Abstract
Backgroud and aims
This study was conducted to reveal the genetic diversity of soybean-nodulating rhizobia in Nepal in relation to climate and soil properties.
Method
A total of 102 bradyrhizobial strains were isolated from the root nodules of soybeans cultivated in 12 locations in Nepal varying in climate and soil properties, and their genetic diversity was examined based on 16S rDNA, ITS regions of 16S–23S rDNA, nodC and nifH. In vitro growth properties of some representative strains were examined to elucidate their characteristic distribution in Nepal.
Results
Four species of the genus Bradyrhizobium were isolated, and B. japonicum dominated at temperate locations, while in subtropical locations, B. elkanii, B. yuanmingense, and B. liaoningense dominated at acidic, moderately acidic, and slightly alkaline soils, respectively. The relative nodule occupancies could not be fully explained by their in vitro growth properties. Similar nodC and nifH genes among the strains suggested co-evolution of these genes also in Nepal, probably through horizontal gene transfer.
Conclusions
The influence of climate and soil pH on diversity at the sub-species level was revealed. It is concluded that the highly diverse climate and soils in Nepal might be conducive for the existence of diverse soybean rhizobial strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean-nodulating rhizobia are genetically diverse and are classified into different genera and species. The slow-growing soybean rhizobia are distributed in five species of Bradyrhizobium: B. japonicum (Jordan 1982), B. elkanii (Kyukendall et al. 1992), B. liaoningense (Xu et al. 1995), B. yuanmingense (Yao et al. 2002), and B. canariense (Vinuesa et al. 2005).Two fast-growing species of Sinorhizobium (Ensifer), namely S. fredii (Chen et al. 1988) and S. xinjiangense (Peng et al. 2002), are also reported to be soybean-nodulating rhizobia. Mesorhizobium tianshanense (Chen et al. 1995), which has varying growth rates, has also been reported as a soybean rhizobial species.
Evidence has suggested that temperature and soil pH influence the genetic diversity of soybean rhizobia at the species level in a particular environment. Among slow growers, B. japonicum and B. elkanii have been found in various climates across the world (Ando and Yokoyama 1999; Chen et al. 2000; Li et al. 2011); Man et al. 2008; Risal et al. 2010; Saeki et al. 2006; Suzuki et al. 2008; van Berkum and Fuhrmann 2000; Vinuesa et al. 2008; Wasike et al. 2009; Yang and Zhou 2008, and B. japonicum was dominant in the soils of cooler environments in Japan and Nepal, as reported (Suzuki et al. 2008; Vinuesa et al. 2008). The other three slow-growing species (B. liaoningense, B. yuanmingense and B. canariense) have not been surveyed worldwide, probably due to their newness as separate species compared to B. japonicum and B. elkanii, or to their biogeographic specificities. B. liaoningense, which is a relatively extra-slow-growing species, has been isolated from alkaline soils in temperate to subtropical climates of south and south-east Asia (Xu et al. 1995; Saeki et al. 2005; Appunu et al. 2008; Vinuesa et al. 2008; Yang and Zhou 2008; Han et al. 2009; Li et al. 2011). The occurrence of B. yuanmingense in India (Appunu et al. 2008), Kenya (Wasike et al. 2009) and Nepal (Risal et al. 2010) suggests its preference for warm climates, although B. yuanmingense is reported in the alkaline-saline soils of temperate China Li et al. (2011). B. canariense, which was first reported in soybean nodules in north and northeast China (Yang and Zhou 2008), is characterized as more acid tolerant than B. japonicum (Vinuesa et al. 2005). Fast-growing S. fredii and S. xinjiangense are acid-producing rhizobia (Chen et al. 1988) and have been isolated in saline-alkaline soils in China, Vietnam and Japan (Appunu et al. 2009; Han et al. 2009; Li et al. 2011); Man et al. 2008; Peng et al. 2002; Saeki et al. 2005; Suzuki et al. 2008. The isolation of M. tianshanense from arid and saline soils has also shown its specificity, but this isolation has only been reported in a study in the Xinjiang Province of China (Chen et al. 1995).
Nepal, a small Himalayan country, varies greatly in topography, climate and vegetation; the elevation ranges from 68 to 8,848 m in a mere 150 to 250-km south–north transect. Soybean is one of the major summer legumes in Nepal and is considered to have been domesticated in the first millennium AD (Smartt and Hymowitz 1985). It is cultivated from southern warm plains to northern mountains up to 2,000 m in elevation in Nepal. However, less is known about the soybean’s nodule microsymbionts in an ecological context despite its long history of cultivation and wide distribution across the country. Among a few studies on soybean rhizobial diversity in Nepal, some genetically distinct B. japonicum strains compared to those in other Asian countries have been reported in a soil in the Kathmandu Valley (1,300 m) (Vinuesa et al. 2008). Recently, the diversity of soybean bradyrhizobia was assessed in five mountain soils of Nepal ranging from 1,500 to 2,600 m in elevation with soil pH levels ranging from 5.2 to 6.2, and a dominant presence of B. elkanii with minor populations of B. japonicum and B. yuanmingense was reported (Risal et al. 2010). However, these reports do not provide information on the diversity of soybean rhizobia in the southern warmer lowlands in Nepal. In addition, they used a soil trap culture method to isolate rhizobia under laboratory conditions in which the possible influence of environmental factors on rhizobial diversity in nodules might be undermined. In the present study, we isolated indigenous soybean rhizobia from root nodules collected from fields ranging over a wider climatic region (225–1,950 m in elevation) and with a wider range of soil properties (pH 3.6–7.4) in Nepal. We found higher genetic diversity among the representative isolates than was found in the previously reported Nepalese bradyrhizobia. In addition, we examined the in vitro growth properties of some representative isolates in response to ranges of temperature and pH to elucidate why one genotype dominated in a particular type of climate and soil pH in Nepal.
Materials and methods
Nodule sampling and isolation of rhizobia
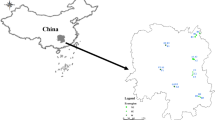
During the summer of 2007, root nodule samples were collected from the 12 soybean-growing locations in Nepal, six from each temperate and subtropical region (Fig. 1 and Table 1). The altitude of the temperate locations varies from 900 to 1,950 m, and that of the subtropical locations from 225 to 625 m. The average minimum and maximum monthly temperatures at the highest location (Charikot, Dolakha) are −1 and 22°C, respectively, while at the lowest location (Rampur, Chitwan), they are 8 and 35°C, respectively (www.mfd.gov.np). None of the sampling sites had a known history of rhizobial inoculation. Representative nodule samples were collected at each site and kept in a preservation vial until the isolation of the rhizobia. Soil samples were also collected and analyzed for pH using the 1:1 soil-to-water (w/v) method, for organic matter by the Walkley and Black method (Walkley and Black 1934), for available P by Olsen’s bicarbonate method (Olsen et al. 1954), and for available K by the flame photometer method after extracting with 1 N ammonium acetate (Knudsen et al. 1982). For the isolation of rhizobia, the nodules were surface-sterilized in 95% ethanol for 1 min, and then in 1% sodium hypochlorite for 50 min. After the nodules were washed in sterile water, each nodule was crushed, streaked onto a yeast extract-mannitol agar (YMA) (Vincent 1970) plate, and incubated in the dark at 25°C. After 5–12 days, a single colony per plate was isolated and maintained for further study.
Map of Nepal showing sampling locations in different climatic regions and regions with different soil pH levels. The location numbers from 1 to 12 correspond with Table 1
DNA preparation and phylogenetic analysis of isolated rhizobia
Genomic DNA was extracted and purified as described previously (Saeki et al. 2005). The targeted genes for PCR amplification and DNA sequencing were (1) 16S rRNA, (2) the 16S–23S rRNA internal transcribed spacer (ITS) region, (3) nodC and (4) nifH. PCR amplification of 16S rDNA was accomplished by the primers fD1 and rP2 designed for most eubacteria (Weisburg et al. 1991). The primers 1512f and LS23r (Suzuki et al. 2008) were used to amplify the ITS region. Similarly, for nodC and nifH, nodCf2 (5′ CTS AAC GTC GAY TCS GAT AC 3′) and nodCr2 (5′ AAG GTA CTY CGY GCC CAV C 3′), and nifHf2 (5′ CTC GAG GAC GTR ATG AAG GT 3′) and nifHr2 (5′ GCA CRA AGT AGA TCA GYT GRG 3′), were designed, respectively. The PCR mixture was prepared by mixing reaction buffer, dNTP, Taq DNA polymerase (BIONEER, Seoul, Korea) and primers together with the template DNA. The PCR cycle for 16S rDNA and ITS consisted of a pre-run at 94°C for 2 min, denaturation at 94°C for 30 s, annealing at 50°C for 30 s and extension at 72°C for 90 s. The extension time for nodC and nifH was set to 1 min. The cycle was repeated a total of 30 times and then followed by a final post-run at 72°C for 90 s.
The successful PCR product revealed by gel electrophoresis was mixed with SOPE resin (Edge Biosystems, Gaithersburg, USA) at the ratio of 1:5, and column-purified with a Performa DTR Gel Filtration Cartridge (Edge Biosystems). Partial fragments of the purified PCR products were cycle sequenced using the corresponding forward and/or reverse primers. Sequence PCR was performed for 1 min at 96°C and then 25 cycles of 10 s at 96°C, 5 s at 50°C and 4 min at 60°C. The products purified with Performa DTR Gel Filtration Cartridges were precipitated by ethanol, dissolved in Hidi-formamide (Applied Biosystems, Tokyo, Japan), and then denatured at 96°C for 2 min and ice chilled. The sequencing analysis was carried out on an ABI Prism, 3100-Avant Genetic Analyzer (Hitachi, Tokyo, Japan).
Taxonomic nomenclature for each isolate was designated based on the homology of their 16S rDNA and/or ITS to known soybean-nodulating rhizobial genera and species in the database (www.ddbj.nig.ac.jp) by performing a BLAST (Altschul et al. 1997) search. Multiple sequence alignments of all 16S rDNA, ITS, nodC and nifH genes were constructed with ClustalW1.83 (Thompson et al. 1994). Alignments were manually edited and their phylogenetic trees with the related reference strains were constructed using ClustalW1.83.
Growth properties of rhizobia in vitro
The growth of a total of eight strains, each representing one of the dominant and minor species isolated from locations having different soil pH values and climates, was examined at temperatures from 15 to 40°C and pHs from 4.5 to 7.0 in YM liquid medium. Temperatures for strains belonging to temperate locations were 15–30°C and those for subtropical locations were 20–40°C. The selected strains included two strains each of B. japonicum and B. yuanmingens, and four of B. elkanii. Among the selected strains, B. japonicum Cha-4 and Kha-1 were dominant and B. elkanii Cha-2 and Kha-8 were minor at the temperate locations Charikot (soil pH 3.4) and Khadichaur (soil pH 4.8), respectively. Likewise, B. yuanmingense Mug-2 and B. elkanii Man-8 were dominant and B. elkanii Mug-1 and B. yuanmingense Man-14 were minor at the subtropical locations Muglin (soil pH 6.4) and Mangalpur (soil pH 5.0), respectively. The growth medium was buffered either with 6 mM of MES (pH 4.5–6.0) or HEPES (pH 7.0). A constant pH of 6.0, which was optimum for the growth of most of the isolates, was used for the temperature experiment, and similarly, the most favorable temperature of 30°C for most of the strains was maintained for the pH experiment. The experiments were carried out in triplicate by shaking 5 mL of culture in 20-mL test tubes reciprocally at 150 rpm in the dark. The initial cell number in each test tube was adjusted to 3.8 × 106 mL−1 with preincubated cells in YM medium for 7 days at 25°C on the basis of microscopic counts after ethidium bromide staining (Someya et al. 2003). Optical density (OD) at 600 nm of the bacterial culture was checked daily until 8 days of incubation, corresponding to the late log phase at the most favorable pH (6.0) and temperature (30°C).
Nucleotide sequence accession numbers
All sequences have been deposited in the DNA Data Bank of Japan under accession numbers AB565790–AB565891 for the 16S rDNA, AB565892–AB565940 for the ITS, AB565941–AB565989 for nifH, and AB565990–AB566039 for nodC.
Results
Isolation of soybean nodule rhizobia
Six to 16 rhizobial strains per location, making a total of 102 strains, were isolated and examined (Table 1). The isolates were designated by the first three letters of the name of the corresponding location followed by a number in the order of the isolation from that location. For example, ‘Cha-1’ stands for the first strain isolated from the location ‘Charikot’. Accordingly, the second strain isolated from the same location was named ‘Cha-2’, and so on.
Phylogenetic analysis of 16S rRNA genes
Phylogenetic analysis based on a 623-bp fragment of 16S rRNA genes divided the strains into eight main groups (Fig. 2). Among them, 38 strains in groups I, II and III had sequences that were 99.7–100% similar to those of the B. elkanii reference strains, and thus will be referred to as B. elkanii hereafter. The other five groups (IV, V, VI, VII and VIII) consisted of 64 isolates, and their sequences were 99.7–100% similar to those of B. japonicum, B. liaoningense, B. yuanmingense and B. canariense, but were not distinguished at a species level. The number of strains in each of the eight groups varied from 4 to 33.
Phylogenetic tree based on the 623-bp alignment of the nucleotide sequences of 16S rRNA genes. The scale bar indicates the number of substitutions per site. Three-letter location codes (used to designate strains), with the number of strains in parentheses, shown in bold and plain letters represent strains that were isolated from temperate and subtropical locations, respectively. Those with underlines, intact, bars and double bars indicate soil pHs of 3.6–4.3, 4.8–5.1, 5.8–6.4 and 7.4, respectively. Isolates grouped into eight clusters (I–VIII) are marked by braces on the right. The accession numbers of the sequences obtained in this study are AB565790–AB565891. Bootstrap values greater than 50% are indicated in the corresponding nodes
Among the three groups of B. elkanii strains, groups I and III belonged to both temperate and subtropical locations and are indicated by bold and plain letters in Fig. 2, respectively. Sequences of strains in groups I and III were most homologous with those of B. elkanii 76T and USDA94, respectively. On the other hand, the group II strains belonging to three subtropical locations had less homologous sequences in the database. Among the eight locations where B. elkanii strains were isolated, higher diversity among the strains was observed at Mangalpur and Rampur, as revealed by their presence in all three groups followed by Khadichaur and Pachkhal in two groups. A slight effect of soil pH on the diversity among B. elkanii strains was observed, as strains in groups I and II belonged to locations with a narrower range of soil pH (4.8–5.1) compared to the wider range (pH 3.6–6.4) for group III, although the number of strains was higher in group III.
Group IV included isolates from Ghorahi, a subtropical location with moderately acidic soil (pH 5.8). Group V comprised strains from one subtropical and two temperate locations. In this group, strains isolated from a subtropical location (Mangalpur) had sequences that were identical with those of B. japonicum USDA124 and B. liaoningense CCBAU33113, and those from temperate locations (Khumaltar and Khadichaur) with soil pH levels that were similar to one another (pH 4.8–5.1) had sequences that were with that of B. japonicum USDA110. Sequences of the group VI strains isolated from two subtropical locations, Deukhuri (soil pH 7.4) and Muglin (soil pH 6.4), were identical with those of some reference strains of B. japonicum, B. yuanmingense and B. canariense. Thirty-three strains from five temperate locations with a wide range of soil pH (3.6–4.8) comprised group VII, and their sequences were 99.8–100% similar to that of B. japonicum USDA6T. Group VIII included five strains from a subtropical location, Deukhuri (soil pH 7.4), with sequences identical to those of B. japonicum USDA135, B. liaoningense LMG18230T and B. canariense MH547, and one strain from a temperate location, Khadichaur (soil pH 4.8), with a sequence identical to that of B. japonicum RLA12. Within Groups IV–VIII, the strains of Khadichaur and Deukhuri appeared in three and two groups, respectively, while the strains from the rest of the locations showed their presence in only one phylogenetic group.
Diversity of 16S–23S rDNA ITS region
The use of the 16S rRNA gene sequence to delineate the species within the genera Bradyrhizobium is considered insufficient. To obtain higher phylogenetic resolution, a total of 48 strains isolated from different locations representing different clusters (Fig. 2) in the phylogenetic tree of 16S rRNA genes were selected for phylogenetic analysis based on the ITS region. As shown in Fig. 3, the B. elkanii clade was clustered into four groups labeled a to d. Strains in groups a, b and d belonged to both temperate and subtropical locations. Locations corresponding to groups a and b had wider ranges of soil pH (4.3–5.1, and 3.6–5.8, respectively), whereas those in group d showed a relatively narrower range (pH 4.8–5.1). On the other hand, group c had strains from two subtropical locations that were close to one another in terms of soil pH (5.0 and 5.1) and also in distance (locations 10 and 11 in Fig. 1). The sequences of groups b, c and d were most homologous with those of B. elkanii strains USDA94, USDA61 and USDA 76T, respectively, while no sequence homologous to those of the group a strains was present in the database. The clustering pattern in ITS was different from the 16S rRNA gene phylogeny. Strains in the group III phylotype of 16S rRNA gene formed four separate clusters in ITS, one of which (group a) included strains from groups I, II and III. Variation in both the 16S rRNA gene and ITS phylogenies among B. elkanii strains was unrelated to soil properties.
Phylogenetic tree based on 622–798-bp alignment of nucleotide sequences of the 16S–23S rDNA internal transcribed spacer (ITS) regions. The scale bar indicates the number of substitutions per site. Strains shown in bold and plain letters were isolated from temperate and subtropical locations, respectively. Those with underlines, intact, bars and double bars indicate soil pHs of 3.6–4.3, 4.8–5.1, 5.8–6.4 and 7.4, respectively. Isolates grouped into 12 clusters (a–l) are marked by braces on the right of the tree followed by their taxonomic nomenclature and corresponding 16S rRNA gene types (Fig. 2) with brackets. The accession numbers of the sequences obtained in this study are AB565892–AB565940. Bootstrap values greater than 50% are indicated in the corresponding nodes
The strains other than B. elkanii, which were not clearly delineated at the species level by 16S rRNA genes, were distinctly separated into B. japonicum, B. yuanmingense and B. liaoningense in the ITS phylogeny. B. japonicum strains were further divided into groups e, k and l. Similarly, strains in four groups (f–i) were designated B. yuanmingense and a single group (j) B. liaoningense. Isolates belonging to each of the three species formed lineages based on the climatic region and soil pH of the locations where they were isolated. Groups e and k belonged to three temperate locations, Khumaltar, Khadichaur and Pokhara, with a narrow range of soil pH (4.8–4.9), with the exception of a strain from Naudada (soil pH 4.1), while group l belonged to three other temperate locations (Charikot, Naudada and Godawari) with much lower soil pH (3.6–4.3). Despite the higher homology among the sequences of the group k and l strains, the 48-bp shorter ITS of the latter clustered them into a completely different group from the former (data not shown). The strains of groups f to i appeared in a common node belonging to four subtropical locations, Deukhuri (soil pH 7.4), Muglin (soil pH 6.4), Ghorahi (soil pH 5.8) and Mangalpur (soil pH 5.0), respectively. B. yuanmingense strain CCBAU10071 also appeared in the common node; however, there was no sequence of the previously reported soybean rhizobia that was homologous to those of the strains in any of the groups from f to i. Two representative strains from Deukhuri (soil pH 7.4) separated distinctly in group j were recognized as B. liaoningense based on their sequence that was closer to the type strain B. liaoningense LMG18230 and their extra-slow growing properties (data not shown). No strain belonged to B. canariense, although strains in groups VI and VIII of 16S rRNA gene phylogeny co-clustered with some reference strains of this species.
Phylogeny of symbiotic genes
Strains chosen for studying ITS phylogeny were also used for the phylogenetic analysis of two symbiotic genes, nodC and nifH. Phylogenetic analysis of nodC and nifH genes divided the isolates into five major clusters (groups 1–5) (Fig. 4). Group 1 strains corresponded to all three 16S rRNA and four ITS phylotypes of B. elkanii, and were isolated from soils with a wide range of soil pH (3.6–6.4) and a wide variety of climates (225–1,950 m). Strains in groups 2–5 were common between the nodC and nifH phylogenies in each of the corresponding clusters, including the reference strains. Strains in group 2 consisted of all of the B. japonicum and B. liaoningense strains, although they were distinctly separated in the phylogenetic tree of the ITS (groups e, j, k, l) and 16S rRNA gene (group V, VII and VIII). In contrast, the nodC and nifH genes of B. yuanmingense strains varied greatly, as shown by their three distinct lineages (groups 3–5) corresponding to the 16S rRNA gene clusters IV, V and VI. However, except for the strains in group 5, most homologous reference strains for groups 3 and 4 in the database were B. japonicum and B. liaoningense, respectively. Group 3 (Man-5, Man-8 and Man-14) comprised strains isolated from Mangalpur (a subtropical location with soil pH 5.0), and group 4 comprised strains isolated from soil with a slightly higher soil pH (5.8) in Ghorahi. Strains isolated from Deukhuri and Muglin with near neutral soil pH (6.4 and 7.4) shared a common node (group 6) even with about 150 km distance between them (Fig. 1).
Phylogenetic trees of nodC (left) and nifH (right) genes based on the 353-bp and 311-bp alignments of nucleotide sequences, respectively. The scale bar indicates the number of substitutions per site. Strains shown in bold and plain letters were isolated from temperate and subtropical locations, respectively. Those with underlines, intact, bar and double bars indicate soil pHs of 3.6–4.3, 4.8–5.1, 5.8–6.4 and 7.4, respectively. Isolates grouped into five clusters are marked by braces with their corresponding 16S rDNA (Fig. 2) and ITS types (Fig. 3) on the right. The accession numbers of the nodC and nifH sequences obtained in this study are AB565990–AB566039 and AB565941–AB565989, respectively. Bootstrap values greater than 50% are indicated in the corresponding nodes
Biogeographical distribution of rhizobial strains
The population of each of the four species differed greatly along with the climatic regions and soil pH (Table 1; Fig. 5). In the temperate locations, strains of B. elkanii and B. japonicum were isolated, and the population of the latter dominated the former. In the present study, 43 out of 50 isolates were B. japonicum. Among six temperate locations, B. elkanii strains were isolated only from Khadichaur (900 m), Godawari (1,400 m) and Charikot (1,950 m).
At subtropical locations, the isolated strains belonged to B. elkanii, B. yuanmingense and B. liaoningense, but none of the strains were B. japonicum. Variation in the populations of these three species at different locations related significantly with soil pH, while no correlation was observed with the other soil properties listed in Table 1 (data not shown). Among the locations with relatively lower soil pH (4.8–5.1), B. elkanii strains were isolated at Pachkhal (soil pH 4.8) and Rampur (soil pH 5.1), while B. yuanmingense strains were isolated at only one location (Mangalpur, soil pH 5.0). In contrast, at Dang and Muglin, which had higher soil pH (5.8 and 6.4, respectively), B. yuanmingense was dominant over B. elkanii. At a location with slightly alkaline soil (Deukhuri, soil pH 7.4), B. liaoningense strains were observed to be dominant over B. yuanmingense.
The number of 16S rRNA gene clusters also varied along with the corresponding climate and soil pH of the isolated strains (Table 2). As the strains used for analyzing ITS and symbiotic gene sequences were not selected randomly, they were not considered for this comparison. Seven 16S rRNA gene phylotypes were observed in subtropical locations compared to only five in temperate locations. Groups, II, IV and VI were absent in temperate locations, as was group VII in subtropical locations. In temperate locations, three phylotypes (groups I, V and VIII) disappeared as the soil pH decreased, and only two phylotypes (III and VII) remained in strongly acidic soils (pH 3.6–4.3). In subtropical locations, on the other hand, the dominant species changed along with the change in soil pH.
Growth in YM liquid medium
The growth properties of some representative strains at different temperatures and soil pH levels were examined to determine whether these properties could explain the effect of temperature and pH on the relative nodule occupancies of different species in different climates and at different soil pH levels as shown in Fig. 5. The tested strains responded differently at various temperatures (Fig. 6). Comparisons were made on the basis of the average minimum and maximum temperatures of the corresponding locations during the soybean cultivation time: 15–25°C at Charikot, 20–30°C at Khadichaur, and 25–35°C at Muglin and Mangalpur. The results showed that the growth of the representative dominant strain of Charikot (B. japonicum Cha-4) was the same as that of the minor strain (B. elkanii Cha-2) at 15 and 20°C, while at 25°C, the growth of the minor strain was superior to that of the dominant strain. In contrast, the growth of the minor strain of Khadichaur (B. elkanii Kha-8) was superior to that of the dominant strain at the lower temperature (i.e., 20°C), and that of the dominant strain was superior to that of the minor strain at 25 and 30°C. For the strains from subtropical locations, the growth of dominant strains was superior to that of the minor strains as the temperature increased. In addition, the dominant strain of Muglin (B. yuanmingense Mug-2) grew even at 40°C, while the minor strain (B. elkanii Mug-1) could not grow (data not shown).
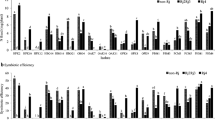
The OD600 of the culture of representative dominant and minor bradyrhizobial strains isolated from the root nodules of soybeans cultivated at different locations in Nepal in response to the temperatures prevalent at the respective locations during soybean cultivation. Each value represents the mean of a triplicate. Vertical bars denote standard deviations. pH was set at 6.0
Although the soil pH of the corresponding four locations ranged from 3.6–6.4, no growth of the strains was observed at pH 4.0 in YM liquid medium. The data on the growth of strains at pH 4.5 to 7.0 are presented in Fig. 7. At pH 4.5, there was no growth of the dominant strain of Charikot (soil pH 3.6), but its minor counterpart grew successfully. Similarly, the growth of the dominant strain of Khadichaur (soil pH 4.8), was greatly reduced at pH 4.5 compared to the minor strain. However, at pH 5.0 and 6.0, the growth of the dominant strain was superior to that of the minor strain. The growth of the dominant strain of Muglin (soil pH 6.4) was superior at all conditions (pH 5.0, 6.0 and 7.0). Between the strains of Mangalpur (soil pH 5.0), the growth of the dominant strain was superior at lower pH (4.5 and 5.0), while at pH 6.0, both strains showed similar growth initially, but the minor strain became superior to the dominant strain as the number of incubating days increased.
The OD600 of the culture of representative bradyrhizobial strains isolated from the root nodules of soybeans cultivated at locations varying in climate and soil pH in Nepal in response to different pH levels. Each value represents the mean of a triplicate. Vertical bars denote standard deviations. Temperature was set at 30°C
Among the three species used for the growth test, all four B. elkanii strains showed growth stability at varying levels of pH and temperatures. In contrast, the growth of both B. japonicum and one of the B. yuanmingense strains was either retarded or completely ceased at pH 4.5 (data not shown). In addition, one of the B. japonicum strains could not grow at 35°C (data not shown), while the other strains could.
Discussion
Previous studies on the diversity of soybean-nodulating rhizobia in Nepal have shown that the native B. japonicum strains are genetically distinct from strains from other countries (Vinuesa et al. 2008) and have also shown the presence of B. elkanii, B. japonicum and B. yuanmingense (Risal et al. 2010). However, both of these previous studies were conducted using only mountain soils without due consideration of climate and soil factors. In addition, these previous researchers isolated rhizobial strains under laboratory conditions using a plant trap culture with collected soil samples. On the other hand, we isolated rhizobia from the nodules of soybeans grown in temperate mountains as well as subtropical plains, and found genetically and physiologically diverse B. elkanii, B. japonicum, B. yuanmingense and B. liaoningense strains in Nepal. In particular, we found B. liaoningense in subtropical alkaline soil in Nepal for the first time.
Differences in the relative occupancies in soybean nodules of each species of native rhizobia were closely related with the climate and soil pH of the locations where they were isolated. For instance, more than 85% of the isolates in the temperate region were B. japonicum (Fig. 5). Similar dominance of B. japonicum in cooler regions has also been reported in Japan (Saeki et al. 2006; Suzuki et al. 2008). Our results, however, did not match the previous findings of the dominance of B. elkanii in temperate locations in Nepal (Risal et al. 2010). This mismatch might be due to the in vitro cultivation of the host soybean in that study, as mentioned above. Growing soybean plants under laboratory conditions using soil samples diluted with vermiculite (Risal et al. 2010) might undermine the possible effects of soil properties and climate on the rhizobial nodulating properties. In a study in Brazil, Alberton et al. (2006) reported decrease in the diversity index of soybean nodule rhizobia in diluted soils compared to field soils and suggested change in soil pH was the possible reason for the decrease.
In subtropical locations, none of the isolates belonged to B. japonicum. The absence of B. japonicum in the similar climate of India, a southern neighboring country of Nepal, has been reported (Appunu et al. 2008, 2009). The lack of in vitro growth of B. japonicum Cha-4 at 35°C in this study is consistent with the absence of this species in warmer climates (data not shown). Among the species isolated from subtropical locations, B. liaoningense was dominant over B. yuanmingense in a location with slightly alkaline soil (pH 7.4). As the pH was decreased to 6.4 or 5.8 in two locations, B. yuanmingense dominated the population and B. elkanii appeared as a minor species. With a further decrease in soil pH (4.8–5.1) as observed in three locations, the population of B. elkanii far exceeded that of B. yuanmingense. The effect of pH on the occurrence of different species in China has been reported (Li et al. 2011), where B. japonicum and B. elkanii was observed only in neutral to slightly alkaline soils, but with an increase in soil pH, the rhizobial population was dominated by B. yuanmingense, B. liaoningense and Sinorhizobium fredii.
For most rhizobia, the optimum temperature range for growth in culture is 28–31°C, with variations in tolerance and survivability among the strains (Graham 1992; Brockwell et al. 1995). We checked the correlation between the in vitro growth properties at different temperatures and pH levels and the relative nodule occupancies of some dominant and minor strains in various habitats. However, the differences in in vitro growth properties at different temperatures between B. japonicum and B. elkanii strains isolated from temperate locations could not explain the relative nodule occupancy. Also, the lack of growth of a dominant strain (B. japonicum Cha-4) isolated from a strongly acid soil (Charikot, pH 3.6) compared to the successful growth of its minor competent (B. elkanii Cha-2) at pH 4.5 suggested that the dominant strain may not necessarily grow in a liquid medium having a temperature or pH similar to that in the soil from which it was isolated. It is also reported that Rhizobium strains that survived in an acid soil could not grow on a nutrient medium with a pH as low as that of the soil from which the strains were isolated (Asanuma and Ayanaba 1990). In vitro growth properties did not reflect the growth properties in a soil environment in which the physical and nutritional conditions are different. Lower survival rate of rhizobia in YM broth than in soil has also been reported previously (Appunu and Dhar 2006). In a similar study, no correlation was found between the growth ability of soybean rhizobial strains on an agar medium and their nodulation potentials in soil (LaFavre and Eaglesham 1986). Therefore, the nodulation behavior of rhizobia in soil may also have affected the nodule occupancy in our study.
Nevertheless, the relatively higher growth rate of some of the representative strains of B. elkanii at pH 4.5 compared to its counterparts was consistent with its dominance in acidic soils in subtropical locations (Fig. 7). The superior growth might have led to the greater population of this species in the soils, resulting in its higher relative proportions in the nodules. The influence of soil pH on soybean rhizobial diversity has also been reported previously up to the genus level. The dominance of S. fredii in many alkaline soils has been reported (Han et al. 2009; Saeki et al. 2005; Suzuki et al. 2008). The abundance of B. liaoningense in alkaline soils has also been reported previously (Appunu et al. 2008; Han et al. 2009; Li et al. 2011), which is consistent with the higher nodule occupancy of this species in a slightly alkaline soil (pH 7.4) in this study. The relatively stable in vitro growth of B. elkanii compared to other species across the tested ranges of temperature and pH was consistent with its occurrence in a wider range of climates and soil pH. In a study in Okinawa, a subtropical place in Japan, the dominance of B. elkanii was reported in an acidic (pH 4.7–6.1) soil as well as in slightly alkaline (pH 7.5) soils (Suzuki et al. 2008).
In this study, soil pH also affected the diversity at a subspecies level. For example, despite the homologous 16S rRNA gene among the group VII phylotypes of B. japonicum strains, these strains had two contrasting ITS sequences (clusters k and l) chiefly relating to the soil pH of the corresponding locations. The soil pH influenced the nodC and nifH types found within B. yuanmingense strains at the subspecies level (Fig. 4). The effect of soil pH on genetic diversity was further revealed by the disappearance of some phylotypes with the decrease in soil pH in temperate locations and the occurrence of new phylotypes with the change in soil pH in subtropical locations (Table 2). Soil acidity was reported to restrict the survival and persistence of rhizobia and to reduce their nodulation (Brockwell et al. 1991; Graham et al. 1982). Lower bacterial diversity in acidic soils has also been reported (Fierer and Jackson 2006; Anyango et al. 1995).
The incongruence of the nodC and nifH genes of some of the strains with those of 16S rDNA and ITS adds to the previously reported evidence showing that distinct rhizobial strains can share similar symbiotic genes (Laguerre et al. 2001; Laranjo et al. 2001). On the other hand, the close similarity between the phylogenies of the nodC and nifH genes of strains isolated in this study and some reference strains in the database is consistent with the finding that the nod and nif genes are often tightly linked in rhizobia (Haukka et al. 1998), revealing the co-evolution of these two symbiotic genes worldwide, including in Nepal. Horizontal gene transfer of Sym genes is the most plausible hypothesis explaining this phylogenetic incongruence (Martinez-Romero and Caballero-Mellado 1996; Young and Haukka 1996).
Conclusions
This study shows that the highly diverse climate and soils in Nepal might be conducive to the existence of diverse soybean rhizobial strains. The relative populations of strains of four Bradyrhizobium species in the nodules varied along with climate and soil pH. The in vitro growth properties of some of the strains further explained the influence of temperature and pH on the relative proportions of different species in Nepalese soils. Some mismatches, however, suggested a need to examine the occupancies of different strains in response to various temperatures and pH levels for a clearer explanation of the correlation.
References
Alberton O, Kaschuk G, Hungria M (2006) Sampling effects on the assessment of genetic diversity of rhizobia associated with soybean and common bean. Soil Biol Biochem 38:1298–1307
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Ando S, Yokoyama T (1999) Phylogenetic analyses of Bradyrhizobium strains nodulating soybean (Glycine max) in Thailand with reference strains of Bradyrhizobium. Can J Microbiol 45:639–645
Anyango B, Wilson KJ, Beynon JL, Giller KE (1995) Diversity of rhizobia nodulating Phaseolus vulgaris L. in two Kenyan soils with contrasting pHs. Appl Environ Microbiol 61:4016–4021
Appunu C, Dhar B (2006) Symbiotic effectiveness of acid-tolerant Bradyrhizobium strains with soybean in low pH soil. Afr J Biotech 5:842–845
Appunu C, N’Zoue A, Laguerre G (2008) Genetic diversity of native Bradyrhizobia isolated from soybeans (Glycine max L.) in different agricultural-ecological-climatic regions of India. Appl Environ Microbiol 74:5991–5996
Appunu C, Sasirekha N, Prabavathi VR, Nair S (2009) A significant proportion of indigenous rhizobia from India associated with soybean (Glycine max L.) distinctly belong to Bradyrhizobium and Ensifer genera. Biol Fertil Soils 46:57–63
Asanuma S, Ayanaba A (1990) Variation in acid-Al tolerance of Bradyrhizobium japonicum strains from African soils. Soil Sci Plant Nutr 36:309–317
Brockwell J, Pilka A, Holliday RA (1991) Soil pH is a major determinant of the numbers of naturally-occurring Rhizobium meliloti in non-cultivated soils of New South Wales. Aust J Exp Agric 31:211–219
Brockwell J, Bottomley PJ, Thies JE (1995) Manipulation of rhizobia microflora for improving legume productivity and soil fertility: a critical assessment. Plant Soil 174:143–180
Chen WX, Yan GH, Li JL (1988) Numerical taxonomic study of fast-growing soybean rhizobia and a proposal that Rhizobium fredii be assigned to Sinorhizobium gen. nov. Int J Syst Bacteriol 38:392–397
Chen W, Wang E, Wang S, Li Y, Chen X, Li Y (1995) Characteristics of Rhizobium tianshanense sp. nov., a moderately and slowly growing root nodule bacterium isolated from an arid saline environment in Xinjiang, People’s Republic of China. Int J Syst Evol Microbiol 45:153–159
Chen LS, Figueredo A, Pedrosa FO, Hungria M (2000) Genetic characterization of soybean rhizobia in Paraguay. Appl Environ Microbiol 66:5099–5103
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103:626–631
Graham PH (1992) Stress tolerance in Rhizobium and Bradyrhizobium, and nodulation under adverse soil conditions. Can J Microbiol 38:475–484
Graham PH, Viteri SE, Mackie F, Vargas AT, Palacios A (1982) Variation in acid soil tolerance among strains of Rhizobium phaseoli. Field Crops Res 5:121–128
Han LL, Wang TT, Han TX, Liu J, Sui XH, Chen WF, Chen WX (2009) Unique community structure and biogeography of soybean rhizobia in the saline-alkaline soils of Xinjiang, China. Plant Soil 324:291–305
Haukka K, Lindstrom K, Young JPW (1998) Three phylogenetic groups of nodA and nifH genes in Sinorhizobium and Mesorhizobium isolates from leguminous trees growing in Africa and Latin America. Appl Environ Microbiol 64:419–426
Jordan DC (1982) Transfer of Rhizobium japonicum Buchanan 1980 to Bradyrhizobium gen. nov., a genus of slow-growing, root nodule bacteria from leguminous plants. Int J Syst Bacteriol 32:136–139
Knudsen D, Peterson GA, Pratt PF (1982) Lithium, sodium and potassium. In: Page et al (eds) Methods of soil analysis, part 2: chemical and microbiological properties - agronomy monograph no. 9, 2nd edn. American Society of Agronomy, Soil Science Society of America, Wisconsin, pp 225–246
Kyukendall LD, Saxena B, Devine TE, Udell SE (1992) Genetic diversity in Bradyrhizobium japonicum Jordan 1982 and a proposal for Bradyrhizobium elkanii sp. nov. Can J Microbiol 38:501–505
LaFavre AK, Eaglesham ARJ (1986) The effects of high temperature on soybean nodulation and growth with different strains of bradyrhizobia. Can J Microbiol 32:22–27
Laguerre G, Nour SM, Macheret V, Sanjuan J, Drouin P, Amarger N (2001) Classification of rhizobia based on nodC and nifH gene analysis reveals a close phylogenetic relationship among Phaseolus vulgaris symbionts. Microbiology 147:981–993
Laranjo M, Rodrigues R, Alho L, Oliveira S (2001) Rhizobia of chickpea from southern Portugal: symbiotic efficiency and genetic diversity. J Appl Microbiol 90:662–667
Li QQ, Wang ET, Zhang YZ, Zhang YM, Tian CF, Sui XH, Chen WF, Chen WX (2011) Diversity and biogeography of rhizobia isolated from root nodules of Glycine max grown in Hebei Province, China. Microb Ecol 61:917–931
Man CX, Wang H, Chen WF, Sui XH, Wang ET, Chen WX (2008) Diverse rhizobia associated with soybean grown in the subtropical and tropical regions of China. Plant Soil 310:77–87
Martinez-Romero E, Caballero-Mellado J (1996) Rhizobium phylogenies and bacterial genetic diversity. Crit Rev Plant Sci 15:113–140
Olsen SR, Cole CV, Watanabe FS, Dean LA (1954) Estimation of available phosphorus in soils by extraction with sodium bicarbonate. U.S. Department of Agriculture Circ. No. 939
Peng GX, Tan ZY, Wang ET, Reinhold-Hurek B, Chen WF, Chen WX (2002) Identification of isolates from soybean nodules in Xinjiang Region as Sinorhizobium xinjiangense and genetic differentiation of S. xinjiangense from Sinorhizobium fredii. Int J Sys Evol Microbiol 52:457–462
Risal CP, Yokoyama T, Ohkama-Ohtsu N, Djedidi S, Sekimoto H (2010) Genetic diversity of native soybean bradyrhizobia from different topographical regions along the southern slopes of the Himalayan Mountains in Nepal. Syst Appl Microbiol 33:416–425
Saeki Y, Kaneko A, Hara T, Suzuki K, Yamakawa T, Nguyen MT, Nagatomo Y, Akao S (2005) Phylogenetic analysis of soybean-nodulating rhizobia isolated from alkaline soils in Vietnam. Soil Sci Plant Nutr 51:1043–1052
Saeki Y, Naoto A, Tsukamoto S, Yamakawa T, Nagatomo Y, Akao S (2006) Diversity and geographical distribution of indigenous soybean-nodulating bradyrhizobia in Japan. Soil Sci Plant Nutr 52:418–426
Smartt J, Hymowitz T (1985) Domestication and evolution of grain legumes. In: Summerfield RJ, Roberts EH (eds) Grain legume crops. Collins, London, pp 32–72
Someya T, Wang X, Gong C, Koshida K, Hakagawa H, Tanaka C, Ishibashi M, Yokobori K, Inoue K (2003) Rapid and specific detection and enumeration of microorganisms in soil and manure. Soil Microorg 57:115–123
Suzuki K, Oguro H, Yamakawa T, Yamamoto A, Akao S, Saeki Y (2008) Diversity and distribution of indigenous soybean-nodulating rhizobia in the Okinawa islands, Japan. Soil Sci Plant Nutr 54:237–246
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
van Berkum P, Fuhrmann JJ (2000) Evolutionary relationships among the soybean bradyrhizobia reconstructed from 16S rRNA gene and internally transcribed spacer region sequence divergence. Int J Syst Evol Microbiol 50:2165–2172
Vincent JM (1970) A manual for the practical study of root nodule bacteria. IBP Handbook No. 15. Blackwell, Oxford
Vinuesa P, Leon-Barrios M, Silva C, Willema A, Jarabo-Lorenzo A, Perez-Galdona R, Warner D, Martinez-Romero E (2005) Bradyrhizobium canariense sp. nov., an acid-tolerant endosymbiont that nodulates endemic genistoid legumes (Papilionoideae:Genisteae) growing in the Canary Islands, along with B. japonicum bv. Genistearum, Bradyrhizobium genospecies α and Bradyrhizobium genospecies β. Int J Syst Evol Microbiol 55:569–575
Vinuesa P, Rojas-Jimenez K, Contreas-Moreira B, Mahna SK, Prasad BN, Moe H, Selvaraju SB, Theirfelder H, Werner D (2008) Multilocus sequence analysis for assessment of the biogeography and evolutionary genetics of four Bradyrhizobium species that nodulate soybeans in the Asiatic Continent. Appl Environ Microbiol 74:6987–6996
Walkley A, Black IA (1934) An examination of Degtjareff method for determining organic carbon in soils: effect of variations in digestion conditions and of inorganic soil constituents. Soil Sci 63:251–263
Wasike VW, Lesueur D, Wachira FN, Mungai NW, Mumera LM, Sanginga N, Mburu HN, Mugadi D, Wango P, Vanlauwe B (2009) Genetic diversity of indigenous Bradyrhizobium nodulating promiscuous soybean [Glycine max (L) Merr.] varieties in Kenya: impact of phosphorus and lime fertilization in two contrasting sites. Plant Soil 322:151–163
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Xu LM, Ge C, Cui Z, Li J, Fan H (1995) Bradyrhizobium liaoningense sp. nov., isolated from the root nodules of soybeans. Int J Syst Bacteriol 45:706–711
Yang JK, Zhou JC (2008) Diversity, phylogeny and host specificity of soybean and peanut bradyrhizobia. Biol Fertil Soils 44:843–851
Yao ZY, Kan FL, Wang ET, Wei GH, Chen WX (2002) Characterization of rhizobia that nodulate legume species of the genus Lespedeza and description of Bradyrhizobium yuanmingense sp. nov. Int J Syst Evol Microbiol 52:2219–2230
Young JPW, Haukka KE (1996) Diversity and phylogeny of rhizobia. New Phytol 133:87–94
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Katharina Pawlowski.
Rights and permissions
About this article
Cite this article
Adhikari, D., Kaneto, M., Itoh, K. et al. Genetic diversity of soybean-nodulating rhizobia in Nepal in relation to climate and soil properties. Plant Soil 357, 131–145 (2012). https://doi.org/10.1007/s11104-012-1134-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-012-1134-6