Abstract
The WRKY transcription factor family is involved in responding to biotic and abiotic stresses. Its members contain a typical WRKY domain and can regulate plant physiological responses by binding to W-boxes in the promoter regions of downstream target genes. We identified the sweet sorghum SbWRKY50 (Sb09g005700) gene, which encodes a typical class II of the WRKY family protein that localizes to the nucleus and has transcriptional activation activity. The expression of SbWRKY50 in sweet sorghum was reduced by salt stress, and its ectopic expression reduced the salt tolerance of Arabidopsis thaliana plants. Compared with the wild type, the germination rate, root length, biomass and potassium ion content of SbWRKY50 over-expression plants decreased significantly under salt-stress conditions, while the hydrogen peroxide, superoxide anion and sodium ion contents increased. Real-time PCR results showed that the expression levels of AtSOS1, AtHKT1 and genes related to osmotic and oxidative stresses in over-expression strains decreased under salt-stress conditions. Luciferase complementation imaging and yeast one-hybrid assays confirmed that SbWRKY50 could directly bind to the upstream promoter of the SOS1 gene in A. thaliana. However, in sweet sorghum, SbWRKY50 could directly bind to the upstream promoters of SOS1 and HKT1. These results suggest that the new WRKY transcription factor SbWRKY50 participates in plant salt response by controlling ion homeostasis. However, the regulatory mechanisms are different in sweet sorghum and Arabidopsis, which may explain their different salt tolerance levels. The data provide information that can be applied to genetically modifying salt tolerance in different crop varieties.
Key message
(1) Sweet sorghum SbWRKY50 is negatively involved in salt response.
(2) Over-expression of SbWRKY50 in A. thaliana affects plant growth, ROS and the ion contents.
(3) SbWRKY50 could directly bind to the upstream promoter of the SOS1 gene in A. thaliana and the promoter of SOS1 and HKT1 in sweet sorghum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salt stress is an important factor that seriously affects crop growth and yield (Asano et al. 2012; Liu et al. 2017a, b). Salt stress can inhibit the growth and development of plants, disrupt the permeability of cell membranes and biologically functional molecules in cells, and seriously affect the physiological and metabolic processes of plants (Syvertsen and Garcia-Sanchez 2014; Reginato et al. 2014; Wu et al. 2014; Han et al. 2014; Liu et al. 2015; Sui et al. 2017, 2018). Soil salinization is a worldwide problem, especially in arid and semi-arid areas. China has severe soil salinization, affecting tens of millions of hectares (Li et al. 2014a, b). Moreover, owing to the improper development and utilization of irrigation measures, a large area of secondary salinized soil has been produced. Most crops do not grow normally on saline–alkali soil (Li et al. 2012). Therefore, studying the molecular mechanisms of salt resistance is necessary for increasing salt tolerance and utilizing salinized soil.
To maintain the normal physiological metabolism of cells under salt-stress conditions, plants undergo a series of physiological and biochemical reactions, including ion transport and absorption (Shao et al. 2014), the synthesis of osmotic regulators (Li et al. 2014; Wei et al. 2017; You et al. 2019) and the enhancement of antioxidant enzyme levels (Guo et al. 2017). Additionally, plants induce the expression levels of genes that have different functions in response to salt stress. Regulating the expression levels of functional genes through transcription factors is a key mechanism of plant responses to stress. At present, there are many transcription factor families responding to stress, such as bZIP, WRKY, AP2/EREBP and MYB and so on.
The WRKY transcription factor is a supergene family named for a highly conserved WRKY domain consisting of 60 amino acids that are found in the family members (Rinerson et al. 2015). However, there are exceptions. In some WRKY transcription factors, WRKY residues in the WRKY domain are replaced by WRRY, WSKY, WKRY, WVKY or WKKY motifs. The N-terminal of the domain has a conservative WRKYGQK core sequence, and the C-terminal has a zinc-finger structure, C2–H2 or C2–HC. WRKY domain proteins bind specifically to W-boxes. The conserved sequence of the W-box is TTGACC or TTGACT, and the four middle bases TGAC are the core W-box sequence. The selection of a W-box by a WRKY transcription factor protein depends on the two flanking bases. At present, WRKY transcription factors are divided into three protein subfamilies, classes I, II and III.
Since the first WRKY gene was cloned in sweet potatoes (Ishiguro and Nakamura 1994), it has been found that WRKY is a ubiquitous class of transcription factors in plants especially in some important economic crops such as soybean (Zhou 2008; Wang et al. 2015), potato (Liu et al. 2017a, b), cotton (Si et al. 2017), tomato (Wang et al. 2017), maize (Zhang et al. 2017), rice (Lee et al. 2018) and wheat (Gao et al. 2018). The WRKY protein is involved in abiotic stress responses (Ali et al. 2018; López-Galiano et al. 2018), plant disease resistance (Lan et al. 2013), plant development and material metabolism (Yang et al. 2016; Sun et al. 2003), seed dormancy (Ding et al. 2014), plant senescence (Potschin et al. 2014), hormone signaling (Birkenbihl et al. 2015; Tian et al. 2017) and other processes (Chen et al. 2012). WRKY transcription factor is a new participant in signal transduction network. The interaction between downstream target genes and upstream regulators constitutes a complex WRKY transcription factor regulatory network, which is a new research field. At present, the regulatory network of WRKY transcription factors has been studied in some plant species (Zou et al. 2004; Kim et al. 2008; Jiang et al. 2016), but the regulatory network in sweet sorghum is the first introduction.
Sweet sorghum [Sorghum bicolor (L.) Moench], known as the "second generation sugarcane", is a kind of crop with high sugar content and high yield (Sui et al. 2015; Schnippenkoetter et al. 2017). It is an important food, feed and energy crop in the world. Sweet sorghum not only has a very high biomass but also has salt and alkali tolerant. Many unique genes were found to be related to salt tolerance of sweet sorghum (Zheng et al. 2011; Yang et al. 2018). Therefore, research on these unique genes and the related characteristics of sweet sorghum can be applied to the genetic modification of other crops, allowing us to obtain stress-tolerant crops, and manage the growth and development of plants in saline environments.
In this study, we isolated SbWRKY50 from our former transcriptomic data, because the gene was down regulated in M81-E and was not induced in Roma by salt stress (Yang et al. 2017, 2018). We found that SbWRKY50 down-regulated under salt treatment. After over-expressing SbWRKY50 in Arabidopsis thaliana, the related physiological indexes were determined, and yeast one-hybrid and luciferase complementation imaging assays were performed. SbWRKY50 can negatively regulate salt responses by altering ion homeostasis in both sweet sorghum and A. thaliana. But the regulating mechanism is different in sweet sorghum and A. thaliana.
Results
SbWRKY50 is a typical WRKY transcription factor, containing the WRKYGQ(K)K motif
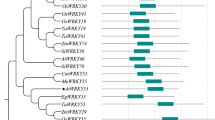
The coding sequence (CDS) of SbWRKY50 contains 621 bases and encodes 206 amino acids. There is a typical WRKY domain between amino acids 109 and 168 in SbWRKY50, and a zinc-finger structure, C2H2, at the C-terminus (Fig. S1A, B). Using the BLAST function of NCBI, 14 sequences with high homology to SbWRKY50 were identified (Fig. S1C). WRKYGQ(K)K and C2–H2 are highly conserved in the WRKY family. The phylogenetic tree showed that the SbWRKY50 protein had high homology with Panicum hallii WRKY51 (Fig. S1D).
SbWRKY50 is a negative regulatory transcription factor expressed in the nucleus
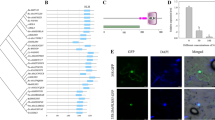
To test if SbWRKY50 is located in the nucleus as a transcription factor, we performed the transient transformation in tobacco with SbWRKY50: GFP. GFP was expressed in cell membrane and nucleus, but GFP fused with SbWRKY50 was located in nucleus (Fig. 1A). This indicates that SbWRKY50 is a nuclear localization protein.
Subcellular localization, expression pattern and transcriptional activity analyses of SbWRKY50. A Localization of SbWRKY50 in tobacco cells (35S: GFP as control group, 35S:WRKY50-GFP as experimental group). B The expression levels of SbWRKY50 in roots and leaves under 0-, 50-, 100-, 150- and 200 mM salt conditions. C Analysis of transcriptional activity of SbWRKY50 using a yeast system
To elucidate the function of SbWRKY50 under salt condition, we performed salt treatment experiments in sweet sorghum. As shown in Fig. 1B, during the 50 mM NaCl treatment, the expression of SbWRKY50 increased in shoot, but decreased sharply during the 100 mM NaCl treatment, and reached its lowest level during the 200 mM NaCl treatment. In roots, its expression level significantly decreased with the increase of salt concentration, reaching its lowest level during the 200 mM NaCl treatment (Fig. 1B). Therefore, these results support that SbWRKY50 can be induced by salt and might be a negative regulatory transcription factor.
SbWRKY50 is a transcription factor with transcriptional activity
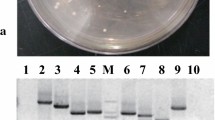
We analyzed the transcriptional activity of SbWRKY50 using a yeast system to investigate the potential role of SbWRKY50. On tryptophan deficient medium (SD/−Trp), both the experimental and control groups grew normally. On tryptophan, adenine and histidine (SD/−Trp − Ade − His) and the same medium supplemented with 5-bromo-4-chloro-3-indole-a-d-galactoside (X-a-gal; SD/−Trp − Ade − His + X-a-gal), the control group did not grow normally, while the experimental group grew normally and colonies turned blue (Fig. 1C). These results indicate that SbWRKY50 has a transactivational capacity.
Over-expression of SbWRKY50 enhances plant sensitivity to salt stress
To further demonstrate that SbWRKY50 acts as a negatively regulated transcription factor in salt tolerance, SbWRKY50 was derived by CaMV35s promoter and was transformed into A. thaliana. We generated two transgenic lines W50-12 and W50-13. The expression level of SbWRKY50 was 7.84 (W50-12) and 9.67 (W50-13) times higher than wild type (WT) Arabidopsis (Fig. 2A), which proved that SbWRKY50 had been successfully transformed into Arabidopsis. Furthermore, we measured the biomass of over-expression and WT plants in the treatment of 0 and 100 mM NaCl. The dry weight and fresh weight of each Arabidopsis line decreased significantly, but the decline degree of over-expressed lines was significantly higher than that of WT (Fig. 2B, C). This suggests that over-expression of SbWRKY50 decreased plant biomass.
The determination of physiological parameters in Arabidopsis wild-type (WT) and SbWRKY50 over-expression lines. A Expression of SbWRKY50 in over-expression plants. B and C Fresh weights (B) and dry weights (C) of WT and over-expression plants under different salt conditions. D Under salt treatment, the phenotypes of WT and over-expressed A. thaliana (W50-12, W50-13) at seedling stage, with six seedlings in each group. E and F Germination rates (E) and root lengths (F) of WT and over-expression plants under different salt conditions. Each column represents the means ± SDs of five measurements. Means identified by different letters are significantly different at P < 0.05
We also measured seed germination and root length. There were no differences in the seed germination rates and root lengths of the lines not subjected to NaCl treatment. As the salt concentration increased, the seed germination rates and root length of each strain decreased significantly (Fig. 2D). When treated with 50 and 100 mM NaCl, the germination rates and root lengths of the over-expression lines were significantly lower than those of the WT. When germinated on 150 mM NaCl, reduction in seed germination and root length was observed both in WT and over-expression plants, however reduction was significantly higher in overexpression lines as compared to WT plants (Fig. 2E, F). This indicates that the over-expression of SbWRKY50 affected seed germination and root length.
Under salt-stress conditions, sodium ions (Na+) are the main ions that cause plant salt damage, while potassium ions (K+) are an essential element and important osmotic adjustment substance for plant growth. The level of Na+ has a significant inhibitory effect on the absorption of K+. To detect ion metabolism in plants, we determined the contents of Na+ and K+ in over-expression and WT lines. Under 0 mM NaCl conditions, there were no significant differences in the Na+ and K+ contents in the leaves and roots of transgenic Arabidopsis compared with WT plants. However, under 100 mM NaCl conditions, compared with WT plants, the Na+ content in SbWRKY50 over-expression plants increased significantly, while the K+ content decreased significantly. As a result, Na+/K+ increased significantly in the over-expression plants (Fig. 3A–C). This suggests that the over-expression of SbWRKY50 reduces the ability of Arabidopsis to resist salinity by increasing the Na+ content and decreasing the K+ content.
Changes in ions and reactive oxygen species. A–C The Na+ and K+ contents and Na+/K+ ratio in roots of WT and over-expression plants during different salt treatments. Each column represents the means ± SDs of five measurements. Means identified by different letters are significantly different at P < 0.05. D DAB staining. E NBT staining. Each staining experiment was repeated five times
Under salt-stress conditions, plants can produce hydrogen peroxide (H2O2), superoxide anions (O2−.) and other substances owing to oxidation. DAB solution can react with plant endogenous H2O2 to form reddish brown spots, which reflect the H2O2 contents in plants. Similarly, nitro blue tetrazolium (NBT) solution can be reduced to a water-insoluble blue substance under the action of O2−., which reflects the O2−. contents in plants. Under 0 mM NaCl conditions, the degrees of DAB and NBT staining of each strain were not significantly different. However, under 100 mM NaCl conditions, the degrees of both DAB and NBT staining in over-expression strains were deeper than in WT (Fig. 3D, E), which indicated that under saline conditions, the over-expression of SbWRKY50 made Arabidopsis plants more sensitive to oxidative stress than WT plants.
SbWRKY50 regulates the expression of stress-related genes
To identify the target genes of SbWRKY50, we assessed the transcriptional changes of some genes in response to osmotic (SOD and APX), oxidative (RD29B and P5CS1) and ion stresses (CLC-C, SOS1 and HKT1) using qRT-PCR. Under salt-stress conditions, the expression levels of these genes were induced, but the levels of induction were less in the SbWRKY50 over-expression strains than in WT strains (Fig. 4A). Thus, SbWRKY50 may play an important role in regulating the expression levels of stress-responsive genes. WRKY transcription factors regulate downstream genes by binding with W-boxes. To further screen the target genes of SbWRKY50, we analyzed the promoters of these genes (Table S1), and only the promoters of A. thaliana salt overly sensitive 1 (AtSOS1, Na+/H+ reverse transporter), sweet sorghum SbSOS1 and high-affinity potassium transporter (SbHKT1) contained W-boxes. Therefore, we speculate that the SOS1 and HKT1 genes might be targets of SbWKY50.
Expression levels of stress-related genes and interactions between SbWRKY50 and its downstream target genes. A The expression levels of genes responding to osmotic, stress and ionic stresses in SbWRKY50 over-expression strains compared with wild-type strains under salt-stress conditions. Each column represents the means ± SDs of five measurements. Means identified by different letters are significantly different at P < 0.05. B and C The interactions of SbWRKY50 with proAtSOS1 and proAtHKT1 were validated using yeast one-hybrid and luciferase complementation imaging assays
SbWRKY50 directly binds to the AtSOS1, SbSOS1 and SbHKT1 promoters
To determine whether SbWRKY50 binds with the promoters of AtSOS1, SbSOS1 and SbHKT1, we performed yeast one-hybrid and luciferase complementation imaging assays. The yeast one-hybrid assays showed that only when proAtSOS1-pLacZi2u, proSbSOS1-pLacZi2u and proSbHKT1-pLacZi2u were independently co-transformed with GAD-SbWRKY50, LacZ expression was activated and colonies appeared blue on SD/−Trp − Ura + X-a-gal (Figs. 4B, 5A). Similarly, when proAtSOS1-pGreenII-0800, proSbSOS1-pGreenII-0800 and proSbHKT1-pGreenII-0800 were independently co-transformed with SbWRKY50-pMWB110, strong in vivo fluorescence signals were observed using the imaging system (Figs. 4C, 5B). No fluorescence signals were observed in the co-transformation of SbWRKY50-pMWB110 and proAtHKT1-pGreenII-0800. This indicates that SbWRKY50 can regulate plant salt tolerance by interacting directly with AtSOS1. However, in sweet sorghum, SbWRKY50 can regulate salt tolerance by directly binding the promoters of SbSOS1 and SbHKT1.
Discussion
SbWRKY50 protein belongs to class II of the WRKY family with the N-terminus containing a conserved WRKY domain, and the C-terminus containing a C2–H2 zinc-finger structure (Fig. S1A–C). The transient transformation of tobacco indicated that WRKY50 was expressed in the nucleus, and the transcriptional activity of WRKY50 was verified using a yeast system (Fig. 1A, C). Under salt-stress conditions, the transcriptional level of WRKY50 in sweet sorghum decreased with the increase of salt concentration (Fig. 1B). These results suggest that the SbWRKY50 gene is constitutively expressed, and its expression is inhibited by salt stress, which indicate that SbWRKY50 might be a negatively regulation factor.
Seed germination is the first stage of plant growth, and roots are the main organ involved in plants perceiving environmental changes (Almansouri et al. 2001; Guo et al. 2018; Li et al. 2018). When plants are subjected to adverse conditions, germination rate and root length are the first characteristics to be affected. Jafarzadeh and Aliasgharzad (2007) found that with an increasing salt content, the yield of sugar beet decreased. The salt level had a significant effect on the seed germination rates and root lengths of four sugar beet varieties (pp22, ic2, pP36 and 7233). Cai et al. (2014) showed that the over-expression of WRKY58 in rice results in delayed germination and inhibited post-germination development. Xu et al. (2018a, b) found that the over-expression of Glycine max WRKY49 significantly increased the germination rates, survival rates, root lengths, rosette diameters, relative electrolyte leakage levels and proline contents in composite seedlings and transgenic Arabidopsis than in WT. In this study, the germination rate, root length and the biomass were significantly inhibited in over-expression SbWRKY50 plants than in WT.
Under salt stress, Na+ enters cells in large quantities, resulting in the reduction in K+ uptake by cells and the destruction of the ion balance in plant cells (Wei et al. 2015). High salt concentrations also destroy plant defense systems, and the accumulation of reactive oxygen species (H2O2 and O2−.) leads to membrane lipid peroxidation, which results in a series of physiological metabolic disorders (Li et al. 2000; Song et al. 2019). In this study, the K+ content of SbWRKY50 over-expression plants were significantly decreased under high-salt conditions compared with WT, while the Na+ content and reactive oxygen species accumulation increased, affecting the ion homeostasis. These results indicate that the over-expression of SbWRKY50 in A. thaliana can inhibit plant growth, and dry matter and reactive oxygen species accumulation, leading to oxidative stress, and reduce the abilities of plants to maintain their ion balance. Thus, plant salt tolerance was reduced.
Salt stress also can induce some stress related genes expression. In this study, we determined the expression levels of some stress-related genes, RD29B, P5CS1, SOD, APX, CLC-C, SOS1 and HKT1. Previous research shows that the osmotic stress-related gene RD29B is induced by salt and drought and activates the Arabidopsis MAKK or MAPK protein (Hu et al. 2006). Deletions of the P5CS1 gene in A. thaliana result in accumulations of reactive oxygen species (Szekely et al. 2008). The over-expression of P5CS1 results in a higher biomass and a higher root growth rate during early development (Ibragimova et al. 2015). Oxidative stress-related genes SOD, APX are induced by salt and play key roles in preventing excessive ROS accumulation and protecting cells against ROS caused by salt stress (Ma 2002; Wang et al. 2004; Pedro et al. 2013). OsCLC1 plays an important role in regulating Cl− homeostasis in rice under salt stress (Diedhiou and Golldack 2006). SOS1 can regulate the excretion of Na+ in root cells (Ding and Zhu 1997; Shi et al. 2002, 2003). HKT1 plays an important role in ion redistribution. Our results also show that the over-expression of SbWRKY50 can reduce the expression levels of RD29B, P5CS1, SOD, APX, CLC-C, SOS1 and HKT1 under salt stress conditions.
The WRKY transcription factor usually combines with the W-box in the promoters of target genes (Bakshi and Oelmüller 2014). In this study, we identified W-boxes only in the promoters of Arabidopsis SOS1 and sweet sorghum SOS1 and HKT1, while there were no binding sites in the promoters of RD29B, P5CS1, SOD, APX and CLC-C. Furthermore, luciferase complementation and yeast one-hybrid assays showed that SbWRKY50 directly binds to the promoters of AtSOS1, SbSOS1 and SbHKT1. SOS1 is mainly responsible for the efflux of Na+ from roots (Wu et al. 1996). The over-expression of SOS1 results in an increase in the salt tolerance of A. thaliana (Yang et al. 2009) and decreases the Na+ accumulation and increases K+ in tobacco (Yue et al. 2012). Soybean SOS1 improves salt tolerance by limiting the accumulation of Na+ and increasing the activity of antioxidant enzymes (Zhao et al. 2017). Similarly, HKT1 is a high affinity K+ transporter responsible for transporting aboveground Na+ back to the root. The over-expression of the soybean HKT1 gene in tobacco can increase the K+ content in transgenic plants and decrease the Na+ content under salt-stress conditions. Thus, it plays an important role in regulating the Na+/K+ ratio in roots (Chen et al. 2014). After silencing the HKT1 in tomato, the Na+/K+ ratio in the leaves changed, and the plants were susceptible to salt-related damage (Jaime-Pérez et al. 2017). In this study, the expression of SOS1 and HKT1 decreased more in transgenic plants. Corresponding with this result, the content of Na+ increased and the K+ content deceased in over-expression SbWRKY50 plants relative to WT under salt stress. This suggests that over-expression of SbWRKY50 can change ion homeostasis by directly regulating the expression of SOS1 and HKT1.
In conclusion, SbWRKY50 can regulate plant salt tolerance mainly by adjusting the Na+ and K+ content in roots. In A. thaliana, SbWRKY50 can bind to the promoter of AtSOS1, while in sweet sorghum, SbWRKY50 can directly bind to the promoters of SbSOS1 and SbHKT1, which might explain why sweet sorghum is more tolerant to salt stress than A. thaliana. In addition, we speculate that SbWRKY50 regulates the expression of osmotic stress- and oxidative stress-related genes by some indirectly pathways.
Materials and methods
Plant materials
Seeds of sweet sorghum M-81E were rinsed with running water for 2 h and placed in a wet dish for germination. After germination, the seedlings were transferred to a hydroponic pot and irrigated with 1/2 Hoagland’s nutrient solution. When sorghum had grown to the three-leaf stage, they were treated with 0, 50, 100, 150 mM NaCl in Hoagland’s nutrient solution for 48 h, and then the root tissues was removed and stored at − 80℃.
Full-length CDS cloning and bioinformatics analysis of the SbWRKY50 gene
The primers SbW50 (Table S2) were designed using the CDS of SbWRKY50 found on NCBI. The total RNA of the sweet sorghum root was extracted and reverse transcribed. The obtained cDNA was subjected to PCR amplification to obtain the desired target fragment. The ligation reaction was performed using the pEASY-Blunt Simple Cloning Kit Vector, and then positive clones were screened for sequencing. Bioinformatics analyses of SbWRKY50 were performed using NCBI databases, DNAMAN and MEGA5.0.
Expression pattern of the SbWRKY50 gene
The primers WRKY50 (Table S2) were designed according to the sequence of SbWRKY50. The shoot and root RNA of sweet sorghum were used independently as templates for real-time PCR.
The subcellular localization of SbWRKY50
The target fragment and 1300-GFP vector were double-digested with KpnI and BamHI and ligated. The ligation was verified to be correct, and the resulting plasmid was transformed into Agrobacterium tumefaciens EHA105. Then, the Agrobacterium transformation with WRKY50-1300-GFP was used to infect tobacco. An Agrobacterium transformation with only 1300-GFP was used as the control. After 2 days under normal culture conditions, the transformants were observed and photographed using two-photon confocal microscopy.
Transcriptional activation analysis in yeast cells
The target fragment and the pGBKT7 vector were digested with EcoRI and BamHI, then ligated and transformed into Escherichia coli. The AH109 yeast strain was transformed after correct digestion. The yeast strain transformed into pGBKT7 was used as the control. The transformed yeast solution was diluted by 1×, 10 × and 100×. Then, 10 µL of solution was dropped on SD/−Trp, SD/−Trp − Ade − His and SD/−Trp − Ade − His + X-a-gal, respectively. The samples were cultured at 28℃ for 48–96 h for observation and imaging.
Expression of SbWRKY50 in transgenic Arabidopsis
The target fragment and pROKII vector were double digestion with KpnI and BamHI, ligated, and then transformed into E. coli. After confirming the correct construction, A. tumefaciens strain GV3101 was transformed. Homozygous Arabidopsis strains over-expressing SbWRKY50 were obtained by infecting the inflorescence of Arabidopsis with Agrobacterium GV3101 transformed with WRKY50-pROKII. RNA was extracted from the roots of homozygous plants for reverse transcription, and the cDNA was used as the template for quantitative fluorescence PCR. The actin2 gene in Arabidopsis was used as an internal reference. The primer sequences are shown in Table S2.
Determination of the seed germination rates and main root length
The seeds of WT and over-expression plants were washed three times with 75% alcohol for 3 to 4 min each time. They were sown on medium containing 1/2 MS, 1/2 MS + 50 mM NaCl, 1/2 MS + 100 mM NaCl and 1/2 MS + 150 mM NaCl. After vernalization at a low temperature for 3 days, seeds were cultured in a tissue culture room. After 24 h, the germination rate was calculated using the following formula:
Then, the lengths of the main roots were photographed and measured.
Biomass determination
The seedlings of WT, W50-12 and W50-13 were independently treated with 0 and 100 mM NaCl in Hoagland’s nutrient solution for 7 days. Then, the seedlings were removed from the soil, and washed with clean water and deionized water. The weights of the seedlings were recorded as fresh weights. The seedlings were then placed in an oven at 70℃ until their weight did not change. These weights were recorded as the dry weights. There were five replicates per treatment.
Determination of sodium ions (Na+) and potassium ions (K+) Contents
The seedlings of WT and over-expression plants were treated with 0 and 100 mM NaCl in Hoagland’s nutrient solution. After 7 days, 0.1 g leaves and 0.1 g roots were sampled and placed into test tubes. Then, 5 mL of ddH2O was added, and tubes were placed in boiling water for 3 h. Afterward, samples were filtered and brought to 10 mL. Then, the Na+ and K+ contents in leaves and roots were determined using a flame spectrophotometer. Each strain had six replicates per treatment.
DAB and NBT staining
The rosette leaves of WT and SbWRKY50 over-expression plants of similar size were cut into EP tubes, and 2 mL DAB or NBT staining solution was added to the tubes to soak the leaves. Samples were placed in the dark for more than 12 h. Diaminobenzidine (DAB) or NBT dyes were replaced with bleaching solution (3:1:1 ethanol:acetic acid:glycerol). Bathed in boiling water for 10–15 min, then the bleaching solution was replaced and samples were incubated for 30 min. The color changes of leaves were observed and images taken. The experimental group receiving DAB staining was treated with 2 mL 10 mM Na2HPO4 as the control group, and the experimental group receiving NBT staining was treated with 2 mL 10 mM phosphate buffer as the control group. There were three repetitions in each group.
Quantitative real-time PCR of stress-related genes
To assess the effects of SbWRKY50 expression, we investigated the expression profiles of several stress-response genes in WT and over-expression lines after 48 h of salt stress. Real-time PCR analyses were performed using cDNAs of WT and over-expression plants as templates. We studied the expression levels of the stress-related genes AtRD29B, AtP5CS1, AtSOD, AtAPX, AtCLC-C, AtSOS1 and AtHKT1. Primers are shown in Table S2.
Yeast one-hybrid assays
In the pLacZi2u vector, EcoRI and XhoI sites were used for the insertion of promoter sequences. Full-length SbWRKY50 was ligated into the EcoRI and XhoI sites of the GAD vector. The co-transformation of the two plasmids into yeast strain EGY48 were cultured on a SD/−Trp − Ura. The positive clones were cultured on SD/−Trp − Ura + X-a-gal. The results were observed and photographed after 3 to 5 days.
Luciferase complementation imaging assays
The target promoter and pGreen II-0800 vector were double-digested with KpnI and XhoI, and transformed into E. coli. The correct construct was transformed into A. tumefaciens GV3103 (psoup). The Agrobacterium solution containing pGreenII-0800 vector with a promoter and pMWB110 vector harboring SbWRKY50 were mixed one to one and injected into the back of tobacco leaves. Agrobacterium containing pGreenII-0800 with a promoter and pMWB110 were mixed as the control. After 2 days of culturing under normal conditions, transformants were observed and live imaged by Lumina II (Xenogen, America).
References
Ali MA, Azeem F, Nawaz MA, Acet T, Abbas A, Imran QM, Shah KH (2018) Transcription factors WRKY11 and WRKY17 are involved in abiotic stress responses in, Arabidopsis J Plant Physiol 226:12–21
Almansouri M, Kinet JM, Lutts S (2001) Effect of salt and osmotic stresses on germination in durum wheat (Triticum durum Desf.). Plant Soil 231:243–254
Asano T, Hayashi N, Kobayashi M, Aoki N, Miyao A, Mitsuhara I, Ichikawa H, Komatsu S, Hirochika H, Kikuchi S, Ohsugi R (2012) A rice calcium-dependent protein kinase OsCPK12 oppositely modulates salt-stress tolerance and blast disease resistance. Plant J 69:26–36
Bakshi M, Oelmüller R (2014) WRKY transcription factors: jack of many trades in plants. Plant Signal Behav 9:e27700
Birkenbihl RP, Kracher B, Somssich IE, Ziegler J, Liu S (2015) Negative regulation of ABA signaling by WRKY33 is critical for Arabidopsis immunity towards Botrytis cinerea 2100. e-Life 4:e07295
Cai R, Zhao Y, Wang Y, Lin Y, Cheng B (2014) Overexpression of a maize WRKY58 gene enhances drought and salt tolerance in transgenic rice. Plant Cell Tissue Org 119:565–577
Chen L, Song Y, Li S, Zhang L, Zou C, Yu D (2012) The role of WRKY transcription factors in plant abiotic stresses. Biochim Biophys Acta 1819:120–128
Chen HT, Chen X, Gu HP, Wu BY, Zhang HM, Yuan XX, Cui XY (2014) GmHKT1;4, a novel soybean gene regulating Na+/K+ ratio in roots enhances salt tolerance in transgenic plants. Plant Growth Regul 73:299–308
Diedhiou C, Golldack D (2006) Salt-dependent regulation of chloride channel transcripts in rice. Plant Sci 170:793–800
Ding L, Zhu JK (1997) Reduced Na+ uptake in the NaCl-hypersensitive SOS1 mutant of Arabidopsis thaliana Plant Physiol 113:795–799
Ding ZJ, Yan JY, Li GX, Wu ZC, Zhang SQ, Zheng SJ (2014) WRKY41 controls Arabidopsis seed dormancy via direct regulation of ABI3 transcript levels not downstream of ABA. Plant J 79:810–823
Gao HM, Wang YF, Xu P, Zhang ZB (2018) Overexpression of a wrky transcription factor TaWRKY2 enhances drought stress tolerance in transgenic wheat. Front Plant Sci 9:997
Guo YY, Tian SS, Liu SS, Wang WQ, Sui N (2017) Energy dissipation and antioxidant enzyme system protect photosystem II of sweet sorghum under drought stress. Photosynthetica 56:861–872
Guo YY, Song YS, Zheng HX, Zhang Y, Guo JR, Sui N (2018) NADP-malate dehydrogenase of sweet sorghum improves salt tolerance of Arabidopsis thaliana J Agric Food Chem 66:5992–6002
Han GL, Wang MJ, Yuan F, Sui N, Song J, Wang BS (2014) The CCCH zinc finger protein gene atzfp1 improves salt resistance in Arabidopsis thaliana Plant Mol Biol 86:237–253
Han Y, Fan T, Zhu X, Wu X, Cao S (2019) WRKY12 represses GSH1 expression to negatively regulate cadmium tolerance in Arabidopsis Plant Mol Biol 99:149–159
Hichri I, Muhovski Y, Žižková E, Dobrev PI, Gharbi E, Jose M, Franco-Zorrilla JM, Lopez-Vidriero I, Solano R, Clippe A, Errachid A, Motyka V, Lutts S (2017) The Solanum lycopersicum wrky3 transcription factor slwrky3 is involved in salt stress tolerance in tomato. Front Plant Sci 8:1343
Hu ZM, Yang X, Fromm ME (2006) Activation of the NaCl and drought-induced RD29A and RD29B promoters by constitutively active Arabidopsis MAPKK or MAPK proteins. Plant Cell Environ 29:1761–1770
Ibragimova SM, Trifonova EA, Filipenko EA, Shymny VK (2015) Evaluation of salt tolerance of transgenic tobacco plants bearing the P5CS1 gene of Arabidopsis thaliana Russ J Genet 51:1181–1188
Ishiguro S, Nakamura K (1994) Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 5' upstream regions of genes coding for sporamin and betaamylase from sweet potato. Mol Gen Genet 244:563–571
Jafarzadeh AA, Aliasgharzad N (2007) Salinity and salt composition effects on seed germination and root length of four sugar beet cultivars. Biologia 62:562–564
Jaime-Pérez N, Pineda B, García-Sogo B, Atares A, Athman A, Byrt CS, Belver A (2017) The sodium transporter encoded by the HKT1; 2 gene modulates sodium/potassium homeostasis in tomato shoots under salinity. Plant Cell Environ 40:658–671
Jiang YZ, Guo L, Liu R, Jiao B, Zhao X, Ling ZY, Luo K (2016) Overexpression of poplar ptrwrky89 in transgenic Arabidopsis leads to a reduction of disease resistance by regulating defense-related genes in salicylate- and jasmonate-dependent signaling. PLoS ONE 11:e0149137
Kim KC, Lai Z, Fan B, Chen Z (2008) Arabidopsis WRKY38 and WRKY62 transcription factors interact with histone deacetylase 19 in basal defense. Plant Cell Online 20:2357–2371
Lan A, Huang J, Zhao W, Peng Y, Chen Z, Kang D (2013) A salicylic acid-induced rice (Oryza sativa L.) transcription factor OsWRKY77 is involved in disease resistance of Arabidopsis thaliana Plant Biol 15:452–461
Lee H, Cha J, Choi C, Choi N, Ji HS, Park SR, Lee S, Hwang DJ (2018) Rice WRKY11 plays a role in pathogen defense and drought tolerance. Rice 11:5
Li J, Yan XF, Zu YG (2000) Generation of activated oxygen and change of cell defense enzyme activity in leaves of Korean pine seedling under low temperature. J Integr Plant Biol 42:148–152
Li JG, Pu LJ, Zhu M, Zhang R (2012) The present situation and hot issues in the salt-affected soil research. Acta Geogr Sin 67:1233–1245
Li JG, Pu LJ, Han MF, Zhu M, Zhang RS, Xiang YZ (2014a) Soil salinization research in China: advances and prospects. J Geogr Sci 24:943–960
Li MF, Guo SJ, Xu Y, Meng QW, Li G, Yang XH (2014b) Glycine betaine-mediated potentiation of HSP gene expression involved calcium signaling pathways in tobacco exposed to NaCl stress. Physiol Plant 150:63–75
Li YB, Li T, Han YY, Fan SX (2017) Cloning and function analysis of heat-shock-protein LsHsp70-2711 gene under high temperature stress in leaf lettuce (Lactuca sativa L.). Sci Agric Sin 8:12
Li J, Wang YQ, Yu B, Song QP, Liu Y, Chen THH, Li G, Yang XH (2018) Ectopic expression of StCBF1 and ScCBF1 have different functions in response to freezing and drought stresses in Arabidopsis Plant Sci 270:221–233
Liu W, Li RJ, Han TT, Cai W, Fu ZW, Lu YT (2015) Salt stress reduces root meristem size by nitric oxide-mediated modulation of auxin accumulation and signaling in Arabidopsis Plant Physiol 168:343–356
Liu QN, Liu Y, Xin ZZ, Zhang DZ, Ge BM, Yang RP, Wang ZF, Yang L, Tang BP, Zhou CL (2017a) Genome-wide identification and characterization of the WRKY gene family in potato (Solanum tuberosum). Biochem Syst Ecol 71:212–218
Liu S, Wang WQ, Li M, Wan SB, Sui N (2017b) Antioxidants and unsaturated fatty acids are involved in salt tolerance in peanut. Acta Physiol Plant 39:207
López-Galiano MJ, González-Hernández AI, Crespo-Salvador O, Rausell C, Real MD, Escamilla M, García-Robles I (2018) Epigenetic regulation of the expression of WRKY75 transcription factor in response to biotic and abiotic stresses in Solanaceae plants. Plant Cell Rep 37:167–176
Ma CL (2002) cDNA cloning and gene expression of APX in Suaeda salsa in response to salt stress. Acta Pharmacol Sin 28:261–266
Pedro DV, Mohamed F, Gregorio BE, Lydia F, Cesar P, José AH, Lorenzo B (2013) Ectopic expression of cytosolic superoxide dismutase and ascorbate peroxidase leads to salt stress tolerance in transgenic plums. Plant Biotechnol J 11:976–985
Potschin M, Schlienger S, Bieker S, Zentgraf U (2014) Senescence networking: WRKY18 is an upstream regulator, a downstream target gene, and a protein interaction partner of WRKY53. J Plant Growth Regul 33:106–118
Reginato MA, Castagna A, Furlan A, Castro S, Ranieri A, Luna V (2014) Physiological responses of a halophytic shrub to salt stress by Na2SO4 and NaCl: oxidative damage and the role of polyphenols in antioxidant protection. AoB Plants 6:plu042
Rinerson CI, Rabara RC, Tripathi P, Shen QJ, Rushton PJ (2015) The evolution of WRKY transcription factors. BMC Plant Biol 15:66
Schnippenkoetter W, Lo C, Liu G, Dibley K, Chan WL, White J, Milne R, Zwart A, Kwong E, Keller B, Godwin I, Krattinger S, Lagudah ES (2017) The wheat Lr34 multipathogen resistance gene confers resistance to anthracnose and rust in sorghum. Plant Biotechnol J 15:1387–1396
Shao Q, Han N, Ding TL, Zhou F, Wang BS (2014) SsHKT1;1 is a potassium transporter of the C3 halophyte Suaeda salsa that is involved in salt tolerance. Funct Plant Biol 41:790
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na/H antiporter sos1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477
Shi H, Lee BH, Wu SJ, Zhu JK (2003) Overexpression of a plasma membrane Na+/H+ antiporter gene improves salt tolerance in Arabidopsis thaliana Nat Biotechnol 21:81
Si AJ, Yu Y, Chen H, Tian Q (2017) Functional prediction of stress response GhWRKY2 gene in cotton (Gossypium hirsutum). J Agric Biotechnol 25:222–230
Song YS, Li JL, Liu ML, Meng Z, Liu KC, Sui N (2019) The role of nitrogen on maize seedlings under drought stress. Funct Plant Biol 46:350–359
Sui N, Yang Z, Liu ML, Wang BS (2015) Identification and transcriptomic profiling of genes involved in increasing sugar content during salt stress in sweet sorghum leaves. BMC Genomics 16:534
Sui N, Tian SS, Wang WQ, Wang MJ, Fan H (2017) Overexpression of glycerol-3-phosphate acyltransferase from Suaeda salsa improves salt tolerance in Arabidopsis Front Plant Sci 8:1337
Sui N, Wang Y, Liu SS, Yang Z, Wang F, Wan SB (2018) Transcriptomic and physiological evidence for the relationship between unsaturated fatty acid and salt stress in peanut. Front Plant Sci 9:1–12
Sun C, Palmqvist S, Olsson H, Mats Borén Ahlandsberg S, Jansson C (2003) A novel WRKY transcription factor, SUSIBA2, participates in sugar signaling in barley by binding to the sugar-responsive elements of the iso1 promoter. Plant Cell 15:2076–2092
Syvertsen JP, Garcia-Sanchez F (2014) Multiple abiotic stresses occurring with salinity stress in citrus. Environ Exp Bot 103:128–137
Szekely G, Abraham E, Cseplo A, Rigo G, Zsigmond L, Csisza J, Ayaydin F, Strizhov N, Jasik J, Schmelzer E, Koncz C, Szabados L (2008) Duplicated P5CS genes of Arabidopsis play distinct roles in stress regulation and developmental control of proline biosynthesis. Plant J 53:18
Tian XJ, Li XF, Zhou WJ, Ren YK, Wang ZY, Liu ZQ, Tang JQ, Tong HN, Fang J, Bu QY (2017) Transcription factor OsWRKY53 positively regulates brassinosteroid signaling and plant architecture. Plant Physiol 175:1337–1349
Ülker B, Mukhtar MS, Somssich IE (2007) The WRKY70 transcription factor of Arabidopsis influences both the plant senescence and defense signaling pathways. Planta 226:125–137
Wang Y, Ying Y, Chen J, Wang X (2004) Transgenic Arabidopsis overexpressing Mn–SOD enhanced salt-tolerance. Plant Sci 167:671–677
Wang F, Chen HW, Li QT, Wei W, Li W, Zhang WK, Ma B, Bi YD, Lai YC, Liu XL, Man WQ, Zhang JS, Chen SY (2015) GmWRKY27 interacts with GmMYB174 to reduce expression of GmNAC29 for stress tolerance in soybean plants. Plant J 83:224–236
Wang L, Zhang XL, Wang L, Tian Y, Jia N, Chen S, Shi NB, Huang XM, Zhou C, Yu YW, Zhang ZQ, Pang XQ (2017) Regulation of ethylene-responsive SlWRKYs involved in color change during tomato fruit ripening. Sci Rep 7:16674
Wei DD, Cheng D, Liu WB, Liu T, Yang XH, Zheng YH (2015) Adequate potassium application enhances salt tolerance of moderate-halophyte Sophora alopecuroides Plant Soil Environ 61:364–370
Wei DD, Zhang W, Wang CC, Meng QW, Li G, Chen TH, Yang XH (2017) Genetic engineering of the biosynthesis of glycine betaine leads to alleviate salt-induced potassium efflux and enhances salt tolerance in tomato plants. Plant Sci 257:74–83
Wu SJ, Ding L, Zhu JK (1996) SOS1, a genetic locus essential for salt tolerance and potassium acquisition. Plant Cell 8:617–627
Wu H, Shabala L, Zhou M, Shabala S (2014) Durum and bread wheat differ in their ability to retain potassium in leaf mesophyll: implications for salinity stress tolerance. Plant Cell Physiol 55:1749–1762
Xu ZL, Raza Q, Xu L, He XL, Huang YH, Yi JX, Zhang DY, Shao HB, Ma HX, Ali Z (2018a) GmWRKY49, a salt-responsive nuclear protein, improved root length and governed better salinity tolerance in transgenic Arabidopsis Front Plant Sci 9:809
Xu Z, Raza Q, Xu L, He X, Huang Y, Yi J, Zhang D, Shao HB, Ma H, Ali Z (2018b) GmWRKY49, a salt-responsive nuclear protein, improved root length and governed better salinity tolerance in transgenic Arabidopsis Front Plant Sci 9:809
Yang Q, Chen ZZ, Zhou XF, Yin HB, Li X, Xin XF, Gong Z (2009) Overexpression of SOS (salt overly sensitive) genes increases salt tolerance in transgenic Arabidopsis Mol Plant 2:22–31
Yang Y, Chi Y, Wang Z, Zhou Y, Fan B, Chen Z (2016) Functional analysis of structurally related soybean GmWRKY58 and GmWRKY76 in plant growth and development. J Exp Bot 67:4727–4742
Yang Z, Wang Y, Wei XC, Zhao X, Wang BS, Sui N (2017) Transcription profiles of genes related to hormonal regulations under salt stress in sweet sorghum. Plant Mol Biol Rep 35:1–14
Yang Z, Zheng HX, Wei XC, Song J, Wang BS, Sui N (2018) Transcriptome analysis of sweet sorghum inbred lines differing in salt tolerance provides novel insights into salt exclusion by roots. Plant Soil 430:423–439
You LL, Song QP, Wu YY, Li SC, Jiang CM, Chang L, Yang XH, Zhang J (2019) Accumulation of glycine betaine in transplastomic potato plants expressing choline oxidase confers improved drought tolerance. Planta 249:1963–1975
Yue Y, Zhang M, Zhang J, Duan L, Li Z (2012) SOS1 gene overexpression increased salt tolerance in transgenic tobacco by maintaining a higher K+/Na+ ratio. J Plant Physiol 169:255–261
Zhang T, Tan D, Zhang L, Zhang X, Han Z (2017) Phylogenetic analysis and drought-responsive expression profiles of the wrky transcription factor family in maize. Agri Gene 3:99–108
Zhao D, Zhao JR, Huang X, Li N, Liu Y, Huang ZJ, Zhang ZY (2011) Functional analysis of TNBL1 gene in wheat defense response to barley yellow dwarf virus using BSMV-VIGS technique. J Crops 37:2106–2110
Zhao X, Wei P, Liu Z, Yu B, Shi H (2017) Soybean Na+/H+ antiporter GmsSOS1 enhances antioxidant enzyme activity and reduces Na+ accumulation in Arabidopsis and yeast cells under salt stress. Acta Physiol Plant 39:19
Zheng LY, Guo XS, He B, Sun L, Peng Y, Dong S, Liu T, Jiang S, Ramachandran S, Liu C, Jing H (2011) Genome-wide patterns of genetic variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol 12:R114
Zhou QY (2008) Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY 21, and GmWRKY 54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol J 6:486–503
Zhou L, Wang N, Gong S, Lu R, Li Y, Li X (2015) Overexpression of a cotton (Gossypium hirsutum) WRKY gene, GhWRKY34, in Arabidopsis enhances salt-tolerance of the transgenic plants. Plant Physiol Biochem 96:311–320
Zou X, Seemann JR, Neuman D, Shen QJ (2004) A WRKY gene from creosote bush encodes an activator of the abscisic acid signaling pathway. J Biol Chem 279:55770–55779
Acknowledgements
We are grateful for financial support from the National Natural Science Research Foundation of China (31871538), the National Key R&D Program of China (2018YFD1000700, 2018YFD1000704), the Major Program of Shandong Provincial Natural Science Foundation (2017C03), the Opening Foundation of Shandong Provincial Key Laboratory of Crop Genetic Improvement, Ecology and Physiology (SDKL2018008-3). We thank Lesley Benyon, PhD, from Liwen Bianji, Edanz Group China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
Yushuang Song, Jinlu Li and Yi Sui imitated the manuscript. Yushuang Song and Jinlu Li performed experiments; Yushuang Song, Guoliang Han, Yi Zhang and Shangjing Guo collected data and carried out all analyses; Na Sui and Yi Sui conceptualized the idea and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2020_966_MOESM1_ESM.tif
Supplementary material 1 Fig. S1 Bioinformatics analysis of SbWRKY50. A Protein sequence analysis of SbWRKY50. B Structural domain analysis of SbWRKY50. C and D Homologous sequence alignment (C) and evolutionary analysis (D) of SbWRKY50 (AtArabidopsis thaliana, BeBambusa emeiensis, DoDichanthelium oligosanthes, LpLolium perenne, SiSetaria italica, ZmZea mays, ObOryza brachyantha, TaTriticum aestivum, TuTriticum urartu) (TIF 1225 kb)
11103_2020_966_MOESM2_ESM.docx
Supplementary material 2 Table S1 Number of W-boxes in promoters of genes. Table S2 Primers for amplification and qPCR gene expression analysis (DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Song, Y., Li, J., Sui, Y. et al. The sweet sorghum SbWRKY50 is negatively involved in salt response by regulating ion homeostasis. Plant Mol Biol 102, 603–614 (2020). https://doi.org/10.1007/s11103-020-00966-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-020-00966-4