Abstract
p53 is a tumor suppressor protein that regulates estrogen receptor 1 (ESR1) expression. To investigate the mechanism of ESR1 gene regulation by p53, chromatin immunoprecipitation was applied to assess the binding of p53, DNMT1, HDAC1 and MeCP2 to both silenced ESR1 promoter in MDA-MB-468 cells and active ESR1 promoter in MCF-7 breast cancer cells. The results of chromatin immunoprecipitation experiments showed that p53 protein binds to both unmethylated CpG island of the ESR1 promoter in the ER-positive MCF-7 and the hypermethylated ESR1 promoter in the ER-negative MDA-MB-468 cells. However, repression complex including DNMT1, HDAC1 and MeCP2 is only associated with silenced ESR1 in ER-negative MDA-MB-468 human breast cancer cells. In addition, ectopically expressed wild type p53 failed to reactivate the ESR1 gene in these cells. These results suggest that specific p53 mutations may contribute to loss of estrogen receptor α expression in breast tumors and also support the hypothesis that mutant p53 is likely to impact DNA methylation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a tumor suppressor protein, p53 functions are based on its ability to up- or downregulate the expression of many genes involved in cell growth, cell cycle progression, DNA repair, cellular senescence, autophagy, metabolism and p53 regulation [1, 2]. These functions are essential in preventing malignant transformation of cells. p53 predominantly functions as a sequence-specific DNA-binding transcription factor to activate transcription following the appropriate signals [3, 4]. However, it can also repress gene expression indirectly by association with histone deacetylases or the basal transcriptional machinery and interference with transcriptional initiation [5, 6]. Inactivating mutations in the p53 gene allows the cells to evade pro-apoptotic signals, thus promoting tumorigenesis [7]. Mutations in the p53 gene most often occur in its sequence-specific DNA-binding region [8].
One of the genes which are regulated by p53 is the estrogen receptor 1 (ESR1) gene [9]. ESR1 transcriptional regulation is poorly defined. It has been shown that p53 upregulates, ESR1 gene expression by increasing its transcription [9]. Numerous studies have demonstrated that hypermethylation of CpG islands located in the promoter regions of tumor suppressor genes including ESR1 is an important mechanism for gene inactivation in breast cancer [10–12]. Gene encoding estrogen receptor α (ERα) in humans (designated ESR1) is involved in cell signaling [12]. Therefore, CpG island methylation of ESR1 gene promoter disrupts its cell growth regulatory effect in breast cancers. ESR1 hypermethylation occurs early in tumorogenesis that results in disruption of pathways that may predispose the cells to malignant transformation [13]. It is known that both promoter methylation and histone deacetylation is required to reach estrogen receptor silencing. Several studies have demonstrated the role of DNA methyltransferase 1 (DNMT1), histone deacetylase 1 (HDAC1) and certain methyl binding proteins to the methylated CpG dinucleotides such as MeCP2 in the repression of ESR1 transcription in ER- negative breast cancer cell lines [14–16]. Intriguingly, most of the breast tumors with p53 mutations are ER- negative [17]. These studies suggest that specific p53 mutations may play a role in the progression of breast cancer from ER-positive to ER-negative and hormone-independent tumors. Loss of ERα expression in ER-negative tumors means that breast cancer cells cannot be stopped by hormone therapy which results in aggressive tumor and poor prognosis [18]. How breast cancer cells with p53 mutation acquire such epigenetic changing of ESR1 promoter is not clear.
In this study, we examined the association of p53, DNMT1, HDAC1 and MeCP2 proteins with ESR1 promoter in two breast cancer cell lines, MDA-MB-468 with and MCF-7 without methylated ESR1 promoter. Then, we studied whether ectopically expressed wild type p53 is capable of reactivating the ESR1 gene in ER-negative MDA-MB-468 cells.
Materials and Methods
Cell Culture
The ER- negative MDA-MB-468 cells and ER-positive MCF-7 cells were obtained from national cell bank of Iran (Pasture Institute, Iran). The human breast cancer cell lines, MDA-MB-468 and MCF-7, were grown in RPMI 1640 (Biosera, UK) supplemented with L-glutamine to 2 mM and 10% fetal calf serum (Cinagen, Iran) and subcultured 2–3 times weekly.
Chromatin Immunoprecipitation (Chip) Assay
ChIP assays were performed on MDA-MB-468 and MCF-7 cells using the ChIP kit (ab500; Abcam, Canada). 1 × 106 cells were grown on 10 Cm plates and formaldehyde was added directly to culture medium to a final concentration of 1% and incubated for 10 min at 37°C to cross-link histones to DNA. The reaction was stopped by adding glycine (0.125 M). After washing and lysing cells as instructed by kit manufacture, mixtures were sonicated on ice three times for 10 s each to shear genomic DNA to an optimal size of 0.2–1 kb. The cell debris was pelleted by centrifugation (14000 × g for 5 min at 4°C). The supernatants were diluted in ChIP dilution buffer and supplemented with protease inhibitors according to the manufacturer’s instructions. ChIPs were carried out overnight at 4°C with rotation using 5 μg of anti-p53 (ab17990; Abcam, Canada), anti-DNMT1 (ab13537; Abcam, Canada), anti-HDAC1 (ab51846; Abcam, Canada) and anti-MeCP2 (ab2828; Abcam, Canada) polyclonal antibodies. Anti-histone H3 (ab1791; Abcam, Canada) antibody was used as a positive control. The antibody/DNA-histone complex was collected with 42 μl of protein A-agarose beads for 2 h. After washing the beads, DNA was decross-linked and purified according to the manufacturer’s recommendation. Immunoprecipitated DNA was analyzed by PCR for the presence of ESR1 gene exon 1 sequence. PCR reactions were carried out in a total volume of 25 μl containing 2 μl of immunoprecipitated or total input, 0.5 μM of each primer (Cinnagen, Iran), 1.5 mM MgCl2, 0.2 mM deoxynucleotid triphosphate mixture (Fermentas, Iran), 1× PCR buffer and 1.25 U of Taq DNA polymerase (Fermentas, Iran). PCR primers were blasted for ESR1 gene (GenBank accession number NC_000006) amplified a region encompassing the NotI site within the ESR1 CpG island at ±311 bp relative to transcription start codon: 5'- TGA ACC GTC CGC AGC TCA AGA TC-3' and 5'-GTC TGA CCG TAG ACC TGC GCG TTG-3' [16]. Amplification of this gene was performed under the following condition: 30 cycles of 94°C for 30 s, 56°C for 30 s and 72°C for 50 s, and a final extension of 10 min at 72°C. The PCR product of 150 bp was electrophoresed on 1.5% agarose gel and analyzed by ethidium bromide staining. All ChIP assays were performed at least twice with similar results.
Western Blot Analyses
The MCF7 and MDA-MB-468 cells which had been solubilized in a lysis buffer containing 10 mM Tris–HCl pH 7.4, 0.825 M NaCl and 1% NP-40 were sonicated and cleared by centrifugation. Fifty-microgram protein samples including the cell lysate proteins were electrophoresed on 12% SDS–PAGE gel and electrophoretically transferred to a nitrocellulose membrane (Amersham Biosciences, US) following the standard protocol. Immunoblotting was performed with anti-p53 antibody (ab17990; Abcam, Canada), anti-DNMT1 (ab13537; Abcam, Canada) and anti-actin (ab1801; Abcam, Canada). A horseradish peroxidase-conjugated anti-rabbit or anti-mouse secondary antibody (Sigma, USA) and chemiluminescence substrates (ECL; Amersham Bioscience AB) were used to detect the immuno-labeled bands.
Plasmid and siRNA Transfections and Reverse Transcription-PCR (RT-PCR) Analysis
A human pcDNA-p53 was kindly provided by Dr. A. Turnell (University of Birmingham, UK). The plasmid contains the full-length coding region of the wild type p53 cDNA (1.3 kb, exons 2–11). MDA-MB-468 cells were seeded at a density of 1 × 106, trasfected with 1 μg of pcDNA3 vector or pcDNA3-p53 vector using lipofectamine (Invitrogen, UK); 48 h after transfection cells were harvested. The presence of exogenous p53 cDNA and its integrity were confirmed by direct sequencing of both strands of the p53 cDNA in total DNA extract by the use of an ABI Prism 3100 genetic analyzer.
siRNA targeting DNMT1 [19] was used. The siRNA sequences were 5′-CGGUGCUCAUGCUUACAACT-3′ (sence) and 5′- GUUGUAAGCAUGAGCACCGTT-3′ (antisence). A non-silencing siRNA (UUCUCCGAACGUGUCACGUTT), with no known homology to any human gene, was used as a negative control. All siRNAs were obtained from Eurofins MWG Operon, Germany. The sence and antisence oligonucleotides were annealed to make double-stranded siRNA. MDA-MB-468 cells (5 × 105) were transfected with 10 nM of siRNAs by electroporation (975 μF and 220 V in 4 mm cuvettes). The cells were grown and harvested 72 h after transfection. Our previous studies revealed that siRNA transfection by electroporation was more efficient than lipofectamine (data not shown). To ascertain the specificity of siRNA, the target protein was monitored by Western blotting.
Total RNA was extracted using the RNXTM plus solution (Fermetas, Iran), according to the manufacturer’s instructions. Three microgram of each RNA were retrotrascribed using RevertAid TM First-Strand cDNA Synthesis Kit using the oligo-d (T) primers (Fermetas, Iran). Semi quantitative PCRs were performed after normalizing all the cDNAs for beta actin control. RT-PCR primers that amplify between exons 7 and 8 of ESR1 (5′-GCTGCTGGCTACATCATC-3′, 5′-AGGACTCGGTGGATATGG-3′) and beta actin (5′-AGTCTTCCTTCCTGGGCAT-3′, 5′-CAGGAGGAGCAATGATCT-3′) were designed. The annealing temperatures used for the ER and actin primers were 60°C and 64°C, respectively. The PCR product of 150 bp was subjected to electrophoresis on 1.5% agarose gel.
Results
p53 is Associated with ESR1 Promoter cpG Island along with DNMT1, HDAC1 and MeCP2 in MDA-MB-468 Breast Cancer Cells
The ER- negative MDA-MB-468 cells with a hypermethylated ESR1 promoter and the ER-positive MCF-7 cells with an unmethylated ESR1 promoter [20] were used in ChIP assays. Anti-p53, anti-DNMT1, anti-HDAC1 and anti-MeCP2 antibodies were used for immunoprecipitation of formaldehyde cross-linked protein-chromatin complexes from MCF-7 and MDA-MB-468 cells. In parallel, anti-histone H3 antibody which is enriched at chromatin was used as a positive control. Immunoprecipitated DNA was analyzed for the presence of the ESR1 gene by PCR using primers spanning a CpG island in its first exon region, where it has been revealed that its methylation is most closely associated with ESR1 gene expression [14, 21]. Figure 1 shows that p53 is associated with both the silenced ESR1 promoter in MDA-MB-468 cells and active ESR1 promoter in MCF-7 breast cancer cells. Histone H3 which was used as positive control was observed at the ESR1 promoter in both MCF-7 and MDA-MB-468 cells. To gain a deeper understanding of the binding proteins along with p53 protein to CpG island of ESR1 gene, the recruitment of binding proteins that recognize methylated DNA was investigated. As shown in Fig. 1, ChIP experiments with anti-DNMT1, anti-HDAC1 and anti-MeCP2 antibodies showed that those proteins are associated with the silenced ER promoter along with p53 in MDA-MB-468 cells whereas the active ER promoter in MCF-7 cells is not associated with these proteins.
ChIP analysis for the presence of p53, DNMT1, HDAC1, MeCP2 and histone-H3 on the human ESR1 CpG island. Cross-linked chromatin-protein complexes from ER-positive MCF-7 and ER-negative MDA-MB-468 human breast cancer cells were immunoprecipitated with antibodies indicated. The immunoprecipitates were analyzed by PCR for ESR1 promoter CpG island. Negative controls had no antibody; aliquots of chromatin taken before immunopercipitation were used as input controls. Data are representative of three independent experiments
Evaluation of the p53 in MCF-7 and MDA-MB- 468 Cells and Expression of Wild Type p53 in MDA-MB-468 Cells
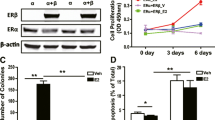
Lysates of MCF-7 and MDA-MB-468 cells were used for immunoblotting for detection of p53, using anti-p53 antibody (Fig. 2a). A band of approximately 53 kD was observed in MDA-MB-468 cells (Fig. 2a). However, p53 protein was not recognized in MCF-7 cells (Fig. 2a). MDA-MB-468 breast cancer cells contain a hemizygous mutant p53 gene that overexpresses a mutant p53 protein [22]. But, MCF-7 human breast cancer cells have normal p53 function. Since MCF-7 cells express wild type (wt) p53, the low amount of p53 proteins in these cells is difficult to measure.
Endogenous p53 expression and effect of ectopically expressed wt p53 in MDA-MB-468 cells. a Evaluation of the p53 protein in MCF-7 and MDA-MB- 468 cells by western blot analysis. 50 μg of whole cell lysates from MCF7 and MDA-MB-468 cells were separated by 12% SDS-PAGE and subjected to Western blot analysis with ab17990. b Effect of wt p53 expression on ER expression. Transfections were done with 1 μg of empty and p53 expression vectors and RNA was prepared at 48 h. RNA extracted from MCF-7 cells was used as an ER positive control. RT-PCR analysis of ESR1 gene and β-Actin internal control was performed
Given the importance of the significant role of tumor suppressor function of p53, in this study, we examined that ectopically expressed wt p53 is capable of activating the ESR1 gene in MDA-MB-468 cells which are hemizygous for a mutant p53 [22]. The full-length coding region of human wt p53 inserted into pcDNA3 and empty vector as a negative control were used for transfection. The cells were examined for ER mRNA expression by RT-PCR; β-actin was used as internal standard. As shown in Fig. 2b, wt p53 did not induce ESR1 gene expression in this cell line as compared with the negative control. This finding indicates that the exogenous wt p53 did not impair the suppression complex on ESR1 gene promoter in this cell line.
siRNA against DNMT1 Reactivates the ESR1 Gene in MDA-MB-468 Cells
To study whether demethylation induces the ESR1 gene in ER negative breast cancer cells, MDA-MB-468 cells were treated with the siRNA against DNMT1 and expression status of ESR1 gene was analyzed by RT-PCR; β-actin was used as internal standard (Fig. 3). Upon introduction of DNMT1 siRNA, DNMT1 proteins get reduced, resulting in functional analysis of this protein in MDA-MB-468 cells. Cell extracts were monitored for DNMT1 expression by using Western blot analysis (Fig. 3a). As shown in Fig. 3b, RNAi-mediated endogenous DNMT1 knockdown restored the expression of ESR1 gene. This result confirmed pervious findings [14, 21] that DNA methylation of the ESR1 gene plays a functional role in silencing of ER expression in ER-negative breast cancer cells.
Effect of DNMT1 siRNA on DNMT1and ER expression in MDA-MB-468 cells. a Western blot analysis of DNMT1 protein in siRNAs treated cells. 50 μg of whole cell lysates from non-silencing or DNMT1 siRNA trasfected cells were separated by 8% SDS-PAGE and subjected to Western blot analysis with anti-DNMT1. β-Actin was probed as a protein loading control. b ER reexpression is induced by DNMT1 siRNA. RT- PCR analysis showed ER mRNA reexpression after treatment with DNMT1 siRNA. RNA extracted from MCF-7 cells was used as an ER positive control. RT-PCR analysis of ESR1 gene and β-Actin internal control was performed. Non-sil., non-silencing RNA oligonucleotides; DNMT1i refers to a situation in which DNMT1 expression was ablated by siRNA
We next determined how the levels of ER reactivation in MDA-MB-468 cells compared with endogenous ER expression in the ER positive MCF7 cells. DNMT1 siRNA induced ER transcripts in MDA-MB-468 cells to 22% of the expression level found in MCF7 cells. It shows that the effect of RNAi-mediated DNMT1 knockdown on the expression of ESR1 gene is more than that of using DNMT1 inhibitors such as 5′-aza-2′-deoxycytidine which only induced ER mRNA in MDA-MB-231 cells to 4% of the MCF7 expression levels [23].
Discussion
In an attempt to better understanding of the epigenetic events on ESR1 promoter, the binding of p53, DNMT1, HDAC1 and MeCP2 proteins to this promoter in the ER- negative MDA-MB-468 cells with a hypermethylated ESR1 promoter and the ER-positive MCF-7 cells with an unmethylated ESR1 promoter [20] were examined. Our ChIP data showed that p53 protein binds to ESR1 promoter CpG island in both the ER-positive MCF-7 and the ER- negative MDA-MB-468 cells. However, DNMT1, HDAC1 and MeCP2 proteins are associated with the silenced ER promoter along with p53 in MDA-MB-468 cells. Western blot analyses revealed that MDA-MB-468 breast cancer cells overexpress the p53 protein, but MCF-7 breast cancer cells express wt p53. To our knowledge, this is the first report on the association of p53 with silent ESR1 promoter in MDA-MB-468 cell line.
ESR1 promoter is regulated by epigenetic modification and its epigenetic lesion by aberrant methylation is of key importance in the development of breast cancer [10–12]. It has been reported that hypermethylation of CpG island of tumor suppressor genes is not a random event [13] and it may occur by loss of specific protection factor of the CpG islands [24]. p53 is a regulatory protein and elucidates its biological functions as both transcription factor [3, 4] and repressor [25–28]. The ability of p53 to regulate gene expression is accompanied by binding to DNA either directly [29, 30] or by its interaction with other transcription factors [31]. One of the genes regulated by p53 is the ESR1 gene [9]. It has been shown that p53 upregulates the expression of this gene in human breast cancer cell line MCF-7. However, it has been demonstrated that p53 ability to upregulate ESR1 gene expression is not dependent on its binding directly to DNA but on interaction with other proteins on ESR1 promoter [9]. These results suggest that specific p53 mutations in breast tumors may contribute to loss of ERα expression and progression of breast cancer to an aggressive tumor [9]. In addition, it has been reported that p53 binds to ESR1 promoter and inhibits [32] as well as maintains the transcriptional activity of ESR1 gene in MCF7 cells [33]. A recent experiment has shown that p53 binds to ESR1 promoter along with CARM1, CBP, c-jun and Sp1 in MCF7 cells and upregulates its expression [33]. Our ChIP experiments confirmed that p53 protein binds to the ESR1 promoter CpG island in the ER-positive MCF-7 cells. However, we found that this protein also binds to ESR1 promoter CpG island along with DNMT1, HDAC1 and MeCP2 proteins in the ER- negative MDA-MB-468 cells. This region includes a critical NotI site in the ESR1 gene, the region where it is methylated in multiple ER- negative breast cancer cell lines. ESR1 gene is unmethylated at the NotI site in the CpG island in all ER-positive cell lines [21]. Therefore, the NotI site methylation in the ESR1 gene is most closely associated with ERα gene expression [14]. MCF-7 cells have normal p53 function and are the ER-positive cells with an unmethylated ESR1 promoter. However, MDA-MB-468 breast cancer cells are hemizygous for p53 gene, containing a single point mutation at codon 273 in the remaining allele and overexpressing a transcribtionaly active mutant p53 protein [22]. These cells are the ER- negative cells with a hypermethylated ESR1 promoter. We showed that the silenced ER promoter in ER-negative MDA-MB-468 cells has a repressive chromatin structure associated with DNMT1, HDAC1 and MeCP2 proteins. Our results are consistent with in vivo studies demonstrating that the ESR1 promoter is silenced by recruitment of DNMT1, HDAC1 and MeCP2 proteins in MDA-MB-231 breast cancer cells [14]. These findings also showed that mutation of p53 in MDA-MB-468 cells does not prevent its binding to ESR1 promoter. However, p53 mutation may change its normal function on this promoter in favor of making different complexes with proteins that result in epigenetic changes on ESR1 promoter. p53 binding to DNMT1 through amino acids 323–393 [34] would not be affected by p53 mutation in this cell line. Mutation of p53 is common in sporadic breast cancer [35]. Intriguingly, most of the breast tumors with p53 mutation are ER-negative [17] and positive for ESR1 methylation [36]. But, how p53 mutation can play a role in the progression of breast cancer from ER-positive to ER-negative is unknown. It had been previously shown that both p53 mutation and aberrant DNA methylation silenced gene expression through independent, but interrelated, mechanisms of transcriptional control [37]. In addition, other studies demonstrated that overexpression of mutated p53 is associated with the silencing of CCN5 gene during progression of cancer [38]. Our data support these findings and hypothesize that ESR1 is one of the genes which are affected by p53 mutation and mutant p53 is likely to impact DNA methylation. However, in an attempt to correct p53 function in MDA-MB-468 cells, ectopically expressed wt p53 did not reactivate ESR1 gene. It has been also reported that the mutant p53 proteins inhibit wt p53 functions [39]. Nevertheless, further studies are demanded to elucidate the molecular mechanism of p53 mutation in ESR1 promoter.
It has been reported that treatment of ER-negative cells with inhibitors of DNMT1 and HDAC1 enzymes results in dissociation of these proteins from the ESR1 promoter and reactivation of this gene [14]. We also demonstrated that RNAi-mediated DNMT1 knockdown restored the expression of ESR1 gene. This result confirms that DNA methylation of the ESR1 gene plays a functional role in silencing of ER expression in ER-negative breast cancer cells.
The role of wt p53 in controlling apoptosis in response to genotoxic stress is essential in breast cancer chemotherapy [40]. It has been demonstrated that down–regulation of mutant p53 in MDA-MB-468 cells inhibits cell proliferation [41]. However, introduction of wt p53 into these cells did not result in cellular apoptosis [22]. While mutant p53 proteins which are highly expressed in one third of breast tumors impair the therapeutic response [39], study of other target proteins might provide novel diagnosis and therapeutic insights.
In conclusion, we revealed that p53 protein binds to the promoter of ESR1 in the ER-positive MCF-7 and along with DNMT1, HDAC1 and MeCP2 proteins in the ER- negative MDA-MB-468 cells. Binding of p53 to ESR1 promoter of the ER- negative MDA-MB-468 cells can be explained by the fact that mutant p53 protein gains a new function in regulation of ESR1 promoter. This should be more elucidated in future works.
Abbreviations
- ESR1 :
-
estrogen receptor 1
- DNMT1:
-
DNA methyltransferase 1
- HDAC1:
-
histone deacetylase 1
- MeCP2:
-
methyl-CpG-binding protein
- ER:
-
estrogen receptor
- ChIP:
-
Chromatin immunoprecipitation
- RT-PCR:
-
reverse transcription-PCR
- siRNA:
-
small interfering RNA
References
Millau JF, Bastien N, Drouin R (2009) p53 transcriptional activities: a general overview and some thoughts. Mutat Res 681:118–133
Olsson A, Manzl C, Strasser A et al (2007) How important are post-translational modifications in p53 for selectivity in target-gene transcription and tumour suppression? Cell Death Differ 14:1561–1575
Harms K, Nozell S, Chen X (2004) The common and distinct target genes of the p53 family transcription factors. Cell Mol Life Sci 61:822–842
Helton ES, Chen X (2007) p53 modulation of the DNA damage response. J Cell Biochem 100:883–896
Murphy M, Ahn J, Walker KK et al (1999) Transcriptional repression by wild-type p53 utilizes histone deacetylases, mediated by interaction with mSin3a. Genes Dev 13:2490–2501
Seto E, Usheva A, Zambetti GP et al (1992) Wild-type p53 binds to the TATA-binding protein and represses transcription. Proc Natl Acad Sci U S A 89:12028–12032
Amaral JD, Xavier JM, Steer CJ et al (2010) The role of p53 in apoptosis. Discov Med 9:145–152
Olivier M, Eeles R, Hollstein M et al (2002) The IARC TP53 database: new online mutation analysis and recommendations to users. Hum Mutat 19:607–614
Angeloni SV, Martin MB, Garcia-Morales P et al (2004) Regulation of estrogen receptor-alpha expression by the tumor suppressor gene p53 in MCF-7 cells. J Endocrinol 180:497–504
Luczak MW, Jagodzinski PP (2006) The role of DNA methylation in cancer development. Folia Histochem Cytobiol 44:143–154
Miremadi A, Oestergaard MZ, Pharoah PD, et al. (2007) Cancer genetics of epigenetic genes. Hum Mol Genet 16 Spec No 1:R28-R49.
Parrella P, Poeta ML, Gallo AP et al (2004) Nonrandom distribution of aberrant promoter methylation of cancer-related genes in sporadic breast tumors. Clin Cancer Res 10:5349–5354
Costello JF, Fruhwald MC, Smiraglia DJ et al (2000) Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat Genet 24:132–138
Sharma D, Blum J, Yang X et al (2005) Release of methyl CpG binding proteins and histone deacetylase 1 from the Estrogen receptor alpha (ER) promoter upon reactivation in ER-negative human breast cancer cells. Mol Endocrinol 19:1740–1751
Yan L, Nass SJ, Smith D et al (2003) Specific inhibition of DNMT1 by antisense oligonucleotides induces re-expression of estrogen receptor-alpha (ER) in ER-negative human breast cancer cell lines. Cancer Biol Ther 2:552–556
Yang X, Ferguson AT, Nass SJ et al (2000) Transcriptional activation of estrogen receptor alpha in human breast cancer cells by histone deacetylase inhibition. Cancer Res 60:6890–6894
Berns EM, Klijn JG, Smid M et al (1996) TP53 and MYC gene alterations independently predict poor prognosis in breast cancer patients. Genes Chromosomes Cancer 16:170–179
Giacinti L, Claudio PP, Lopez M et al (2006) Epigenetic information and estrogen receptor alpha expression in breast cancer. Oncologist 11:1–8
Suzuki M, Sunaga N, Shames DS et al (2004) RNA interference-mediated knockdown of DNA methyltransferase 1 leads to promoter demethylation and gene re-expression in human lung and breast cancer cells. Cancer Res 64:3137–3143
Lapidus RG, Nass SJ, Butash KA et al (1998) Mapping of ER gene CpG island methylation-specific polymerase chain reaction. Cancer Res 58:2515–2519
Yan L, Yang X, Davidson NE (2001) Role of DNA methylation and histone acetylation in steroid receptor expression in breast cancer. J Mammary Gland Biol Neoplasia 6:183–192
Vinyals A, Peinado MA, Gonzalez-Garrigues M et al (1999) Failure of wild-type p53 gene therapy in human cancer cells expressing a mutant p53 protein. Gene Ther 6:22–33
Yang X, Phillips DL, Ferguson AT et al (2001) Synergistic activation of functional estrogen receptor (ER)-alpha by DNA methyltransferase and histone deacetylase inhibition in human ER-alpha-negative breast cancer cells. Cancer Res 61:7025–7029
Esteller M (2002) CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene 21:5427–5440
Ho JS, Ma W, Mao DY et al (2005) p53-Dependent transcriptional repression of c-myc is required for G1 cell cycle arrest. Mol Cell Biol 25:7423–7431
Imbriano C, Gurtner A, Cocchiarella F et al (2005) Direct p53 transcriptional repression: in vivo analysis of CCAAT-containing G2/M promoters. Mol Cell Biol 25:3737–3751
Scoumanne A, Chen X (2006) The epithelial cell transforming sequence 2, a guanine nucleotide exchange factor for Rho GTPases, is repressed by p53 via protein methyltransferases and is required for G1-S transition. Cancer Res 66:6271–6279
Sengupta S, Shimamoto A, Koshiji M et al (2005) Tumor suppressor p53 represses transcription of RECQ4 helicase. Oncogene 24:1738–1748
Horvath MM, Wang X, Resnick MA et al (2007) Divergent evolution of human p53 binding sites: cell cycle versus apoptosis. PLoS Genet 3:e127
Wei CL, Wu Q, Vega VB et al (2006) A global map of p53 transcription-factor binding sites in the human genome. Cell 124:207–219
Koutsodontis G, Tentes I, Papakosta P et al (2001) Sp1 plays a critical role in the transcriptional activation of the human cyclin-dependent kinase inhibitor p21(WAF1/Cip1) gene by the p53 tumor suppressor protein. J Biol Chem 276:29116–29125
Akaogi K, Nakajima Y, Ito I et al (2009) KLF4 suppresses estrogen-dependent breast cancer growth by inhibiting the transcriptional activity of ERalpha. Oncogene 28:2894–2902
Shirley SH, Rundhaug JE, Tian J et al (2009) Transcriptional regulation of estrogen receptor-alpha by p53 in human breast cancer cells. Cancer Res 69:3405–3414
Esteve PO, Chin HG, Pradhan S (2005) Human maintenance DNA (cytosine-5)-methyltransferase and p53 modulate expression of p53-repressed promoters. Proc Natl Acad Sci U S A 102:1000–1005
Petitjean A, Mathe E, Kato S et al (2007) Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat 28:622–629
Li SY, Rong M, Iacopetta B (2006) Germ-line variants in methyl-group metabolism genes and susceptibility to DNA methylation in human breast cancer. Oncol Rep 15:221–225
Oshiro MM, Watts GS, Wozniak RJ et al (2003) Mutant p53 and aberrant cytosine methylation cooperate to silence gene expression. Oncogene 22:3624–3634
Dhar G, Banerjee S, Dhar K et al (2008) Gain of oncogenic function of p53 mutants induces invasive phenotypes in human breast cancer cells by silencing CCN5/WISP-2. Cancer Res 68:4580–4587
Goh AM, Coffill CR, Lane DP (2011) The role of mutant p53 in human cancer. J Pathol 223:116–126
Bertheau P, Espie M, Turpin E et al (2008) TP53 status and response to chemotherapy in breast cancer. Pathobiology 75:132–139
Avila MA, Velasco JA, Cansado J et al (1994) Quercetin mediates the down-regulation of mutant p53 in the human breast cancer cell line MDA-MB468. Cancer Res 54:2424–2428
Acknowledgements
This work was supported by grants No: 87–4153 and 89–5176 from Shiraz University of Medical Sciences, Iran for Mr. Zahed Khatooni’s thesis and Mrs Rita Arabsolghar’s thesis, respectively.
Conflict of interest
None of authors has any conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rasti, M., Arabsolghar, R., Khatooni, Z. et al. p53 Binds to Estrogen Receptor 1 Promoter in Human Breast Cancer Cells. Pathol. Oncol. Res. 18, 169–175 (2012). https://doi.org/10.1007/s12253-011-9423-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-011-9423-6