Abstract
Mitochondrial Complex I [NADH Coenzyme Q (CoQ) oxidoreductase] is the least understood of respiratory complexes. In this review we emphasize some novel findings on this enzyme that are of relevance to the pathogenesis of neurodegenerative diseases. Besides CoQ, also oxygen may be an electron acceptor from the enzyme, with generation of superoxide radical in the mitochondrial matrix. The site of superoxide generation is debated: we present evidence based on the rational use of several inhibitors that the one-electron donor to oxygen is an iron-sulphur cluster, presumably N2. On this assumption we present a novel mechanism of electron transfer to the acceptor, CoQ. Complex I is deeply involved in pathological changes, including neurodegeneration. Complex I changes are involved in common neurological diseases of the adult and old ages. Mitochondrial cytopathies due to mutations of either nuclear or mitochondrial DNA may represent a useful model of neurodegeneration. In this review we discuss Parkinson’s disease, where the pathogenic involvement of Complex I is better understood; the accumulated evidence on the mode of action of Complex I inhibitors and their effect on oxygen radical generation is discussed in terms of the aetiology and pathogenesis of the disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The Mitochondrial Generation of Reactive Oxygen Species
The products of partial reduction of molecular oxygen and their derivatives are cumulatively designated as Reactive Oxygen Species (ROS). There are several reactions in cells that are able to give rise to superoxide anion radical and/or to hydrogen peroxide; the latter can react with a reduced metal ion (such as Fe2+ or Cu+) and generate the most aggressive hydroxyl radical. Within a cell, mitochondria largely contribute to the production of ROS via the respiratory chain [1–4]. The relevance of mitochondrial production of ROS within a cell is indirectly revealed by the results of deficiency of mitochondrial antioxidant enzymes. Mitochondria contain an isozyme of superoxide dismutase (SOD-2) and glutathione peroxidase (GPx). The lack of SOD-2 [5] and of mitochondrial GPx [6] is deleterious to cells.
Although the first product of oxygen reduction by the respiratory chain is superoxide [7, 8], this radical has a short life and is rapidly converted into hydrogen peroxide by mitochondrial SOD or by spontaneous disproportionation, or can attack other molecules, such as lipids, before being able to escape the mitochondrion. Although hydrogen peroxide is also removed by GPx, it is much more stable than superoxide, so that some molecules can escape the organelle and be detected outside [7]. Most of superoxide is generated at the matrix side of the inner membrane, as appears from the observation that superoxide is detected in submitochondrial particles which are inside-out with respect to mitochondria, while, in intact mitochondria, only hydrogen peroxide was detected. A study with suitable spin traps, however, demonstrated the formation of superoxide radical in mitoplasts [9] indicating that a significant aliquot of this species is released at the outer face of the inner membrane [10]. It is likely that Complex I releases ROS in the matrix while Complex III in the intermembrane space. The superoxide anion released at the intermembrane space may be exported to the cytoplasm through an anion channel related to VDAC [11]; the hypothesis that superoxide in the matrix is exported to the intermembrane space via an anion channel in uncoupling proteins (UCPs) has been excluded [12].
It is worth noting that mitochondria from different tissues may vary conspicuously in their capacity to produce ROS using different substrates [13], and this capacity may also be related to animal species and age.
The first site of damage of the ROS produced by mitochondria is the mitochondrion itself: superoxide dismutates to H2O2 and the hydroxyl radical produced by H2O2 in presence of reduced metal ions may damage several biomolecules in the inner membrane and in the matrix [1, 14]. Another important effect of ROS is induction of permeability transition by opening the cyclosporin-sensitive pore (permeability transition pore) in the inner membrane [15].
An important question is whether free radicals produced by mitochondria are also physiologically released to the cytosol. Staniek and Nohl [16] applied a non-invasive detecting system for hydrogen peroxide and found that isolated intact rat heart mitochondria do not produce detectable H2O2, unless when using succinate in presence of antimycin. Korshunov et al. [17] also found no hydrogen peroxide formation by intact rat heart mitochondria, unless pretreated in such a way to deplete them of endogenous antioxidants. It may be inferred that under normal conditions ROS are not exported out of mitochondria. There is however overwhelming evidence that ROS production detected in different cells and acting as signal transducers has a mitochondrial origin [18].
It is not easy, however, to demonstrate that ROS detected in cells are produced by mitochondria; the effect of respiratory inhibitors appears to be the best way to discriminate between mitochondrial and non-mitochondrial ROS. However, the effect of inhibitors is ambiguous. Although antimycin is usually found to stimulate ROS production [19] in intact cells, as it does in mitochondria, the effect of rotenone is contradictory. Some studies showed that rotenone enhances ROS production in intact cells [20–22] whereas others showed inhibition of cellular ROS production by the same inhibitor [23–25]. Since rotenone decreases ROS production by Complex III while enhancing ROS production by Complex I, and the relative contribution of the two Complexes to ROS production may vary in different cells, if ROS production by Complex III is relatively high, rotenone inhibition would decrease total ROS production, whereas, if ROS production by Complex III is low, then enhancement of ROS release by Complex I would prevail and total ROS would be increased. Since ROS production by reverse flux of electrons is decreased by rotenone [26], another critical point is represented by membrane potential and the contribution of reverse electron transfer in Complex I.
Mitochondrial ROS production is enhanced in State 4 and when the rate of electron transfer is lowered [27]. The rationale is in a more reduced state of the respiratory carriers capable of donating electrons to oxygen. To this purpose uncoupling and release of excessive membrane proton potential may protect mitochondria from damage due to excessive free radical production. In rat hepatocytes the futile cycle of proton pumping and proton leak may be responsible for 20–25% of respiration [28]; in perfused rat muscle the value is even greater. Uncoupling may be obtained by activating proton leak through UCPs [8]. In such way a tissue may dissipate a conspicuous part of the energy conserved by its mitochondria, however it keeps the mitochondrial respiratory chain under more oxidized conditions preventing the formation of damaging free radicals. Indeed, superoxide activates proton transport through UCPs [12].
The major sites of superoxide formation in the respiratory chain are within respiratory complexes I and III [29]. Further sites, however, may have importance and physiological relevance, such as Complex II [30], glycerophosphate dehydrogenase [31], dihydroorotate dehydrogenase [32]. Recently, an additional source of ROS in mitochondria (directly in the form of hydrogen peroxide) has been demonstrated in the p66Shc protein: a fraction of p66Shc has a mitochondrial localization in the intermembrane space and has been demonstrated to directly produce hydrogen peroxide by accepting electrons from reduced cytochrome c [33]
In this review we emphasize some novel findings on mitochondrial Complex I [NADH Coenzyme Q (CoQ) oxidoreductase], that are of relevance to the pathogenesis of neurodegenerative diseases. In particular we will consider the mechanism by which Complex I reduces oxygen to superoxide radical.
Structure and Function of Complex I: General Aspects
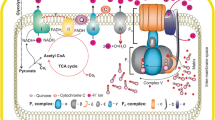
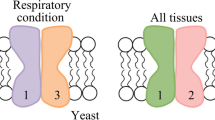
Most organisms possess Complex I, a very large enzyme catalyzing the first step of the mitochondrial electron transport chain [34]. The enzyme oxidizes NADH in the mitochondrial matrix and reduces Ubiquinone (CoQ), a lipid soluble electron carrier embedded in the lipid bilayer of the inner mitochondrial membrane. The total number of subunits in the bovine enzyme is 46 [35] or 45 [36] for a molecular mass of about 1,000 kDa. Seven subunits are the products of the mitochondrial genome [37, 38] and correspond to hydrophobic components named ND1-ND6 and ND4L. The minimal active form of the enzyme is that found in bacteria, composed of 14 subunits, all of which are homologous to their mitochondrial counterparts, while all other “accessory” subunits still have an undefined role. From structural and phylogenetic considerations, the enzyme is envisaged to consist of three different sectors: a dehydrogenase unit and a hydrogenase-like unit, constituting the peripheral arm protruding into the matrix, and a transporter unit deeply embedded in the membrane and involved in proton translocation [39–41] (Fig. 1).
Schematic representation of Complex I formed by the apposition of three different modules: a dehydrogenase module, where NADH is oxidized, containing FMN and iron sulphur clusters N1a, N1b, N3, N4 and N5; a hydrogenase module where CoQ is reduced, containing iron-sulphur clusters N6a, N6b and N2; and a transporter module containing no prosthetic groups and involved in proton translocation
Several prosthetic groups contribute to electron transfer within the enzyme: FMN is the entry point for electrons that are then transferred to a series of iron-sulphur clusters [42]. Enzymes from different sources have different numbers of iron-sulphur clusters, most of which share the same midpoint potential. Two clusters present different characteristics: N1a, that is of the kind Fe2S2, has the lowest midpoint potential (E m = –370 mV), while N2, that is of the kind Fe4S4 and resides at the interface between the PSST and the 49 kDa subunits [43], has the highest midpoint potential (E m between −150 mV and −50 mV), presenting EPR magnetic interactions with the ubisemiquinone radicals; for these reasons it is considered to be the direct electron donor to ubiquinone [44]. N2 iron-sulphur cluster is most likely located in the connection between the peripheral and the membrane arm. The magnetic interaction with the semiquinone radical, corresponding to a distance of about 10 Å [45, 46], suggests that the ubiquinone headgroup could somehow reach up into the peripheral arm as recently assumed by Brandt et al. [47], who have hypothesized an amphipathic “ramp” guiding ubiquinone into the catalytic site. Recently the arrangement of iron-sulphur clusters in the hydrophilic domain of Complex I from T. thermophilus has been determined by x-ray crystallography, showing a linear chain of all clusters except N1a and N7 [48, 49].
Complex I is inhibited by more than 60 different families of compounds [50] starting from rotenone, the prototype of this series, to a number of synthetic insecticides/acaricides. These inhibitors were grouped into three classes based on their effects on the kinetic behaviour of the enzyme, having as prototypes piericidin A, rotenone, and capsaicin, respectively. Nevertheless kinetic studies did not allow to assign different binding sites for these three classes of inhibitors: it is commonly accepted that they share the same hydrophobic large pocket in the enzyme [51].
Complex I is also involved in the formation of the trans-membrane proton gradient with a stoichiometry of 4H+/2e−. The limited knowledge about the mechanism of electron transfer of Complex I makes it difficult to predict the mechanism by which this respiratory chain complex uses redox energy to translocate protons across the inner mitochondrial membrane (for reviews see [52–54]).
Besides its well known redox role in the electron transport chain, Complex I is considered one of the main sites of production of ROS: electron leaks at Complex I can release single electrons to oxygen and give rise to superoxide anion. The mechanism of superoxide production by Complex I is not clear, probably for lack of knowledge of the exact sequence of the electron carriers and how electron transfer is coupled to proton translocation. The major sites of ROS production in the mitochondrial electron transport chain have been localized in Complex I and Complex III [3, 29, 55]; while the site of electron escape in Complex III has been identified in the so-called centre “o”, the direct oxygen reductant site in Complex I is not yet known with certainty.
The notion of Complex I as an individual enzyme stems out of its isolation as a discrete lipoprotein unit by detergent fractionation [56]. Recent structural and kinetic evidence, however, strongly suggests that Complexes I and III form stable functional supercomplexes [57–59].
ROS Production by Complex I
Complex I is generally considered as the major enzyme contributing to generation of ROS in mitochondria [4]; the site of univalent oxygen reduction in Complex I is still controversial and the reason is in part in the scant knowledge of the mechanism of electron transfer within the enzyme prosthetic groups. The physiological relevance of ROS generation by Complex I as well as by different mitochondrial sites is still uncertain and is even questioned by some investigators [60].
Early experiments proved the involvement of Complex I in ROS production [61]; addition of either NADH at low concentration or NADPH, which feeds the electrons at decreased rate into the Complex, led to copious ROS production detected by lipid peroxidation; addition of NADH at high concentration, but in presence of rotenone, also induced peroxidation. Water-soluble CoQ homologs used as electron acceptors from isolated Complex I stimulated H2O2 production whereas CoQ6 and CoQ10 were inactive [62]. More recent studies confirmed that Complex I is a major source of superoxide production in several types of mitochondria [29, 63] and localized the oxygen reducing site between the ferricyanide and the quinone reduction sites [63, 64].
The superoxide production by Complex I is higher during the reverse electron transport from succinate to NAD+ [17, 65–69], whereas during the forward electron transport it is much lower. Reverse electron transfer-supported ROS production requires high membrane potential and is inhibited by uncouplers and by processes dissipating membrane potential [26, 68–70]. Rotenone has been found to enhance ROS formation during forward electron transfer [63, 64] and to inhibit it during reverse electron transfer [46, 66, 71].
The identification of the oxygen reducing site has been the subject of extensive investigation, and several prosthetic groups in the enzyme have been suggested to be the direct reductants of oxygen.
Ubisemiquinone
On the basis of the significant differences found in the stimulating effects of rotenone, piericidin and myxothiazol on ROS production by Complex I, Brand [66] excluded any site upstream of the quinone/semiquinone couple itself: since all these are inhibitors of the CoQ site, the sites upstream of CoQ should have been affected to the same extent by the different inhibitors. However, using the same reasoning, it is not possible to exclude as the site of superoxide generation the electron donor(s) to CoQ, such as iron-sulphur cluster N2, that share the acceptor pocket with the quinone itself. Ohnishi et al. [46] reached similar conclusion from the differential effects of rotenone and piericidin in both forward and reverse electron transfer, and concluded that cluster N2 and/or ubisemiquinones bound to cluster N2 may be the electron donor(s) to oxygen. From the EPR data reported by the Ohnishi group [45] it appears that Complex I inhibitors such as rotenone and piericidin A turn off the EPR signal from the semiquinone species. Unfortunately there is no available evidence about the effects of the other Complex I inhibitors on the EPR semiquinone signals. From our unpublished results on ROS production it appears that inhibitors known to shut down the semiquinone signal are also most efficient in the direct transfer of electrons to molecular oxygen. These results would suggest that the endogenous semiquinone formed during the redox cycle of the enzyme is not involved in ROS production. This conclusion is also in line with a previous report showing that in CoQ-depleted mitochondria Complex I is able to produce oxygen radicals at a rate comparable with the enzyme in non-extracted mitochondria [64].
Flavin Mononucleotide
A major candidate as the electron donor to oxygen has been proposed to be FMN [71–73]; the rationale for such identification has been that diphenylene iodonium (DPI), an inhibitor of Complex I at the FMN region, blocks reverse electron transfer-supported ROS formation [72]; however, DPI also inhibits NADH-supported ROS formation [29, 72]. On the other hand, Ohnishi and co-workers [46] showed that DPI enhances ROS production in the reverse electron transfer, while inhibiting it in the forward electron transfer. The loss of ROS detection in presence of DPI seems to exclude any involvement of FMN in ROS production to advantage of a direct involvement of iron-sulphur clusters. In fact DPI inhibits the iron-sulphur clusters reduction while the reduced state of protein-bound FMN is stabilized [74]. Indeed the FMN involvement in ROS production still remains an open question and the discrepancies in the literature should be at least in part ascribed to difficulty in achieving complete inhibition of the enzyme: the inhibition of Complex I activity was never more than 80–85%, allowing a residual electron flux to iron-sulphur clusters. Herrero and Barja [63] found that ROS production in forward electron transfer in Complex I was also inhibited by ethoxyformic anhydride, an inhibitor of iron-sulphur clusters, clearly excluding FMN as the site of oxygen reduction. In addition, the studies by Lambert and Brand [66] and by Ohnishi et al. [46] also exclude FMN as the reductant of oxygen, pinpointing a site close to or coincident with the CoQ-binding site (see above).
These findings are in contrast with findings in isolated Complex I [75, 76] where FMN is considered the major electron donor to oxygen to form superoxide anion. Galkin and Brandt [75] showed that ROS production was still present in complex I from a mutant of Yarrowia lipolytica lacking iron-sulphur cluster N2, concluding that FMN is directly involved in this activity. Accurate redox titrations of the electron donor and an EPR study of the different redox centres [76] appeared however to exclude either FMN semiquinone or any FeS cluster as the source of superoxide, suggesting that the fully reduced flavin delivers an electron to oxygen and the other one to the chain of iron-sulphur clusters.
The identification of flavin as the site of oxygen reduction would be incompatible with our finding that two classes of inhibitors both acting downstream of the iron-sulphur clusters in the enzyme have opposite effects, in that rotenone enhances superoxide production whereas stigmatellin inhibits it [29]. A possible explanation is that two sites for oxygen reduction exist in the complex, represented by flavin and an iron-sulphur cluster; the latter site would be predominant in membrane particles whereas the former one might be made better available after Complex I isolation. The role of super-complex organization in shielding/opening different sites in the enzyme cannot be overlooked.
Nevertheless, a major role can be envisioned for FMN in the formation of radical species by Complex I in the presence of physiological hydrophilic quinones (i.e. cathecolamine-derived oxidative products). The mechanism through which adrenochrome was shown to enhance the formation of ROS by Complex I is a multiple-step process involving a site situated upstream in the redox-active chain of the enzyme, likely coincident with a FMN, since the reaction is insensitive to both rotenone and p-hydroxymercuribenzoate [77].
Iron-Sulphur Clusters
Another major candidate as the direct oxygen reductant is the iron-sulphur cluster N2; according to Brandt and colleagues [78, 79] this site is localized at the interface between the matrix site and the membranous part of the enzyme. The recent crystallographic identification of the steric location of all iron-sulphur clusters of the bacterial enzyme [48, 49] allows to locate N2 more precisely, closer to the membrane sector of the enzyme than previously suggested. Because of its midpoint potential higher than that of the other clusters, N2 is considered as the direct electron donor to ubiquinone. It is commonly accepted that Complex I inhibitors share the same hydrophobic large pocket binding site in the enzyme [51] and, according to the structural model proposed by Brandt [53], this pocket could be the amphipathic “ramp” guiding ubiquinone into the catalytic site. In this picture rotenone and related inhibitors would prevent the quinone access to the catalytic site, but would not prevent the reduction of N2 cluster.
The electron transfer from NADH to ubiquinone in Complex I requires the presence of at least eight iron-sulphur clusters, seven of which are well protected from reacting with oxygen with the exception of N2. From structural and functional studies the iron-sulphur cluster N2 seems to be localized in a region that should be accessible to protein bound ubisemiquinones, to H+ ions and to water, hence this region should be also accessible to molecular oxygen. On the other hand the midpoint potential of cluster N2 is around –0.15 to −0.05 V [80] and therefore it is compatible with the reduction of oxygen to superoxide anion (mid point potential for the couple superoxide/oxygen is −0.14 V). The correct value of the midpoint potential for the superoxide/oxygen couple [46] makes less stringent the identification of a group having lower potential such as cluster N1a [26] and flavin itself (see above).
We have exploited the ability of Complex I to transfer electrons directly to molecular oxygen with the aim to elucidate not only the site of electron escape in Complex I but also the mechanism of electron transfer inside the enzyme [29]. To this purpose we have tested the effects of different inhibitors on the radical production from Complex I. The findings provide evidence on a strikingly differential effect of two classes of Complex I inhibitors, based on their ability to affect oxygen radical production by the enzyme. Class A inhibitors induce a strong increase in the ROS production from Complex I, whereas Class B inhibitors completely prevent ROS production from the enzyme.
Class A inhibitors include rotenone, piericidin A, rolliniastatin-1 and -2, but also myxothiazol, while Class B includes stigmatellin, capsaicin, mucidin at high concentration, and also short ubiquinone analogues such as CoQ2. Accurate controls have excluded for these compounds a generic effect as free radical scavengers.
Starting from available knowledge from the literature and from the results described in this work, we proposed that Class A inhibitors prevent access of the physiological CoQ10 to its reduction site, thus allowing the reductant of CoQ to release one electron to oxygen instead, while Class B inhibitors directly act on the site of oxygen reduction.
Our results agree with cluster N2 being the direct reductant of molecular oxygen. Anyway during normal redox cycle the electron leak from Complex I is very low: it can be increased by the presence of Class A inhibitors while it is not related to the reduced state of the enzyme. In fact in presence of 1.8 μM mucidin, that inhibits Complex III and prevents radical formation from it without affecting the Complex I activity, and at saturating concentrations of NADH (condition that allows the full reduction of all redox centres in Complex I as well as the reduction of the quinone pool [81], the superoxide production was not significantly enhanced. On the other hand, when mucidin was used at 60 μM concentration, we could achieve full inhibition of the Complex I activity together with a full inhibition of ROS production even in presence of Class A inhibitors. These results suggest that to rise up the electron escape from Complex I, Class A inhibitors are necessary. It might be guessed that they induce in the enzyme a conformational change that makes the reducing centre more accessible to molecular oxygen, whereas Class B inhibitors would either directly block this reducing centre, or induce a conformational change making it less accessible.
These findings have allowed to get a deeper insight into the mechanism of electron transfer of Complex I to the CoQ acceptor. This issue will be developed in the next section.
Mechanism of Electron Transfer in Complex I
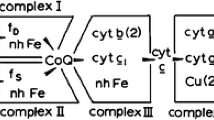
The primary acceptor of electrons from NADH is FMN bound to the 51 kDa subunit [82]; since iron-sulphur cluster N1a has a very negative potential and is situated too far from the other iron-sulphur clusters [48], it is not considered to reside in the main pathway of electrons [83]. Thus electrons would flow from FMN to N3 in the same 51 kDa subunit, and to N4 and N5 in the 75 kDa subunit [80, 84, 85], and then to N6a and N6b in the TYKY subunit [86] and to N2 in PSST subunit [39, 87, 88] but shared with the 49 kDa subunit [89]. N2 is the direct electron donor to bound ubiquinone [80] and probably this step is linked to proton translocation [44], although the mechanism is still debated [52, 53, 90–93]. A recent view favours a conformational mechanism [47], since all redox groups in the enzyme appear to be located in the hydrophilic arm or at least at the interface with the hydrophobic arm.
The mechanism of CoQ reduction is particularly intriguing, since more than one bound quinone species has been assigned to the enzyme; three ubisemiquinone EPR signals are detectable in the enzyme [42, 80].
The findings in our laboratory that two different classes of inhibitors have opposite effects on oxygen reduction to superoxide during forward electron transfer allow to draw two minimal schemes of electron transfer in Complex I (Fig. 2). In a linear scheme the electron donor to oxygen is presumably FeS cluster N2, whose reduction would be inhibited by stigmatellin while its reoxidation would be inhibited by rotenone. This scheme is not compatible with the notion that the stigmatellin inhibition site is downstream with respect to the rotenone site, since the behaviour of stigmatellin as an inhibitor is shared by reduced quinone analogs [50]. On the other hand, in the bifurcated scheme shown in the figure, an iron-sulphur cluster located upstream N2 centre acts as a “switch” for electron delivery. In absence of quinone in the active site of the enzyme (e.g. in presence of Class A inhibitors) the iron-sulphur clusters chain is completely connected and electrons flow directly to N2 centre. In presence of quinone, however, the chain is interrupted at a level of the “switch” that gives the first electron to quinone. The resulting semiquinone allows a conformational change connecting the “switch” to the downstream Fe-S clusters, inducing the complete reduction of semiquinone to quinol via N2 centre. Class B inhibitors would prevent the delivery of the second electron to semiquinone without affecting its formation and, acting on N2 centre, would prevent also superoxide formation.
A further confirmation of this scheme derives from the effect of inhibitors on reduction of the acceptor dichlorophenol indophenol (DCIP). Some DCIP is reduced at the level of FMN, since there is a component insensitive to DPI; another component is sensitive to DPI and must be reduced at the level of CoQ. In fact both hydrophilic and hydrophobic quinones enhance DPI-sensitive DCIP reduction. The reduction is inhibited by rotenone but only slightly by stigmatellin.
These findings demonstrate that DCIP is reduced at a site situated between the rotenone and the stigmatellin inhibition sites, a further indication for a split pathway of electrons at the CoQ binding site. According to the scheme presented in Fig. 2, DCIP would be reduced by ubisemiquinone, since its formation is rotenone sensitive but stigmatellin insensitive.
The results of this investigation have to be reconciled with the linear pathway of electrons along the series of iron-sulphur clusters as demonstrated by the crystallographic study of Hinchliffe and Sazanov [48]; our interpretation is not in contrast with the existence of a linear pathway, because the two electrons delivered to CoQ for its complete reduction could well be provided alternatively by two different clusters if a suitable conformational change occurs after the first electron delivery in order to provide a gating mechanism for the second electron.
Complex I in Pathology
Mitochondrial Cytopathies as a Model of Neurodegeneration
Mitochondrial diseases comprise a heterogeneous group of disorders characterized by impairment of mitochondrial oxidative phosphorylation; muscle and brain are mostly affected, probably because of their high dependence on oxidative metabolism [94], but the term mitochondrial encephalomyopathies is not totally correct because of the systemic involvement of the whole organism: for this reason the term mitochondrial cytopathies is to be preferred. Overall, mitochondrial cytopathies have an incidence of about 1 in 7600–10,000 [95, 96].
A genetic classification of mitochondrial cytopathies [97, 98] distinguishes disorders due to defects of the mitochondrial genome and those due to nuclear DNA mutations. Only one-third of the over 150 pathogenic mtDNA mutations concerns structural genes, the others are either deletions or rearrangements or they affect mitochondrial tRNA or rRNA genes. A more recent and broader classification distinguishes four categories of mitochondrial disorders [99]: those due to mutations in respiratory chain subunits, those due to mutations affecting respiratory complexes assembly, those due to alteration of mitochondrial DNA (mtDNA) translation or its integrity, and those due to mutations affecting mitochondrial morphology and motility. The heterogeneity of the clinical patterns of mtDNA defects is related to the complexity of mitochondrial genetics [100]: the degree of heteroplasmy usually differs in different tissues due to mitotic segregation and other less known phenomena; in addition threshold effects [101] allow normal biochemical phenotype until a well-defined threshold (usually high, up to 90% mutated mtDNA with respect to wild type) is reached.
The phenotypic threshold may be explained by complementation of the altered products of mutated mtDNA by the normal products of wild-type mtDNA at different levels: transcription, translation, enzyme complex assembly, biochemical level and cellular level. The possibility of mitochondrial trans-complementation is controversial [102–105]. Three mechanisms may underlie the biochemical threshold [101]: an excess of active oxidative phosphorylation complexes, the presence of inactive complexes that are activated when the oxidative phosphorylation level becomes insufficient, and an increased turnover of the active complexes due to regulation mechanisms. Flux control analysis [106] has been critical for the understanding of the biochemical threshold.
Several mtDNA point mutations of structural genes have been associated with Leber’s Hereditary Optic Neuropathy (LHON) [107] (Table 1); here we survey some biochemical aspects of the LHON syndrome due to the three primary mutations in Complex I ND subunits.
Leber’s Hereditary Optic Neuropathy is due to three main mutations in genes for Complex I subunits affecting subunits ND1, ND4, ND6. The clinical syndrome is characterized by retinal ganglion cells and optic nerve degeneration with sudden blindness. The disease mainly affects individuals with homoplasmic mutations, but not all subjects harbouring the pathogenic mutations are affected, suggesting that other genetic and/or environmental factors are required for the development of the disease. The three pathogenic mutations of complex I [108, 109] occur at positions G11778A/ND4, G3460A/ND1, and T14484C/ND6.
Biochemical investigations of the three most frequent mutations revealed some subtle biochemical changes in Complex I function [110]. Only the 3460/ND1 mutation showed a consistent reduction in complex I electron transfer activity [111–113], while both 11778/ND4 and 14484/ND6 mutations had normal activities [111, 114–116].
Studies on the sensitivity of Complex I to different inhibitors showed a decreased sensitivity to rotenone and an enhanced sensitivity to quinol product inhibitors [117], while sensitivity to other complex I inhibitors not interfering with the CoQ binding site, such as rolliniastatin-2 or amytal, was unchanged. These results might be interpreted assuming that the mutations interfere with the interaction of complex I with CoQ, suggesting that the CoQ binding site may be affected by the mutations.
The complex I dysfunction in LHON may have three major consequences: (a) the release of quinol product may be affected, thus leading to decreased total respiratory activity; (b) due to alteration of the hydrophobic quinone binding site(s), proton pumping through complex I may be defective thereby affecting energy conservation; (c) an increase of ROS generation may occur as a consequence of altered electron flow, as reported in the case of nuclear complex I mutations [118]. Studies using osteosarcoma-derived cybrids carrying each of the LHON mutations indicate that Complex I-dependent ATP synthesis is reduced by all three mutations, though the inhibiting effect was less severe with the 11778/ND4 mutation. Significantly, this mutation was associated with an uncoupling of the oxidative phosphorylation more than with the reduced electron transport activity of complex I, which in fact appeared to be more effective in the presence of the 3460/ND1 and the 14484/ND6 mutations. The reduced ATP synthesis rate of the mutated cybrids was reflected by the slight reduction of total ATP cellular content observed [107, 119]. Complex II-dependent ATP synthesis does not appear to be significantly affected.
Besides an energy defect, overproduction of ROS may represent a major element in LHON pathophysiology [120–122]. This hypothesis is supported by the increased ROS generation after partial Complex I inhibition (cf. [20] and previous sections). The apoptotic cell death occurring in LHON cybrids when incubated in galactose medium [123] may be the result of both decreased OXPHOS and increased ROS generation. [124].
Several examples of enhanced ROS production in genetic defects of Complex I are known in the literature, particularly for nuclear genes mutations [118, 125, 126], whereas the effect of mitochondrial gene mutations is less clear [107, 127, 128]; recently cybrids carrying the LHON 14487 ND6 mutation were shown to undergo a ROS overproduction [129]. Also physiological states, such as subunit phosphorylation, may modify the ROS generating capacity of Complex I [126, 130, 131]. It is therefore tempting to speculate that endocrine alterations may affect the capacity of ROS formation by hyper- or hypo-phosphorylation of the Complex.
Complex I and Parkinson’s Disease
The incidence of Parkinson’s disease (PD) is estimated as 8–18 per 100,000 person-years, and the prevalence is approximately 0.3% of the entire population: PD affects more than 1% of those older than 60 years and up to 4% of those older than 80 years [132].
Epidemiological studies reveal that <10% of PD has a strict familial aetiology while the majority of cases are sporadic. The discovery of genes linked to rare familial forms of PD during the last decade confirmed the role of genetics in development of PD, and provided vital clues in understanding molecular pathogenesis of the common sporadic illness [133].
The neuropathological hallmarks are characterized by progressive and profound loss of neuromelanin-containing dopaminergic neurons in the substantia nigra pars compacta (SN) with presence of eosinophilic, intracytoplasmic, proteinaceous inclusions termed as Lewy bodies (LB) and dystrophic Lewy neurites in surviving neurons [134].
Expeditiously, after the identification of mutations in the gene encoding the protein α-synuclein (α-syn) in kindreds with PD [135], it was determined that filamentous α-syn is the major component of Lewy pathology [136–140].
Markers of oxidative stress, such as products of lipid peroxidation [141, 142], protein oxidation [143–146] and oxidation of mtDNA and cytoplasmic RNA [147], are increased in dopaminergic neurons of PD brains. Increased oxidation in the SN of PD patients also may be partially due to the reported accumulation of iron [148–152], which in the form of Fe2+ can catalyze the formation of strong oxidants. The presence of advanced glycation products and 3-nitrotyrosine in Lewy pathology and the demonstration that α-syn is a specific target of nitration [153–157] suggests that oxidative damage may be involved in the formation of these inclusions [158].
Numerous studies indicated the involvement of ROS and oxidative stress in PD pathogenesis, including reduced amounts of the thiol-reducing agent glutathione [159, 160] and elevated concentrations of iron [161] in SN of PD patients. Loss of neuromelanin-containing DAergic cells is characteristic for PD and the dark brown pigment neuromelanin attracted attention to the auto-oxidation of dopamine (DA), as it consists primarily of products of DA redox chemistry [162]. Normal metabolism of DA, partly accomplished by monoamine oxidases, produces hydrogen peroxide (H2O2) [163]. From this reaction alone, DAergic neurons are exposed to oxidative stress. In addition, DA can be oxidized to a dopamine quinone. This oxidation occurs spontaneously, is accelerated by the presence of transition metal ions, or can be enzyme-catalysed. The resulting dopamine quinone covalently modifies cellular macromolecules, which may serve as a mechanism for dopamine-induced neurotoxicity [164, 165]. Oxidation of DA to dopamine quinone by superoxide may trigger a vicious cycle of oxidative stress through the reduction of the quinone by mitochondrial Complex I to its semiquinone form and its reoxidation by oxygen to form additional superoxide [166, 167]. Indeed this mechanism has been proven for an adrenaline/adrenochrome cycle in isolated mitochondria [77, 168].
Moreover, multiple lines of evidence suggest a pathogenic role of oxidative damage and mitochondrial dysfunction in causing PD [169]. The direct relation between mitochondrial dysfunction and PD came from the post-mortem description of complex I deficiency in the SN of patients with PD [170]. Subsequently, the deficiency was also seen in the skeletal muscle and platelets [171, 172] and there was a decrease in complex I proteins in the SN of patients with PD [173]. Consistent deficits in the subunits and activity of mitochondrial complex I of the electron transport chain in blood platelets and SN of PD patients is a prominent phenomenon [174, 175]. Reduced complex I activity is also seen in cytoplasmic hybrid (cybrid) cell lines containing mtDNA from PD patients [176]. The complex I deficiency in the substantia nigra and platelets implies that it is a systemic defect in a proportion of cases (~25% on the basis of platelet activities) and this might be due to genetic or environmental (endogenous or exogenous) causes. The complex I defect in patients with PD lowers the threshold of apoptosis mediated by the mitochondria—through a decrease in ATP production and by the generation of free radicals—and sensitises cells to the proapoptotic protein Bax [177]. The specificity of the complex I defect in the brains of patients with PD and its relation with oxidative stress have been supported by the recent finding that this is the only respiratory chain protein complex that is affected by endogenous oxidative damage and has reduced structural stability [174].
The connection between mitochondria and PD has been reinforced by the finding that several of the genes that cause familial PD encode mitochondrial proteins and that mitochondrial toxins can cause PD in animals [178].
Mutations or polymorphisms in both mtDNA and nuclear DNA were implicated in causing PD or in affecting PD risk [179]. To this purpose, mtDNA mutations may be involved in the aetiology and predisposition to the idiopathic disease, since cybrids containing mitochondria from Parkinson’s patients exhibit a reduced activity of Complex I [180] and generate Lewy inclusion bodies [181]. However, exhaustive sequencing of mtDNA has not yet revealed mutations that consistently associate with PD [182]; nevertheless, somatic deletions of mtDNA are found more frequently in SN from PD patients [183, 184].
Mitochondrial dysfunction has long been implicated in PD pathogenesis; this hypothesis arose with the discovery that 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) produced PD-like symptoms in designer drug abusers [185]. Its metabolite, 1-methyl-4-phenylpyridinium (MPP+), is actively transported into DAergic neurons by the dopamine transporter. Within these neurons MPP+ enters mitochondria, and selectively inhibits mitochondrial respiration at complex I of the electron transport chain [186, 187]. Chronic infusion of rotenone, a highly selective complex I inhibitor, also reproduced behavioural and neuropathological features of PD in rats [188, 189]. These neurotoxins and neurotoxic animal models of PD renewed interest in possible environmental causes of PD, as similar compounds in the environment might play a causative role in the disease [190–192]. Furthermore, the environmental toxins causing parkinsonism identified thus far are all inhibitors of Complex I, e.g. MPTP, rotenone and annonacin [190].
In addition, genetic defects causing familial forms of PD have been identified in the last decade. Despite the rarity of these familial forms of PD the identification of PD-linked genes has fuelled our understanding of possible pathogenic mechanisms of PD, and placed ubiquitin–proteasome system dysfunction, oxidative stress and mitochondrial dysfunction at centre stage [192]. The discovery of complex I deficiency in PD and the role of mitochondria in PD has been enhanced by the subsequent identification of mutations in genes encoding mitochondrial proteins, e.g. PINK1 and DJ1 as causes of autosomal recessive PD, and by the mitochondrial abnormalities associated with α-synuclein and parkin expression.
A major step in our understanding of the aetiopathogenesis of the disease came when mutations were identified in α-synuclein in 1997, followed by mutations in parkin a year after that [135, 193]. The demonstration that α-syn is the main constituent of Lewy bodies in the same year suggested a primary role for α-syn aggregation, however, later studies revealed close interplay between α-syn aggregation and oxidative stress in the pathogenesis of PD [136, 194, 195]. The identification of mutations in DJ1 [PD (autosomal recessive, early onset), a possible redox sensor] in 2003 and phosphatase and tensin homologue (PTEN)-induced kinase 1 (PINK1, a mitochondrial kinase) in 2004 provided strong evidence that mitochondrial dysfunction and oxidative stress might have a primary role in the pathogenesis of PD, although how mutations in these genes cause neuronal degeneration is still unclear [196, 197]. The recent observation [198] that α-syn can be imported into mitochondria and inhibit Complex I inducing enhancement of ROS production and that these effects have an earlier onset with mutated α-syn is strongly relevant to the role and interplay of all these factors in the pathogenesis of PD.
Thus, although classically regarded as an archetypical non-genetic disease due to the high proportion of sporadic cases, hugely significant advances in our understanding of PD have stemmed directly from the study of these genes associated with a small proportion of familial cases. [199].
A precise role of mitochondrial Complex I in the formation of Lewy bodies through α-synuclein aggregation is not yet defined: nevertheless the hypothesis is tenable that a primary mitochondrial dysfunction may lead to enhanced ROS production [200], triggering cell death mechanisms in dopaminergic cells [201] by an interplay of different endogenous and exogenous factors: indeed neurotoxins inducing parkinsonism, such as MPP+ and rotenone, stimulate ROS production by Complex I (see Sect. “ROS Production by Complex I”). Figure 3 schematically depicts possible pathogenetic mechanisms of PD.
A cartoon showing a hypothetical series of events in the pathogenesis of Parkinson’s disease. See text for explanations. The central event in the development of the disease is mitochondrial Complex I deficiency that can be promoted by a number of different causes either genetic or due to xenobiotic exposure. α-Synuclein aggregation is also a necessary prerequisite for PD development, induced by either mutations or post-translational modifications caused by ROS
Abbreviations
- CoQ (Q):
-
Coenzyme Q ubiquinone
- Cyt. c:
-
Cytochrome c
- EPR:
-
Electron paramagnetic resonance
- FeS:
-
Iron-sulphur cluster
- GPx :
-
Glutathione peroxidase
- DA:
-
Dopamine
- DCIP:
-
Dichlorophenol indophenol
- DPI:
-
Diphenylene iodonium
- Gpx :
-
Glutathione peroxidase
- LB:
-
Lewy bodies
- LHON:
-
Leber’s hereditary optic neuropathy
- MPP+ :
-
1-Methyl-4-phenylpyridinium
- MPTP:
-
1-Methyl-4-phenyl-1, 2, 3, 6-tetrahydropyridine
- PD:
-
Parkinson’s disease
- PTP:
-
Permeability transition pore
- ROS:
-
Reactive oxygen species
- SMP:
-
Submitochondrial particles
- SN:
-
Substantia nigra
- SOD:
-
Superoxide dismutase
- α-syn:
-
α-Synuclein
- UCP:
-
Uncoupling protein
- VDAC:
-
Voltage-dependent anion channel
References
Lenaz G (1998) Role of mitochondria in oxidative stress and ageing. Biochim Biophys Acta 1366:53–67
Lenaz G (2001) The mitochondrial production of reactive oxygen species: mechanisms and implications in human pathology. IUBMB Life 52:159–164
Andreyev AY, Kushnareva YE, Starkov AA (2005) Mitochondrial metabolism of reactive oxygen species. Biochemistry (Mosc) 70:200–214
Adam-Vizi V, Chinopoulos C (2006) Bioenergetics and the formation of mitochondrial reactive oxygen species. Trends Pharmacol Sci 27:639–645
Melov S, Coskun P, Patel M, Tuinstra R, Cottrell B, Jun AS, Zastawny TH, Dizdaroglu M, Goodman SI, Huang TT, Miziorko H, Epstein CJ, Wallace DC (1999) Mitochondrial disease in superoxide dismutase 2 mutant mice. Proc Natl Acad Sci USA 96:846–851
Esposito LA, Kokoszka JE, Waymire KG, Cottrell B, MacGregor GR, Wallace DC (2000) Mitochondrial oxidative stress in mice lacking the glutathione peroxidase-1 gene. Free Radic Biol Med 28:754–766
Chance B, Sies H, Boveris A (1979) Hydroperoxide metabolism in mammalian organs. Physiol Rev 59:527–605
Casteilla L, Rigoulet M, Penicaud L (2001) Mitochondrial ROS metabolism: modulation by uncoupling proteins. IUBMB Life 52:181–188
Han D, Williams E, Cadenas E (2001) Mitochondrial respiratory chain-dependent generation of superoxide anion and its release into the intermembrane space. Biochem J 353:411–416
St-Pierre J, Buckingham JA, Roebuck SJ, Brand MD (2002) Topology of superoxide production from different sites in the mitochondrial electron transport chain. J Biol Chem 277:44784–44790
Han D, Antunes F, Canali R, Rettori D, Cadenas E (2003) Voltage-dependent anion channels control the release of the superoxide anion from mitochondria to cytosol. J Biol Chem 278:5557–5563
Echtay KS, Murphy MP, Smith RA, Talbot DA, Brand MD (2002) Superoxide activates mitochondrial uncoupling protein 2 from the matrix side. Studies using targeted antioxidants. J Biol Chem 277:47129–47135
Kwong LK, Sohal RS (1998) Substrate and site specificity of hydrogen peroxide generation in mouse mitochondria. Arch Biochem Biophys 350:118–126
Lee HC, Wei YH (2007) Oxidative stress, mitochondrial DNA mutation, and apoptosis in aging. Exp Biol Med (Maywood) 232:592–606
Zorov DB, Juhaszova M, Sollott SJ (2006) Mitochondrial ROS-induced ROS release: an update and review. Biochim Biophys Acta 1757:509–517
Staniek K, Nohl H (2000) Are mitochondria a permanent source of reactive oxygen species? Biochim Biophys Acta 1460:268–275
Korshunov SS, Skulachev VP, Starkov AA (1997) High protonic potential actuates a mechanism of production of reactive oxygen species in mitochondria. FEBS Lett 416:15–18
Finkel T (2001) Reactive oxygen species and signal transduction. IUBMB Life 52:3–6
Boveris A, Oshino N, Chance B (1972) The cellular production of hydrogen peroxide. Biochem J 128:617–630
Barrientos A, Moraes CT (1999) Titrating the effects of mitochondrial complex I impairment in the cell physiology. J Biol Chem 274:16188–16197
Li N, Ragheb K, Lawler G, Sturgis J, Rajwa B, Melendez JA, Robinson JP (2003) Mitochondrial complex I inhibitor rotenone induces apoptosis through enhancing mitochondrial reactive oxygen species production. J Biol Chem 278:8516–8525
Chen Y, Millan-Ward E, Kong J, Israels SJ, Gibson SB (2007) Mitochondrial electron-transport-chain inhibitors of complexes I and II induce autophagic cell death mediated by reactive oxygen species. J Cell Sci 120:4155–4166
Li Y, Trush MA (1998) Diphenyleneiodonium, an NAD(P) H oxidase inhibitor, also potently inhibits mitochondrial reactive oxygen species production. Biochem Biophys Res Commun 253:295–299
McLennan HR, Degli Esposti M (2000) The contribution of mitochondrial respiratory complexes to the production of reactive oxygen species. J Bioenerg Biomembr 32:153–162
Vrablic AS, Albright CD, Craciunescu CN, Salganik RI, Zeisel SH (2001) Altered mitochondrial function and overgeneration of reactive oxygen species precede the induction of apoptosis by 1-O-octadecyl–2-methyl-rac-glycero–3-phosphocholine in p53-defective hepatocytes. FASEB J 15:1739–1744
Kushnareva Y, Murphy AN, Andreyev A (2002) Complex I-mediated reactive oxygen species generation: modulation by cytochrome c and NAD(P) + oxidation-reduction state. Biochem J 368:545–553
Skulachev VP (1996) Role of uncoupled and non-coupled oxidations in maintenance of safely low levels of oxygen and its one-electron reductants. Q Rev Biophys 29:169–202
Brand MD (2000) Uncoupling to survive? The role of mitochondrial inefficiency in ageing. Exp Gerontol 35:811–820
Lenaz G, Fato R, Genova ML, Bergamini C, Bianchi C, Biondi A (2006) Mitochondrial complex I: structural and functional aspects. Biochim Biophys Acta 1757:1406–1420
Zhang L, Yu L, Yu CA (1998) Generation of superoxide anion by succinate-cytochrome c reductase from bovine heart mitochondria. J Biol Chem 273:33972–33976
Drahota Z, Chowdhury SK, Floryk D, Mracek T, Wilhelm J, Rauchova H, Lenaz G, Houstek J (2002) Glycerophosphate-dependent hydrogen peroxide production by brown adipose tissue mitochondria and its activation by ferricyanide. J Bioenerg Biomembr 34:105–113
Forman JH, Kennedy J (1975) Superoxide production and electron transport in mitochondrial oxidation of dihydroorotic acid. J Biol Chem 250:4322–4326
Giorgio M, Migliaccio E, Orsini F, Paolucci D, Moroni M, Contursi C, Pelliccia G, Luzi L, Minucci S, Marcaccio M, Pinton P, Rizzuto R, Bernardi P, Paolucci F, Pelicci PG (2005) Electron transfer between cytochrome c and p66Shc generates reactive oxygen species that trigger mitochondrial apoptosis. Cell 122:221–233
Schultz BE, Chan SI (2001) Structures and proton-pumping strategies of mitochondrial respiratory enzymes. Annu Rev Biophys Biomol Struct 30:23–65
Carroll J, Fearnley IM, Shannon RJ, Hirst J, Walker JE (2003) Analysis of the subunit composition of complex I from bovine heart mitochondria. Mol Cell Proteomics 2:117–126
Carroll J, Fearnley IM, Skehel JM, Shannon RJ, Hirst J, Walker JE (2006) Bovine complex I is a complex of 45 different subunits. J Biol Chem 281:32724–32727
Chomyn A, Mariottini P, Cleeter MW, Ragan CI, Matsuno-Yagi A, Hatefi Y, Doolittle RF, Attardi G (1985) Six unidentified reading frames of human mitochondrial DNA encode components of the respiratory-chain NADH dehydrogenase. Nature 314:592–597
Chomyn A, Cleeter MW, Ragan CI, Riley M, Doolittle RF, Attardi G (1986) URF6, last unidentified reading frame of human mtDNA, codes for an NADH dehydrogenase subunit. Science 234:614–618
Friedrich T, Scheide D (2000) The respiratory complex I of bacteria, archaea and eukarya and its module common with membrane-bound multisubunit hydrogenases. FEBS Lett 479:1–5
Mathiesen C, Hagerhall C (2002) Transmembrane topology of the NuoL, M and N subunits of NADH:quinone oxidoreductase and their homologues among membrane-bound hydrogenases and bona fide antiporters. Biochim Biophys Acta 1556:121–132
Friedrich T, Bottcher B (2004) The gross structure of the respiratory complex I: a Lego System. Biochim Biophys Acta 1608:1–9
Ohnishi T, Sled VD, Yano T, Yagi T, Burbaev DS, Vinogradov AD (1998) Structure-function studies of iron-sulfur clusters and semiquinones in the NADH-Q oxidoreductase segment of the respiratory chain. Biochim Biophys Acta 1365:301–308
Kerscher S, Kashani-Poor N, Zwicker K, Zickermann V, Brandt U (2001) Exploring the catalytic core of complex I by Yarrowia lipolytica yeast genetics. J Bioenerg Biomembr 33:187–196
Yano T, Ohnishi T (2001) The origin of cluster N2 of the energy-transducing NADH-quinone oxidoreductase: comparisons of phylogenetically related enzymes. J Bioenerg Biomembr 33:213–222
Magnitsky S, Toulokhonova L, Yano T, Sled VD, Hagerhall C, Grivennikova VG, Burbaev DS, Vinogradov AD, Ohnishi T (2002) EPR characterization of ubisemiquinones and iron-sulfur cluster N2, central components of the energy coupling in the NADH-ubiquinone oxidoreductase (complex I) in situ. J Bioenerg Biomembr 34:193–208
Ohnishi ST, Ohnishi T, Muranaka S, Fujita H, Kimura H, Uemura K, Yoshida K, Utsumi K (2005) A possible site of superoxide generation in the complex I segment of rat heart mitochondria. J Bioenerg Biomembr 37:1–15
Brandt U, Kerscher S, Drose S, Zwicker K, Zickermann V (2003) Proton pumping by NADH:ubiquinone oxidoreductase. A redox driven conformational change mechanism? FEBS Lett 545:9–17
Hinchliffe P, Sazanov LA (2005) Organization of iron-sulfur clusters in respiratory complex I. Science 309:771–774
Sazanov LA (2007) Respiratory complex I: mechanistic and structural insights provided by the crystal structure of the hydrophilic domain. Biochemistry 46:2275–2288
Degli Esposti M (1998) Inhibitors of NADH-ubiquinone reductase: an overview. Biochim Biophys Acta 1364:222–235
Okun JG, Lummen P, Brandt U (1999) Three classes of inhibitors share a common binding domain in mitochondrial complex I (NADH:ubiquinone oxidoreductase). J Biol Chem 274:2625–2630
Ohnishi T, Salerno JC (2005) Conformation-driven and semiquinone-gated proton-pump mechanism in the NADH-ubiquinone oxidoreductase (complex I). FEBS Lett 579:4555–4561
Brandt U (1997) Proton-translocation by membrane-bound NADH:ubiquinone-oxidoreductase (complex I) through redox-gated ligand conduction. Biochim Biophys Acta 1318:79–91
Sherwood S, Hirst J (2006) Investigation of the mechanism of proton translocation by NADH:ubiquinone oxidoreductase (complex I) from bovine heart mitochondria: does the enzyme operate by a Q-cycle mechanism? Biochem J 400:541–550
Raha S, Robinson BH (2000) Mitochondria, oxygen free radicals, disease and ageing. Trends Biochem Sci 25:502–508
Green DE, Tzagoloff A (1966) The mitochondrial electron transfer chain. Arch Biochem Biophys 116:293–304
Schagger H (2001) Respiratory chain supercomplexes. IUBMB Life 52:119–128
Bianchi C, Genova ML, Parenti Castelli G, Lenaz G (2004) The mitochondrial respiratory chain is partially organized in a supercomplex assembly: kinetic evidence using flux control analysis. J Biol Chem 279:36562–36569
Lenaz G, Fato R, Formiggini G, Genova ML (2007) The role of Coenzyme Q in mitochondrial electron transport. Mitochondrion 7(Suppl):S8–S33
Nohl H, Gille L, Staniek K (2005) Intracellular generation of reactive oxygen species by mitochondria. Biochem Pharmacol 69:719–723
Takeshige K, Minakami S (1979) NADH and NADPH-dependent formation of superoxide anions by bovine heart submitochondrial particles and NADH-ubiquinone reductase preparation. Biochem J 180:129–135
Cadenas E, Boveris A, Ragan CI, Stoppani AO (1977) Production of superoxide radicals and hydrogen peroxide by NADH-ubiquinone reductase and ubiquinol-cytochrome c reductase from beef-heart mitochondria. Arch Biochem Biophys 180:248–257
Herrero A, Barja G (2000) Localization of the site of oxygen radical generation inside the complex I of heart and nonsynaptic brain mammalian mitochondria. J Bioenerg Biomembr 32:609–615
Genova ML, Ventura B, Giuliano G, Bovina C, Formiggini G, Parenti Castelli G, Lenaz G (2001) The site of production of superoxide radical in mitochondrial Complex I is not a bound ubisemiquinone but presumably iron-sulfur cluster N2. FEBS Lett 505:364–368
Turrens JF (2003) Mitochondrial formation of reactive oxygen species. J Physiol 552:335–344
Lambert AJ, Brand MD (2004) Inhibitors of the quinone-binding site allow rapid superoxide production from mitochondrial NADH:ubiquinone oxidoreductase (complex I). J Biol Chem 279:39414–39420
Lambert AJ, Brand MD (2004) Superoxide production by NADH:ubiquinone oxidoreductase (complex I) depends on the pH gradient across the mitochondrial inner membrane. Biochem J 382:511–517
Starkov AA, Fiskum G (2003) Regulation of brain mitochondrial H2O2 production by membrane potential and NAD(P) H redox state. J Neurochem 86:1101–1107
Hansford RG, Hogue BA, Mildaziene V (1997) Dependence of H2O2 formation by rat heart mitochondria on substrate availability and donor age. J Bioenerg Biomembr 29:89–95
Starkov AA, Polster BM, Fiskum G (2002) Regulation of hydrogen peroxide production by brain mitochondria by calcium and Bax. J Neurochem 83:220–228
Vinogradov AD, Grivennikova VG (2005) Generation of superoxide-radical by the NADH:ubiquinone oxidoreductase of heart mitochondria. Biochemistry (Mosc) 70:120–127
Liu Y, Fiskum G, Schubert D (2002) Generation of reactive oxygen species by the mitochondrial electron transport chain. J Neurochem 80:780–787
Vinogradov AD (1998) Catalytic properties of the mitochondrial NADH-ubiquinone oxidoreductase (complex I) and the pseudo-reversible active/inactive enzyme transition. Biochim Biophys Acta 1364:169–185
Majander A, Finel M, Wikstrom M (1994) Diphenyleneiodonium inhibits reduction of iron-sulfur clusters in the mitochondrial NADH-ubiquinone oxidoreductase (Complex I). J Biol Chem 269:21037–21042
Galkin A, Brandt U (2005) Superoxide radical formation by pure complex I (NADH:ubiquinone oxidoreductase) from Yarrowia lipolytica. J Biol Chem 280:30129–30135
Kussmaul L, Hirst J (2006) The mechanism of superoxide production by NADH:ubiquinone oxidoreductase (complex I) from bovine heart mitochondria. Proc Natl Acad Sci USA 103:7607–7612
Genova ML, Abd-Elsalam NM, Mahdy el SM, Bernacchia A, Lucarini M, Pedulli GF, Lenaz G (2006) Redox cycling of adrenaline and adrenochrome catalysed by mitochondrial Complex I. Arch Biochem Biophys 447:167–173
Grgic L, Zwicker K, Kashani-Poor N, Kerscher S, Brandt U (2004) Functional significance of conserved histidines and arginines in the 49-kDa subunit of mitochondrial complex I. J Biol Chem 279:21193–21199
Garofano A, Zwicker K, Kerscher S, Okun P, Brandt U (2003) Two aspartic acid residues in the PSST-homologous NUKM subunit of complex I from Yarrowia lipolytica are essential for catalytic activity. J Biol Chem 278:42435–42440
Ohnishi T (1998) Iron-sulfur clusters/semiquinones in complex I. Biochim Biophys Acta 1364:186–206
Kroger A, Klingenberg M (1973) The kinetics of the redox reactions of ubiquinone related to the electron-transport activity in the respiratory chain. Eur J Biochem 34:358–368
Fecke W, Sled VD, Ohnishi T, Weiss H (1994) Disruption of the gene encoding the NADH-binding subunit of NADH: ubiquinone oxidoreductase in Neurospora crassa. Formation of a partially assembled enzyme without FMN and the iron-sulphur cluster N–3. Eur J Biochem 220:551–558
Videira A, Duarte M (2002) From NADH to ubiquinone in Neurospora mitochondria. Biochim Biophys Acta 1555:187–191
Yagi T, Yano T, Di BS, Matsuno-Yagi A (1998) Procaryotic complex I (NDH–1), an overview. Biochim Biophys Acta 1364:125–133
Yano T, Yagi T, Sled VD, Ohnishi T (1995) Expression and characterization of the 66-kilodalton (NQO3) iron-sulfur subunit of the proton-translocating NADH-quinone oxidoreductase of Paracoccus denitrificans. J Biol Chem 270:18264–18270
Friedrich T, Brors B, Hellwig P, Kintscher L, Rasmussen T, Scheide D, Schulte U, Mantele W, Weiss H (2000) Characterization of two novel redox groups in the respiratory NADH:ubiquinone oxidoreductase (complex I). Biochim Biophys Acta 1459:305–309
Friedrich T (1998) The NADH:ubiquinone oxidoreductase (complex I) from Escherichia coli. Biochim Biophys Acta 1364:134–146
Sousa R, Barquera B, Duarte M, Finel M, Videira A (1999) Characterisation of the last Fe-S cluster-binding subunit of Neurospora crassa complex I. Biochim Biophys Acta 1411:142–146
Kashani-Poor N, Zwicker K, Kerscher S, Brandt U (2001) A central functional role for the 49-kDa subunit within the catalytic core of mitochondrial complex I. J Biol Chem 276:24082–24087
Friedrich T (2001) Complex I: a chimaera of a redox and conformation-driven proton pump? J Bioenerg Biomembr 33:169–177
Dutton PL, Moser CC, Sled VD, Daldal F, Ohnishi T (1998) A reductant-induced oxidation mechanism for complex I. Biochim Biophys Acta 1364:245–257
Vinogradov AD (2001) Respiratory complex I: structure, redox components, and possible mechanisms of energy transduction. Biochemistry (Mosc) 66:1086–1097
Brandt U (2006) Energy converting NADH:quinone oxidoreductase (complex I). Annu Rev Biochem 75:69–92
Filosto M, Tomelleri G, Tonin P, Scarpelli M, Vattemi G, Rizzuto N, Padovani A, Simonati A (2007) Neuropathology of mitochondrial diseases. Biosci Rep 27:23–30
Darin N, Oldfors A, Moslemi AR, Holme E, Tulinius M (2001) The incidence of mitochondrial encephalomyopathies in childhood: clinical features and morphological, biochemical, and DNA anbormalities. Ann Neurol 49:377–383
Skladal D, Halliday J, Thorburn DR (2003) Minimum birth prevalence of mitochondrial respiratory chain disorders in children. Brain 126:1905–1912
DiMauro S, Schon EA (2003) Mitochondrial respiratory-chain diseases. N Engl J Med 348:2656–2668
DiMauro S, Hirano M (2005) Mitochondrial encephalomyopathies: an update. Neuromuscul Disord 15:276–286
Schon EA, DiMauro S (2007) Mitochondrial mutations: genotype to phenotype. Novartis Found Symp 287:214–225
DiMauro S (2004) Mitochondrial diseases. Biochim Biophys Acta 1658:80–88
Rossignol R, Faustin B, Rocher C, Malgat M, Mazat JP, Letellier T (2003) Mitochondrial threshold effects. Biochem J 370:751–762
Attardi G, Enriquez JA, Cabezas-Herrera J (2002) Inter-mitochondrial complementation of mtDNA mutations and nuclear context. Nat Genet 30:360
Enriquez JA, Cabezas-Herrera J, Bayona-Bafaluy MP, Attardi G (2000) Very rare complementation between mitochondria carrying different mitochondrial DNA mutations points to intrinsic genetic autonomy of the organelles in cultured human cells. J Biol Chem 275:11207–11215
Ono T, Isobe K, Nakada K, Hayashi JI (2001) Human cells are protected from mitochondrial dysfunction by complementation of DNA products in fused mitochondria. Nat Genet 28:272–275
D’Aurelio M, Gajewski CD, Lin MT, Mauck WM, Shao LZ, Lenaz G, Moraes CT, Manfredi G (2004) Heterologous mitochondrial DNA recombination in human cells. Hum Mol Genet 13:3171–3179
Kacser H, Burns JA (1979) Molecular democracy: who shares the controls? Biochem Soc Trans 7:1149–1160
Lenaz G, Baracca A, Carelli V, D’Aurelio M, Sgarbi G, Solaini G (2004) Bioenergetics of mitochondrial diseases associated with mtDNA mutations. Biochim Biophys Acta 1658:89–94
Carelli V (2002) Leber’s hereditary optic neuropathy. In: Schapira AHV, Di Mauro S (eds) Mitochondrial disorders in neurology. Butterworth-Heinemann, Boston, pp 115–142
Wallace DC, Singh G, Lott MT, Hodge JA, Schurr TG, Lezza AM, Elsas LJ, Nikoskelainen EK (1988) Mitochondrial DNA mutation associated with Leber’s hereditary optic neuropathy. Science 242:1427–1430
Brown MD (1999) The enigmatic relationship between mitochondrial dysfunction and Leber’s hereditary optic neuropathy. J Neurol Sci 165:1–5
Carelli V, Ghelli A, Ratta M, Bacchilega E, Sangiorgi S, Mancini R, Leuzzi V, Cortelli P, Montagna P, Lugaresi E, Degli Esposti M (1997) Leber’s hereditary optic neuropathy: biochemical effect of 11778/ND4 and 3460/ND1 mutations and correlation with the mitochondrial genotype. Neurology 48:1623–1632
Howell N, Bindoff LA, McCullough DA, Kubacka I, Poulton J, Mackey D, Taylor L, Turnbull DM (1991) Leber hereditary optic neuropathy: identification of the same mitochondrial ND1 mutation in six pedigrees. Am J Hum Genet 49:939–950
Majander A, Huoponen K, Savontaus ML, Nikoskelainen E, Wikstrom M (1991) Electron transfer properties of NADH:ubiquinone reductase in the ND1/3460 and the ND4/11778 mutations of the Leber hereditary optic neuroretinopathy (LHON). FEBS Lett 292:289–292
Carelli V, Ghelli A, Bucchi L, Montagna P, De NA, Leuzzi V, Carducci C, Lenaz G, Lugaresi E, Degli Esposti M (1999) Biochemical features of mtDNA 14484 (ND6/M64 V) point mutation associated with Leber’s hereditary optic neuropathy. Ann Neurol 45:320–328
Cock HR, Cooper JM, Schapira AH (1995) The 14484 ND6 mtDNA mutation in Leber hereditary optic neuropathy does not affect fibroblast complex I activity. Am J Hum Genet 57:1501–1502
Majander A, Finel M, Savontaus ML, Nikoskelainen E, Wikstrom M (1996) Catalytic activity of complex I in cell lines that possess replacement mutations in the ND genes in Leber’s hereditary optic neuropathy. Eur J Biochem 239:201–207
Degli Esposti M, Carelli V, Ghelli A, Ratta M, Crimi M, Sangiorgi S, Montagna P, Lenaz G, Lugaresi E, Cortelli P (1994) Functional alterations of the mitochondrially encoded ND4 subunit associated with Leber’s hereditary optic neuropathy. FEBS Lett 352:375–379
Pitkanen S, Robinson BH (1996) Mitochondrial complex I deficiency leads to increased production of superoxide radicals and induction of superoxide dismutase. J Clin Invest 98:345–351
Baracca A, Solaini G, Sgarbi G, Lenaz G, Baruzzi A, Schapira AH, Martinuzzi A, Carelli V (2005) Severe impairment of complex I-driven adenosine triphosphate synthesis in Leber hereditary optic neuropathy cybrids. Arch Neurol 62:730–736
Carelli V, La MC, Iommarini L, Carroccia R, Mattiazzi M, Sangiorgi S, Farne’ S, Maresca A, Foscarini B, Lanzi L, Amadori M, Bellan M, Valentino ML (2007) Mitochondrial optic neuropathies: how two genomes may kill the same cell type? Biosci Rep 27:173–184
Yen MY, Wang AG, Wei YH (2006) Leber’s hereditary optic neuropathy: a multifactorial disease. Prog Retin Eye Res 25:381–396
Carelli V, Ross-Cisneros FN, Sadun AA (2002) Optic nerve degeneration and mitochondrial dysfunction: genetic and acquired optic neuropathies. Neurochem Int 40:573–584
Ghelli A, Zanna C, Porcelli AM, Schapira AH, Martinuzzi A, Carelli V, Rugolo M (2003) Leber’s hereditary optic neuropathy (LHON) pathogenic mutations induce mitochondrial-dependent apoptotic death in transmitochondrial cells incubated with galactose medium. J Biol Chem 278:4145–4150
Carelli V, Rugolo M, Sgarbi G, Ghelli A, Zanna C, Baracca A, Lenaz G, Napoli E, Martinuzzi A, Solaini G (2004) Bioenergetics shapes cellular death pathways in Leber’s hereditary optic neuropathy: a model of mitochondrial neurodegeneration. Biochim Biophys Acta 1658:172–179
Robinson BH (1998) Human complex I deficiency: clinical spectrum and involvement of oxygen free radicals in the pathogenicity of the defect. Biochim Biophys Acta 1364:271–286
Scacco S, Petruzzella V, Bertini E, Luso A, Papa F, Bellomo F, Signorile A, Torraco A, Papa S (2006) Mutations in structural genes of complex I associated with neurological diseases. Ital J Biochem 55:254–262
Wong A, Cavelier L, Collins-Schramm HE, Seldin MF, McGrogan M, Savontaus ML, Cortopassi GA (2002) Differentiation-specific effects of LHON mutations introduced into neuronal NT2 cells. Hum Mol Genet 11:431–438
Howell N (2003) LHON and other optic nerve atrophies: the mitochondrial connection. Dev Ophthalmol 37:94–108
Gonzalo R, Garcia-Arumi E, Llige D, Marti R, Solano A, Montoya J, Arenas J, Andreu AL (2005) Free radicals-mediated damage in transmitochondrial cells harboring the T14487C mutation in the ND6 gene of mtDNA. FEBS Lett 579:6909–6913
Raha S, Myint AT, Johnstone L, Robinson BH (2002) Control of oxygen free radical formation from mitochondrial complex I: roles for protein kinase A and pyruvate dehydrogenase kinase. Free Radic Biol Med 32:421–430
Maj MC, Raha S, Myint T, Robinson BH (2004) Regulation of NADH/CoQ oxidoreductase: do phosphorylation events affect activity? Protein J 23:25–32
de Lau LM, Breteler MM (2006) Epidemiology of Parkinson’s disease. Lancet Neurol 5:525–535
Thomas B, Beal MF (2007) Parkinson’s disease. Hum Mol Genet 16(2):R183–R194
Forno LS (1996) Neuropathology of Parkinson’s disease. J Neuropathol Exp Neurol 55:259–272
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A, Pike B, Root H, Rubenstein J, Boyer R, Stenroos ES, Chandrasekharappa S, Athanassiadou A, Papapetropoulos T, Johnson WG, Lazzarini AM, Duvoisin RC, Di IG, Golbe LI, Nussbaum RL (1997) Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276:2045–2047
Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M (1997) Alpha-synuclein in Lewy bodies. Nature 388:839–840
Spillantini MG, Crowther RA, Jakes R, Hasegawa M, Goedert M (1998) Alpha-Synuclein in filamentous inclusions of Lewy bodies from Parkinson’s disease and dementia with Lewy bodies. Proc Natl Acad Sci USA 95:6469–6473
Baba M, Nakajo S, Tu PH, Tomita T, Nakaya K, Lee VM, Trojanowski JQ, Iwatsubo T (1998) Aggregation of alpha-synuclein in Lewy bodies of sporadic Parkinson’s disease and dementia with Lewy bodies. Am J Pathol 152:879–884
Wakabayashi K, Hayashi S, Kakita A, Yamada M, Toyoshima Y, Yoshimoto M, Takahashi H (1998) Accumulation of alpha-synuclein/NACP is a cytopathological feature common to Lewy body disease and multiple system atrophy. Acta Neuropathol 96:445–452
Irizarry MC, Growdon W, Gomez-Isla T, Newell K, George JM, Clayton DF, Hyman BT (1998) Nigral and cortical Lewy bodies and dystrophic nigral neurites in Parkinson’s disease and cortical Lewy body disease contain alpha-synuclein immunoreactivity. J Neuropathol Exp Neurol 57:334–337
Yoritaka A, Hattori N, Uchida K, Tanaka M, Stadtman ER, Mizuno Y (1996) Immunohistochemical detection of 4-hydroxynonenal protein adducts in Parkinson disease. Proc Natl Acad Sci USA 93:2696–2701
Dexter DT, Carter CJ, Wells FR, Javoy-Agid F, Agid Y, Lees A, Jenner P, Marsden CD (1989) Basal lipid peroxidation in substantia nigra is increased in Parkinson’s disease. J Neurochem 52:381–389
Alam ZI, Daniel SE, Lees AJ, Marsden DC, Jenner P, Halliwell B (1997) A generalised increase in protein carbonyls in the brain in Parkinson’s but not incidental Lewy body disease. J Neurochem 69:1326–1329
Duda JE, Giasson BI, Chen Q, Gur TL, Hurtig HI, Stern MB, Gollomp SM, Ischiropoulos H, Lee VM, Trojanowski JQ (2000) Widespread nitration of pathological inclusions in neurodegenerative synucleinopathies. Am J Pathol 157:1439–1445
Butterfield DA, Kanski J (2001) Brain protein oxidation in age-related neurodegenerative disorders that are associated with aggregated proteins. Mech Ageing Dev 122:945–962
Beal MF (2002) Oxidatively modified proteins in aging and disease. Free Radic Biol Med 32:797–803
Zhang J, Perry G, Smith MA, Robertson D, Olson SJ, Graham DG, Montine TJ (1999) Parkinson’s disease is associated with oxidative damage to cytoplasmic DNA and RNA in substantia nigra neurons. Am J Pathol 154:1423–1429
Molina-Holgado F, Hider RC, Gaeta A, Williams R, Francis P (2007) Metals ions and neurodegeneration. Biometals 20:639–654
Dexter DT, Carayon A, Javoy-Agid F, Agid Y, Wells FR, Daniel SE, Lees AJ, Jenner P, Marsden CD (1991) Alterations in the levels of iron, ferritin and other trace metals in Parkinson’s disease and other neurodegenerative diseases affecting the basal ganglia. Brain 114(Pt 4):1953–1975
Dexter DT, Wells FR, Lees AJ, Agid F, Agid Y, Jenner P, Marsden CD (1989) Increased nigral iron content and alterations in other metal ions occurring in brain in Parkinson’s disease. J Neurochem 52:1830–1836
Dexter DT, Wells FR, Agid F, Agid Y, Lees AJ, Jenner P, Marsden CD (1987) Increased nigral iron content in postmortem parkinsonian brain. Lancet 2:1219–1220
Good PF, Olanow CW, Perl DP (1992) Neuromelanin-containing neurons of the substantia nigra accumulate iron and aluminum in Parkinson’s disease: a LAMMA study. Brain Res 593:343–346
Reynolds MR, Berry RW, Binder LI (2007) Nitration in neurodegeneration: deciphering the “Hows” “nYs”. Biochemistry 46:7325–7336
Castellani R, Smith MA, Richey PL, Perry G (1996) Glycoxidation and oxidative stress in Parkinson disease and diffuse Lewy body disease. Brain Res 737:195–200
Good PF, Hsu A, Werner P, Perl DP, Olanow CW (1998) Protein nitration in Parkinson’s disease. J Neuropathol Exp Neurol 57:338–342
Giasson BI, Duda JE, Murray IV, Chen Q, Souza JM, Hurtig HI, Ischiropoulos H, Trojanowski JQ, Lee VM (2000) Oxidative damage linked to neurodegeneration by selective alpha-synuclein nitration in synucleinopathy lesions. Science 290:985–989
Munch G, Luth HJ, Wong A, Arendt T, Hirsch E, Ravid R, Riederer P (2000) Crosslinking of alpha-synuclein by advanced glycation endproducts—an early pathophysiological step in Lewy body formation? J Chem Neuroanat 20:253–257
Giasson BI, Ischiropoulos H, Lee VM, Trojanowski JQ (2002) The relationship between oxidative/nitrative stress and pathological inclusions in Alzheimer’s and Parkinson’s diseases. Free Radic Biol Med 32:1264–1275
Bharath S, Hsu M, Kaur D, Rajagopalan S, Andersen JK (2002) Glutathione, iron and Parkinson’s disease. Biochem Pharmacol 64:1037–1048
Pearce RK, Owen A, Daniel S, Jenner P, Marsden CD (1997) Alterations in the distribution of glutathione in the substantia nigra in Parkinson’s disease. J Neural Transm 104:661–677
Kienzl E, Puchinger L, Jellinger K, Linert W, Stachelberger H, Jameson RF (1995) The role of transition metals in the pathogenesis of Parkinson’s disease. J Neurol Sci 134((Suppl)):69–78
Wakamatsu K, Fujikawa K, Zucca FA, Zecca L, Ito S (2003) The structure of neuromelanin as studied by chemical degradative methods. J Neurochem 86:1015–1023
Maker HS, Weiss C, Silides DJ, Cohen G (1981) Coupling of dopamine oxidation (monoamine oxidase activity) to glutathione oxidation via the generation of hydrogen peroxide in rat brain homogenates. J Neurochem 36:589–593
Mori F, Nishie M, Kakita A, Yoshimoto M, Takahashi H, Wakabayashi K (2006) Relationship among alpha-synuclein accumulation, dopamine synthesis, and neurodegeneration in Parkinson disease substantia nigra. J Neuropathol Exp Neurol 65:808–815
Stokes AH, Hastings TG, Vrana KE (1999) Cytotoxic and genotoxic potential of dopamine. J Neurosci Res 55:659–665
Spencer JP, Jenner P, Daniel SE, Lees AJ, Marsden DC, Halliwell B (1998) Conjugates of catecholamines with cysteine and GSH in Parkinson’s disease: possible mechanisms of formation involving reactive oxygen species. J Neurochem 71:2112–2122
Zoccarato F, Toscano P, Alexandre A (2005) Dopamine-derived dopaminochrome promotes H(2) O(2) release at mitochondrial complex I: stimulation by rotenone, control by Ca(2+), and relevance to Parkinson disease. J Biol Chem 280:15587–15594
Bindoli A, Deeble DJ, Rigobello MP, Galzigna L (1990) Direct and respiratory chain-mediated redox cycling of adrenochrome. Biochim Biophys Acta 1016:349–356
Schapira AH (2008) Mitochondria in the aetiology and pathogenesis of Parkinson’s disease. Lancet Neurol 7:97–109
Schapira AH, Cooper JM, Dexter D, Jenner P, Clark JB, Marsden CD (1989) Mitochondrial complex I deficiency in Parkinson’s disease. Lancet 1:1269
Bindoff LA, Birch-Machin M, Cartlidge NE, Parker WD Jr, Turnbull DM (1989) Mitochondrial function in Parkinson’s disease. Lancet 2:49
Parker WD Jr, Boyson SJ, Parks JK (1989) Abnormalities of the electron transport chain in idiopathic Parkinson’s disease. Ann Neurol 26:719–723
Mizuno Y, Ohta S, Tanaka M, Takamiya S, Suzuki K, Sato T, Oya H, Ozawa T, Kagawa Y (1989) Deficiencies in complex I subunits of the respiratory chain in Parkinson’s disease. Biochem Biophys Res Commun 163:1450–1455
Keeney PM, Xie J, Capaldi RA, Bennett JP Jr (2006) Parkinson’s disease brain mitochondrial complex I has oxidatively damaged subunits and is functionally impaired and misassembled. J Neurosci 26:5256–5264
Beal MF (2005) Mitochondria take center stage in aging and neurodegeneration. Ann Neurol 58:495–505
Swerdlow RH, Parks JK, Miller SW, Tuttle JB, Trimmer PA, Sheehan JP, Bennett JP Jr, Davis RE, Parker WD Jr (1996) Origin and functional consequences of the complex I defect in Parkinson’s disease. Ann Neurol 40:663–671
Hartmann A, Michel PP, Troadec JD, Mouatt-Prigent A, Faucheux BA, Ruberg M, Agid Y, Hirsch EC (2001) Is Bax a mitochondrial mediator in apoptotic death of dopaminergic neurons in Parkinson’s disease? J Neurochem 76:1785–1793
von Bohlen Und Halbach O (2004) Synucleins and their relationship to Parkinson’s disease. Cell Tissue Res 318:163–174
Bogaerts V, Theuns J, van BC (2008) Genetic findings in Parkinson’s disease and translation into treatment: a leading role for mitochondria? Genes Brain Behav 7:129–151
Swerdlow RH, Parks JK, Cassarino DS, Binder DR, Bennett JP Jr, Di IG, Golbe LI, Parker WD Jr (2001) Biochemical analysis of cybrids expressing mitochondrial DNA from Contursi kindred Parkinson’s subjects. Exp Neurol 169:479–485
Trimmer PA, Borland MK, Keeney PM, Bennett JP Jr, Parker WD Jr (2004) Parkinson’s disease transgenic mitochondrial cybrids generate Lewy inclusion bodies. J Neurochem 88:800–812
Pankratz N, Foroud T (2007) Genetics of Parkinson disease. Genet Med 9:801–811
Bender A, Krishnan KJ, Morris CM, Taylor GA, Reeve AK, Perry RH, Jaros E, Hersheson JS, Betts J, Klopstock T, Taylor RW, Turnbull DM (2006) High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat Genet 38:515–517
Kraytsberg Y, Kudryavtseva E, McKee AC, Geula C, Kowall NW, Khrapko K (2006) Mitochondrial DNA deletions are abundant and cause functional impairment in aged human substantia nigra neurons. Nat Genet 38:518–520
Langston JW, Ballard P, Tetrud JW, Irwin I (1983) Chronic Parkinsonism in humans due to a product of meperidine-analog synthesis. Science 219:979–980
Richardson JR, Caudle WM, Guillot TS, Watson JL, Nakamaru-Ogiso E, Seo BB, Sherer TB, Greenamyre JT, Yagi T, Matsuno-Yagi A, Miller GW (2007) Obligatory role for complex I inhibition in the dopaminergic neurotoxicity of 1-methyl–4-phenyl–1, 2, 3, 6-tetrahydropyridine (MPTP). Toxicol Sci 95:196–204
Krueger MJ, Singer TP, Casida JE, Ramsay RR (1990) Evidence that the blockade of mitochondrial respiration by the neurotoxin 1-methyl–4-phenylpyridinium (MPP +) involves binding at the same site as the respiratory inhibitor, rotenone. Biochem Biophys Res Commun 169:123–128
Betarbet R, Sherer TB, MacKenzie G, Garcia-Osuna M, Panov AV, Greenamyre JT (2000) Chronic systemic pesticide exposure reproduces features of Parkinson’s disease. Nat Neurosci 3:1301–1306
Sherer TB, Kim JH, Betarbet R, Greenamyre JT (2003) Subcutaneous rotenone exposure causes highly selective dopaminergic degeneration and alpha-synuclein aggregation. Exp Neurol 179:9–16
Liu Y, Yang H (2005) Environmental toxins and alpha-synuclein in Parkinson’s disease. Mol Neurobiol 31:273–282
Panov A, Dikalov S, Shalbuyeva N, Taylor G, Sherer T, Greenamyre JT (2005) Rotenone model of Parkinson disease: multiple brain mitochondria dysfunctions after short term systemic rotenone intoxication. J Biol Chem 280:42026–42035
Betarbet R, Canet-Aviles RM, Sherer TB, Mastroberardino PG, McLendon C, Kim JH, Lund S, Na HM, Taylor G, Bence NF, Kopito R, Seo BB, Yagi T, Yagi A, Klinefelter G, Cookson MR, Greenamyre JT (2006) Intersecting pathways to neurodegeneration in Parkinson’s disease: effects of the pesticide rotenone on DJ–1, alpha-synuclein, and the ubiquitin-proteasome system. Neurobiol Dis 22:404–420
Kitada T, Asakawa S, Hattori N, Matsumine H, Yamamura Y, Minoshima S, Yokochi M, Mizuno Y, Shimizu N (1998) Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392:605–608
Tofaris GK, Spillantini MG (2007) Physiological and pathological properties of alpha-synuclein. Cell Mol Life Sci 64:2194–2201
Conway KA, Rochet JC, Bieganski RM, Lansbury PT Jr (2001) Kinetic stabilization of the alpha-synuclein protofibril by a dopamine-alpha-synuclein adduct. Science 294:1346–1349
Bonifati V, Rizzu P, van Baren MJ, Schaap O, Breedveld GJ, Krieger E, Dekker MC, Squitieri F, Ibanez P, Joosse M, van Dongen JW, Vanacore N, van Swieten JC, Brice A, Meco G, van Duijn CM, Oostra BA, Heutink P (2003) Mutations in the DJ–1 gene associated with autosomal recessive early-onset parkinsonism. Science 299:256–259
Valente EM, bou-Sleiman PM, Caputo V, Muqit MM, Harvey K, Gispert S, Ali Z, Del TD, Bentivoglio AR, Healy DG, Albanese A, Nussbaum R, Gonzalez-Maldonado R, Deller T, Salvi S, Cortelli P, Gilks WP, Latchman DS, Harvey RJ, Dallapiccola B, Auburger G, Wood NW (2004) Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 304:1158–1160
Devi L, Raghavendran V, Prabhu BM, Avadhani NG, Anandatheerthavarada HK (2008) Mitochondrial import and accumulation of {alpha}-synuclein impair complex I in human dopaminergic neuronal cultures and Parkinson disease brain. J Biol Chem 283:9089–9100
Abou-Sleiman PM, Muqit MM, Wood NW (2006) Expanding insights of mitochondrial dysfunction in Parkinson’s disease. Nat Rev Neurosci 7(3):207–219
Sherer TB, Betarbet R, Testa CM, Seo BB, Richardson JR, Kim JH, Miller GW, Yagi T, Matsuno-Yagi A, Greenamyre JT (2003) Mechanism of toxicity in rotenone models of Parkinson’s disease. J Neurosci 23:10756–10764
Tretter L, Sipos I, dam-Vizi V (2004) Initiation of neuronal damage by complex I deficiency and oxidative stress in Parkinson’s disease. Neurochem Res 29:569–577
Acknowledgements
The experimental work from our laboratory, which has been reported in this paper, was supported by MIUR-Rome (Italy).
Author information
Authors and Affiliations
Corresponding author
Additional information
Special issue article in honor of Dr. Anna Maria Giuffrida-Stella.
Rights and permissions
About this article
Cite this article
Fato, R., Bergamini, C., Leoni, S. et al. Generation of Reactive Oxygen Species by Mitochondrial Complex I: Implications in Neurodegeneration. Neurochem Res 33, 2487–2501 (2008). https://doi.org/10.1007/s11064-008-9747-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-008-9747-0