Abstract
DNA methylation and histone deacetylation are two epigenetic mechanisms involved in the lack of estrogen receptor (ER) expression. Our previous studies demonstrated that mutant p53 along with repression complex proteins including DNMT1, HDAC1 and MeCP2 is associated with ER-negative promoter in MDA-MB-468 cells. To elucidate the molecular mechanism of estrogen receptor 1 (ESR1) gene silencing in these cells, we down-regulated DNMT1 and HDAC1 expression using siRNAs and studied the ability of DNMT1, HDAC1, MeCP2 and p53 in binding to ESR1 promoter CpG island. Our results showed that DNMT1 or HDAC1 down-regulation disassembled the repression complex proteins and mutant p53 from ER-negative promoter. The partial demethylation of ESR1 promoter and ER re-expression in down-regulated cells supports these findings. In vivo binding studies demonstrated that mutation of p53 protein in this cell line did not affect its binding capacity to DNMT1, HDAC1 and MeCP2 proteins. Our observations suggest that not only histone deacetylase activity of HDAC1 contributes to inactivation of methylated ESR1 gene but also HDAC1 presence on ESR1 promoter is important for assembly of DNMT1 in repression complex. In addition, our data revealed that mutant p53 protein binds to the promoter of ESR1 through direct interaction with HDAC1 and indirect interaction with DNMT1, MeCP2 proteins in the ER-negative MDA-MB-468 cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Estrogen and estrogen receptor (ER)-α play important roles in development and growth of invasive breast cancer [1]. The tumor expression of ER-α is a prognostic factor for hormonal therapies of breast cancer [2]. However, some breast tumors lose ER-α expression during tumor progression or lack ER-α at the time of diagnosis [3]. Many studies have shown that the epigenetic alterations in mammalian cells including hypermethylation of CpG islands within the estrogen receptor 1 (ESR1) promoter are involved in its gene silencing [2].
Over the last decade, many studies have reported that a repression complex that includes DNA methyl transferase family (DNMT1 and DNMT3b), histone deacetylase (HDAC1) and methyl CpG-binding proteins regulates ESR1 promoter transcription in ER-negative breast cancer cells [4, 5]. DNMT1 is the most abundant and catalytically active DNA methyltransferases at silenced ESR1 promoter [6]. It is a primary maintenance methylase which establishes methylation patterns from the parent strand to the daughter during cell division [7]. But, DNMT3b is a de novo DNA methyltransferase [8]. Down-regulation of DNMT1 by antisense oligonucleotides [6] or treatment with the DNA methyltransferase inhibitor, 5-aza-2 deoxycytidine (5-aza-dC), results in re-expression of ER-α protein in ER-negative human breast cancer cells [9]. HDAC1 is involved in histone modification and binds to the N-terminus of DNMT1, thereby forming a transcriptional repression complex which cooperatively initiates and sustains epigenetic gene silencing [10]. In addition, HDAC inhibitors such as trichostatin A (TSA), scriptaid and LBH589 can induce ER-α protein expression [11–13]. In vivo experimental evidence has documented the association of methyl-CpG-binding domain (MBD) proteins such as MeCP2 (methyl-CpG-binding protein) to the ESR1 promoter in ER-negative breast cancer cell lines [5]. MeCP2 has been identified to recruit histone deacetylases (HDAC1 and HDAC2) for gene silencing [14, 15]. It also binds to N-terminal domain of DNMT1 [16, 17].
Several studies have shown that p53 is also involved in ESR1 transcription regulation [18]. p53 is one of the most widely studied tumor suppressor proteins due to its crucial role in apoptosis, senescence or a reversible and protective cell cycle arrest [19]. It binds to ESR1 promoter [20–22] and up-regulates ER-α expression [18, 21]. Since most breast tumors with p53 mutation are ER-negative, it has been suggested that p53 mutations in these tumors may contribute to loss of ER-α expression [18]. Our recent studies have shown that mutant p53 protein interacts with ESR1 promoter CpG island along with DNMT1, HDAC1 and MeCP2 proteins in ER-negative MDA-MB-468 cells [22]. As mutant p53 proteins which are highly expressed in one-third of breast tumors impair chemotherapy responses [19], study of other target proteins might provide novel diagnosis and therapeutic insights.
To elucidate the molecular mechanism of ESR1 gene silencing in ER-negative MDA-MB-468 cells, we down-regulated DNMT1 and HDAC1 expression using siRNAs and studied the ability of DNMT1, HDAC1, MeCP2 and p53 in binding to ESR1 promoter CpG island. In addition, we compared the binding capacity of wild type (WT) p53 in MCF7 cells with mutated one in MDA-MB-468 cells through interaction with DNMT1, HDAC1 and MeCP2 proteins.
Materials and methods
Cell culture and siRNA transfections
The human breast cancer cell lines, MDA-MB-468 cells (ER-negative) and MCF-7 cells (ER-positive) were prepared from national cell bank of Iran (Pasture Institute, Iran). The cell lines were grown in RPMI 1640 (Biosera, UK) supplemented with l-glutamine to 2 mM and 10 % fetal calf serum (Cinagen, Iran). For siRNA transfections, MDA-MB-468 cells at 50–70 % confluence were diluted to the density of 5 × 105 cells and transfected with 10 nM of siRNA by electroporation (975 μF and 220 V in 4 mm cuvettes). The cells were grown and harvested 72 h after transfection. The siRNAs targeting DNMT1 [23] and HDAC1 [24] transcripts were used. The DNMT1 siRNAs sequences were 5′-CGG UGC UCA UGC UUA CAA CTT-3′ (sense), 5′-GUU GUA AGC AUG AGC ACC GTT-3′ (antisense) and the HDAC1 siRNA sense sequence was 5′-AAG CCG GUC AUG UCC AAA GUA TT-3′, antisense sequence was 5′-UAC UUU GGA CAU GAC CGG CUU-3′. A non-silencing siRNA (UUCUCCGAACGUGUCACGUTT), with no known homology to any human gene, was used as the negative control. All siRNAs were synthesized by Eurofins MWG Operon, Germany. The sense and antisense oligonucleotides were annealed to make double-stranded siRNA.
Chromatin immunoprecipitation assay
Chromatin immunoprecipitation experiments were performed as previously described, using 1 × 106 cells equivalent of chromatin for each immunoprecipitation [22]. Five micrograms of anti-p53 (ab17990; Abcam, USA), anti-DNMT1 (ab13537; Abcam, USA), anti-HDAC1 (ab51846; Abcam, USA), and anti-MeCP2 (ab2828; Abcam, USA) polyclonal antibodies were applied. In addition, anti-histone H3 (ab1791; Abcam, USA) antibody was used as a positive control. The immunoprecipitated DNA from test and total DNA of input samples were purified and subjected to PCR analysis for the presence of ESR1 gene exon 1 sequence. This region includes a critical NotI site in the ESR1 gene, the region where it is methylated in multiple ER-negative breast cancer cell lines. ESR1 gene is unmethylated at the NotI site in the CpG island in all ER-positive cell lines [4]. PCR mixture contained 2 μl of the purified input or test DNA, 0.5 μM of each primer (Cinnagen, Iran), 1.5 mM MgCl2, 0.2 mM deoxynucleotide triphosphate mixture (Fermentas, Iran), 1× PCR buffer and 1.25 U of Taq DNA polymerase (Fermentas, Iran) in a total volume of 25 μl. The primers 5′-TGA ACC GTC CGC AGC TCA AGA TC-3′ and 5′-GTC TGA CCG TAG ACC TGC GCG TTG-3′ were used for ESR1 gene analysis. PCR condition was as previously described [22], 30 cycles of 94 °C for 30 s, 56 °C for 30 s and 72 °C for 50 s, and a final extension of 10 min at 72 °C. The PCR products (150 bp) were electrophoresed on 1.5 % agarose gel and analyzed by ethidium bromide staining. All ChIP experiments were performed at least three times.
GST pull-down analysis
The cDNA of WT p53 was sub-cloned from a human pcDNA-p53 (kindly provided by Dr. A. Turnell, University of Birmingham, UK) to a pGEX-4T-1 vector (Amersham Biosciences, UK) for bacterial expression. Glutathione S-transferase (GST) fusion protein was expressed and purified as described previously [25]. The purified proteins were dialyzed against a buffer containing 50 mM Tris–HCl, (pH 7.4), 0.15 M NaCl, 1 mM dithiothreitol, and 10 % (vol/vol) glycerol. The MCF7 and MDA-MB-468 cell lysates were prepared by solubilization in a lysis buffer containing 10 mM Tris–HCl pH 7.4, 0.825 M NaCl and 1 % NP-40 and then clarified by sonication and centrifugation. For GST pull-downs, 50 μg of GST-p53 or GST proteins was mixed with 5 mg of either the MCF7 or MDA-MB-468 cell lysates for 2 h at 4 °C. The pull-downed complexes with GST and GST-fusion proteins were eluted from the glutathione-agarose by 25 mM reduced glutathione in 50 mM Tris–HCl solution pH 8.0. The samples were resolved by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and then subjected to Western blot analysis with appropriate antibody.
Immunoprecipitation of proteins
The cells were lysed in buffer containing 10 mM Tris–HCl pH 7.4, 0.825 M NaCl and 1 % NP-40. Cell lysates were sonicated and cleared by centrifugation. Immunocomplexes were precipitated from 5 mg of protein lysate, using 5 μg of appropriate primary antibody. After 2 h mixing at 4 °C on a rotator, 30 μl of packed protein G agarose beads (Sigma, UK) was added into the protein-antibody complex and mixed for a further 1 h. Immunocomplexes bound to the beads were then spun and washed, prior to re-suspending in the SDS sample buffer for SDS-PAGE analysis.
Western blotting
Immunoprecipitates or GST pull-down samples with 50 μg proteins of appropriate lysates were electrophoresed on 8 or 12 % SDS-PAGE gels for detection of DNMT1 or ER-α, β-actin, MeCP2 and HDAC1 proteins. Following electrophoresis, cell lysates were transferred to a nitrocellulose membrane (Amersham Biosciences, USA) using the standard protocol. Immunoblotting was performed with anti-DNMT1 (ab13537; Abcam, USA), anti-HDAC1 (ab51846; Abcam, USA), anti-MeCP2 (ab2828; Abcam, USA), anti-ER-α (ab32063; Abcam, USA), anti-p53 (ab17990; Abcam, USA), and anti-β-actin (ab1801; Abcam, USA). A horseradish peroxidase-conjugated anti-rabbit or anti-mouse secondary antibody (Sigma, USA) and chemiluminescence substrates (ECL; Amersham Bioscience AB) were used to detect the immuno-labeled bands.
DNA extraction and methylation-specific PCR (MSP)
Genomic DNA was extracted by the standard phenol/chloroform procedure. DNA from MCF-7 and MDA-MB-468 cells as well as siRNAs transfected MDA-MB-468 cells were treated with sodium bisulfate according to a previously published protocol [26]. Methylation status of bisulfite-modified DNA using primers encompassing nucleotides +44 to +236 bp within the ESR1 CpG island was characterized by PCR as described previously (25). The following primers were used: ESR1 unmethylated (U) 5′-TTT TGG GAT TGT ATT TGT TTT TGT TG-3′ and 5′-AAA CAA AAT ACA AAC CAT ATC CCC A-3′; ESR1 methylated (M) 5′-TTT TGG GAT TGT ATT TGT TTT CGT C-3′ and 5′-AAC AAA ATA CAA ACC GTA TCC CCG-3′ [27]. The amplified products were loaded onto a 2 % agarose gel and visualized under UV illumination. The PCR for those samples demonstrating methylation was repeated to confirm the reproducibility of the results.
Results
Down-regulation of DNMT1 or HDAC1 expression induced ER-α expression
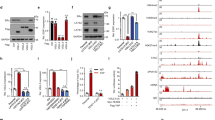
The ER-negative cell line MDA-MB-468 was used as a cell model to investigate whether knock-down of either DNMT1 or HDAC1 expression by RNAi could reactivate ER-α expression. As shown before, 10 mM siRNA against DNMT1 could down-regulate DNMT1 expression and re-express ER-α mRNA [22]. Our Western blot analysis also confirmed that DNMT1 siRNA treatment of MDA-MB-468 cells led to increased ER-α protein expression (Fig. 1a, b). Again, 10 mM siRNA against HDAC1 not only down-regulated HDAC1 expression but also reactivated ER-α protein expression (Fig. 1a, b).
Requirement for DNMT1 and HDAC1 in ESR1 repression. a Down-regulation of DNMT1 or HDAC1 expression. MDA-MB-468 Cell line was transfected with 10 mM of siRNA against DNMT1 or HDAC1 and their protein expression was analyzed by Western blotting. β-Actin was probed as a protein loading control. b Effect of DNMT1 or HDAC1 knock-down upon ER protein expression. Whole cell lysates from siRNA transfected cells were analyzed for ER-α protein expression by Western blotting. MCF7 was used as a positive control for ER expression. β-Actin expression was used as a protein loading control. Non-sil. non-silencing RNA oligonucleotides, DNMT1i and HDAC1i refer to situations where DNMT1 or HDAC1 expression was ablated by siRNAs
Down-regulation of DNMT1 or HDAC1 expression alters the association of DNMT1, HDAC1 and MeCP2 with ESR1 promoter.
We next determined the effect of down-regulation of DNMT1 or HDAC1 expression on the binding of DNMT1, HDAC1 and MeCP2 proteins to ESR1 promoter. We already showed that DNMT1, HDAC1 and MeCP2 proteins are associated with the silenced ESR1 promoter in MDA-MB-468 breast cancer cell line, whereas they were not observed on active ESR1 promoter in MCF7 cells [22]. As shown in Fig. 2a, ChIP analysis with anti-HDAC1 and anti-MeCP2 antibodies revealed that down-regulation of DNMT1 expression abolished the association of HDAC1 and MeCP2 with the ESR1 promoter. Thus, recruitment of HDAC1 and MeCP2 to methylated ESR1 promoter was not observed upon demethylation of the ESR1 promoter.
Dissociation of DNMT1, HDAC1 and MeCP2 proteins from ESR1 promoter after down-regulation of DNMT1 or HDAC1. ChIP analysis was used for immunoprecipitation of Cross-linked chromatin-protein complexes with indicated antibodies in ER-negative MDA-MB-468 human breast cancer cells. The immunoprecipitates were analyzed by PCR for ESR1 promoter CpG island. Negative controls had no antibody; aliquots of chromatin taken before immunoprecipitation were used as input; DNMT1i and HDAC1i refer to situations where DNMT1 or HDAC1 expression was ablated by siRNAs. Data are representative of three independent experiments
In addition, ChIP assays with anti-DNMT1 and anti-MeCP2 antibodies showed that down-regulation of HDAC1 expression also prevents the association of these proteins with the ESR1 promoter (Fig. 2b). Therefore, deacetylation of chromatin dissociates DNMT1 and MeCP2 proteins from silent ESR1 promoter.
Methylation status of ESR1 gene is changed by siRNAs against DNMT1 or HDAC1
To show whether reactivation of ESR1 by knocking-down of DNMT1 or HDAC1 expression is associated with demethylation of the promoter, methylation-specific PCR (MSP) analysis was performed on transfected MDA-MB-468 cells with siRNAs against DNMT1 or HDAC1 transcript. As shown in Fig. 3, the methylated ESR1 gene promoter was partially demethylated by down-regulation of DNMT1 or HDAC1 expression. The MSP method is a non-quantitative method and it can be seen from Fig. 3 that the non-silenced MDA-MB-468 cells have only methylated band, but down regulation of either DNMT1 or HDAC1 expression made the unmethylated band in addition to the methylated one. MSP analysis was applied using primers encompassing nucleotides +44 to +236 bp related to the exon 1 of ESR1 gene. The CpG island in this region has been shown to be methylated in MDA-MB-468 cells [4]. These results suggest that down-regulation of DNMT1 or HDAC1 expression activates silenced ESR1 gene through DNA demethylation and histone deacetylation in the ESR1 CpG island.
ESR1 gene methylation status after siRNA treatment. MSP analysis of ESR1 CpG island was performed after siRNAs treatments in MDA-MB-468 cells. Normal tissue (N) and ER-positive MCF7 cells line were used as an unmethylated control, in vitro-methylated DNA (IVD) and non-silencing RNA oligonucleotides (Non-sil) treated MDA-MB-468 cells were used as methylated control. U Unmethylated, M methylated, H 2 O without DNA, DNMT1i and HDAC1i refer to situations where DNMT1 or HDAC1 expression was ablated by siRNAs
Mutant p53 associates with ESR1 promoter in the presence of DNMT1 and HDAC1
We showed that DNMT1 and HDAC1 proteins are important in repression of ESR1 promoter in ER-negative MDA-MB-468 cell line. To elucidate the role of mutant p53 in repression complex on ESR1 promoter [22], we performed ChIP assays with antibody against p53 in knocking down either DNMT1 or HDAC1 cell lines. Immunoprecipitated complex was examined for ESR1 gene region in treated and untreated cells by PCR. As shown in Fig. 4, mutant p53 was only found to be associated with the ESR1 promoter on the untreated MDA-MB-468 cells. Thus, mutant p53 did not interact with ESR1 promoter in the absence of DNMT1 or HDAC1 protein in ER-negative MDA-MB-468 cell line.
Effect of DNMT1 or HDAC1down-regulation upon mutant p53 binding to ESR1 promoter. MDA-MB-468 cells were treated with siRNAs against DNMT1 and HDAC1. Cross-linked chromatin-protein complex was immunoprecipitated by ChIP with antibodies indicated. ESR1 promoter CpG island was detected by PCR. Negative controls had no antibody; aliquots of chromatin taken before immunoprecipitation were used as the input; DNMT1i and HDAC1i refer to situations where DNMT1 or HDAC1 expression was ablated by siRNAs. Data are representative of three independent experiments
Mutant and WT p53 associate to DNMT1, HDAC1 and MeCP2 in MDA-MB-468 and MCF7 cells
We further tested the ability of mutant p53 for interaction with partner proteins in repression complex of ESR1 promoters. Immunoprecipitation of mutant p53 from MDA-MB-468 cells was compared to WT p53 in MCF7 cells. Western blotting for bound p53 revealed that p53 mutant similar to WT p53 could bind to DNMT1, HDAC1 and MeCP2 in vivo (Fig. 5a). In vivo binding studies presented within this report suggested that mutation at codon 273 of p53 protein did not affect its binding capacity to DNMT1, HDAC1 and MeCP2 proteins.
Interaction between p53 and DNMT1, HDAC1 and MeCP2 in vitro and in vivo. a Immunoprecipitation of p53 in vivo. p53 complexes were immunoprecipitated from MCF7 and MDA-MB-468 cellular lysates with anti-DNMT1 and anti-p53. The coexistence of p53 with DNMT1, HDAC1 and MeCP2 in the immunocomplex was revealed by Western analysis with anti-p53, anti-HDAC1 and anti-MeCP2. b Physical interaction between GST-p53 and DNMT1, HDAC1 and MeCP2. GST-p53 was used to pull-down protein partners in MCF7 and MDA-MB-468 cell extracts. Western blot analysis with anti-DNMT1, anti-HDAC1 and anti-MeCP2 was performed.WCE whole-cell extract, I·P. immunoprecipitante
To investigate the in vitro binding capacity of the WT p53 for DNMT1, HDAC1 and MeCP2 proteins, GST-p53 fusion protein was expressed and purified as described in Materials and Methods section. The results from GST-pull down experiments indicated that the GST-p53 fusion protein retained its binding capacity for HDAC1 but could not bind to DNMT1 and MeCP2 in lysates of MDA-MB-468 and MCF7 cell lines (Fig. 5b).
Discussion
In breast cancer, it has been shown that loss of ER-α expression is as a result of the epigenetic changes of CpG island in ESR1 promoter and its first exon regions [4]. Many studies have documented that aberrant methylation of ESR1 CpG island is occurred during breast cancer progression [28, 29]. In vivo experiments have shown that the ESR1 promoter is silenced by recruitment of repression complex proteins including DNMT1, HDAC1 and MeCP2 proteins in MDA-MB-231 and MDA-MB-468 breast cancer cells [5]. Histone deacetylation and DNA demethylation of ER-negative promoter have been used as potential targets in order to re-expression of ER-α protein which is necessary for response to endocrine therapies in treatment of breast cancers [30]. The molecular mechanism which regulates gene silencing of ESR1 gene and reactivates this gene in ER-negative breast cancers is not understood in detail. In this study, MDA-MB-468 cell line which was previously shown to have densely methylated ESR1 CpG island [4, 27] and has a hemizygous mutant p53 [31] was used as a model for study of the epigenetic regulation of ESR1 promoter. To elucidate if DNMT1 and HDAC1 binding to ER-negative promoter are associated together or act separately, we down-regulated DNMT1 and HDAC1 transcripts in MDA-MB-468 cell line. ChIP results indicated that down-regulation of DNMT1 in MDA-MB-468 cells dissociated the repression complex proteins including HDAC1 and MeCP2 at CpG island on ESR1 promoter. Indeed, depletion of DNMT1 via siRNA not only contributed to DNA partial demethylation but also decreased recruitment of HDAC1 and MeCP2 to methylated ESR1 promoter. This result was consistent with previous report showing that MDA-MB-231 cells treatment with 5-aza-dC reduced the level of association of DNMT1 and MeCP2 with the ER-negative promoter and decreased the DNMT1 protein expression in MDA-MB-231 cell line [5]. We also obtained the same results in the case of using RNAi-mediated HDAC1 knockdown through depletion of DNMT1 and MeCP2 binding to ESR1 promoter. Therefore, deacetylation of chromatin dissociates these proteins from the silent ESR1 promoter. In contrast, TSA treatment of MDA-MB-231 cells was demonstrated to reduce only MeCP2 binding to the ESR1 promoter in MDA-MB-231 cells [5].
In addition, our data showed that down-regulation of each of these two proteins resulted in re-expression of ER-α protein by loss of DNA hypermethylation. Partially methylated pattern in RNAi-mediated DNMT1 or HDAC1 cells may be due to contribution of DNMT3a and 3b to the methylation status of the entire human genome [32]. 5-aza-dC or TSA treatment of ER-negative breast cancer cells could lead to partial demethylation and without alteration in the methylation status of the ESR1 CpG island, respectively [5, 33]. Our results confirm pervious findings that DNMT1 and HDAC1 proteins play a functional role in silencing of ER-α expression in ER-negative breast cancer cells [5]. Our observations suggest that there is a connection between DNMT1 and HDAC1 at CpG island of ER-negative promoter. In addition, our studies showed that not only histone deacetylase activity of HDAC1 contributes to inactivation of methylated ESR1 gene but also HDAC1 presence on ESR1 promoter is important for assembly of DNMT1 in repression complex. HDAC1 is a critical component of ESR1 gene silencing in order to inactivate chromatin structure [5]. It binds to the DNMT1 [14, 16] and MeCP2 at CpG island via a corepressor, mSin3A [34] to sustain its epigenetic silencing. Therefore, we suggest that binding of HDAC1 to DNMT1 is as important as its histone deacetylase activity on ER-negative promoter in MDA-MB-468 breast cancer cells.
Our previous report demonstrated that mutant p53 protein in MDA-MB-468 cells is associated with repressor complex including DNMT1, HDAC1 and MeCP2 proteins on silenced ESR1 promoter [22]. Many studies have shown that WT p53 is involved in ER-α transcriptional regulation in MCF7 cell line [18, 20, 21]. The WT p53 up-regulates ER-α expression and is associated with activator complex on ESR1 promoter in ER-positive cell lines [18, 21]. The WT p53 mediates its effects on target genes regulation as both activator and repressor [35, 36]. The role of mutant p53 on silenced ESR1 promoter in MDA-MB-468 cells is not yet established. We showed that mutant p53 was not able to bind ESR1 promoter in the absence of DNMT1 and HDAC1 in MDA-MB-468 cells. It implies that mutant p53 in MDA-MB-468 cell line binds to this promoter through interaction with these proteins. WT p53 binds to DNMT1 through amino acids 323–393 [37] and the HDAC1 [38]. To elucidate how mutant p53 protein contributes to the repression complex on ER-negative promoter, we examined the ability of mutant p53 for interaction with partner proteins in repression complex of ESR1 promoters in ER-negative cell line in comparison to the WT p53 in MCF7 cells. In vivo binding studies suggested that mutation of p53 protein in MDA-MB-468 cells did not affect its binding capacity to studied proteins. In addition, mutant p53 may stabilize the DNMT1-MeCP2-HDAC1-p53 complex on silenced ESR1 promoter, by the ability of direct interaction with HDAC1 and indirect interaction with DNMT1 and MeCP2 proteins. In vitro studies confirmed that GST-p53 has a strong binding capacity for HDAC1 in extracts of MDA-MB-468 and MCF7 cells. It has been reported that interaction of p53 and HDAC1 deacetylates p53 and, therefore, down regulates its transcriptional activity [38]. Since previous report showed that DNMT1 binds physically to GST-p53 [37], the low level of this protein in our study may be the reason for our in vitro result. While there has been no report on p53 interaction with MeCP2 so far, our data suggested that either the post-translational modification of p53 or an associated protein affect the p53 association with MeCP2 in vivo.
In response to cellular stress and DNA damage, WT p53 is stabilized and accumulated in cells [19]. Thus, it is able to repress some p53-responsive genes upon association with their promoters along with other repressor complex proteins including DNMT1, HDAC1 [37, 39]. It has been reported that WT p53 knockdown induced DNMT1 and HDAC1 proteins as well as increased promoter methylation of some p53-activator genes [40]. On the other hand, mutant p53 may have a dominant negative or a gain of function effect in tumorigenesis [19]. It has been demonstrated that mutant p53 in MDA-MB-468 cells has an active function in mediating the survival of breast cancer cells [41]. Another study also showed that over-expression of mutant p53 is associated with silencing of CCN5 gene during progression of cancer [42]. Therefore, p53 either by post-translational modification in normal pathways or by mutation in cancer cells is able to bind to repression complex proteins and repress gene expression. These results indicate that specific p53 mutations may stabilize the repression complex on ER-negative promoter and contribute to loss of estrogen receptor α expression in breast tumors and that mutant p53 is likely to impact DNA methylation.
It has been shown that ESR1 promoter is silenced by recruitment of MBD1, MBD2, DNMT3b as well as the MeCP2, DNMT1 and HDAC1 in MDA-MB-231 cells [5]. WT p53 is able to bind to Sp1 (Sephacryl and phosphocellulose purification 1) and impairs its transactivator function in some promoters [43]. However, Sp1 was reported to be recruited to ESR1 promoter to up-regulate the ER-α expression in a regulator complex with CARM1, CBP, c-jun and WT p53 in MCF7 cells [21]. The effects of these proteins in binding of mutant p53 to ER-negative promoter in MDA-MB-468 cells should be more elucidated in future studies.
In conclusion, our results confirm pervious findings indicating that DNMT1 and HDAC1 play a functional role in silencing of ER-α expression in ER-negative breast cancer cells. We also showed that the binding of HDAC1 to ER-negative promoter is as important as its histone deacetylase activity for ESR1 gene repression. In addition, our data revealed that mutant p53 protein binds to the promoter of ESR1 through association with DNMT1, HDAC1 and MeCP2 proteins in the ER-negative MDA-MB-468 cells. Binding of mutant p53 to ESR1 promoter of the MDA-MB-468 cells may stabilize the repression complex proteins on ER-negative promoter.
References
Sui M, Zhang H, Fan W (2011) The role of estrogen and estrogen receptors in chemoresistance. Curr Med Chem 18:4674–4683
Giacinti L, Claudio PP, Lopez M, Giordano A (2006) Epigenetic information and estrogen receptor alpha expression in breast cancer. Oncologist 11:1–8
Lo PK, Sukumar S (2008) Epigenomics and breast cancer. Pharmacogenomics 9:1879–1902
Yan L, Yang X, Davidson NE (2001) Role of DNA methylation and histone acetylation in steroid receptor expression in breast cancer. J Mammary Gland Biol Neoplasia 6:183–192
Sharma D, Blum J, Yang X et al (2005) Release of methyl CpG binding proteins and histone deacetylase 1 from the Estrogen receptor alpha (ER) promoter upon reactivation in ER-negative human breast cancer cells. Mol Endocrinol 19:1740–1751
Yan L, Nass SJ, Smith D et al (2003) Specific inhibition of DNMT1 by antisense oligonucleotides induces re-expression of estrogen receptor-alpha (ER) in ER-negative human breast cancer cell lines. Cancer Biol Ther 2:552–556
Tajima S, Suetake I (1998) Regulation and function of DNA methylation in vertebrates. J Biochem 123:993–999
Chedin F, Lieber MR, Hsieh CL (2002) The DNA methyltransferase-like protein DNMT3L stimulates de novo methylation by Dnmt3a. Proc Natl Acad Sci USA 99:16916–16921
Ferguson AT, Lapidus RG, Baylin SB, Davidson NE (1995) Demethylation of the estrogen receptor gene in estrogen receptor-negative breast cancer cells can reactivate estrogen receptor gene expression. Cancer Res 55:2279–2283
Dobosy JR, Selker EU (2001) Emerging connections between DNA methylation and histone acetylation. Cell Mol Life Sci 58:721–727
Yang X, Ferguson AT, Nass SJ et al (2000) Transcriptional activation of estrogen receptor alpha in human breast cancer cells by histone deacetylase inhibition. Cancer Res 60:6890–6894
Keen JC, Yan L, Mack KM et al (2003) A novel histone deacetylase inhibitor, scriptaid, enhances expression of functional estrogen receptor alpha (ER) in ER negative human breast cancer cells in combination with 5-aza 2′-deoxycytidine. Breast Cancer Res Treat 81:177–186
Zhou Q, Atadja P, Davidson NE (2007) Histone deacetylase inhibitor LBH589 reactivates silenced estrogen receptor alpha (ER) gene expression without loss of DNA hypermethylation. Cancer Biol Ther 6:64–69
Robertson KD, Ait-Si-Ali S, Yokochi T et al (2000) DNMT1 forms a complex with Rb, E2F1 and HDAC1 and represses transcription from E2F-responsive promoters. Nat Genet 25:338–342
Rountree MR, Bachman KE, Baylin SB (2000) DNMT1 binds HDAC2 and a new co-repressor, DMAP1, to form a complex at replication foci. Nat Genet 25:269–277
Qin W, Leonhardt H, Pichler G (2011) Regulation of DNA methyltransferase 1 by interactions and modifications. Nucleus 2:392–402
Kimura H, Shiota K (2003) Methyl-CpG-binding protein, MeCP2, is a target molecule for maintenance DNA methyltransferase, Dnmt1. J Biol Chem 278:4806–4812
Angeloni SV, Martin MB, Garcia-Morales P et al (2004) Regulation of estrogen receptor-alpha expression by the tumor suppressor gene p53 in MCF-7 cells. J Endocrinol 180:497–504
Goh AM, Coffill CR, Lane DP (2011) The role of mutant p53 in human cancer. J Pathol 223:116–126
Akaogi K, Nakajima Y, Ito I et al (2009) KLF4 suppresses estrogen-dependent breast cancer growth by inhibiting the transcriptional activity of ERalpha. Oncogene 28:2894–2902
Shirley SH, Rundhaug JE, Tian J et al (2009) Transcriptional regulation of estrogen receptor-alpha by p53 in human breast cancer cells. Cancer Res 69:3405–3414
Rasti M, Arabsolghar R, Khatooni Z, Mostafavi-Pour Z (2012) p53 Binds to estrogen receptor 1 promoter in human breast cancer cells. Pathol Oncol Res 18:169–175
Suzuki M, Sunaga N, Shames DS et al (2004) RNA interference-mediated knockdown of DNA methyltransferase 1 leads to promoter demethylation and gene re-expression in human lung and breast cancer cells. Cancer Res 64:3137–3143
Keshelava N, Davicioni E, Wan Z et al (2007) Histone deacetylase 1 gene expression and sensitization of multidrug-resistant neuroblastoma cell lines to cytotoxic agents by depsipeptide. J Natl Cancer Inst 99:1107–1119
Rasti M, Grand RJ, Mymryk JS et al (2005) Recruitment of CBP/p300, TATA-binding protein, and S8 to distinct regions at the N terminus of adenovirus E1A. J Virol 79:5594–5605
Rasti M, Tavasoli P, Monabati A, Entezam M (2009) Association between HIC1 and RASSF1A promoter hypermethylation with MTHFD1 G1958A polymorphism and clinicopathological features of breast cancer in Iranian patients. Iran Biomed J 13:199–206
Lapidus RG, Nass SJ, Butash KA et al (1998) Mapping of ER gene CpG island methylation-specific polymerase chain reaction. Cancer Res 58:2515–2519
Hoque MO, Prencipe M, Poeta ML et al (2009) Changes in CpG islands promoter methylation patterns during ductal breast carcinoma progression. Cancer Epidemiol Biomarkers Prev 18:2694–2700
Zhao L, Wang L, Jin F et al (2009) Silencing of estrogen receptor alpha (ERalpha) gene by promoter hypermethylation is a frequent event in Chinese women with sporadic breast cancer. Breast Cancer Res Treat 117:253–259
Saxena NK, Sharma D (2010) Epigenetic reactivation of estrogen receptor: promising tools for restoring response to endocrine therapy. Mol Cell Pharmacol 2:191–202
Vinyals A, Peinado MA, Gonzalez-Garrigues M et al (1999) Failure of wild-type p53 gene therapy in human cancer cells expressing a mutant p53 protein. Gene Ther 6:22–33
Okano M, Bell DW, Haber DA, Li E (1999) DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99:247–257
Yang X, Phillips DL, Ferguson AT et al (2001) Synergistic activation of functional estrogen receptor (ER)-alpha by DNA methyltransferase and histone deacetylase inhibition in human ER-alpha-negative breast cancer cells. Cancer Res 61:7025–7029
Ng HH, Bird A (1999) DNA methylation and chromatin modification. Curr Opin Genet Dev 9:158–163
Harms K, Nozell S, Chen X (2004) The common and distinct target genes of the p53 family transcription factors. Cell Mol Life Sci 61:822–842
Ho JS, Ma W, Mao DY, Benchimol S (2005) p53-Dependent transcriptional repression of c-myc is required for G1 cell cycle arrest. Mol Cell Biol 25:7423–7431
Esteve PO, Chin HG, Pradhan S (2005) Human maintenance DNA (cytosine-5)-methyltransferase and p53 modulate expression of p53-repressed promoters. Proc Natl Acad Sci USA 102:1000–1005
Juan LJ, Shia WJ, Chen MH et al (2000) Histone deacetylases specifically down-regulate p53-dependent gene activation. J Biol Chem 275:20436–20443
Le Gac G, Esteve PO, Ferec C, Pradhan S (2006) DNA damage-induced down-regulation of human Cdc25C and Cdc2 is mediated by cooperation between p53 and maintenance DNA (cytosine-5) methyltransferase 1. J Biol Chem 281:24161–24170
Lai JC, Cheng YW, Goan YG et al (2008) Promoter methylation of O(6)-methylguanine-DNA-methyltransferase in lung cancer is regulated by p53. DNA Repair (Amst) 7:1352–1363
Lim LY, Vidnovic N, Ellisen LW, Leong CO (2009) Mutant p53 mediates survival of breast cancer cells. Br J Cancer 101:1606–1612
Dhar G, Banerjee S, Dhar K et al (2008) Gain of oncogenic function of p53 mutants induces invasive phenotypes in human breast cancer cells by silencing CCN5/WISP-2. Cancer Res 68:4580–4587
Böhlig L, Rother K (2011) One function-multiple mechanisms: the manifold activities of p53 as a transcriptional repressor. J Biomed Biotechnol 2011:464916
Acknowledgments
This work was supported by Grants No: 89-5176 and 90-5620 from the Shiraz University of Medical Sciences, Iran for Ms Rita Arabsolghar’s thesis and Ms Tayebe Azimi’s thesis, respectively. The authors would like to thank the School of Advanced Medical Sciences and Technologies for their support in performing this project.
Conflict of interest
None of the authors has any conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Rita Arabsolghar and Tayebeh Azimi contributed equally to this study and both are considered first author.
Rights and permissions
About this article
Cite this article
Arabsolghar, R., Azimi, T. & Rasti, M. Mutant p53 binds to estrogen receptor negative promoter via DNMT1 and HDAC1 in MDA-MB-468 breast cancer cells. Mol Biol Rep 40, 2617–2625 (2013). https://doi.org/10.1007/s11033-012-2348-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2348-7