Abstract
Key message
Two intercistronic regions were identified as functional intercistronic expression elements (IEE) for the simultaneous expression of aphA-6 and gfp in a synthetic operon in the chloroplast of C. reinhardtii.
Abstract
Chlamydomonas reinhardtii, a biflagellate photosynthetic microalga, has been widely used in basic and applied science. Already three decades ago, Chlamydomonas had its chloroplast genome transformed and to this day constitutes the only alga routinely used in transplastomic technology. Despite the fact that over a 100 foreign genes have been expressed from the chloroplast genome, little has been done to address the challenge of expressing multiple genes in the form of operons, a development that is needed and crucial to push forward metabolic engineering and synthetic biology in this organism. Here, we studied five intercistronic regions and investigated if they can be used as intercistronic expression elements (IEE) in synthetic operons to drive the expression of foreign genes in the chloroplast of C. reinhardtii. The intercistronic regions were those from the psbB-psbT, psbN-psbH, psaC-petL, petL-trnN and tscA-chlN chloroplast operons, and the foreign genes were the aminoglycoside 3′-phosphotransferase (aphA-6), which confers resistance to kanamycin, and the green fluorescent protein gene (gfp). While all the intercistronic regions yielded lines that were resistant to kanamycin, only two (obtained with intercistronic regions from psbN-psbH and tscA-chlN) were identified as functional IEEs, yielding lines in which the second cistron (gfp) was translated and generated GFP. The IEEs we have identified could be useful for the stacking of genes for metabolic engineering or synthetic biology circuits in the chloroplast of C. reinhardtii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic engineering in the chloroplast of the green alga Chlamydomonas reinhardtii has been used to express foreign genes to produce a wide range of recombinant therapeutic proteins (Almaraz-Delgado et al. 2014; Dyo and Purton 2018), including antibodies (Mayfield et al. 2003; Tran et al. 2009), immunotoxins (Tran et al. 2013a, b), antigens (Dreesen et al. 2010; Michelet et al. 2011; Jones et al. 2013), toxins (Kang et al. 2017) and growth factors (Rasala et al. 2010; Wannathong et al. 2016). However, the number of reports applying metabolic engineering in the chloroplast of C. reinhardtii is still scarce (Fukusaki et al. 2003; Gangl et al. 2015; Zedler et al. 2015). Nevertheless, the potential is enormous, novel or endogenous biosynthetic pathways can be introduced or modified (Gimpel et al. 2015) to obtain high value molecules such as carotenoids (Harada et al. 2014), as well as molecules that could be used as a renewable source for biofuel production such as lipids (Scranton et al. 2015) and to improve biohydrogen production (Wu et al. 2011). Despite the biotechnological potential of genetic engineering in the chloroplast of C. reinhardtii, tools that allow for a more systematic manipulation of the metabolism, by introducing and expressing entire pathways for instance, have not been developed (Scaife et al. 2015), resulting in a technology that is lagging behind in development compared to the advances that have been made in plant chloroplast biotechnology (Fuentes et al. 2018).
For many years it has been clear that some genes in the chloroplast of C. reinhardtii are expressed as bicistrons or polycistrons (Rochaix 1996; Harris 2009). While some of these polycistronic transcripts (psbB-psbT-psbH) are post transcriptionally processed to yield monocistronic units, others (psbF-psbL, ycf4-ycf3 and atpA-psbI-cemA-atpH) seem to remain as translatable bicistronic or polycistronic units (Mor et al. 1995; Boudreau et al. 1997; Drapier et al. 1998). The attention paid to the characterization of these operons has been somewhat uneven, being the transcripts for psbB-psbT-psbH and petA-petD the most widely studied while the transcripts for rps7-atpE and psbZ-psbM have been perhaps the least studied. More recently, two research groups have found that polycistronic transcription of genes in the chloroplast of C. reinhardtii seems to be more common than what was previously thought. Using high-throughput sequencing of small and long RNAs, Cavaiuolo et al. (2017) have reported that 84 of the 109 genes from C. reinhardtii chloroplast are arranged in 22 polycistronic units, while Gallaher et al. (2017) have identified 16 clusters of transcribed genes. Polycistronic transcription of genes in the chloroplast is a feature that could be exploited for the design and construction of synthetic polycistronic operons; genes from a particular multi-gene metabolic pathway could be stacked, making the introduction and expression of entire foreign pathways in the chloroplast of C. reinhardtii somewhat easier.
Transcript stability and translation initiation depends largely on the nature of untranslated regions (UTR) at the 5′- and 3′-ends (Stern et al. 2010; Rasala et al. 2011) as well as in nucleus-encoded trans-acting protein factors (Johnson et al. 2010). In polycistronic mRNAs, intergenic regions act as 3′UTR of the first cistron and as 5′UTR of the second cistron, containing elements for mRNA processing and mRNA stability (Vaistij et al. 2000a, b; Stoppel and Meurer 2013). It has been proposed that processing of polycistronic transcripts to monocistronic mRNAs is initiated through non-specific cleavages by endonucleases RNase E and RNase J (Pfalz et al. 2009; Luro et al. 2013). Maturation of the 5´-end termini is then achieved as the result of the activity of exonucleases (for example, RNAse J), which degrade the mRNA until encountering a sequence-specific RNA-binding protein. In plants and in C. reinhardtii, these proteins have been demonstrated to be members of the helical-hairpin-repeat protein families: PPR (pentatricopeptide repeat), TPR (tetratricopeptide repeat) and OPR (octotricopeptide repeat) proteins. Proteins of these families have also been implicated in the maturation of the transcript 3′-end in plants but not in Chlamydomonas, where the presence of a stem-loop seems to be the structure that predominantly prevents exonuclease activity. Helical-hairpin-repeat proteins bind on a sequence-specific manner to mRNAs and act as trans-acting factors that control splicing, editing, stabilization, turn over, processing and translation activation (Auchincloss et al. 2002; Loiselay et al. 2008; Stern et al. 2010; Eberhard et al. 2011; Rahire et al. 2012; Wang et al. 2015; Douchi et al. 2016). The absence of these regulatory elements, cis and trans-acting factors, brings a rapid degradation or allows only for low accumulation of primary transcripts (Salvador et al. 2004) and, in some cases, a reduction in translation efficiency is seen (Trösch et al. 2018). There is also evidence that coding regions might have a significant contribution to transcript stability. When the 5´UTR of the chloroplast rbcL was used to drive the expression of a foreign gene, the mRNA generated was susceptible to rapid degradation upon exposure to light. This susceptibility was eliminated with the addition of a segment of the rbcL coding region (Salvador et al. 1993). Transcript processing and stability are, not surprisingly, complex and multi factor events.
In tobacco chloroplasts a small element, termed the intercistronic expression element (IEE), with origin on the psbT-psbH intercistronic region from the psbB operon, was used for bicistronic expression of nptII and yfp (Zhou et al. 2007). While the presence of the IEE was not required for processing and maturation of the first cistron (nptII) it was required for stability and translation of the second cistron (yfp). A few years later, the IEE was used to express three genes involved in the tocochromanol pathway for vitamin E production in tobacco and tomato (Lu et al. 2013). The plants expressing the three genes, stacked one after the other using the IEE in between them, accumulated more tocochromanol than the lines where the genes were expressed using a bacterial type operon or independently. More recently, Fuentes et al. (2016) used IEEs to express the core biosynthetic pathway of artemisinic acid production in tobacco chloroplasts, making evident again that this element can be used for metabolic engineering in this plant. The functionality of the tobacco IEE has been attributed to the fact that it contains the target sequence for the half-a-tetratricopeptide (HAT) repeat RNA-binding protein HCF107 (Hammani et al. 2012), which stabilizes the transcript and makes translation more efficient (Zhou et al. 2007; Lu et al. 2013; Bock 2013). In plants, the HAT protein HCF107 is required for the stability of the psbH transcripts (Felder et al. 2001; Hammani et al. 2012).
Even though the tobacco IEE has been demonstrated to be sufficiently robust, recently, Legen et al. (2018) have shown that when it is used to stabilize the transcripts of foreign genes, a negative effect is seen on the stabilization of the endogenous psbH transcript. Stabilization of the foreign transcripts, achieved by the binding of the HCF107 HAT protein to the target site contained in the IEE, yields a reduction on the accumulation of the psbH transcript. There does not seem to be enough HCF107 to serve target sites in excess. To overcome this problem, Legen et al. (2018) proposed the use of binding sites for PPR proteins as IEE-like elements. They have showed that by placing target sites for PPR10, HCF152, CCR2 and the binding site of a not yet identified protein, located upstream of the rpl12 reading frame, as IEEs in a synthetic neo-egfp bicistron in tobacco chloroplasts, GFP accumulates to various levels. These results indicate that alternative sequences, to the already popular IEE, can be developed and used as modulators and enhancers of genes expression in polycistronic constructs.
In this regard, identification of IEEs for C. reinhardtii chloroplasts is strongly needed to speed up and improve metabolic engineering. IEEs would facilitate the stacking of genes and consequently make the expression of entire biosynthetic pathways in the chloroplast of a biotechnology relevant organism a reality. Here, we have examined different intercistronic regions from the chloroplast genome of C. reinhardtii to determine their usefulness as IEEs in the construction of synthetic operons. We show that the intercistronic regions from the psbN-psbH and tscA-chlN bicistrons can be placed between the coding sequences of two foreign genes and yield monocistronic translatable units that produce the aminoglycoside phosphotransferase enzyme (APH(3′)-VI) and the green fluorescent protein (GFP). The IEEs that we have identified could be used for the stacking and co-expression of genes in synthetic operons in C. reinhardtii.
Materials and methods
Algal strain and culture conditions
Chlamydomonas reinhardtii wild-type CC-125 (mt+) was obtained from the Chlamydomonas Resource Center (University of Minnesota, USA). Wild-type and transformed cells were grown in Tris-acetate-phosphate (TAP) medium (Gorman and Levine 1965) at 25 °C under a photoperiod (16 h /8 h, light/dark) with white LED lighting (10,000–12,500 lx). For transformed lines, TAP solid medium was supplemented with spectinomycin (150 µg/mL) and kanamycin (100 µg/mL) while for liquid cultures, the antibiotics were used at concentrations of 50 µg/mL for spectinomycin and 20 µg/mL for kanamycin.
Plasmid construction
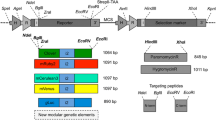
Expression cassettes were first assembled in the pJ248/aphA-6/gfp vector. This vector carries the aphA-6 gene (Bateman and Purton 2000) and a codon-optimized gfp gene (DNA 2.0, now AUTUM, USA; Supplementary Data 1). The genes are under the control of the rbcL promoter (PrbcL) and terminator (TrbcL). The aphA-6 gene contains a HindIII site at the 3′-end while the gfp gene contains a NcoI site at the 5′-end for convenient insertion of the IEEs in between them (Fig. 1a). All endogenous IEEs (IEE-1 to IEE-5) were amplified by PCR using C. reinhardtii total DNA as template, IEE-6 (T7 g10) was amplified from the previously reported tobacco chloroplast transformation vector pZSJH1-PrrnIp24 (Zhou et al. 2008). Primers were designed to introduce HindIII and NcoI sites at the 5′- and 3′-ends respectively (Supplementary Table 1). The expression cassettes PrbcL/aphA-6/IEE/gfp/TrbcL were transferred to the BamHI site of C. reinhardtii chloroplast transformation vector p320 (Guzmán-Zapata et al. 2016) to generate p320-IEE vectors (p320-IEE1, p320-IEE2, p320-IEE3, p320-IEE4, p320-IEE5, p320-IEE6). Chloroplast transformation vector p320 targets the insertion of foreign genes to the intergenic region between exon 5 of psbA and 5S rRNA by homologous recombination in the C. reinhardtii chloroplast genome (Fig. 1b).
Generation of transplastomic C. reinhardtii lines with a synthetic operon using different intercistronic regions. a Chloroplast transformation vector p320. This vector targets the insertion of foreign genes to the intergenic region between 23S-5S rRNA and exon 5-intron 4 of psbA. b and c Physical map of the synthetic bicistronic operon and origin of the intercistronic regions used, respectively. In the synthetic operon aphA-6/IEE/gfp genes are under the control of the rbcL promoter (PrbcL) and terminator (TrbcL). The intercistronic regions indicated were used to generate vectors p320-IEE1 to p320-IEE6. d Physical map of the transformed genome of lines obtained with vectors of the series p320-IEE. Introns and exons of the chloroplast genome are represented in filled black and white boxes, respectively. Foreign genes are represented with light grey boxes while the intercistronic region is represented by a line filled box. A DNA fragment generated with SacII and PstI restriction enzymes was used for restriction fragment length polymorphism analysis and is represented as a black line with borders. Probes used for Southern and northern blotting are indicated with bold black lines. Primers are indicated with arrows and their respective names
Chloroplast transformation, selection of transformed lines and PCR analysis
Chloroplast transformation was carried out by particle bombardment using the PDS-1000/He Particle Delivery System (Bio-Rad, USA) as described by Guzmán-Zapata et al. (2016). Briefly, a 250 µL aliquot of C. reinhardtii cellular culture (1 × 108 cell/mL) was placed in a Petri dish with solid TAP medium supplemented with antibiotics, and then bombarded with M-10 (0.7 µm) tungsten particles (Bio-Rad) coated with one microgram of each plasmid. After transformation, Petri dishes were incubated overnight in the dark, and then placed under a photoperiod for 4 weeks until antibiotic resistant colonies were visualized. Chloroplast transformation vector p228 (Chlamydomonas Resource Center, University of Minnesota), which by introducing a point mutation in the 16S rRNA gene confers resistance to spectinomycin, was used for co-transformation with the p320-IEEs vectors mentioned above. Selection of primary transformed cells was carried out in spectinomycin (150 µg/mL). Selection for kanamycin-resistant transformed lines was carried out in TAP media supplemented with 100 µg/mL kanamycin for solid medium and 20 µg/mL for liquid media. Kanamycin-resistant lines were verified by colony PCR using the resin Chelex-100 (Bio-Rad) as described previously (Cao et al. 2009; Guzmán-Zapata et al. 2016). To identify transformed lines with the transformation cassette, both gfp and aphA-6 genes were amplified by PCR using primers pairs qFJAB7/qRJAB7 and qFJAB8/qRJAB8, respectively (Supplementary Table 1). For detection of the mRNA transcripts by RT-PCR we first synthesized the cDNA using a random hexamer and M-MuLV Reverse Transcriptase (New England Biolabs, USA) following the manufacturer’s instructions. Samples were treated with DNase I (New England Biolabs).
DNA isolation and Southern blot
Total DNA was extracted from 15 mL (1 × 105 cell/mL) of liquid culture using the Wizard Genomic DNA Purification Kit (Promega, USA) following the manufacturer’s instructions for plant DNA extraction. Briefly, cells were recovered by centrifugation at 3500×g and then resuspended in Nuclei Lysis Solution and incubated for 15 min at 65 °C. RNase Solution was added and the sample incubated for 15 min at 37 °C. Proteins were eliminated by protein precipitation with Protein Precipitation Solution and DNA was purified by ethanol precipitation and then resuspended with nuclease-free water.
For Southern blotting, 10 µg of total DNA were digested with restriction enzymes SacII and PstI and then separated by electrophoresis in a 0.6% agarose gel. DNA was transferred to a Zeta-probe nylon membrane (Bio-Rad) by capillary blotting following a standard protocol (Reed and Mann 1985). The psbA probe (1 343 bp), comprising part of the psbA gene, was generated by PCR using primers NVDF354 and NVDF355 (Supplementary Table 1). The probe was labeled with [α-32P]-dATP by random priming (Sambrook and Russell 2001). Hybridization was carried out at 65 °C following a standard protocol (Sambrook and Russell 2001). Membranes were exposed to a storage phosphor imaging screen, followed by scanning with a Personal Molecular Imager System (Bio-Rad) and analyzed with Quantity One 1-D analysis software (Bio-Rad).
RNA isolation and northern blot
Total RNA was isolated from a 50-mL (1 × 105 cell/mL) culture with Tri-Reagent (Sigma-Aldrich, USA). Fifteen micrograms of RNA were denatured, separated by electrophoresis in a denaturing, formaldehyde-containing, 2% agarose gel and blotted onto a Zeta-Probe nylon membrane (Bio-Rad) by capillary blotting. Membranes were stained with methylene blue solution to verify RNA transference. Probes for aphA-6 and gfp were generated by PCR using primers NVDF233/234 and NVDF235/236 (Supplementary Table 1), respectively, using plasmid pJ248/aphA-6/gfp as a template (Fig. 1b). After purification by agarose gel electrophoresis, probes were labeled with [α-32P]-dATP by random priming and hybridization was carried out at 65 °C following a standard protocol (Sambrook and Russell 2001). Northern blots were exposed to a storage phosphor imaging screen followed by scanning with a Personal Molecular Imager System (Bio-Rad) and analyzed with Quantity One 1-D analysis software (Bio-Rad).
Protein extraction and immunoblot analyses
Total soluble protein was extracted as described previously (Bertalan et al. 2015). A 15-mL aliquot of cell culture was harvested by centrifugation (3000×g, 5 min). The pellet was washed once with 1 mL of solution A (0.1% Na2CO3) and then resuspended in 300 µL of solution A, 200 µL of solution B (5% SDS, 30% sucrose) and 25 µL ß-mercaptoethanol (14.3 M) and agitated with a vortex for 25 min at room temperature. Cell debris was removed by centrifugation for 10 min at 16,000×g at 4 °C. The supernatant was recovered, and total cellular protein was quantified by Bradford assay with Quick Start Bradford 1× Dye Reagent (Bio-Rad). Proteins (30 µg) were separated by electrophoresis in 15% SDS-Polyacrylamide gels, and subsequently transferred to Immun-Blot PVDF Membranes (Bio-Rad). Membranes were probed with a chicken anti-GFP primary antibody (1:5000, Aves Lab Inc., USA) followed by probing with a goat anti-chicken antibody conjugated to horseradish peroxidase (1:5000, Abcam, UK). Immunobiochemical detection was carried out using the Western Lightning Plus-ECL reagent (Perkin Elmer Inc., USA) following the manufacturer’s instructions and recorded using a ChemiDoc MP Imaging System (Bio-Rad). Purified recombinant GFP (rGFP) was obtained by overexpressing the gfp gene using the pET28-b vector in E. coli BL21 Star (Invitrogen, USA) and subsequently purified using Ni–NTA agarose following standard protocols (Invitrogen).
Microscopy
Subcellular localization of GFP was determined using a Zeiss Confocal Laser Scanning Multiphoton LSM-710NLO microscope system (Carl Zeiss, Germany), equipped with a plan-apochromat 63×/1.4 oil objective. GFP fluorescence was visualized using a 468-nm excitation filter and a 510-nm emission filter. Chlorophyll autofluorescence was visualized using a 450-nm and 705-nm excitation and emission filters, respectively. Images were processed and analyzed using the ZEN software (Carl Zeiss).
Circularized RT-PCR
RNA circularization was performed as described previously by Zandueta-Criado and Bock (2004). Briefly, 5 µg of total RNA were self-ligated at 37 °C for 2 h with 10 units of T4 RNA ligase (New England Biolabs) in a final reaction volume of 50 µL. For cDNA synthesis, 1 µg of the circularized RNA, 2 µL (50 µM) of oligonucleotide and 1 µL of dNTPs (10 mM), in a 10-µL total volume reaction, were denatured at 70 °C for 5 min and used for cDNA synthesis with 200 U of M-MuLV Reverse Transcriptase (New England Biolabs) following the manufacturer’s instructions. The qRJAB7 primer was used as a specific oligonucleotide for gfp transcripts. The resulting cDNA of each reaction was used directly as a template for PCR reactions, and amplified following standard protocols using KOD Hot Start polymerase (Merck Millipore, USA) using primer pairs qRJAB7/NVDF267 (Supplementary Table 1). PCR products were cloned in pBlueScript SK II (+) and sequenced with universal primers M13F and M13R.
Results
Selection of intercistronic regions and construction of the synthetic operons
To characterize IEE that can be used for the expression of multiple foreign genes in the chloroplast of C. reinhardtii we followed the next approach. First, we looked for genes that are expressed in a polycistronic transcript, for which there is northern blot evidence, and identified the following that comply with such criteria: petA-petD (Sakamoto et al. 1994; Sturm et al. 1994; Loiselay et al. 2008), psbB-psbT-psbH (Monod et al. 1992; Johnson and Schmidt 1993; Vaistij et al. 2000b; Loizeau et al. 2014), psbD-exon 2 psaA (Herrin and Schmidt 1988; Choquet et al. 1998), rps7-atpE (Robertson et al. 1990), psaC-ORF58-petL (Takahashi et al. 1991, 1996) and tscA-chlN (Rochaix 1996; Glanz et al. 2012). From these, we selected the intercistronic regions that had a length shorter than 1 kb: psbB-psbT-psbH (451 bp and 569 bp, respectively), psaC-petL (607 bp), petL-trnN (334 bp) and tscA-chlN (650 bp), identified hereafter as IEE-1, IEE-2, IEE-3, IEE-4 and IEE-5, respectively. petA-petD and rps7-atpE have a size of 2 531 bp and 1 229 bp respectively. Ideally, IEEs should be short, as this facilitates manipulation and has little impact on the final size of the polycistronic transcript. With this in mind we have left out the intercistronic regions from petA-petD and rps7-atpE.
The five chloroplast intercistronic regions but one, the petL-trnN, come from bicistrons that contain protein coding genes. The petL-trnN is a bicistron that contains a protein coding gene, petL, and a tRNA coding gene, trnN. As such, this intercistronic region would not be useful for the translation of the second cistron and we have included it both as a control and to preliminarily explore the potential of this region for the expression of a foreign protein and an RNA. Additionally, we used the 5′UTR from gene 10 of bacteriophage T7 (Olins et al. 1988; Olins and Rangwala 1989). This 5′UTR, also referred to as 5′UTR T7g10, lacks a bicistronic processing site but contains a ribosome-binding site (RBS) and has been used in plant chloroplasts for the expression of genes and accumulation of proteins at a high level (Oey et al. 2009). We included the 5′UTR T7g10, hereafter identified with the code IEE-6, because to the best of our knowledge, this element has not been tested in the chloroplast of C. reinhardtii.
With the selected intercistronic regions, IEE1–IEE6 (Fig. 1c, Supplementary Data 1), we constructed a series of synthetic bicistronic operons and inserted them in the previously reported chloroplast transformation vector p320 (Guzmán-Zapata et al. 2016). This vector targets the insertion of foreign genes to the intergenic region between 23S rRNA-5S rRNA and exon 5-intron 4 of psbA in the chloroplast genome of C. reinhardtii (Fig. 1a). The synthetic operons comprised the selectable marker aphA-6 (Bateman and Purton 2000) and the codon-optimized reporter gene gfp (Supplementary Data 1), placed under the control of the rbcL promoter and terminator (PrbcL, TrbcL). Each intercistronic region was placed between aphA-6 and gfp (Fig. 1b) to generate vectors p320-IEE1, p320-IEE2, p320-IEE3, p320-IEE4, p320-IEE5, and p320-IEE6 (Fig. 1c). We reasoned that both aphA-6 and gfp would be transcribed from the rbcL promoter as a single mRNA unit and that translation of the first cistron (aphA-6) would occur from cis translation elements present in the 5´UTR of PrbcL (Wang et al. 2008). In turn, translation of the second cistron (gfp) would only occur if the intercistronic region yields a transcript that can either be processed to generate translatable monocistronic transcripts or serve directly as a bicistronic transcript for translation (Stern et al. 2010). In any case, the resultant mRNA would require a certain stability and to contain the necessary cis-elements and the regions for interaction with trans-acting factors, if any. Although the aphA-6 gene, confers resistance to kanamycin and selection of transformed lines could have been carried out directly on this antibiotic, we did not know if translation of the aphA-6 mRNA would be somehow affected by the presence of the IEE, placed downstream of the coding sequence. For this reason, we decided to use vector p228 for co-transformation and primary selection of transformed lines (Newman et al. 1990; Chlamydomonas Resource Center, University of Minnesota). Vector p228 carries a point mutation in the 16S rRNA gene so when the endogenous 16S rRNA is replaced, strains carrying the point mutation become resistance to spectinomycin.
Chloroplast transformation and primary analysis of transplastomic lines
To introduce the synthetic operons in the chloroplast of C. reinhardtii, vectors p320-IEE1, p320-IEE2, p320-IEE3, p320-IEE4, p320-IEE5 and p320-IEE6, along with vector p228, were bombarded using a particle bombardment device. After four events of transformation we recovered a total of 47, 33, 50, 8, 18 and 8 spectinomycin-resistant colonies for vector combinations p320-IEE1/p228, p320-IEE2/p228, p320-IEE3/p228, p320-IEE4/p228, p320-IEE5/p228 and p320-IEE6/p228, respectively (Fig. 2a). Because not all the recovered spectinomycin-resistant lines are the result of transformation with both vectors (they could be the result of transformation with p228 only), we tested each colony to determine if they were resistant to both spectinomycin and kanamycin by growing them in TAP media with these antibiotics (Fig. 2b, Supplementary Fig. 1). With this test we determined that 16, 11, 33, 8, 2 and 4 lines were the successful events of transformation with vectors p320-IEE1, p320-IEE2, p320-IEE3, p320-IEE4, p320-IEE5 and p320-IEE6, respectively (Fig. 2a, Supplementary Fig. 1). The resistance to kanamycin in these lines strongly suggests that the first cistron (aphA-6), independently of the intercistronic region placed between the two cistrons, is translated, producing in turn the detoxifying enzyme APH(3′)-VI.
Transplastomic lines obtained after co-transformation with p320-IEE1-6 and p228. C. reinhardtii lines recovered after transformation with vectors of the series p320-IEE (p320-IEE1–6, Fig. 1c), and vector p288, which contains a fragment of the chloroplast 16S rRNA with a point mutation that confers resistance to spectinomycin. a Total number of primary lines recovered and resistant to spectinomycin (Spec); total number from the primary lines that were also resistant to kanamycin (Km). b Transplastomic lines for each IEEs were grown in TAP media (TAP), TAP media supplemented with spectinomycin (TAP/Spec; 150 µg/mL spectinomycin) and TAP media supplemented with kanamycin (TAP/Km; 100 µg/mL kanamycin)
We selected 2–3 independent lines from each of the strains transformed with p320-IEE1, p320-IEE2, p320-IEE3, p320-IEE4, p320-IEE5 and p320-IEE6, and carried out a more detailed molecular characterization. Here we present the results of these lines, namely CrC-IEE1, CrC-IEE2, CrC-IEE3, CrC-IEE4, CrC-IEE5 and CrC-IEE6. First, we checked by PCR for the presence of both aphA-6 and gfp and detected amplification products of the expected sizes, 248 bp for aphA-6 and 178 bp for gfp, in all transplastomic lines but not in the wild-type strain (Fig. 3a). Then, we performed a restriction fragment length polymorphism (RFLP) using SacII and PstI restriction enzymes (Fig. 1a, d) to determine homoplasmy and that the synthetic operon had integrated in the chloroplast genome in the targeted site. We used a probe that aligns in the psbA gene (Fig. 1d) and detected restriction fragments of 3.7 kb for the wild-type and of 6.6, 6.7, 6.7, 6.5, 6.8 and 6.2 kb for transplastomic lines CrC-IEE1, CrC-IEE2, CrC-IEE3, CrC-IEE4, CrC-IEE5 and CrC-IEE6, respectively (Fig. 3b). This result indicated that the synthetic operon had integrated in the expected site and that no copies of the wild-type genome could be detected (with this method). When copies of the wild-type genome cannot be detected it is acceptable to say that homoplasmy has been reached.
Confirmation of transformation in transplastomic lines by PCR and RFLP. a Amplification of a fragment of aphA-6 and gfp by PCR in the transplastomic lines. b RFLP analysis to determine homoplasmy. DNA samples from wild type and transplastomic lines were extracted, digested with SacII and PstI, separated by electrophoresis in a 0.6% agarose gel, blotted and hybridized to the radiolabeled psbA probe shown in Fig. 1d. The probe detects a 3.7 kb fragment in the wild-type strain and fragments of 6.6 kb, 6.7 kb, 6.7 kb, 6.5 kb, 6.8 kb and 6.2 kb in transplastomic lines CrC-IEE1, CrC-IEE2, CrC-IEE3, CrC-IEE4, CrC-IEE5 and CrC-IEE6 obtained with vectors p320-IEE1, CrC-IEE2, CrC-IEE3, CrC-IEE4, CrC-IEE5 and CrC-IEE6
Analysis of transcript processing in transplastomic lines
To determine the presence of aphA-6 and gfp transcripts we carried out qualitative RT-PCR and northern blot analyses. For RT-PCR, we first synthesized the cDNA as described in the section "Materials and methods". Using specific primers for gfp and aphA-6 we determined, by RT-PCR, that in all lines the aphA-6 and gfp mRNA transcripts were present (Fig. 4a). For all lines, except for the wild-type strain, we detected an amplification product of 248 bp for aphA-6 and a product of 178 bp for gfp. RNA samples, not taken through the cDNA step, were also used as template for the RT-PCR to rule out amplification from DNA contamination (negative control, not shown in Fig. 4a). We used the constitutively expressed rbcL transcript as a control and detected it in all lines including the wild-type strain. Detection of the transcripts by RT-PCR reliably indicates that the transcripts are present but it is not useful to determine the monocistronic or bicistronic nature of the transcripts. Because we were ultimately interested in determining the cleavage pattern of the synthetic operon, as this could help us to better understand the molecular behavior of the IEEs used, we performed a northern blot analysis as is described next.
Analysis by RT-PCR and northern blotting of aphA-6 and gfp transcripts in transplastomic lines. a RT-PCR assays were performed to detect aphA-6 and gfp and rbcL transcripts in wild-type and transplastomic lines after transformation with vectors p320-IEE1, p320-IEE2, p320-IEE3, p320-IEE4, p320-IEE5 and p320-IEE6. b Northern blot analysis of transplastomic lines using a radiolabeled aphA-6 probe (See Fig. 1d). The upper band in all lanes corresponds to the bicistronic form of the transcript composed of aphA-6-IEE-gfp, while the bands indicated with the stars correspond to the monocistronic form of aphA-6. c Northern blot analysis of transplastomic lines using a radiolabeled gfp probe (See Fig. 1d). The upper band in all lanes corresponds to the bicistronic form of the transcript composed of aphA-6-IEE-gfp, while the bands indicated with the stars correspond to the monocistronic form of gfp. Membranes stained with methylene blue are shown
Total RNA was extracted, separated by electrophoresis and blotted onto a nylon membrane. It was then hybridized with radiolabeled probes for both genes in the synthetic operon. First, a probe specific for aphA-6 revealed that larger transcripts with a size of around 2.0 kb for Crc-IEE1, Crc-IEE2, Crc-IEE3 and Crc-IEE5, 1.8 kb for Crc-IEE4 and 1.6 kb for Crc-IEE6 were present in the transformed lines (Fig. 4b). The sizes of these transcripts corresponded to non-cleaved bicistronic units originating from the PrbcL. In wild-type cells, the IEEs that we have used in this study also yield a stable bicistron of the genes from which they come from (Takahashi et al. 1991; Johnson and Schmidt 1993; Rochaix 1996; Vaistij et al. 2000b; Glanz et al. 2012). Our finding indicates then that the IEEs, even in a foreign gene context are sufficient to yield stable bicistronic transcripts. In addition to these bicistronic transcripts, we detected smaller-sized transcripts for all lines except for Crc-IEE6. For lines CrC-IEE1, CrC-IEE3, CrC-IEE4 we detected a discrete, clearly defined transcript of approximately 800 bp, which presumably corresponds to the cleaved and processed monocistronic form of aphA-6 (Fig. 4b, indicated with stars). For lines Crc-IEE2 and Crc-IEE5 we also detected discrete clearly defined bands, however, we detected two in each case: transcripts of 1 kb and 1.1 kb for Crc-IEE2 and transcripts of 1.1 kb and 1.3 kb for Crc-IEE 5 (Fig. 4b, also indicated with stars). Faint bands were also detected for Crc-IEE5, most likely the result of complex processing after cleavage of the stable and highly abundant bicistronic transcript. As mentioned above, we could not detect a monocistronic unit for aphA-6 in line Crc-IEE-6 and at least in this case, we can say that the aphA-6 transcript can be translated from a bicistronic uncleaved aphA-6/gfp mRNA. The presence of the aphA-6 cistron in all lines, either in the bicistronic or monocistronic form, is consistent with the kanamycin resistant phenotype observed.
Then we performed a northern blot assay using a probe for gfp, the second cistron of the synthetic operon. Consistent with what we observed when the aphA-6 probe was used, with the gfp probe we also detected the presence of the bicistronic unprocessed mRNA transcript in all lines (Fig. 4c). The sizes of these bicistronic transcripts are, as mentioned above, 2.0 kb for Crc-IEE1, Crc-IEE2, Crc-IEE3 and Crc-IEE5, 1.8 kb for Crc-IEE4 and 1.6 kb for Crc-IEE6. A monocistronic form of gfp, with sizes 0.7 kb, 1.1 kb, 0.7 kb and 0.9 kb were detected in lines Crc-IEE2, Crc-IEE3, Crc-IEE4 and Crc-IEE-5, respectively (Fig. 4c, indicated with stars). Note that these monocistronic units seem to be less abundant in Crc-IEE3 and Crc-IEE4 than in Crc-IEE2 and Crc-IEE5. Interestingly, for line Crc-IEE-5 additional bands were detected, again suggesting that this IEE is yielding mRNA transcripts with a broad range of sizes (Fig. 4c). Transcripts of various sizes are frequently reported as the result of polycistronic processing (Barkan 1988; Hahn et al. 1998; Monde et al. 2000). This has been attributed to the initial random cleavage by RNases in intercistronic regions (Luro et al. 2013; Pfalz et al. 2009). However, the presence of transcripts of various sizes as the result of mRNA instability and rapid degradation cannot be discarded. For lines Crc-IEE1 and Crc-IEE6 we could not detect a monocistronic unit for gfp. In the case of Crc-IEE1, the bicistron is in fact cleaved, as the aphA-6 transcript was detected but not the gfp transcript, indicating that the resultant cistron containing the gfp unit is highly unstable and shortly degraded after processing of the bicistronic aphA-6-IEE1-gfp transcript. In wild-type cells, in the expression of the psbB-psbT-psbH polycistron only the psbB, psbH and psbB-psbT units but not the psbT monocistronic unit can be detected (Monod et al. 1992; Vaistij et al. 2000a, b). Our IEE1 comes from the psbB-psbT region and it is behaving similarly, we can detect the bicistronic form of the transcript plus the monocistronic form of the first cistron but not the second one. In the case of Crc-IEE6, the bicistron does not seem to be processed whatsoever as neither aphA-6 nor gfp were detected in the monocistronic form. This is not surprising as the 5´UTR T7g10 does not seem to contain a recognizable sequence for chloroplast RNases to initiate cleavage and processing.
Immunodetection and confocal microscopy detection of GFP in transplastomic lines
To determine if the second cistron, corresponding to gfp, was efficiently translated, we checked for the presence of GFP by western blotting and by confocal laser microscopy. For immunodetection, total soluble protein was extracted from lines Crc-IEE1, Crc-IEE2, Crc-IEE3, Crc-IEE4, Crc-IEE5, Crc-IEE6 and used to determine the presence of GFP using a specific polyclonal antibody. It is worth reminding the reader that all the lines were resistant to kanamycin, indirectly indicating that the first cistron of the synthetic operon was translatable. However, when we checked for the presence of GFP we could not detect GFP in CrC-IEE1, CrC-IEE3, Crc-IEE4 and Crc-IEE6 while we were able to detect a positive result in lines CrC-IEE-2 and Crc-IEE-5 (Fig. 5a). Drawing conclusions on why GFP was not detected in lines CrC-IEE1, Crc-IEE4 and Crc-IEE6 is somewhat straightforward. GFP could have only been generated from the bicistronic form of the transcript, as in both cases the gfp monocistronic mRNA was absent (Fig. 4 c). We can conclude then that the gfp cistron cannot be used for translation when located as the second cistron using the IEE regions from the psbB-psbT and T7g10 intercistronic regions. Interestingly, in both cases, the presence of these intercistronic regions in the uncleaved bicistronic mRNA does not seem to affect translation of aphA-6 to produce APH(3′)-VI. In the case of Crc-IEE4 we did not expect translation of the gfp transcripts from either the bicistronic or monocistronic units; the IEE4 comes from petL-trnN, a protein-tRNA intercistronic region and as such lacks the necessary cis elements for translation. However, the usefulness of this IEE could rely on the potential to use it as an element to co-express a protein coding gene along with an RNA coding gene (e.g. a Cas9 protein with a gRNA) eventually needed in the implementation of the CRISPR/Cas9 technology in C. reinhardtii chloroplast. As mentioned above,
Detection and quantification of GFP protein in transplastomic lines. a Total soluble protein was extracted from wild-type and transplastomic lines and 30 µg separated by SDS-PAGE, blotted and incubated with a polyclonal antibody against GFP. The expected size for GFP is of approximately 27 kDa. Recombinant GFP (rGFP) from E. coli was used as a positive control. b Quantification of GFP protein in positive lines Crc-IEE2 and Crc-IEE5. Quantification was performed by comparison with the density of a dilution series of rGFP in the range 5–30 ng. Picture of the staining of the gels with Coomassie brilliant blue to show equal loading of protein are also presented
GFP could only be detected in lines CrC-IEE-2 and Crc-IEE-5, two of the lines where we also detected processing of the bicistronic mRNA to yield monocistronic aphA-6 and gfp. This could tempt us to think that the monocistronic form of gfp is required for translation, however in line Crc-IEE3 the monocistronic for of gfp is also detected, albeit to a lesser extent, which could contradict our presumption, because in line Crc-IEE3 no GFP was detected. An alternative explanation for the accumulation of GFP in these two lines could be that GFP correlates with the accumulation of bicistronic mRNA. CrC-IEE-2 and Crc-IEE-5 accumulate bicistronic mRNA to a greater degree than CrC-IEE-3 which could make the difference in the level of protein accumulation. Monocistronic transcripts are not always required for translation and there is evidence showing that individual cistrons can be translated from bicistronic mRNA (Barkan 1988; Zoschke and Barkan 2015). We cannot say at present if translation is occurring from the mono or bicistronic form of the transcript in lines Crc-IEE2 and Crc-IEE5. However, we can conclusively say that the use of the intercistronic regions from psbN-psbH and tscA-chlN in a synthetic bicistronic construct can efficiently serve to generate APH(3′)-VI and GFP from translatable transcripts.
Having detected the presence of GFP in lines Crc-IEE2 and Crc-IEE5, we then determined the level of accumulation of this protein by densitometry of the western blot membrane and found it to be 0.05–0.1% of total soluble protein (Fig. 5b). We then used confocal laser-scanning microscopy to visualize GFP fluorescence. We screened all lines but as expected, and in accordance with the results of immunodetection, we could only visualize GFP in lines Crc-IEE2 and Crc-IEE5 (Fig. 6). GFP fluorescence in these lines is solid evidence that the gfp transcripts are being translated and that the resulting protein is folded correctly to generate biologically active GFP. We did not observe fluorescence in the wild-type line (Fig. 6) nor in the rest of the lines (pictures not shown).
Detection of GFP fluorescence in transplastomic lines Crc-IEE2 and Crc-IEE5 by confocal laser-scanning microscopy. Wild-type and transplastomic lines were visualized by CLSM using a 468-nm excitation filter and a 510-nm emission filter. Chlorophyll fluorescence was determined using a 450 nm and 705 nm excitation and emission filters. Bright fields and the overlays of the GFP/chlorophyll fluorescence are shown
Mapping of the 5′-end termini in the gfp cistrons
Having demonstrated that the intercistronic regions from psbN-psbH and tscA-chlN (IEE2 and IEE5) can be used as IEEs in synthetic polycistronic operons, as both yielded biologically active APH(3′)-VI and GFP, we then wished to map the 5′-end of the monocistronic gfp transcripts, generated after processing of the bicistronic aphA-6-gfp transcript when this regions are used (Fig. 4c). Total RNA was extracted from lines Crc-IEE3 and Crc-IEE5, circularized using T4 RNA ligase and used for the synthesis of cDNA using primer qRJAB7. The cDNA was then used as template for PCR amplification of the head–tail junction with primers qRJAB7 and NVDF267 (Supplementary Table. 1). PCR products were gel purified, cloned in vector pBlueScript SKII+ and sequenced using universal primers M13F and M13R. For line Crc-IEE2, we observed that the monocistronic transcripts of gfp had three different 5′-ends. This 5′-ends corresponded to positions -286, -59 and -56 with respect to the start codon of the gfp coding region (Fig. 7a). From the sequencing results, those with the ends at -59 and -56 were more abundant than the transcript with the end at position -286 in a ratio of 3:1. The 5′-end located at position -59 has been previously reported by Loizeau et al. (2014). Whereas the 5′-end at positions -56 and -286 had not been reported before. This should be taken with caution though, as we did not detect an additional, clearly defined, band for the transcript with the 5′-termini at position -286 in the northern blot and this could be an intermediate species in the processing and maturation of the mRNA. Interestingly, in the three transcript ends we mapped we could observe the intact target sequence of Mbb1 (Loizeau et al. 2014), a nuclear encoded protein, member of the TPR family (Fig. 7a, underlined). The target sequence of Mbb1 contains an S-box sequence (Fig. 7a, in bold) which has been found to be determinant in the processing and stabilization of the monocistronic forms of psbB and psbH in wild-type C. reinhardtii (Vaistij et al. 2000a; Loizeau et al. 2014). As the gfp transcripts identified in line Crc-IEE2 include the target binding sequence of Mbb1, it is reasonable to believe that this TPR protein could bind and stabilize the gfp transcripts protecting them from exonucleases (Prikryl et al. 2011; Hammani et al. 2012). For line Crc-IEE5, we determined the 5´-end of one monocistronic form of gfp (Fig. 7b), most likely the most abundant form of a ~ 900 band detected in the northern blot (Fig. 4c). This 5′-end was located at position -133 with respect to the start codon in the gfp coding region. In wild-type cells, processing of the bicistronic tscA-chlN, yields at least three different RNA species (Hahn et al. 1998). It is interesting to note that in the 5′end of this transcript, closely related sequences AAGUAAg/AAGUAAu, similar to the S-box sequence AAGUAAA at the core of the Mbb1 footprint are present (Fig. 7b, in bold). However, their potential role, if any, as the target site of RNA binding proteins remains to be determined. It has been suggested that PPR7 might be involved in the processing and stabilization of the tscA-chlN transcript (Jalal et al. 2015), however not much is known on the stabilization on the 5´-end of the monocistronic form of chlN.
Mapping of the 5′-end termini of gfp monocistrons in transplastomic lines Crc-IEE2 and Crc-IEE5. 5′-end termini were determined by sequencing of the circularized gfp cistrons generated after cleavage and processing of the intercistronic regions IEE2 and IEE5 derived from the chloroplast operons psbN-psbH and tscA-chlN, respectively. a The three 5′-end termini mapped for IEE2. The target sequence of the nuclear encoded, member of the TPR (Tetratricopeptide repeat) family, Mbb1 protein is underline while the S-box is in bold. b The only 5′-end termini mapped for IEE5. Closely related sequences to the S-box are shown in bold. In both cases, the start codon of GFP is highlighted in bold uppercase
Discussion
In this study, we have used the intercistronic regions psbB-psbT, psbN-psbH, psaC-petL, petL-trnN, tscA-chlN from the C. reinhardtii chloroplast genome and the T7g10 5′-UTR (corresponding to the 5′UTR sequence from gene 10 of bacteriophage T7) to construct synthetic bicistronic operons with the foreign genes aphA-6 and gfp. We used these constructs to determine if such intercistronic regions could serve as IEEs for the coexpression of foreign genes in the chloroplast of C. reinhardtii. The transplastomic lines obtained, all showed a kanamycin resistant phenotype, independently of the IEE used, an indirect indication that the APH(3′)-VI was accumulating. However, only the lines that harbored the synthetic operon with IEE2 and IEE5 were able to generate a translatable transcript of gfp and accumulate a detectable amount of GFP. These elements, IEE2 and IEE5, come from psbN-psbH and tscA-chlN, respectively. One of these regions, psbN-psbH was in fact the source of the first IEE proposed and developed for tobacco chloroplast (Zhou et al. 2007) over a decade ago. The tobacco chloroplast IEE has since then been used for metabolic engineering (Lu et al. 2013; Bock 2014; Fuentes et al. 2016) but this is the first time that IEEs have been characterized for C. reinhardtii.
In all lines obtained we observed the accumulation of a bicistronic transcript containing the aphA-6 and gfp genes (Fig. 4b, c). All chloroplast intercistronic regions were cleaved to generate monocistronic mRNA units of aphA-6 and gfp, however, in line IEE1 we could not detect the gfp monocistronic transcript by northern blotting, strongly suggesting that the transcript is highly unstable and rapidly degraded. In fact, in wild-type cells, the psbB-psbT bicistron yields a monocistronic form of psbB but not of psbT. Because the ultimate aim of using IEEs is to obtain expression of the two genes (or more), cleavage and processing of the bicistronic transcript is not enough to qualify as a useful element for the construction of synthetic operons. In this regard IEE3, IEE4 and IEE6 are not useful for such purpose. As mentioned above, recently, it has been reported that transcription in the chloroplast of C. reinhardtii generates polycistronic units more abundantly than what was previously thought (Cavaiuolo et al. 2017; Gallaher et al. 2017). We find this interesting, as a few more intercistronic regions could still be characterized in future studies and IEEs added to the two we have identified here. Alternatively, data obtained from sRNA-Seq could serve as the source to identify sRNA footprints, the result of RNA-binding proteins (e.g., PPR, TPR and OPR proteins), that could serve in turn to develop synthetic construct with cis-elements and trans-acting factors that can be incorporated to the 5′-end of transcripts to stabilize and enhance their translation. A similar approach has been successfully used in tobacco chloroplast where the target sequences of various PPR proteins been incorporated in synthetic bicistronic constructs facilitating the expression of the second cistron (Legen et al. 2018).
When we determined the level of GFP accumulation, this was on the limit of what is estimated to be a good level of protein accumulation (> 1%) for a biotechnology application (Rasala et al. 2010). However, a high level of protein accumulation is not always required. For applications that involve the introduction of 2–3 enzymes, involved all in a metabolic pathway, transcript stability and translatability seems to be more important (Fuentes et al. 2016). In one of the IEEs we report here, psbN-psbH, we found that a binding sequence for the nuclear encoded Mbb1 TPR protein is present in the mature translatable form of the gfp transcript, suggesting that this transcript is also being stabilized as it occurs with the psbH transcript in wild-type cell. An exciting area of research will be the characterization of new factors that generate and stabilize transcripts (monocistronic or polycistronic) and how this knowledge could then help to fine tune the expression of foreign genes for metabolic engineering and synthetic biology in C. reinhardtii.
Before this report, there were no studies on the identification and use of IEE for the stacking and expression of foreign genes in the chloroplast of C. reinhardtii. However, there has been some progress made in this regard in the chloroplast of tobacco. In a pioneering work Zhou et al. (2007) studied the psbB operon from tobacco chloroplast, and identified a small intercistronic element, called the IEE, that when included in a synthetic construct to drive the expression of nptII and yfp, proved to be sufficient to mediate processing of polycistronic transcripts into stable and translatable monocistronic mRNAs. Since then, this feature has been used to express polycistronic mRNA of transgenes from a single promoter in a single transformation event (Lu et al. 2013; Bock 2014; Fuentes et al. 2016). For example, the tobacco chloroplast IEE was used for the construction of a synthetic operon for the expression of three key genes (coding for the key enzymes homogentisate phytyltransferase, tocopherol cyclase and c-tocopherol methyltransferase) involved in the biosynthesis of vitamin E (tocochromanol). Expression of the genes containing the IEE resulted in a tenfold increase in the accumulation of tocochromanol, in transplastomic tobacco and tomato plants, compared to the levels obtained when the genes were expressed without the IEE (Lu et al. 2013). These works have shown that IEEs can serve to facilitate and improve the efficient translation of transcripts from synthetic operons in the plastid genome and with this contribute to the development of metabolic engineering at a more accelerated pace.
The IEEs we have identified could be used to construct bi, tri or polycistronic operons for C. reinhardtii chloroplast. These could contain genes that comprise an entire metabolic pathway not present in C. reinhardtii chloroplast, or genes that complement one that is already present but lack the enzymes to divert a certain intermediate for the production of high value molecules such as terpenoids, vitamins or fatty acids.
References
Almaraz-Delgado AL, Flores-Uribe J, Pérez-España VH, Salgado-Manjarrez E, Badillo-Corona JA (2014) Production of therapeutic proteins in the chloroplast of Chlamydomonas reinhardtii. AMB Express 4:1–9. https://doi.org/10.1186/s13568-014-0057-4
Auchincloss AH, Zerges W, Perron K, Girard-Bascou J, Rochaix JD (2002) Characterization of Tbc2, a nucleus-encoded factor specifically required for translation of the chloroplast psbC mRNA in Chlamydomonas reinhardtii. J Cell Biol 157:953–962. https://doi.org/10.1083/jcb.200201060
Barkan A (1988) Proteins encoded by a complex chloroplast transcription unit are each translated from both monocistronic and polycistronic mRNAs. EMBO J 7:2637–2644
Bateman JM, Purton S (2000) Tools for chloroplast transformation in Chlamydomonas: expression vectors and a new dominant selectable marker. Mol Gen Genet 263:404–410
Bertalan I, Munder MC, Weiß C, Kopf J, Fischer D, Johanningmeier U (2015) A rapid, modular and marker-free chloroplast expression system for the green alga Chlamydomonas reinhardtii. J Biotechnol 195:60–66. https://doi.org/10.1016/j.jbiotec.2014.12.017
Bock R (2013) Strategies for metabolic pathway engineering with multiple transgenes. Plant Mol Biol 83:21–31. https://doi.org/10.1007/s11103-013-0045-0
Bock R (2014) Genetic engineering of the chloroplast: novel tools and new applications. Curr Opin Biotechnol 26:7–13. https://doi.org/10.1016/j.copbio.2013.06.004
Boudreau E, Takahashi Y, Lemieux C, Turmel M, Rochaix JD (1997) The chloroplast ycf3 and ycf4 open reading frames of Chlamydomonas reinhardtii are required for the accumulation of the photosystem I complex. EMBO J 16:6095–6104. https://doi.org/10.1093/emboj/16.20.6095
Cao M, Fu Y, Guo Y, Pan J (2009) Chlamydomonas (Chlorophyceae) colony PCR. Protoplasma 235:107–110. https://doi.org/10.1007/s00709-009-0036-9
Cavaiuolo M, Kuras R, Wollman FA, Choquet Y, Vallon O (2017) Small RNA profiling in Chlamydomonas: insights into chloroplast RNA metabolism. Nucleic Acids Res 45:10783–10799. https://doi.org/10.1093/nar/gkx668
Choquet Y, Stern DB, Wostrikoff K, Kuras R, Girard-Bascou J, Wollman F-A (1998) Translation of cytochrome f is autoregulated through the 5′ untranslated region of petA mRNA in Chlamydomonas chloroplasts. Proc Natl Acad Sci USA 95:4380–4385. https://doi.org/10.1073/pnas.95.8.4380
Douchi D, Qu Y, Longoni P, Legendre-Lefebvre L, Johnson X, Schmitz-Linneweber C, Goldschmidt-Clermont M (2016) A nucleus-encoded chloroplast phosphoprotein governs expression of the photosystem I subunit PsaC in Chlamydomonas reinhardtii. Plant Cell 28:1182–1199. https://doi.org/10.1105/tpc.15.00725
Drapier D, Suzuki H, Levy H, Rimbault B, Kindle KL, Stern DB, Wollman F a (1998) The chloroplast atpA gene cluster in Chlamydomonas reinhardtii. Functional analysis of a polycistronic transcription unit. Plant Physiol 117:629–641
Dreesen IA, Charpin-El Hamri G, Fussenegger M (2010) Heat-stable oral alga-based vaccine protects mice from Staphylococcus aureus infection. J Biotechnol 145:273–280. https://doi.org/10.1016/j.jbiotec.2009.12.006
Dyo YM, Purton S (2018) The algal chloroplast as a synthetic biology platform for production of therapeutic proteins. Microbiology. https://doi.org/10.1099/mic.0.000599
Eberhard S, Loiselay C, Drapier D, Bujaldon S, Girard-Bascou J, Kuras R, Choquet Y, Wollman FA (2011) Dual functions of the nucleus-encoded factor TDA1 in trapping and translation activation of atpA transcripts in Chlamydomonas reinhardtii chloroplasts. Plant J 67:1055–1066. https://doi.org/10.1111/j.1365-313X.2011.04657.x
Felder S, Meierhoff K, Sane AP, Meurer J, Driemel C, Plücken H, Klaff P, Stein B, Bechtold N, Westhoff P, Molekularbiologie E (2001) The Nucleus-Encoded HCF107 gene of Arabidopsis provides a link between intercistronic RNA processing and the accumulation of translation-competent psbH transcripts in chloroplasts. Plant Cell 13:2127–2141
Fuentes P, Zhou F, Erban A, Karcher D, Kopka J, Bock R (2016) A new synthetic biology approach allows transfer of an entire metabolic pathway from a medicinal plant to a biomass crop. Elife 5:1–26. https://doi.org/10.7554/eLife.13664
Fuentes P, Armarego-Marriott T, Bock R (2018) Plastid transformation and its application in metabolic engineering. Curr Opin Biotechnol 49:10–15. https://doi.org/10.1016/j.copbio.2017.07.004
Fukusaki E-I, Nishikawa T, Kato K, Shinmyo A, Hemmi H, Nishino T, Kobayashi A (2003) Introduction of the archaebacterial geranylgeranyl pyrophosphate synthase gene into Chlamydomonas reinhardtii chloroplast. J Biosci Bioeng 95:283–287. https://doi.org/10.1016/S1389-1723(03)80030-0
Gallaher SD, Fitz-Gibbon ST, Strenkert D, Purvine SO, Pellegrini M, Merchant SS (2017) High-throughput sequencing of the chloroplast and mitochondrion of Chlamydomonas reinhardtii to generate improved de novo assemblies, analyze expression patterns and transcript speciation, and evaluate diversity among laboratory strains a. Plant J 1–21. https://doi.org/10.1111/tpj.13788
Gangl D, Zedler JAZ, Rajakumar PD, Martinez EMR, Riseley A, Włodarczyk A, Purton S, Sakuragi Y, Howe CJ, Jensen PE, Robinson C (2015) Biotechnological exploitation of microalgae. J Exp Bot 66:6975–6990. https://doi.org/10.1093/jxb/erv426
Gimpel JA, Henríquez V, Mayfield SP (2015) In metabolic engineering of eukaryotic microalgae: Potential and challenges come with great diversity. Front Microbiol 6:1–14. https://doi.org/10.3389/fmicb.2015.01376
Glanz S, Jacobs J, Kock V, Mishra A, Kück U (2012) Raa4 is a trans-splicing factor that specifically binds chloroplast tscA intron RNA. Plant J 69:421–431. https://doi.org/10.1111/j.1365-313X.2011.04801.x
Gorman DS, Levine RP (1965) Cytochrome f and plastocyanin: their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 54:1665–1669. https://doi.org/10.1073/pnas.54.6.1665
Guzmán-Zapata D, Macedo-Osorio K, Almaraz-Delgado AL, Duran-Figueroa N, Badillo-Corona JA (2016) Production of recombinant proteins in the chloroplast of the green alga Chlamydomonas reinhardtii. In: MacDonald J, Kolotilin I, Menassa R (eds) Recombinant proteins from plants; methods and protocols, 2nd edn. Humana Press, New York, pp 69–87
Hahn D, Nickelsen J, Hackert A, Kück U (1998) A single nuclear locus is involved in both chloroplast RNA trans-splicing and 3´ end processing. Plant J 15:575–581
Hammani K, Cook WB, Barkan A (2012) RNA binding and RNA remodeling activities of the half-a-tetratricopeptide (HAT) protein HCF107 underlie its effects on gene expression. Proc Natl Acad Sci USA 109:5651–5656. https://doi.org/10.1073/pnas.1200318109
Harada H, Maoka T, Osawa A, Hattan JI, Kanamoto H, Shindo K, Otomatsu T, Misawa N (2014) Construction of transplastomic lettuce (Lactuca sativa) dominantly producing astaxanthin fatty acid esters and detailed chemical analysis of generated carotenoids. Transgenic Res 23:303–315. https://doi.org/10.1007/s11248-013-9750-3
Harris EH (2009) The Chlamydomonas sourcebook: introduction to Chlamydomonas and its laboratory uses, 2nd edn. Academic Press, San Diego
Herrin DL, Schmidt GW (1988) trans-splicing of transcripts for the chloroplast psaA1 gene. In vivo requirement for nuclear gene products. J Biol Chem 263:14601–14604
Jalal A, Schwarz C, Schmitz-linneweber C, Vallon O, Bohne A (2015) A small multifunctional pentatricopeptide repeat protein in the chloroplast of Chlamydomonas reinhardtii. Mol Plant. https://doi.org/10.1016/j.molp.2014.11.019
Johnson CH, Schmidt GW (1993) The psbB gene cluster of the Chlamydomonas reinhardtii chloroplast: sequence and transcriptional analyses of psbN and psbH. Plant Mol Biol 22:645–658
Johnson X, Wostrikoff K, Finazzi G, Kuras R, Schwarz C, Bujaldon S, Nickelsen J, Stern DB, Wollman F-A, Vallon O (2010) MRL1, a conserved pentatricopeptide repeat protein, is required for stabilization of rbcL mRNA in Chlamydomonas and Arabidopsis. Plant Cell 22:234–248. https://doi.org/10.1105/tpc.109.066266
Jones CS, Luong T, Hannon M, Tran M, Gregory JA, Shen Z, Briggs SP, Mayfield SP (2013) Heterologous expression of the C-terminal antigenic domain of the malaria vaccine candidate Pfs48/45 in the green algae Chlamydomonas reinhardtii. Appl Microbiol Biotechnol 97:1987–1995. https://doi.org/10.1007/s00253-012-4071-7
Kang S, Odom OW, Thangamani S, Herrin DL (2017) Toward mosquito control with a green alga: expression of Cry toxins of Bacillus thuringiensis subsp. israelensis (Bti) in the chloroplast of Chlamydomonas. J Appl Phycol 29:1377–1389. https://doi.org/10.1007/s10811-016-1008-z
Legen J, Ruf S, Kroop X, Wang G, Barkan A, Bock R, Schmitz-linneweber C (2018) Stabilization and translation of synthetic operon-derived mRNAs in chloroplasts by sequences representing PPR protein binding sites. Plant J. https://doi.org/10.1111/tpj.13863
Loiselay C, Gumpel NJ, Girard-bascou J, Watson AT, Purton S, Choquet Y (2008) Molecular identification and function of cis- and trans-acting determinants for petA transcript stability in Chlamydomonas reinhardtii chloroplasts. Mol Cell Biol 28:5529–5542. https://doi.org/10.1128/MCB.02056-07
Loizeau K, Qu Y, Depp S, Fiechter V, Ruwe H, Lefebvre-Legendre L, Schmitz-Linneweber C, Goldschmidt-Clermont M (2014) Small RNAs reveal two target sites of the RNA-maturation factor Mbb1 in the chloroplast of Chlamydomonas. Nucleic Acids Res 42:3286–3297. https://doi.org/10.1093/nar/gkt1272
Lu Y, Rijzaani H, Karcher D, Ruf S, Bock R (2013) Efficient metabolic pathway engineering in transgenic tobacco and tomato plastids with synthetic multigene operons. Proc Natl Acad Sci USA 110:E623–E632. https://doi.org/10.1073/pnas.1216898110
Luro S, Germain A, Sharwood RE, Stern DB (2013) RNase J participates in a pentatricopeptide repeat protein-mediated 5′ end maturation of chloroplast mRNAs. Nucleic Acids Res 41:9141–9151. https://doi.org/10.1093/nar/gkt640
Mayfield SP, Franklin SE, Lerner RA (2003) Expression and assembly of a fully active antibody in algae. Proc Natl Acad Sci USA 100:438–442. https://doi.org/10.1073/pnas.0237108100
Michelet L, Lefebvre-Legendre L, Burr SE, Rochaix J-D, Goldschmidt-Clermont M (2011) Enhanced chloroplast transgene expression in a nuclear mutant of Chlamydomonas. Plant Biotechnol J 9:565–574. https://doi.org/10.1111/j.1467-7652.2010.00564.x
Monde R, Greene JC, Stern DB (2000) Disruption of the petB-petD intergenic region in tobacco chloroplasts affects petD RNA accumulation and translation. Mol Gen Genet 263:610–618
Monod C, Goldschmidt-Clermont M, Rochaix J-D (1992) Accumulation of chloroplast psbB RNA requires a nuclear factor in Chlamydomonas reinhardtii. Mol Gen Genet 449–459
Mor TS, Ohad I, Hirschberg J, Pakrasi HB (1995) An unusual organization of the genes encoding cytochrome b559 in Chlamydomonas reinhardtii: psbE and psbF genes are separately transcribed from different regions of the plastid chromosome. MGG Mol Gen Genet 246:600–604. https://doi.org/10.1007/BF00298966
Newman SM, Boynton JE, Gillham NW, Randolph-Anderson BL, Johnson AM, Harris EH (1990) Transformation of chloroplast ribosomal RNA genes in Chlamydomonas: molecular and genetic characterization of integration events. Genetics 126:875–888
Oey M, Lohse M, Kreikemeyer B, Bock R (2009) Exhaustion of the chloroplast protein synthesis capacity by massive expression of a highly stable protein antibiotic. Plant J 57:436–445. https://doi.org/10.1111/j.1365-313X.2008.03702.x
Olins PO, Rangwala SH (1989) A novel sequence element derived from bacteriophage T7 mRNA acts as an enhancer of translation of the lacZ gene in Escherichia coli. J Biol Chem 264:16973–16976
Olins PO, Devine CS, Rangwala SH, Kavka KS (1988) The T7 phage gene 10 leader RNA, a ribosomal binding site that dramatically enhances the expression of foriegn genes in Escherichia coli. Gene 73:227–235
Pfalz J, Bayraktar OA, Prikryl J, Barkan A (2009) Site-specific binding of a PPR protein defines and stabilizes 5′ and 3′ mRNA termini in chloroplasts. EMBO J 28:2042–2052. https://doi.org/10.1038/emboj.2009.121
Prikryl J, Rojas M, Schuster G, Barkan A (2011) Mechanism of RNA stabilization and translational activation by a pentatricopeptide repeat protein. Proc Natl Acad Sci USA 108:415–420. https://doi.org/10.1073/pnas.1012076108
Rahire M, Laroche F, Cerutti L, Rochaix JD (2012) Identification of an OPR protein involved in the translation initiation of the PsaB subunit of photosystem I. Plant J 72:652–661. https://doi.org/10.1111/j.1365-313X.2012.05111.x
Rasala BA, Muto M, Lee PA, Jager M, Cardoso RMF, Behnke A, Kirk P, Hokanson CA, Crea R, Mendez M, Mayfield SP (2010) Production of therapeutic proteins in algae, analysis of expression of seven human proteins in the chloroplast of Chlamydomonas reinhardtii. Plant Biotechnol J 8:719–733. https://doi.org/10.1111/j.1467-7652.2010.00503
Rasala BA, Muto M, Sullivan J, Mayfield SP (2011) Improved heterologous protein expression in the chloroplast of Chlamydomonas reinhardtii through promoter and 5′ untranslated region optimization. Plant Biotechnol J 9:674–683. https://doi.org/10.1111/j.1467-7652.2011.00620.x
Reed KC, Mann DA (1985) Rapid transfer of DNA from agarose gels to nylon membranes. Nucleic Acids Res 13:7207–7221. https://doi.org/10.1093/nar/13.20.7207
Robertson D, Boynton JE, Gillham NW (1990) Cotranscription of the wild-type chloroplast atpE gene encoding the CF1/CF0 epsilon subunit with the 3′ half of the rps7 gene in Chlamydomonas reinhardtii and characterization of frameshift mutations in atpE. MGG Mol Gen Genet 221:155–163. https://doi.org/10.1007/BF00261715
Rochaix J-D (1996) Post-transcriptional regulation of chloroplast gene expression in Chlamydomonas reinhardtii. Plant Mol Biol 32:327–341. https://doi.org/10.1007/BF00039389
Sakamoto W, Sturm NR, Kindle KL, Stern DB (1994) petD mRNA maturation in Chlamydomonas reinhardtii chloroplasts: role of 5′ endonucleolytic processing. Mol Cell Biol 14:6180–6186. https://doi.org/10.1128/MCB.14.9.6180
Salvador ML, Klein UWE, Bogoradt L (1993) 5′ Sequences are important positive and negative determinants of the longevity of Chlamydomonas chloroplast gene transcripts. Proc Natl Acad Sci USA 90:1556–1560
Salvador ML, Suay L, Anthonisen IL, Klein U (2004) Changes in the 5′-untranslated region of the rbcL gene accelerate transcript degradation more than 50-fold in the chloroplast of Chlamydomonas reinhardtii. Curr Genet 45:176–182. https://doi.org/10.1007/s00294-003-0470-8
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Scaife MA, Nguyen GTDT, Rico J, Lambert D, Helliwell KE, Smith AG (2015) Establishing Chlamydomonas reinhardtii as an industrial biotechnology host. Plant J 82:532–546. https://doi.org/10.1111/tpj.12781
Scranton MA, Ostrand JT, Fields FJ, Mayfield SP (2015) Chlamydomonas as a model for biofuels and bio-products production. Plant J 82:523–531. https://doi.org/10.1111/tpj.12780
Stern DB, Goldschmidt-Clermont M, Hanson MR (2010) Chloroplast RNA metabolism. Annu Rev Plant Biol 61:125–155. https://doi.org/10.1146/annurev-arplant-042809-112242
Stoppel R, Meurer J (2013) Complex RNA metabolism in the chloroplast: an update on the psbB operon. Planta 237:441–449. https://doi.org/10.1007/s00425-012-1782-z
Sturm NR, Kuras R, Buschlen S, Sakamoto W, Kindle KL, Stern DB, Wollman F(1994) The petD gene is transcribed by functionally redundant promoters in Chlamydomonas reinhardtii chloroplasts. 14:6171–6179
Takahashi Y, Soen S, Franzen LG, Rochaix J (1991) Directed chloroplast transformation in Chamydomonas reinhardtii: insertional inactivation of the psaC gene encoding the iron sulfur protein destabilizes photosystem I. EMBO J 10:2033–2040
Takahashi Y, Rahire M, Breyton C, Popot JL, Joliot P, Rochaix JD (1996) The chloroplast ycf7 (petL) open reading frame of Chlamydomonas reinhardtii encodes a small functionally important subunit of the cytochrome b6f complex. Embo J 15:3498–3506
Tran M, Zhou B, Pettersson PL, Gonzalez MJ, Mayfield SP (2009) Synthesis and assembly of a full-length human monoclonal antibody in algal chloroplasts. Biotechnol Bioeng 104:663–673. https://doi.org/10.1002/bit.22446
Tran M, Henry R, Siefker DT, Van C, Newkirk G, Kim J, Jack B, Mayfield SP (2013a) Production of anti-cancer immunotoxins in algae: ribosome inactivating proteins fusion partners. Biotechnol Bioeng 110:2984–2993. https://doi.org/10.1002/bit.24966
Tran M, Van C, Barrera DJ, Pettersson PL, Peinado CD, Bui J, Mayfield SP (2013b) Production of unique immunotoxin cancer therapeutics in algal chloroplasts. Proc Natl Acad Sci USA 110:E15–E22. https://doi.org/10.1073/pnas.1214638110
Trösch R, Barahimipour R, Gao Y, Badillo-Corona JA, Gotsmann VL, Zimmer D, Mühlhaus T, Zoschke R, Willmund F (2018) Commonalities and differences of chloroplast translation in a green alga and land plants. Nat Plants. https://doi.org/10.1038/s41477-018-0211-0
Vaistij FE, Boudreau E, Lemaire SD, Goldschmidt-Clermont M, Rochaix JD (2000a) Characterization of Mbb1, a nucleus-encoded tetratricopeptide-like repeat protein required for expression of the chloroplast psbB/psbT/psbH gene cluster in Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 97:14813–14818. https://doi.org/10.1073/pnas.97.26.14813
Vaistij FE, Goldschmidt-Clermont M, Wostrikoff K, Rochaix JD (2000b) Stability determinants in the chloroplast psbB/T/H mRNAs of Chlamydomonas reinhardtii. Plant J 21:469–482. https://doi.org/10.1046/j.1365-313X.2000.00700.x
Wang X, Brandsma M, Tremblay R, Maxwell D, Jevnikar AM, Huner N, Ma S (2008) A novel expression platform for the production of diabetes-associated autoantigen human glutamic acid decarboxylase (hGAD65). BMC Biotechnol 8:1–13. https://doi.org/10.1186/1472-6750-8-87
Wang F, Johnson X, Cavaiuolo M, Bohne AV, Nickelsen J, Vallon O (2015) Two Chlamydomonas OPR proteins stabilize chloroplast mRNAs encoding small subunits of photosystem II and cytochrome b6f. Plant J 82:861–873. https://doi.org/10.1111/tpj.12858
Wannathong T, Waterhouse JC, Young REB, Economou CK, Purton S (2016) New tools for chloroplast genetic engineering allow the synthesis of human growth hormone in the green alga Chlamydomonas reinhardtii. Appl Microbiol Biotechnol 100:5467–5477. https://doi.org/10.1007/s00253-016-7354-6
Wu S, Xu L, Huang R, Wang Q (2011) Improved biohydrogen production with an expression of codon-optimized hemH and lba genes in the chloroplast of Chlamydomonas reinhardtii. Bioresour Technol 102:2610–2616. https://doi.org/10.1016/j.biortech.2010.09.123
Zandueta-Criado A, Bock R (2004) Surprising features of plastid ndhD transcripts: addition of non-encoded nucleotides and polysome association of mRNAs with an unedited start codon. Nucleic Acids Res 32:542–550. https://doi.org/10.1093/nar/gkh217
Zedler JAZ, Gangl D, Hamberger B, Purton S, Robinson C (2015) Stable expression of a bifunctional diterpene synthase in the chloroplast of Chlamydomonas reinhardtii. J Appl Phycol 27:2271–2277. https://doi.org/10.1007/s10811-014-0504-2
Zhou F, Karcher D, Bock R (2007) Identification of a plastid intercistronic expression element (IEE) facilitating the expression of stable translatable monocistronic mRNAs from operons. Plant J 52:961–972. https://doi.org/10.1111/j.1365-313X.2007.03261.x
Zhou F, Badillo-Corona JA, Karcher D, Gonzalez-Rabade N, Piepenburg K, Borchers AMI, Maloney AP, Kavanagh TA, Gray JC, Bock R (2008) High-level expression of human immunodeficiency virus antigens from the tobacco and tomato plastid genomes. Plant Biotechnol J 6:897–913. https://doi.org/10.1111/j.1467-7652.2008.00356.x
Zoschke R, Barkan A (2015) Genome-wide analysis of thylakoid-bound ribosomes in maize reveals principles of cotranslational targeting to the thylakoid membrane. Proc Natl Acad Sci USA 112:E1678–E1687. https://doi.org/10.1073/pnas.1424655112
Acknowledgement
Research in the laboratory of JABC is supported by Instituto Politécnico Nacional through Secretaría de Investigación y Posgrado (SIP-IPN) with the grant 20170513. A grant to JABC from Secretaría de Ciencia, Tecnología e Inovación (SECITI-CDMX) contributed in part for the development of this project. KMSO and DGZ were financially supported by CONACYT and IPN during their PhD studies. We acknowledge Centro de Nanociencias y Micro y Nanotecnologías-IPN for providing access to the confocal microscope and the technical expertise of Dr. María de Jesús Perea Flores.
Author information
Authors and Affiliations
Contributions
KSMO, VHPE, CGO and JABC conceived and designed experiments. KSMO performed all experiments. KSMO, NVDF, DGZ and JABC analyzed data. JABC and KSMO wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Data 1. Sequences of gfp and the intercistronic regions used in this study (PDF 56 KB)
11103_2018_776_MOESM2_ESM.pdf
Supplementary Figure 1. Selection of transformed lines in spectinomycin and kanamycin. Colonies from lines Crc-IEE1, *Crc-IEE2, Crc-IEE3, Crc-IEE4, Crc-IEE5, Crc-IEE6 recovered after transformation with vectors p320-IEE1/p228, * p320-IEE2/p228, p320-IEE3/p228, p320-IEE4/p228, p320-IEE5/p228, p320-IEE6/p228, respectively. Colonies were grown in TAP media supplemented with spectinomycin (150 µg/mL), to determine successful transformation with vector p228 vector, and in TAP media supplemented with kanamycin (100 µg/mL, Km), to determine transformation with vectors of the series p320IEE. (PDF 306 KB)
Rights and permissions
About this article
Cite this article
Macedo-Osorio, K.S., Pérez-España, V.H., Garibay-Orijel, C. et al. Intercistronic expression elements (IEE) from the chloroplast of Chlamydomonas reinhardtii can be used for the expression of foreign genes in synthetic operons. Plant Mol Biol 98, 303–317 (2018). https://doi.org/10.1007/s11103-018-0776-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-018-0776-z