Abstract
Fusarium wilt of tomato caused by the pathogen Fusarium oxysporum f. sp. lycopersici (Fol) is one of the most devastating soilborne diseases of tomato. To evaluate whether microbial community composition associated with Fol-infected tomato is different from healthy tomato, we analyzed the tomato-associated microbes in both healthy and Fol-infected tomato plants at both the taxonomic and functional levels; both bacterial and fungal communities have been characterized from bulk soil, rhizosphere, rhizoplane, and endosphere of tomatoes using metabarcoding and metagenomics approaches. The microbial community (bacteria and fungi) composition of healthy tomato was significantly different from that of diseased tomato, despite similar soil physicochemical characteristics. Both fungal and bacterial diversities were significantly higher in the tomato plants that remained healthy than in those that became diseased; microbial diversities were also negatively correlated with the concentration of Fol pathogen. Network analysis revealed the microbial community of healthy tomato formed a larger and more complex network than that of diseased tomato, probably providing a more stable community beneficial to plant health. Our findings also suggested that healthy tomato contained significantly greater microbial consortia, including some well-known biocontrol agents (BCAs), and enriched more functional genes than diseased tomato. The microbial taxa enriched in healthy tomato plants are recognized as potential suppressors of Fol pathogen invasion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tomato (Solanum lycopersicum) is one of the most widely cultivated vegetables worldwide, with a global annual yield of more than 177 million tons (FAO, http://www.fao.org/faostat). The production of tomato is often limited by diseases, especially the devastating soilborne disease Fusarium wilt, which is caused by the tomato-specific fungus Fusarium oxysporum f. sp. lycopersici (Fol) [1, 2]. This pathogen can infect tomato plants at all stages of growth and is one of the most devastating diseases of tomato [3]. Disease-resistant cultivars, chemical fungicides, biocontrol agents (BCAs), crop rotation, and soil fumigation are commonly used to manage Fusarium wilt [1, 2]. Crop rotation, however, is not favorable for greenhouse farmers who do not prefer growing less profitable cereals. Methyl bromide (for soil fumigation) is quite effective but has been phased out in the “Montreal Protocol,” due to its considerable environmental pollution and ozone-depleting effect [4]. On the other hand, the BCAs have been increasingly regarded as a green and effective solution and applied in suppressing soilborne disease [5,6,7]. The increasing demand for healthy and productive food crops requires a green and effective strategy for controlling this plant disease.
Soil is a rich bank of potentially beneficial and antagonistic organisms, and the status of its microbial community largely determines the productivity of agroecosystems [8, 9]. Root-associated microorganisms serve as important determinants of plant health and growth by supporting disease resistance, nutrient uptake, and abiotic stress tolerance [10, 11]. The use of naturally occurring microbial communities offers a safe and effective approach for suppressing pathogens that have recently emerged, representing an ideal alternative to chemical control [12]. Several studies have demonstrated that plant endophytic and rhizospheric microbes can enhance disease suppression, and they are thus used for disease suppression [13,14,15,16]. For example, many plant-associated microbes have antagonistic activities against Fol and have been used as BCAs, including Bacillus spp., Pseudomonas spp., Lactobacilli spp., Rhizobia spp., Trichoderma spp., and Gliocladium spp. [9, 17,18,19]. The BCA species have different mechanisms in disease suppression, including competition for nutrition and ecological niches, producing antibiotics components, and induced systemic resistance of plant [10, 13]. High microbial diversity usually could increase resistance to pathogen invasions and plant infestation [10, 20]. For example, Hu et al. [12] reported that enriched Pseudomonas diversity significantly enhanced pathogen suppression through intensified resource competition and interference with the pathogen. Niu et al. [21] showed that a simplified and representative bacterial community had a better disease suppression effect on the Fusarium verticillioides pathogen than did single bacterial species in maize. High diversity of microbial communities also could increase interference competition among pathogens and intensify resource-use efficiency, which could help microbes to better colonize and occupy the root niche [22, 23]. All these studies demonstrated that a more diverse microbial consortium could affect the survival, establishment, and functioning of pathogens and have better effects on disease suppression [10].
Studies on tomato diseases have been mainly focused on tomato bacterial wilt disease caused by Ralstonia solanacearum [12, 13, 24]. Wei et al. [24] found the significant taxonomic and functional differences between R. solanacearum diseased and healthy tomato-associated bacterial communities. The bacterial community of healthy tomato was interacting more often and more predictably than those of bacterial wilted tomato, which could potentially increase the community stability against R. solanacearum invasions [24]. The disease outcomes are affected by the composition and assembly of the plant-associated microbiome and physicochemical properties of the soil [2, 14]. It has, however, remained unclear whether the microbiomes of Fol pathogen diseased tomato plants are different from the microbiomes of remain healthy tomato plants. Moreover, previous studies only demonstrated the bacterial community of healthy and diseased tomato plants, and they excluded fungi and other important microbes [12, 13, 24]. For example, many fungi have been shown to suppress pathogens and affect the composition of root-associated microbial communities [2, 25]. Hence, we need a more comprehensive understanding of the variation between healthy and diseased tomato for both bacterial and fungal communities. High-throughput sequencing (HTS) has provided a powerful approach to investigate the complex interactions between plants and microbial communities in situ [23] and has identified microbial consortium and functions associated with healthy and diseased plants [13, 24]. We investigated the taxonomic and functional differences between healthy- and diseased-tomato-associated microbiomes using both amplicon sequencing (targeting the 16S rRNA of bacterial and ITS region of fungi) and metagenomic sequencing. Additionally, the microbial diversity differences between different tomato root zone areas were also examined to reveal microbial selection by tomato root compartments.
Materials and Methods
Experimental Design and Sampling
In February 2018, a greenhouse of ~ 660 m2 in Luojiazhuang, Shandong Province, China (36° 55′ 26.47″ N 118° 46′ 20.75″ E) was selected for the tomato cultivation experiment. This greenhouse has been used for monoculture tomato planting continuously for 8 years, with two crop seasons (spring season: from February to June; autumn season: from July to November) per year. Surface-sterilized tomato seeds (70% ethanol for 1 min, 1% sodium hypochlorite solution for 15 min) were germinated in sterile moist filter paper and planted into the greenhouse field. All the samples were collected at the spring crop season, and the tomato root and bulk soil samples were collected in June (the end of the growing period). The numbers of healthy and diseased tomato plants were recorded, and about 80% of the total plants showed disease symptoms. The disease indexes were recorded using the following scale: 0, no wilting; 1, about 1–25% leaves wilted; 2, about 25–50% leaves wilted; 3, about 51–75% leaves wilted; and 4, about 76–100% leaves wilted [13]. The tomato plants that showed clear disease symptoms (disease index ≥ 3) and were positive for Fol pathogen isolation from surface-sterilized roots were classified as diseased plants [13], while the tomato plants that showed no wilt symptoms and were negative for Fol pathogen isolation from surface-sterilized roots were classified as healthy plants. Twelve Fol-diseased plants and twelve healthy plants were collected as biological replicates for each root compartment. In brief, samples of bulk soil and root compartments (including the rhizosphere, rhizoplane, and endosphere) were collected according to the methods described by Edwards et al. [26]. Soil physicochemical properties, namely, pH, total carbon, nitrogen, phosphorus, sodium ions, potassium ions, calcium ions, iron ions, and magnesium ions, were measured for each sample using standard soil testing procedures (http://vdb3.soil.csdb.cn). The samples were transported back to the laboratory in iceboxes and stored immediately at − 80 °C until use.
DNA Extraction from the Rhizosphere, Rhizoplane, and Endosphere
For each plant, we extracted total DNA independently from bulk soil, rhizosphere, rhizoplane, and endosphere samples according to the methods described by Edwards et al. [26]. In brief, each tomato root was vigorously shaken by hand until the loose soil was completely removed. The remaining ~ 1 mm layer of soil was washed off with phosphate-buffered saline (PBS) and kept as the rhizosphere sample. The clean root was then washed twice to remove the remaining soil and placed into clean PBS in a 50-ml falcon tube. The rhizoplane sample was collected from the clean root by sonication for approximately 30 min. The sonicated roots were washed with PBS for three times, followed by sterilized in 70% ethanol for 1 min and finally rinsed extensively in sterile water for three times. Then, the entire root tissues were used for endosphere microbial DNA extraction. Total genomic DNA was extracted from each sample using a FastDNA® Spin Kit following the manufacturer’s instructions (MP Biomedicals, Solon, OH, USA). The V3-V4 variable region of the bacterial 16S rRNA gene was amplified using the primers 338F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) [27]. The fungal ITS1 region was amplified using the primers ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS2R (5′-GCTGCGTTCTTCATCGATGC-3′) [28]. The PCR reaction was performed with 0.2 μM forward primer, 0.2 μM reverse primer, 10 ng of template DNA, and 12.5 μl of Phusion High-Fidelity PCR Master Mix (New England Biolabs). The reaction conditions were as follows: 95 °C for 2 min; 25 cycles at 95 °C for 30 s; 55 °C for 30 s; and 72 °C for 60 s; 72 °C for 10 min and held at 4 °C. The second round of PCR amplification was performed using an Ultra DNA Library Prep Kit for Illumina (New England Biolabs, USA) following the manufacturer’s recommendations to add index codes. All PCR products were purified with a GeneJET Gel Extraction Kit (Thermo Scientific) before sequencing. Library quality was checked with a Qubit@ 2.0 Fluorometer (Life Technologies, CA, USA) and an Agilent Bioanalyzer 2100 system (Agilent Technologies, Santa Clara, CA). Finally, paired-end 2 × 300 bp sequencing was performed using Illumina MiSeq technology (Illumina Inc., San Diego, CA, USA) at Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China). For shotgun paired-end metagenomic library construction, DNA was randomly fragmented to an average size of approximately 300 bp by a Covaris M220 ultrasonicator (Gene Company Limited, China). The paired-end Illumina library was prepared by using a TruSeqTM DNA Sample Prep Kit (Illumina, San Diego, CA, USA). Adapters were ligated to the blunt-end fragments by the sequencing primer hybridization sites. Paired-end sequencing was performed at Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China) using the Illumina HiSeq platform (Illumina Inc., San Diego, CA, USA) according to the manufacturer’s instructions.
Quantification of Fol DNA by Quantitative Real-time PCR
The Fol pathogen-specific virulence gene SIX1 sequenced with primers SP1-2F (5-GCTGGCGGATCTGACACTGT-3) and SP1-2R (5-CCTAAACCACATATCT CGTCCAAA-3) was used to quantify the concentrations of Fol pathogen [29]. Real-time PCR was performed by using ChamQ Universal SYBR qPCR Mix and a Bio-Rad CFX96 Real-Time PCR System. The 25 μl reaction mixture contained 12.5 μl of ChamQ Universal SYBR qPCR Mix, 0.5 μl of ROX, 50 ng of template DNA, and 1.0 μl (10 μM/l) each of the forward and reverse primers. The thermocycling conditions were as follows: 95 °C for 2 min; 40 cycles of 95 °C for 10 s, 60 °C for 30 s, and 72 °C for 30 s; and a final elongation step at 72 °C for 10 min. For SYBR Green amplification, a melting step was added to improve amplification specificity. The quantitative PCRs (qPCRs) were performed in triplicate (technical replicates) for each sample, and six independent samples per group were used as biological replicates. Statistical analyses were performed using the GraphPad Prism 8 software with one-way analysis of variance (ANOVA) followed by Tukey’s honestly significant difference (HSD) post hoc test.
Bioinformatic Analysis
Metagenomic Sequencing and Bioinformatics Pipeline
All raw FASTQ data were filter-trimmed using Trimmomatic to remove low-quality (length < 50 bp, quality value < 20, and/or N bases) and adapter sequences [30]. The remaining sequences were compared with the host (Solanum lycopersicum) genome, and the contaminant host reads were comprehensively removed by the Bowtie2 software [31]. In total, 20,627,647 host genomic sequences were detected as contaminants, which accounted for 3.5% (ranging from 0.29 to 4.44%) (Table S1) of the total sequences for all samples. After removing non-microbial sequences, the taxonomic classification of the remaining sequences was performed using MetaPhlAn2 [32] against the Kyoto Encyclopedia of Genes and Genomes (KEGG) Ortholog (KO) database with the “very sensitive” global alignment option [33]. The humann2_renorm_table command was used to normalize the KO abundances within each sample by the median universal single-copy gene abundance. The Statistical Analysis of Metagenomic Profiles (STAMP) software [34] was implemented to detect significant differences in the abundance of functional genes/pathways corresponding to the healthy and diseased groups. Welch’s t test (P < 0.05) and Benjamini-Hochberg’s false discovery rate (FDR) multiple test correction were applied to generate extended error bar plots. The linear discriminant analysis effect size (LEfSe) method was also used to detect the significant functional features. The LEfSe method combines a statistical test (pairwise Wilcoxon test or Kruskal-Wallis test) with linear discriminant analysis (LDA) for feature selection. The threshold logarithmic LDA score for discriminating features was 2.0, and the alpha value employed for the factorial Kruskal-Wallis test was 0.05 [35].
16S rRNA Gene and ITS rRNA Gene Bioinformatics Pipeline
The fungal ITS1 and bacterial 16S rRNA V3-V4 regions were processed using the USEARCH 10.0 and VSEARCH 2.14 software respectively [36, 37]. All 16S rRNA paired-end reads were quality filtered using the fastq_filter command and joined by the fastq_mergepairs command. Chimeric sequences were detected and removed using UCHIME against the Ribosomal Database Project (RDP) Gold database [36]. The non-chimeric sequences were sorted by abundance, dereplicated, and clustered into operational taxonomic units (OTUs) using the UPARSE algorithm (≥ 97%, singletons removed). Then, all non-chimeric sequences were reassigned to formulate a final OTU table using the otutab command. The SILVA 16S rRNA database (132 release) was used to assign taxonomic categories to OTUs using the sintax command at a confidence threshold of 0.8. The fungal ITS1 sequences were processed similarly, with screening and removal of chimeric sequences according to the UNITE CHIME reference database [38]. The taxonomic classification of the representative sequences was performed using the RDP classifier against the Warcup fungal ITS V2 database at a confidence threshold of 0.8 [39].
Diversity and Statistical Analysis
Most diversity and statistical analyses were performed using the vegan package as implemented in R unless stated otherwise [40]. Rarefaction curves of all samples were constructed with the rarecurve function. The OTU tables were rarefied to the smallest number of reads among all samples using the rrarefy command to allow comparison on an equal basis. All bacterial and fungal samples were rarefied to 29,805 sequences and 36,098 sequences, respectively, before calculating the diversity indices. Alpha diversity was calculated based on species richness and the Shannon-Wiener index via the diversity command [40]. ANOVA followed by Tukey’s HSD method was used to explore variation in Shannon diversity and OTU richness among different groups. The beta diversity of the bacterial and fungal communities was calculated using UniFrac distances and Bray-Curtis distances based on a normalized OTU table, respectively [41]. A principal coordinate analysis (PCoA) plot based on UniFrac distance and Bray-Curtis distance matrices was used to visualize the dissimilarities among samples. Differences in microbial community composition between root zone groups were measured by multivariate permutational analysis of variance (PERMANOVA) and analysis of similarities (ANOSIM) with the adonis and anosim commands, respectively. The relationships between environmental factors and microbial community composition were calculated using the envfit command (999 random permutations). The significance of each variable (P < 0.05) was calculated based on Bray-Curtis distances using the capscale command in the vegan package [40]. The negative binomial generalized linear model in the edgeR package was used to identify and visualize significantly (P < 0.05) enriched OTUs in healthy or diseased tomato samples [42]. We used the estimateGLMTagwiseDisp and estimateGLMCommonDisp functions to estimate common and tagwise dispersion, respectively. The OTU read counts were fitted with a negative binomial generalized log-linear model, and differential OTU abundances were evaluated by the glmFit function. Correction for multiple testing was carried out by the false discovery rate (FDR) with a P value of less than 0.05. The ggplot2 package was used to generate all graphs and plots [43].
Co-occurrence Network Construction and Analysis
Co-occurrence networks of microbial communities were constructed based on OTU relative abundance for both healthy and diseased tomato. For all networks, we utilized OTUs with a sequence abundance greater than 0.01% for network construction. We calculated descriptive and topological network properties with the R package igraph [44]. These properties included the sums of network nodes (representing OTUs), sums of edges (significant correlations between OTUs), and co-occurrence degrees (number of direct correlations to a node). For this task, we combined the bacterial and fungal OTUs into one OTU table for both healthy and diseased tomatoes. The pairs of OTUs with a positive Spearman rank correlation value (ρ) > 0.7 and a significant P value (< 0.001) were identified. To explore community structure within the healthy and diseased networks, the network modules with a greater edge density within groups than between groups were identified by using the “greedy optimization of the modularity” algorithm [45]. Ecologically important microbes frequently co-occur with other microbes in co-occurrence networks and potentially play a key role within the microbiome in ecosystems [46, 47]. The keystone taxa in healthy and diseased networks were identified using the NetShift method (https://web.rniapps.net/netshift) based on differences in network interactions between healthy- and diseased-tomato microbiomes [48]. The co-occurrence networks were constructed and visualized using the “Fruchterman-Reingold” layout with 104 permutations in igraph [45].
Data Accessibility
All raw paired-end Illumina sequence data have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive database (https://www.ncbi.nlm.nih.gov/sra/?term=SRP156834) under BioProject no. PRJNA485233.
Results
Data Summary and Variation in Alpha Diversity
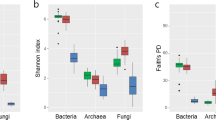
After sequence quality filtering and reference-based chimera removal, approximately 22.9% nonbacterial sequences and 1.7% non-fungal sequences were removed. A total of 1,842,109 fungal ITS1 sequences and 2,148,761 bacterial 16S V3-V4 region sequences remained. All the non-chimeric fungal sequences were clustered into 539 fungal OTUs (singletons removed), with an average of 37,954 sequences per sample. All the non-chimeric prokaryotic sequences were clustered into 3682 bacterial and Archaea OTUs (singletons removed), with an average of 45,763 sequences per sample. Comparative analyses of alpha and beta diversity were performed after fungal and bacterial sequences were normalized to 29,825 and 36,098 reads, respectively. The rarefaction curves of both bacterial and fungal communities approached an asymptote, indicating the sufficiency of our sequencing depths (Fig. S1). The microbial diversity of healthy tomato was significantly higher (Tukey’s HSD test, P < 0.05) than that of diseased tomato in both the bacterial and fungal communities (Fig. 1a–d). The differences in microbial diversity between healthy tomato and diseased tomato indicated that healthy tomato contained significantly (Tukey’s HSD test, P < 0.05) more microbial taxa. In addition, the differences in diversity between root compartments (bulk soil, rhizosphere, rhizoplane, and endosphere) were also significant. For example, bacterial diversity significantly decreased in the order bulk soil > rhizosphere > rhizoplane > endosphere (Tukey’s HSD test, P < 0.05) for both healthy and diseased tomato (Fig. 1a and b ). However, for the fungal community, the healthy tomato plants did not exhibit differences in diversity among bulk soil and the root compartments (Fig. 1c and d ).
The alpha diversity indices of the tomato bacterial and fungal communities. a and b OTU richness and Shannon-Wiener diversity of tomato bacterial communities. c and d OTU richness and Shannon-Wiener diversity of tomato fungal communities. The top and bottom whiskers indicate the 5th and 95th percentiles. The lines inside the boxes represent the median. Means were compared by one-way ANOVA, Tukey’s HSD, and LSD-t comparisons. Different lowercase letters within the panels denote significant differences at the level of P < 0.05. HEPC: healthy-tomato endosphere samples; IEPC: diseased-tomato endosphere samples; HBKS: healthy-tomato bulk soil samples; IBKS: diseased-tomato bulk soil samples; HRPH: healthy-tomato rhizosphere samples; IRPH: diseased-tomato rhizosphere samples; HRPL: healthy-tomato rhizoplane samples; IRPL: diseased-tomato rhizoplane samples
Composition of and Environmental Influence on the Tomato Microbial Community
In the PCoA of the bacterial community, the unweighted and weighted UniFrac distances explained 67.76 and 78.45% of the total variation, respectively (Fig. 2a and b ). ANOSIM revealed significant differences between healthy- and diseased-tomato bacterial communities (r = 0.08287, P = 0.026). Moreover, the bacterial communities of tomato root compartments were significantly (P < 0.001) separated from the bacterial community of the bulk soil (Fig. 2a and b ). In the PCoA of fungal communities, the unweighted and weighted Bray-Curtis distances explained 51.78% and 72.24% of the total variation, respectively (Fig. 2c and d ). The ANOSIM revealed a striking difference between the fungal communities of healthy tomato and diseased tomato (r = 0.3281, P = 0.001). Distance-based redundancy analysis (db-RDA) was employed to illustrate the influence of environmental factors on microbial community composition. However, no significant differences (P > 0.05) were found between the healthy and diseased tomato in terms of soil physicochemical properties (Table S2). This result suggests that abiotic environmental factors have a limited influence on tomato microbial communities. The densities of Fol were one of the most important drivers for causing the tomato Fol disease. To confirm this, the concentration of Fol pathogen was detected and compared among all the samples using qPCR targeting a Fol-specific gene. We extracted the OTUs (OTUs3) which belong to the Fol species and compared the relative abundance for both healthy and diseased tomato groups (Fig. S2a). The results suggested that the relative abundance of Fol in diseased root compartments were significantly (P < 0.001, Student’s t test) higher than the healthy root compartments (Fig. S2a). Moreover, the Fol absolute concentrations of diseased root compartments were also significantly (P < 0.001, Student’s t test) higher than those of healthy root compartments (Fig. S2b). Similar trends were also found for the bulk soils, with the Fol concentrations of diseased-tomato bulk soils being significantly (P < 0.001, Student’s t test) greater than those of healthy-tomato bulk soils (Fig. S2). Crucially, the absolute concentrations of Fol in the rhizosphere of healthy tomato plants were very low (Fig. S2). These results demonstrated that outbreaks of Fol disease are significantly correlated with the concentration of Fol.
Fungal and Bacterial Taxa in Healthy and Diseased Tomato Roots
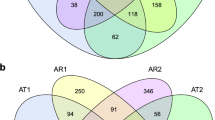
For the bacterial community, the phyla Proteobacteria (28.8%), Actinobacteria (11.7%), Firmicutes (9.4%), Bacteroidetes (8.2%), Acidobacteria (6.3%), Candidatus Saccharibacteria (3.1%), Verrucomicrobia (2.3%), Planctomycetes (1.6%), Chloroflexi (1.5%), and Chlamydiae (1.0%) were most dominant in the tomato roots (Table S3). For the fungal community, the most dominant phyla were mainly Ascomycota (57.5%), Basidiomycota (19.9%), Mortierellomycota (5.4%), Rozellomycota (2.8%), Aphelidiomycota (2.0%), and Olpidiomycota (1.9%) (Table S3). The 16S rRNA analyses showed that the genera Arthrobacter, Pyrenochaeta, Plectosphaerella, Pseudomonas, and Sphingobium were selected from the bulk soil and enriched in the root compartments of both healthy and diseased tomato plants (Fig. 3a and b). We next explored significantly enriched microbial taxa associated with healthy and diseased tomato plants. A total of 186 bacterial and 168 fungal OTUs were found to be significantly enriched in the healthy tomato plants, accounting for 7.44% and 9.54% of the total sequences, respectively (Fig. 4a and b; Table S4). Of these discriminating OTUs, the bacteria associated with healthy tomato with high abundances belonged to Acinetobacter, Aeromonas, Bacillus, Chloroflexus, Lysinibacillus, Paenibacillus, Planococcaceae, Pseudomonas, and Streptomyces. For the fungal community, the taxa associated with healthy tomato with high abundances were members of Aspergillus, Cladosporium, Chaetomium, Cordyceps, Conocybe, Cutaneotrichosporon, Metarhizium, Mortierella, Olpidium, Penicillium, Ramicandelaber, Tuber, Uncinocarpus, and Vermiculariopsiella. On the basis of comparison analysis, we found many enriched taxa of healthy tomato with potential use as BCAs to inhibit Fol disease (Table S4).
Annotated heatmaps of significantly enriched fungal and bacterial OTUs of healthy and diseased tomatoes. a The heatmap representing the bacterial OTUs differed significantly (P < 0.05) in abundance between the healthy and diseased tomato groups. b The heatmap representing the fungal OTUs differed significantly (P < 0.05) in abundance between the healthy and diseased tomato groups. The higher the abundance of OTUs in a sample, the deeper the color in the heatmap. The labels on the bottom of the heatmaps are the names of tomato samples
Network Analysis
Separate co-occurrence networks were constructed for healthy and diseased tomato to explore differences in co-occurrence patterns. The healthy-tomato networks differed profoundly from the diseased-tomato networks, with the healthy-tomato networks characterized by more links between nodes, longer average path lengths, and greater clustering coefficient (Fig. 5a and c; Table S5). The co-occurrence networks of diseased tomato were more isolated and less dense than those of healthy tomato (Table S5). Moreover, the greater complexity of the healthy-tomato networks was reflected by their greater average degree (average 120.92 vs. 13.76) and higher network connectivity (connectance 0.23 vs. 0.06) than observed in the diseased tomato networks (Table S5). According to the taxonomy annotation, the healthy-tomato network was mainly dominated by phyla of Acidobacteria, Actinobacteria, Bacteroidetes, Firmicutes, and Proteobacteria (Fig. 5a). For the diseased tomato, the most dominant phyla for this network include Acidobacteria, Actinobacteria, Ascomycota, Bacteroidetes, Firmicutes, and Proteobacteria (Fig. 5c). The fungal phylum of Ascomycota was markedly increased (P < 0.05, Student’s t test) in the network of diseased tomato than that in the healthy-tomato network. Based on the NetShift analysis, Arthrobacter, Agromyces, Bacillus, Buttiauxella, Microbacterium, Lacibacterium, Methylobacillus, Pedobacter, Pseudomonas, Lentzea, Rhizobium, Streptomyces, and Plectosphaerella genera were identified as potential keystone taxa associated with Fol pathogen suppression in the healthy-tomato roots (Fig. 5b).
Visualization of microbial community co-occurrence network properties between healthy and diseased tomato plants. a Co-occurrence networks of healthy tomato samples. b Co-occurrence networks of diseased tomato. Nodes represent OTUs and colored by fungal and bacterial phylum. Links between the nodes indicate significant correlations. c Potential keystone taxa based on bacterial and fungal network analysis of healthy and diseased tomato. Node sizes are proportional to their scaled NESH score (a score identifying important microbial taxa of microbial association networks). The large red nodes represent particularly important driver taxa of healthy tomato, and these taxa names are shown in bold. Grayed-out nodes represent the ones present in both but directly interacting with the common sub-network in either healthy or diseased tomato microbiomes. Line colors indicate node (taxa) connections as follows: association present only in healthy tomato microbiomes (red edges), association present only in diseased tomato microbiomes (green edges), and association present in both diseased and healthy tomato microbiomes (blue edges)
Structure and Function of Healthy- and Diseased-Tomato Root-Associated Microbiomes
Metagenomic sequencing (MGS) was employed to reveal the functional differences between the healthy- and diseased-tomato microbial communities. In total, approximately 587,047,580 raw paired-end reads were generated from all tomato rhizosphere samples. Of these reads, 566,419,933 clean reads were used for de novo assembly, and we obtained 10,509,668 contigs, with a maximum contig length of 784,595 bp and an N50 of 928 bp (all contigs ≥ 300 bp) (Table S1). In total, 13,678,714 protein-coding genes with an average length of 464 bp were predicted from the assembly. After removing redundant sequences (identity > 95%), 8,598,069 unigenes were obtained. The taxonomic annotations revealed that bacteria were the predominant domain, with smaller fractions of eukaryotes, Archaea, and viruses detected based on the annotated unigenes. For the bacterial community, Proteobacteria (63.78 ± 6.10%), Actinobacteria (22.78 ± 4.12%), Bacteroidetes (5.37 ± 3.88%), and Acidobacteria (1.96 ± 1.06%) were the dominant phyla (relative abundance ≥ 1%) (Fig. S3). For the fungal community, Ascomycota was the most abundant phylum (0.48 ± 0.06%). At the genus level, Pseudomonas, Sphingobium, and Microbacterium were highly abundant, followed by Sphingopyxis, Streptomyces, Sphingomonas, Variovorax, Devosia, Agromyces, and Mesorhizobium (Fig. S4). According to the KEGG level 2 pathway annotations, carbohydrate metabolism, amino acid metabolism, global and overview maps, and energy metabolism were the most abundant functions (Table S6). Statistical analysis by the LEfSe software revealed a set of functional features that were significantly different between the healthy and diseased tomato plants. Compared with the microbes of diseased tomato, those of healthy tomato were associated with more functional genes (Fig. S5). The healthy-tomato microbiomes significantly enriched functions like fatty acid degradation and biofilm formation could respond to the pathogen suppression and nutrition exchange of healthy-tomato plants (Kruskal-Wallis test, P < 0.05) (Fig. S5).
Discussion
Previous studies revealed that the variation in bacterial communities between healthy and diseased tomato may be related to the health of individuals [20, 24]. However, which microbes are enriched and the primary taxonomic and functional differences between tomato plants that remain healthy and those that become diseased are still unclear [49]. According to the CCA test, no significant differences were found in soil physicochemical factors across all healthy and diseased samples. However, the root microbial communities of healthy and diseased tomato were significantly different, suggesting that abiotic environmental factors may not be the determinants of the divergence in the microbial community composition between healthy and diseased tomato in this study.
Plants can achieve specific suppression by selecting and enriching some microbial community members [23, 50]. Due to the close relationship between the microbes and their host plants, certain microbes are enriched from the outer layer of bulk soil to the root compartments, as driven by plant selection [51, 52]. Our results demonstrated that the bulk soil samples had the highest bacterial diversity and that only some of the bulk soil taxa are present in the root compartments. For example, we found that the genera Arthrobacter, Pyrenochaeta, Plectosphaerella, Pseudomonas, and Sphingobium were strongly selected from the bulk soil and enriched in the root compartments. Previous studies suggested that plants lacking genetic resistance to specific pathogens may enrich particular microorganisms for pathogen suppression [13, 50]. Our results demonstrated that microbial diversity was significantly higher in tomato plants that remained healthy than in those that became diseased, while the concentration of Fol pathogen was significantly lower in the healthy tomato than in the diseased tomato (P < 0.001). The differences in Fol pathogen density between healthy and diseased tomato imply that microbial diversity plays an important role in Fol disease outbreaks [24]. Several studies had found that high microbial diversity has been associated with higher resistance to pathogen invasions and plant infestation [10, 12, 20]; for example, increasing Pseudomonas species diversity significantly increased pathogen suppression and decreased the disease incidence through intensified resource competition and interference with the pathogen [12]. Moreover, the healthy tomato plants harbored more known BCA taxa than the diseased tomato plants, and these taxa may inhibit Fol disease [10, 23, 53]. For example, Pseudomonas, Trichoderma, and Bacillus species can suppress the Fol pathogen by producing antibiotics, regulating the jasmonic acid (JA)-/salicylic acid (SA)-related defensive pathways and forming biofilms to protect the plant in vitro or under artificial conditions [25, 54]. These enriched microbial species may help tomato to gain resistance against Fol pathogens. Together, these results suggested that Fol pathogen suppression is driven by microbial consortium rather than individual microorganism [23], and future studies could try to isolate these taxa and verify their disease suppression ability [7, 55].
According to the shotgun metagenomic data, pathways involved in biofilm formation were significantly enriched in healthy tomato compared with diseased tomato, suggesting that the microbial community in healthy tomato is beneficial for biofilm formation, which occupies the root-surface niches and prevents tomato infection by Fol [54]. Previous studies revealed that nutrient resource competition can indirectly impact pathogen inhibition by mediating processes involved in nutrient dynamics [13, 56]. We found several metabolisms and degradation-related pathways were significantly enriched in healthy tomato, suggesting greater access of the microorganisms to nutrient resources in the roots of healthy tomato than in those of diseased tomato, indicating the presence of more active nutrition exchange and microbe-plant interactions in the healthy tomato plants. Our results revealed that the diseased-tomato microbes contained significantly less functional features, such as the relative abundance of amino acid metabolism, fatty acid degradation, and tricarboxylic acid (TCA) cycle functions compared with healthy-tomato microbes. These findings demonstrated the negative effects of Fol on the tomato root zone microbiome at the community function level.
Poudel et al. [49] pointed out that co-occurrence network analysis of plant microbiomes can provide new perspectives for enhancing disease management and identifying candidate microbes affecting plant health. Our data showed that the microbial co-occurrence networks of healthy and diseased tomato were significantly different. In general, the network associated with healthy tomato comprised more connections and associated species, displayed longer average path lengths, and had higher clustering coefficient in structure. Similar to the differences in community composition, diversity, and functions, the network differences between tomato plants that remained healthy and those that became diseased were clear. Previous studies indicated that keystone taxa frequently co-occur with other microbes, and they may play primary roles in ecosystems by determining community dynamics, maintaining network structure, and connecting with other microbes in the network [46, 57, 58]. Based on NetShift analyses, several keystone OTUs in the healthy-tomato networks were characterized. According to the taxonomic annotation, the keystone taxa included Arthrobacter spp., Agromyces spp., Bacillus spp., Buttiauxella spp., Microbacterium spp., Lacibacterium spp., Methylobacillus spp., Pedobacter spp., Pseudomonas spp., Lentzea spp., Rhizobium spp., Streptomyces spp., and unassigned Plectosphaerella. Among these, Arthrobacter, Bacillus, Microbacterium, and Pedobacter [12, 59,60,61] are well known genera that comprise BCAs. Together, these findings suggest that fungal and bacterial communities have more interaction in healthy tomato than in diseased tomato and that the related keystone microbes may be positively correlated with suppression of Fol disease.
Based on these findings, we suggest an approach to controlling Fol soilborne disease by managing the natural fungal and bacterial community as a whole instead of selecting only one or several BCA species for disease suppression. These hypotheses need to be verified by culture-based experiments. Beyond the scope of this study, further studies should try to isolate these keystone taxa, as well as the species enriched by plant root exudates, and evaluate the consistency of disease suppression ability in vivo. Our study provides important baseline information for implementing agricultural management strategies to control Fol wilt disease using microbes that contribute to plant health.
References
Srinivas C, Devi DN, Murthy KN et al (2019) Fusarium oxysporum f. sp. lycopersici causal agent of vascular wilt disease of tomato: biology to diversity-a review. Saudi J Boil Sci 26:1315–1324. https://doi.org/10.1016/j.sjbs.2019.06.002
Gordon TR (2017) Fusarium oxysporum and the Fusarium wilt syndrome. Annu Rev Phytopathol 55:23–39. https://doi.org/10.1146/annurev-phyto-080615-095919
Duffy B, Défago G (1999) Macro-and microelement fertilizers influence the severity of Fusarium crown and root rot of tomato in a soilless production system. HortScience 34(2):287–291. https://doi.org/10.21273/HORTSCI.34.2.287
Gareau BJ (2010) A critical review of the successful CFC phase-out versus the delayed methyl bromide phase-out in the Montreal Protocol. Int Environ Agreements 10:209–231. https://doi.org/10.1007/s10784-010-9120-z
Martin FN (2003) Development of alternative strategies for management of soilborne pathogens currently controlled with methyl bromide. Annu Rev Phytopathol 41(1):325–350. https://doi.org/10.1146/annurev.phyto.41.052002.095514
Giotis C, Markelou E, Theodoropoulou A, Toufexi E, Hodson R, Shotton P, Shiel R, Cooper J, Leifert C (2009) Effect of soil amendments and biological control agents (BCAs) on soil-borne root diseases caused by Pyrenochaeta lycopersici and Verticillium alboatrum in organic greenhouse tomato production systems. Eur J Plant Pathol 123(4):387–400. https://doi.org/10.1007/s10658-008-9376-0
Toju H, Peay KG, Yamamichi M, Narisawa K, Hiruma K, Naito K, Fukuda S, Ushio M, Nakaoka S, Onoda Y, Yoshida K, Schlaeppi K, Bai Y, Sugiura R, Ichihashi Y, Minamisawa K, Kiers ET (2018) Core microbiomes for sustainable agroecosystems. Nat Plants 4(9):733–733. https://doi.org/10.1038/s41477-018-0245-3
Van Der H, Bardgett MG, RD and Van Straalen NM. (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310. https://doi.org/10.1111/j.1461-0248.2007.01139.x
Nuzzo A, Satpute A, Albrecht U, Strauss SL (2020) Impact of soil microbial amendments on tomato rhizosphere microbiome and plant growth in field soil. Microb Ecol. https://doi.org/10.1007/s00248-020-01497-7
Mendes R, Kruijt M, de Bruijn I, Dekkers E, van der Voort M, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332(6033):1097–1100. https://doi.org/10.1126/science.1203980
Vandenkoornhuyse P, Quaiser A, Duhamel M, Van L, Dufresne A (2015) The importance of the microbiome of the plant holobiont. New Phytol 206:1196–1206. https://doi.org/10.1111/nph.13312
Hu J, Wei Z, Friman VP, Gu SH, Wang XF, Eisenhauer N, Yang TJ, Ma J, Shen QR, Xu YC, Jousset A (2016) Probiotic diversity enhances rhizosphere microbiome function and plant disease suppression. Mbio 7(6). https://doi.org/10.1128/mBio.01790-16
Kwak M, Kong H, Choi K et al (2018) Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat Biotechnol 36:1100–1109. https://doi.org/10.1038/nbt.4232
Carrión VJ, Perez-Jaramillo J, Cordovez V, Tracanna V, de Hollander M, Ruiz-Buck D, Mendes LW, van Ijcken WFJ, Gomez-Exposito R, Elsayed SS, Mohanraju P, Arifah A, van der Oost J, Paulson JN, Mendes R, van Wezel GP, Medema MH, Raaijmakers JM (2019) Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366(6465):606–612. https://doi.org/10.1126/science.aaw9285
Chialva M, Zhou Y, Spadaro D, Bonfante P (2018) Not only priming: soil microbiota may protect tomato from root pathogens. Plant Signal Behav 13(8):1–9. https://doi.org/10.1080/15592324.2018.1464855
Finkel OM, Castrillo G, Paredes SH, González IS, Dangl JL (2017) Understanding and exploiting plant beneficial microbes. Curr Opin Plant Biol 38:155–163. https://doi.org/10.1016/j.pbi.2017.04.018
Larkin RP, Fravel DR (1998) Efficacy of various fungal and bacterial biocontrol organisms for control of Fusarium wilt of tomato. Plant Dis 82(9):1022–1028. https://doi.org/10.1094/PDIS.1998.82.9.1022
Fravel DR (2005) Commercialization and implementation of biocontrol. Annu Rev Phytopathol 43:337–359. https://doi.org/10.1146/annurev.phyto.43.032904.092924
Ousley MA, Lynch JM, Whipps JM (1993) Effect of Trichoderma on plant growth: a balance between inhibition and growth promotion. Microb Ecol 26:277–285. https://doi.org/10.1007/BF00176959
Zhou D, Feng H, Schuelke T, de Santiago A, Zhang Q, Zhang J, Luo C, Wei L (2019) Rhizosphere microbiomes from root knot nematode non-infested plants suppress nematode infection. Microb Ecol 78:470–481. https://doi.org/10.1007/s00248-019-01319-5
Niu B, Paulson JN, Zheng X, Kolter R (2017) Simplified and representative bacterial community of maize roots. PNAS 114(12):E2450–E2459. https://doi.org/10.1073/pnas.1616148114
Coyte KZ, Schluter J, Foster KR (2015) The ecology of the microbiome: networks, competition, and stability. Science 350(6261):663–666. https://doi.org/10.1126/science.aad2602
Gómez Expósito R, de Bruijn I, Postma J, Raaijmakers JM (2017) Current insights into the role of rhizosphere bacteria in disease suppressive soils. Front Microbiol 8(2529). https://doi.org/10.3389/fmicb.2017.02529
Wei Z, Gu Y, Friman VP, Kowalchuk GA, Xu Y, Shen Q, Jousset A (2019) Initial soil microbiome composition and functioning predetermine future plant health. Sci Adv 5(9):eaaw0759. https://doi.org/10.1126/sciadv.aaw0759
Martínez-Medina A, Fernandez I, Lok GB, Pozo MJ, Pieterse CMJ, Van Wees SCM (2017) Shifting from priming of salicylic acid-to jasmonic acid-regulated defences by Trichoderma protects tomato against the root knot nematode Meloidogyne incognita. New Phytol 213:1363–1377. https://doi.org/10.1111/nph.14251
Edwards J, Johnson C, Santos-Medellin C et al (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. PNAS 112(8):E911–E920. https://doi.org/10.1073/pnas.1414592112
Xu N, Tan G, Wang H, Gai X (2016) Effect of biochar additions to soil on nitrogen leaching, microbial biomass and bacterial community structure. Eur J Soil Biol 74:1–8. https://doi.org/10.1016/j.ejsobi.2016.02.004
Adams R, Miletto M, Taylor J et al (2013) Dispersal in microbes: fungi in indoor air are dominated by outdoor air and show dispersal limitation at short distances. ISME J 7:1262–1273. https://doi.org/10.1038/ismej.2013.28
Inami K, Yoshioka C, Hirano Y, Kawabe M, Tsushima S, Teraoka T, Arie T (2010) Real-time PCR for differential determination of the tomato wilt fungus, Fusarium oxysporum f. sp. lycopersici, and its races. J Gen Plant Pathol 76:116–121. https://doi.org/10.1007/s10327-010-0224-7
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Langmead B, Salzberg S (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Segata N, Waldron L, Ballarini A, Narasimhan V, Jousson O, Huttenhower C (2012) Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods 9:811–814. https://doi.org/10.1038/nmeth.2066
Abubucker S, Segata N, Goll J, Schubert AM, Izard J, Cantarel BL, Rodriguez-Mueller B, Zucker J, Thiagarajan M, Henrissat B, White O, Kelley ST, Methé B, Schloss PD, Gevers D, Mitreva M, Huttenhower C (2012) Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput Biol 8(6):e1002358. https://doi.org/10.1371/journal.pcbi.1002358
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30(21):3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Edgar R (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Nilsson RH, Tedersoo L, Ryberg M, Kristiansson E, Hartmann M, Unterseher M, Porter TM, Bengtsson-Palme J, Walker DM, de Sousa F, Gamper HA, Larsson E, Larsson KH, Kõljalg U, Edgar RC, Abarenkov K (2015) A comprehensive, automatically updated fungal ITS sequence dataset for reference-based chimera control in environmental sequencing efforts. Microbes Environ 30(2):145–150. https://doi.org/10.1264/jsme2.ME14121
Deshpande V, Wang Q, Greenfield P, Charleston M, Porras-Alfaro A, Kuske CR, Cole JR, Midgley DJ, Tran-Dinh N (2016) Fungal identification using a Bayesian classifier and the Warcup training set of internal transcribed spacer sequences. Mycologia 108(1):1–5. https://doi.org/10.3852/14-293
Oksanen AJ, Blanchet FG, Friendly M, et al (2016) Vegan: community ecology package. https://github.com/vegandevs/vegan
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71(12):8228–8235. https://doi.org/10.1128/AEM.71.12.8228-8235.2005
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer, New York
Csardi G, Nepusz T (2006) The igraph software package for complex network research. InterJournal Complex Systems 1695
Clauset A, Newman ME, Moore C (2004) Finding community structure in very large networks. Phys Rev E 70:066111. https://doi.org/10.1103/PhysRevE.70.066111
Agler MT, Ruhe J, Kroll S, Morhenn C, Kim ST, Weigel D, Kemen EM (2016) Microbial hub taxa link host and abiotic factors to plant microbiome variation. PLoS Biol 14(1):e1002352. https://doi.org/10.1371/journal.pbio.1002352
Van Der Heijden MG, Hartmann M (2016) Networking in the plant microbiome. PLoS Biol 14(2):e1002378. https://doi.org/10.1371/journal.pbio.1002378
Kuntal BK, Chandrakar P, Sadhu S, Mande SS (2019) ‘NetShift’: a methodology for understanding ‘driver microbes’ from healthy and disease microbiome datasets. ISME J 13:442–454. https://doi.org/10.1038/s41396-018-0291-x
Poudel R, Jumpponen A, Schlatter D et al (2016) Microbiome networks: a systems framework for identifying candidate microbial assemblages for disease management. Phytopathology 106(10):1083–1096. https://doi.org/10.1094/PHYTO-02-16-0058-FI
Santhanam R, Luu VT, Weinhold A, Goldberg J, Oh Y, Baldwin IT (2015) Native root-associated bacteria rescue a plant from a sudden-wilt disease that emerged during continuous cropping. PNAS 112(36):E5013–E5020. https://doi.org/10.1073/pnas.1505765112
Hirsch PR, Miller AJ, Dennis PG (2013) Do root exudates exert more influence on rhizosphere bacterial community structure than other rhizodeposits? In Molecular Microbial Ecology of the Rhizosphere, F.J. de Bruijn (Ed.). https://doi.org/10.1002/9781118297674.ch22
Reinhold-Hurek B, Bünger W, Burbano CS, Sabale M, Hurek T (2015) Roots shaping their microbiome: global hotspots for microbial activity. Annu Rev Phytopathol 53:403–424. https://doi.org/10.1146/annurev-phyto-082712-102342
Renault D, Laparie M, McCauley SJ, Bonte D (2018) Environmental adaptations, ecological filtering, and dispersal central to insect invasions. Annu Rev Entomol 63:345–368. https://doi.org/10.1146/annurev-ento-020117-043315
Chen Y, Yan F, Chai Y, Liu H, Kolter R, Losick R, Guo JH (2013) Bacillus subtilis and plant biocontrol. Environ Microbiol 15:848–864. https://doi.org/10.1111/j.1462-2920.2012.02860.x
Rahman A, Sharifah FS, Singh E, Pieterse CM, Schenk PM (2018) Emerging microbial biocontrol strategies for plant pathogens. Plant Sci 267:102–111. https://doi.org/10.1016/j.plantsci.2017.11.012
Hu L, Robert CAM, Cadot S, Zhang X, Ye M, Li B, Manzo D, Chervet N, Steinger T, van der Heijden MGA, Schlaeppi K, Erb M (2018) Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat Commun 9:2738. https://doi.org/10.1038/s41467-018-05122-7
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198. https://doi.org/10.1016/j.soilbio.2016.03.017
Banerjee S, Schlaeppi K, Van Der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576. https://doi.org/10.1038/s41579-018-0024-1
Colombo EM, Kunova A, Pizzatti C, Saracchi M, Cortesi P, Pasquali M (2019) Selection of an endophytic Streptomyces sp. strain DEF09 from wheat roots as a biocontrol agent against Fusarium graminearum. Front Microbiol 10(2356). https://doi.org/10.3389/fmicb.2019.02356
Das K, Prasanna R, Saxena AK (2017) Rhizobia: a potential biocontrol agent for soilborne fungal pathogens. Folia Microbiol 62:425–435. https://doi.org/10.1007/s12223-017-0513-z
Berendsen RL, Vismans G, Yu K, Song Y, de Jonge R, Burgman WP, Burmølle M, Herschend J, Bakker PAHM, Pieterse CMJ (2018) Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J 12:1496–1507. https://doi.org/10.1038/s41396-018-0093-1
Acknowledgments
We thank Dr. Wei Liu and Dr. Shenyong Fu (Institute of Microbiology, CAS) for the greenhouse experiments and sample collections, and Dr. Junmin Liang (Institute of Microbiology, CAS) for the editorial support of the manuscript.
Funding
This study was financially supported by NSFC 31725001. X. Zhou received financial support for his studentship (QYZDB-SSW-SMC044). C.K. Tsui acknowledges CAS153211KYSB20160029 for supporting his visit to Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
L. Cai planned and supervised this research; X. Zhou performed the experiments with the assistant of J.T. Wang; X. Zhou conducted the bioinformatic analyses and mainly wrote the manuscript. W.H. Wang and C. K. Tsui contributed to the data analysis and revision of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Fig. S1
Rarefaction curves of tomato bacterial and fungal OTUs. a) Rarefaction curves of tomato bacterial OTUs. b) Rarefaction curves of tomato fungal OTUs. (EPS 835 kb)
Fig. S2
Concentrations of Fol in healthy and diseased tomato plants. a) Comparison of relative abundance of Fol in healthy and diseased tomato groups. b) Comparison of absolute abundance of Fol in healthy and diseased tomato groups. (EPS 757 kb)
Fig. S3
Relative abundance of different bacterial and fungal taxa at the phylum level as revealed by shotgun metagenomics. (EPS 3112 kb)
Fig. S4
Relative abundance of different bacterial and fungal taxa at the genus level as revealed by shotgun metagenomics. (EPS 2394 kb)
Fig. S5
Distinct active functions identified using the LEfSe method. The functional annotations were obtained using HUMAnN2 against the KO database. LDA scores showed significant functional differences between the healthy and diseased groups. Green represents the functions enriched in healthy tomato samples, and the red represents the functions enriched in diseased tomato samples. (EPS 1953 kb)
ESM 6
(XLSX 63 kb)
Rights and permissions
About this article
Cite this article
Zhou, X., Wang, JT., Wang, WH. et al. Changes in Bacterial and Fungal Microbiomes Associated with Tomatoes of Healthy and Infected by Fusarium oxysporum f. sp. lycopersici. Microb Ecol 81, 1004–1017 (2021). https://doi.org/10.1007/s00248-020-01535-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01535-4