (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488:91–95
Caporaso G, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Cardon Z, Whitbeck J (2007) The rhizosphere: an ecological perspective, 1st edn. Academic, San Diego, p 232pp
Chu H, Fierer N, Lauber CL, Caporaso JG, Knight R, Grogan P (2010) Soil bacterial diversity in the Arctic is not fundamentally different from that found in other biomes. Environ Microbiol 12:2998–3006
Ciampi-Panno L, Fernandez C, Bustamante P, Andrade N, Ojeda S, Conteras A (1989) Biological control of bacterial wilt of potatoes caused by Pseudomonas solanacearum. Am Potato J 66:315–332
Cook RJ, Thomashow LS, Weller DM, Fujimoto D, Mazzola M, Bangera G, Kim DS (1995) Molecular mechanisms of defense by rhizobacteria against root disease. Proc Natl Acad Sci U S A 92:4197–4201
Crawford DL (1978) Lignocellulose decomposition by selected Streptomyces strains. Appl Environ Microbiol 6:1041–1045
DeAngelis KM, Brodie EL, DeSantis TZ, Andersen GL, Lindow SE, Firestone MK (2009) Selective progressive response of soil microbial community to wild oat roots. ISME J 3:168–178
Diaz PI, Dupuy AK, Abusleme L, Reese B, Obergfell C, Choquette L, Dongari-Bagtzoglou A, Peterson DE, Terzi E, Strausbaugh LD (2012) Using high throughput sequencing to explore the biodiversity in oral bacterial communities. Mol Oral Microbiol 27:182–201
Dick RP (1994) Soil enzyme activities as indicators of soil quality. In: Doran JW, Coleman DC, Bezdicek DF, Stewart BA (eds) Defining soil quality for a sustainable environment. SSSA special publication no. 35. Soil Science Society of America Inc, Madison, pp 107–124
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Fierer N, Hamady M, Lauber CL, Knight R (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 105:17994–17999
Fitter AH, Gilligan CA, Hollingworth K, Kleczkowski A, Twyman RM, Pitchford JW (2005) Biodiversity and ecosystem function in soil. Funct Ecol 19:369–377
French E, Gutarra L, Aley P, Elphinstone J (1995) Culture media for Pseudomonas solanacearum isolation, identification and maintenance. Fitopatol 30:126–130
Garland JL, Mills AL (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl Environ Microbiol 57:2351–2359
Glenn TC (2011) Field guide to next generation DNA sequencers. Mol Ecol Resour 15:759–769
Guan SY (1986) Soil enzyme and research methods. China Agriculture Press, Beijing, pp 123–339
Haack SK, Garchow H, Klug MJ, Forney LJ (1995) Analysis of factors affecting the accuracy, reproducibility, and interpretation of microbial community carbon source utilization patterns. Appl Environ Microbiol 61:1458–1468
Harvell CD, Kim K, Burkholder JM, Colwell RR, Epstein PR, Grimes DJ, Hofmann EE, Lipp EK, Osterhause ADME, Overstreet RM, Porter JW, Smith GW, Vasta GR (1999) Emerging marine disease climate links and anthropomorphic factors. Science 285:1505–1510
Hayward AC (1995) Pseudomonas solanacearum. In: Singh US, Singh RP, Kohmoto K (eds) Pathogenesis and host specificity in plant disease: histopathological, biochemical, genetic and molecular bases, vol 1. Elsevier, Tarrytown, pp 139–151
Ishak HD, Plowes R, Sen R, Kellner K, Meyer E, Estrada DA, Dowd SE, Mueller UG (2011) Bacterial diversity in solenopsis invicta and solenopsis geminata ant colonies characterized by 16S amplicon 454 pyrosequencing. Microb Ecol 61:821–831
Kamilova F, Kravchenko LV, Shaposhnikov AI, Azarova T, Makarova N, Lugtenberg B (2006) Organic acids, sugars, and l-tryptophan in exudates of vegetables growing on stonewool and their effects on activities of rhizosphere bacteria. Mol Plant Microbe Interact 19:250–256
Kandeler E, Kampichler C, Horak O (1996) Influence of heavy metals on the functional diversity of soil microbial communities. Biol Fertil Soils 23:299–306
Kang YG, Chung YH, Yu YH (2004) Relationship between the population of Ralstonia solanacearum in soil and the incidence of bacterial wilt in the naturally infested tobacco fields. Plant Pathol J 20:289–292
Kidwell KK, Osborn TC (1992) Simple plant DNA isolation procedures. In: Beckman JS, Osborn TC (eds) Plant genomes: methods for genetic and physical mapping. Kluwer Academic Publishers, Dordrecht, The Netherlands, pp 1–13
Kielak A, Pijl AS, van Veen JA, Kowalchuk GA (2008) Differences in vegetation composition and plant role of HLB in microbial communities species identity lead to only minor changes in soil-borne microbial communities in a former arable field. FEMS Microbiol Ecol 63:372–382
Konopka A (2009) What is microbial community ecology? ISME J 3:1223–1230
Lagomarsino A, Moscatelli MC, Di Tizio A, Mancinelli R, Grego S, Marinari S (2009) Soil biochemical indicators as a tool to assess the short-term impact of agricultural management on changes in organic C in a Mediterranean environment. Ecol Indic 9:518–527
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120
Lugtenberg B, Kamilova F (2009) Plant-growth promoting rhizobacteria. Annu Rev Microbiol 63:541–556
Lundberg DS, Lebeis SL, Herrera-Paredes S, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrekston A, Kunin V, Glavina del Rio T, Edgar R, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90
Marschner P, Yang CH, Lieberei R (2001) Soil and plant specific effects on bacterial community composition in the rhizosphere. Soil Biol Biochem 33:1437–1445
McSpadden Gardener BB, Weller DM (2001) Changes in populations of rhizosphere bacteria associated with take-all disease of wheat. Appl Environ Microbiol 67:4414–4425
Mendes R, Kruijt M, De Bruijn I, Dekkers E, Van der Voort M, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Montesinos E, Bonaterra A, Badosa E, Frances J, Alemany J, Llorente I, Moragrega C (2002) Plant-microbe interactions and the new biotechnological methods of plant disease control. Int Microbiol 5:169–175
Nannipieri P, Ascher J, Ceccherini MT, Landi L, Pietramellara G, Renella G (2003) Microbial diversity and soil functions. Eur J Soil Sci 54:655–670
Phillips DA, Fox TC, King MD, Bhuvaneswari TV, Teuber LR (2004) Microbial products trigger amino acid exudation from plant roots. Plant Physiol 136:2887–2894
Price MN, Dehal PS, Arkin A (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650
Reiter B, Pfeifer U, Schwab H, Sessitsch A (2002) Response of endophytic bacterial communities in potato plants to infection with Erwinia carotovora subsp. atroseptica. Appl Environ Microbiol 68:2261–2268
Rosario K, Iverson SL, Henderson DA, Chartrand S, McKeon C, Glenn EP, Maier RM (2007) Bacterial community changes during plant establishment at the San Pedro River mine tailings site. J Environ Qual 36:1249–1259
Tabatabai MA (1994) Soil enzymes. In: Weaver RW, Angle JR, Bottomley PS (eds) Methods of soil analysis. Part 2: microbiological and biochemical properties. Soil Science Society of America, Madison, pp 775–833
Tian L, Dell E, Shi W (2010) Chemical composition of dissolved organic matter in agroecosystems: correlations with soil enzyme activity and carbon and nitrogen mineralization. Appl Soil Ecol 46:426–435
Trivedi P, Duan Y, Wang N (2010) Huanglongbing, a systemic disease, restructures the bacterial community associated with citrus roots. Appl Environ Microbiol 76:3427–3436
Trivedi P, Spann TM, Wang N (2011) Isolation and characterization of beneficial bacteria associated with citrus roots in Florida. Microb Ecol 62:324–336
Trivedi P, He Z, Van Nostrand JD, Albrigo G, Zhou JZ, Wang N (2012) Huanglongbing alters the structure and functional diversity of microbial communities associated with citrus rhizosphere. ISME J 6:363–383
van der Putten WH, Klironomos JN, Wardle DA (2007) Microbial ecology of biological invasions. ISME J 1:28–37
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wiedenbeck J, Cohan FM (2011) Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol Rev 35:957–976
Xia WW, Zhang CX, Zeng XW, Feng YZ, Weng JH, Lin XG, Zhu JG, Xiong ZQ, Xu J, Cai ZC, Jia ZJ (2011) Autotrophic growth of nitrifying community in an agricultural soil. ISME J 5:1226–1236
Yang CH, Crowley DE, Menge JA (2001) 16S rDNA fingerprinting of rhizosphere bacterial communities associated with healthy and Phytophthora infected avocado roots. FEMS Microbiol Ecol 35:129–136
Ye L, Zhang T (2013) Bacterial communities in different sections of a municipal wastewater treatment plant revealed by 16S rDNA 454 pyrosequencing. Appl Microbiol Biotechnol 97:2681–2690
Yergeau E, Schoondermark-Stolk SA, Brodie EL, Dejean S, Desantis TZ, Goncalves O, Piceno YM, Andersen GL, Kowalchuk GA (2009) Environmental microarray analyses of Antarctic soil microbial communities. ISME J 3:340–351
Acknowledgments
This research was supported by national natural science foundation of China (41201241), China postdoctoral science foundation (2013 M531412), and innovation key program of the Chinese academy of sciences (KSCX-EW-B-6).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Simon Jeffery.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
(DOC 41 kb)
Rights and permissions
About this article
Cite this article
Li, JG., Ren, GD., Jia, ZJ. et al. Composition and activity of rhizosphere microbial communities associated with healthy and diseased greenhouse tomatoes. Plant Soil 380, 337–347 (2014). https://doi.org/10.1007/s11104-014-2097-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-014-2097-6