Abstract
microRNAs are short, non-coding RNAs, that exert a posttranscriptional control on protein synthesis by mRNA interference. They are involved in normal and pathological embryologic development, as well as in adult life pathology, from myocardial infarction to cancer. There are several brain-specific species of microRNA, showing time-dependent pattern of expression, selectivity for neuronal population, significant roles in correct cellular differentiation and system development. The growing interest in microRNAs extended also in the area of neurodegeneration, some of brain-restricted microRNAs being reported to associate with disorders such as Alzheimer’s disease, Parkinson’s disease or Huntington’s disease. The microRNAs research in the last 3 years offered a considerable amount of information that needs to be integrated in the vast machinery of cellular biology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
MicroRNAs: what are they?
A short introduction on microRNAs
It is now known that less than 2% of human DNA is translated into proteins and the fraction of protein-coding DNA in the genome decreases with increasing organism complexity. In bacteria, about 90% of the genome codes for proteins, whereas in yeast, nematodes and mammals, the percentage is progressively restricted to 68, 23–24, and 1.5–2%, respectively [1]. Yet more than half of the human genome is transcribed, meaning that most of the transcripts are non-coding RNAs. It is now clear that this genetic information is not a redundant activity, but is rather a way for the cell to regulate its phenotype, as important as posttranslational events of protein synthesis [2]. Non-coding RNAs (ncRNAs) include several species that share only one common feature-the absence of a reading frame to initiate polypeptide synthesis: ribosomal RNA (rRNA), transport RNA (tRNA), small nucleolar RNAs (snoRNAs), small nuclear RNAs (snRNAs), microRNAs (miRNAs) and Piwi-interacting RNAs (piRNAs), some genetic mobile elements (a subset of retrotransposons) and other isolated subspecies of RNAs, unclassified so far. ncRNAs diversity has been demonstrated and accepted only recently and until 1999 only several hundred species were described. However, in the following 5 years, the ncRNA list expanded up to 5,000 species and several new categories were added to formerly known ones [3]. By 2010, the database “miRNABase: the home of microRNA data” (accessible at http://microrna.sanger.ac.uk/sequences/) registered more than 15,000 entries, of which over 1,000 for Homo sapiens [4–6].
The ncRNAs control gene expression through a variety of mechanisms and act in distinct stages of ontogenetic development. The idea that RNAs might be involved in posttranscriptional regulatory mechanisms first rose in the late 1960s, when it was assumed that some RNA types are responsible for the active or inactive state of a gene by direct complementary association [7]. The newly discovered transcription factors unjustly dismissed this theory until few decades later, when it was convincingly argued that RNAs are not just simple intermediaries between genes and proteins. Experiment in plants in the late 1980s and early 1990s reported an occasional protein downregulation, following transfection or induced overexpression of certain genes. These results were further confirmed in Caenorhabditis elegans, by key experiments that proved that double-stranded RNA (dsRNA) triggers messenger RNA (mRNA) destruction and this dsRNA is further converted into smaller species that mediate mRNA silencing. Two studies reported independently, in 1993, the first cloned microRNA–lin-14, a small RNA involved in repressing lin-14 (lin 14) gene expression, gene encoding a protein involved in regulating developmental timing of C. elegans larvar lineages [8, 9]. Lee et al. reported two definitory elements for microRNAs: (i) the increase of transcriptome expression correlates with downregulation of protein expression and (ii) the gene transcript contains base sequences complementary to 3′untranslated region (UTR) of the mRNA of the downregulated protein.
Up-to-date the interest taken in microRNA biology grew almost exponentially, more than 1,800 papers being published only in 2009, bringing new information on their structure and biological function and demonstrating their involvement in different pathological mechanisms.
MicroRNAs biogenesis
MicroRNAs are single stranded RNA molecules (ssRNA) of approximately 22 nucleotides, partially complementary to the 3′UTR of a mRNA. At first it was hypothesized that coding sequences for microRNAs lie between protein and coding genes, but soon after, intronic microRNA were described, located within either coding or noncoding transcriptional units, respecting the transcription pattern of the host gene [10]. Intergenic microRNAs are independent transcription units, with their own transcriptional regulatory elements and many of them are clustered in the genome [11]. Intronic or exonic genes can also be grouped and translated as polycistronic RNA. These functional units usually contain 2 or 3 genes, but may go up to 7 genes [12, 13]. Clustering of microRNAs may have structural semblance or may be functionally related, targeting mRNAs of proteins from the same metabolic chain. In humans, all chromosomes contain microRNAs genes, except the Y chromosome [14].
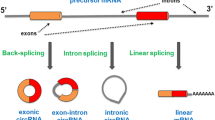
MicroRNAs usually are transcribed as long pri-microRNA precursors, usually several kilobases, that contain stem-loop structures [15], that will be processed in the nucleus by RNase III complex, mainly composed of a processing enzyme (Drosha) and a RNA binding protein (DGCR8/Pasha) [16]. The enzymatic complex interacts with the pri-microRNA at the base of the stem, positioning the enzyme in the correct spatial configuration for asymmetric cleavage. Drosha will sever one dsRNA spiral away from the attachment site (11 bps), releasing a pre-microRNA containing a two nucleotide 3′overhang, essential for nuclear export by exportin 5 (Fig. 1) [17].
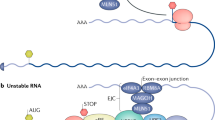
The first step of microRNA biogenesis is the transcription of a long pri-microRNA that contains loop-stem structures. These loop-stem structures will be further cleaved-off from the primary transcript by the RNAse III Drosha, a processing step that also takes place in the nucleus. The cleavage product is called pre-microRNA and has a typical two nucleotide 3′overhang, essential for nuclear export by exportin 5. The next enzymatic processing takes place into the cytoplasm and it is performed by another Rnase III–Dicer, specialized to bind RNA ends, especially duplex ends with short (~2 nt) 3′ overhangs. Dicer releases a 22-nucleotide mature double-stranded microRNA that will be further incorporated into a RNA induced silencing complex (RISC), unwinded by a helicase, and cleaved in two ssRNAs, of which only one will stably associate with RNA binding proteins called Argonaute proteins (Ago). RISC will further interact with 3′UTRs of mRNAs, to either degrade target mRNAs or to inhibit translation and protein synthesis
In the cytoplasm it will be further processed by another Rnase III complex–Dicer, specialized to bind RNA ends, especially duplex ends with short (~2 nt) 3′ overhangs. Dicer releases a 22-nucleotide double-stranded microRNA with a 5′phosphate end and a 2-nucleotide 3′overhang [18]. The dsRNA will be incorporated into a RNA induced scilencing complex (RISC), unwinded by a helicase, and cleaved in two ssRNAs, of which only one will stably associate with RNA binding proteins [19]. The key player of RISC is RNA binding protein Argonaut (Ago), which, in mammals, forms a trimeric complex with Dicer and its RNA binding partner—TRBP. Such an association is capable of rapid generation of mature duplexes from pre-microRNAs and further guidance to target [20]. The microRNA-RISC forms the so-called “effector complex”, or microribonucleoprotein complex (miRNP), responsible for the microRNAs biological actions. MicroRNAs will bind to a specific site of a mRNA, in the 3′UTR region, site usually present in multiple copies. Most microRNAs bind imperfectly to their target, although a key feature of recognition involves Watson–Crick base pairing between the microRNA “seed region”-nucleotides 2–8 and mRNA 3′UTR region. Based on this “seed region”, microRNA targets can be predicted using algorithms, such as TargetScanS algorithm, that identifies conserved Watson–Crick (W–C) matches to microRNA bases 2–7, flanked by either a Watson–Crick match to the m8 position of the microRNA or a conserved adenosine in the t1 position of the target, designated as the t1A anchor [21]. Several computational prediction programs are available for online target searches and the use of bioinformatic analysis revealed that one microRNA may actually interact with hundreds of mRNAs and one mRNA 3′UTR may have multiple binding site for different microRNAs. Pairing prediction cannot be made only by perfect matching of seed region to mRNA 3′UTR and sequences outside the seed area are important to specifying targets or optimizing the silencing [22].
Recently, mature microRNAs were also identified inside cell’s nucleus [23, 24], raising the possibility of epigenetic regulation of gene expression [25].
MicroRNAs roles in normal embryological development
Since the identification of the first microRNA and its role in normal development of C. elegans, it was only logical that interest for this new ncRNA species should focus on embryological timing development and subsequent regulatory (either activatory, either inhibitory) mechanisms. Indeed, many reports confirm the role of microRNAs in normal tissue differentiation, and there are reports of different expression patterns of the same microRNA between embryologic and adult life and also between different tissue types. [26–29]
MicroRNA expression seems to be tightly regulated at each step and varies with the tissue type and ontogenetic period In the same tissue, during embryonic development some species might be expressed at very high levels (up to 10,000 molecules/cell), while others barely detectable, where as in adult life, the report can be reversed [30]. A rough approximate shows that more than 5,000 genes are controlled by microRNAs, with a ratio between microRNA: transcripts of 1:200. The role in cell differentiation is elegantly illustrated by Lim et al. who successfully modified the cell’s faith by microRNAs overexpression. Hence, overexpression or miRNA-214 in HeLa cells modifies the phenotype into a nervous-like one, while miRNA-1 in a muscle-like one [31]. In both cases, approximately 100 microRNAs were repressed, suggesting a tissular profile responsible for the cellular phenotype. To enunciate only a few roles of microRNAs in normal development, one can think at developmental timing and correct larval stages development in invertebrates [8, 9], balance between apoptosis and proliferation [32, 33], fat storage [34], sensorial specialization of nervous structures [35, 36], hematopoietic differentiation [37], stem cells asymmetric division [38, 39] and malignancy [40, 41].
Significant functional data regarding microRNAs function were obtained by Drosha or Dicer knock-out (KO) animal models, at first in C. elegans and later on in insects and mammals.
Drosha KO does not seem to modify the phenotype significantly, therefore most results involved Dicer1 (Dcr1) KO. In Dcr1 KO, offsprings were infertile, so heterozygous mothers were used. The main concern with this model was that there was always a possibility for cytoplasmic transfer from the mother, which would have masked the early phenotype of the KO. Indeed, the Drosophila studies provided a complex phenotype, similar to ones seen in lin-4 and let-7 KO mutants [42]. The zebrafish Dicer KO showed an early phase of microRNAs accumulation that faded in the following days, finally leading to embryo death around day 10, with morphogenesis anomalies during gastrulation, the brain, heart and somatic appearance being the most affected [43]. Dicer1 mutant mice die in utero around day 7, 5 with gastrulation and axis defects. Interestingly, they do not seem to have Oct 4 positive stem cells. Furthermore, Dicer was shown to be absolutely necessary for in vitro embryonic stem cells differentiation. Ago proteins KO models also proved useful in C. elegans leading to severely impaired embryologic development as well [44].
To study a single microRNA, one may use conditional KO techniques in animals and/or tissue analysis at various stages of development. To insure a higher specificity and lack of interference with other Dicer intracellular pathways, the KO models were generated using complementary constructs, either 2′O (ribose)-methyl antisense nucleotides, either locked nucleic acids (LNA). LNA is a modified ribonucleic acid, in which the ribose ring is stiffened by a methylene bridge between 2′O and 4′C. Such experiments offered valuable information on particular microRNA species, for example miRNA-302 was detected only in stem cells and proposed as a novel stem cell marker [42].
Overall, the first studies involving the first microRNAs (lin-4 and let-7), discovered in C. elegans and later on in other species, showed beyond doubt the role of these ncRNAs in correct cellular and tissular differentiation and organogenesis.
MicroRNAs and central nervous system embryological development
Narrowing the field of research to central nervous system (CNS) ontogenetic and phylogenetic development, the same features apply for brain specific microRNA species: time-dependent pattern of expression, selectivity for certain cellular populations, significant roles in correct cellular differentiation and system development. Dicer KO zebrafish shows abnormal CNS morphology and neuronal maturation defects [43]. Dicer mutants mice die in utero, before neurulation [45]. Conditional Dicer KO in cell lines such as dopaminergic neurons, Purkinje cerebellar neurons, cortical neurons or hippocampal neurons offered more information on miRNAs roles in neuronal development and function. They seem to interfere with key regulatory processes such as neuronal stem cells differentiation, neurites outgrowth and synapse formation. In C. elegans, microRNAs are necessary for correct specification and maintenance of taste neurons, in Drosophila, for maintenance of correct differentiation of photoreceptors and in vertebrae, microRNAs are involved in much more complex mechanisms such as synaptic plasticity and learning, acting at synaptic level ([46] and citations herein). It is currently known that long-term potentiation needs local synthesis of key constituents, which implies microRNA transport to designated peripheral locations, in an inactive form as a ribonucleic complex called “neuronal RNA granules” [47]. A subset of these RNAs can be found along the dendrites and in dendritic spines. In synaptosomes were detected both microRNAs and pre-microRNAs, along with Dicer and Dicer-associated proteins. Enzyme activation occurs locally by neuronal stimulation, through calpain-controlled cleavage. Thus, posttranslational control mechanisms seem to be regulated by synaptic transmission [48]. Involvement of microRNAs in synaptic protein synthesis was further proved by study of miRNA-134 in mice hippocampal cultures. miRNA-134 colocalizes at synaptic level with Limk1 (Limdomaincontaining protein kinase 1), a protein involved in actin filaments dynamics and dendritic spine morphology [49]. Limk-1mRNA suppression leads to restriction of dendritic spines formation and, subsequently, excitatory synapses. Interestingly, Limk1 mRNA -miRNA-134 interaction is suppressed by brain-derived neurotrophic factor (BDNF)—a neurotrophin associated with molecular mechanisms of learning [50]. Another microRNA present in the dendritic compartment is miRNA-138, a synaptic plasticity inhibitor by alteration of Acyl protein thioesterase 1 (APT1) -mediated palmitoylation of certain membrane proteins. Although not a direct inhibitor of mRNA translation of α subunit of heterotrimeric G proteins, its microRNA-mediated depalmitoylation is responsible, according to Christensen, for the miRNA-138 phenotype [51].
Several neuronal specific microRNA species were described in CNS embryological development and differentiation studies in mice: miRNA-125, miRNA-128, miRNA-133, miRNA-134, miRNA-388, miRNA-9 and miRNA-124. The last two are especially related to neurogenesis, their overexpression leading to poor astrocytic differentiation in vitro, whereas miRNA-9 suppression or co-suppression of both leads to a reduction in neuronal number. In mice, miRNA-9 is abundantly expressed in cerebral cortex during embryonic life, involved in correct differentiation of Cajal-Retzius cells at hippocampal level, differentiation mediated at least partially by STAT3 phosphorylation to TYR705 [52]. In zebrafish, the same microRNA is involved in delimitation of rombencephalon-mesencephalon boundary, an active zone of neural tube organization and, by contrast with other CNS structures, is absent in this particular area. Its local induction promotes local neurogenesis and loss of zone characteristics by suppression of FGF8, FGFR1 and Her 5 and 9–signaling molecules with inhibitory roles in neuronal differentiation [53]. miRNA-124 is one of the most abundantly expressed microRNAs in the CNS. In both humans and mice, 3 genes coding for miRNA-124 were identified, each on a different chromosome. In mouse, the three genes are translated concomitantly, with overlapping patterns. miRNA-124 can be initially detected in neuronal precursors [54] and persists in terminally differentiated cells and acts by interaction with RE-1 silencing transcription factor (also known as neuron-restrictive silencing factor/RE-1 silencing transcription factor-NRSF/REST) that suppresses neuron-specific genes in non-neuronal cells. Furthermore, miRNA-124 postranslationally represses RNA binding protein PTBR1 that acts as an inhibitor of alternative splicing that generates neuronal-specific transcripts. Besides its role in promoting expression of neuronal-specific genes, it is also involved in neurites outgrowth and neuronal differentiation of embryonic carcinoma in mice, as proven by Yu et al. in P19 cell line, by transient overexpression of neural specific basic helix–loop–helix (bHLH) transcription factors [55].
As for miRNA-125, an enhancement of miRNA-125a and -b expression is observed in neuronal differentiation of embryonic carcinoma. In mammals, target for both miRNA-125s is Lin-28, a protein recently involved in stem reprogramming of somatic cells and also a target of let-7, one of the first microRNAs to be described. A new target of miRNA-125 is Mlin41, the homolog of lin41 of C. elegans, a target gene for let-7. Mice with mutated Mlin41 show neural tube closure defects and embryonic lethality [56].
miRNA-132 interferes with p250GAP mRNA translation, a GTPase-Activating-protein important for neurites outgrowth and neurogenesis. In adult mammalian hippocampus, this mRNA is usually found at very low levels [57].
miRNA-133b controls dopaminergic neurons differentiation, raising the possibility of pathogenic implication in Parkinson’s Disease [58]. Animal models of PD indicated also miRNA-64 and miRNA-65 to be underexpressed in this form of neurodegeneration and thus, one more possible pathogenic link [59].
miRNA-134 is involved in dendritic spines formation by regulating Limk1 protein expression in hippocampal neurons, as previously described.
miRNA-388 is coexpressed during CNS ontogenesis with apoptosis-associated tyrosine kinase (AATK), a kinase essential for neuronal differentiation and neurites outgrowth. miRNA-388 gene is actually an intronic sequence of AATK gene and falls under the regulation of AATK gene expression and are both required for proper differentiation. Furthermore, miRNA-388 acts as a suppressor of neuronal differentiation inhibitors [47].
Interestingly, although some of microRNA species are highly conserved, their expression pattern might be completely different between species, as Bak et al. exemplifies by a comparative study of microRNAs in CNS of zebrafish and mice. A total of 44 species were expressed at three folds, not in generalized manner, but with regional specificity. Although the mature microRNAs had an almost 100% sequence match between the two animal models and the seed region was 100% conserved in 40 out of 44 microRNAs studied, in about 20 cases the expression pattern was divergent. In adult mouse, and correlated with literature results, Bak et al. reported elevated expression of miRNA-9, miRNA-124a, miRNA-125b, miRNA-127, miRNA-128, and let-7 family members. miRNA-195, miRNA-497, and miRNA-30b were elevated in cerebellum, brain stem was characterized by increased levels of miRNA-34a, miRNA-451, miRNA-219, miRNA-338, miRNA-10a, and miRNA-10b and hipothalamus-miRNA-7 and miRNA-7b. Hippocampus was rich in miRNA-218, miRNA-221, miRNA-222, miRNA-26a, miRNA-128a/b, miRNA-138 and let-7c and spinal cord in miRNA-10a and miRNA-10b [60].
MicroRNAs in neurodegeneration
Neurodegeneration implies a vast spectrum of pathologies and ethiopathogenies. Neurodenerative diseases affect different subsets of neurons, but a common risk factor of most pathologies is advanced age. Early on-sets or aggressive evolution are rare and usually genetically determined, in which the mutant allele is expressed in a dominant manner or inherited as a homozygotic gene. The most frequent neurodegenerative disorders are Alzheimer’s (AD), Parkinson’s disease (PD), amyotrophic lateral sclerosis (ALS) and Huntington’s disease (HD). The growing interest in microRNAs extended in the area of neurodegeneration as well, especially following identification of CNS specific species. Until 2009 more than 400 species of microRNAs were identified in the human and chimpanzee brain and it is estimated that the human brain expresses over 1,000 species [61]. Interestingly, Berezikov presents some of these microRNAs as newly acquired phylogenetically, as they do not seem to be conserved between primate families.
Of all brain specific microRNAs, only a few have been so far associated with neurodegeneration (see Table 1): miRNA-133b [62], miRNA-433 [63], miRNA-64, miRNA-65 [59] in PD, miRNA-9 in HD [64, 65], miRNA-146a promotes specific aspects of inflammatory neurodegeneration [66, 67], miRNA-132, miRNA-124a, miRNA-125b, miRNA-107 miRNA-219 and miRNA-128, in AD [68–71].
One of the first reports on altered microRNA brain profile in AD patients belongs to Walter Lukiw, who presented a comparative analysis between fetal, adult and AD hippocampus [68]. He limited his investigation at 13 brain specific microRNAs, expressed in hippocampus of AD patients, as compared to normal fetal and adult hippocampus. His results indicated abnormally high levels of miRNA-9, miRNA-125b and miRNA-128 in AD hippocampus. miRNA-124a was detected at lower levels in AD hippocampus as compared to adult and fetal controls, but this modification in expression did not reach statistical significance (P < 0.05). Lukiw follows the line of investigation and identifies new microRNA species in AD hippocampal specimens as compared to healthy age-matched subjects. The results of his and P.Sethi’s study showed that human neurons use only a limited set of microRNAs, including the ones previously reported (miRNA-9, miRNA-125b, miRNA-132, miRNA-146a, miRNA-183). The short half-life of microRNA makes them very volatile postmortem, predisposing to erroneous data interpretation due to delays in sample prelevation. Taking this aspect into consideration and adjusting the results accordingly, Lukiw and Sethi reported specifically increased levels of miRNA-9, miRNA-125b and miRNA-146a in temporal cortex of AD patients, as compared to PD, ALS or schizophrenia patients [69].
A study by Wang et al. on brain slices of AD patients, as compared to healthy or non-demented patients, showed a statistically significant decrease of miRNA-107, even in early stages. The AD cases were classified in four groups, according to neuropsychiatric assessment and autopsy results: (1) non-demented, without/with few anatomopathological elements of AD; (2) non-demented but with histopathological signs of early AD; (3) MCI with histopathological elements of moderate AD and (4) AD. Approximately 200 microRNAs were modified, but only 70 at statistically significant levels. Only miRNA-107 showed a statistically significant decrease from one group to another, even between the group 1 and 2, making it useful for screening in early stages. Wang et al. proved that the 3′UTR of BACE1 mRNA is a possible target for miRNA-107, linking this miRNA to the ethiopathogeny of AD. Moreover, beside the temporal cortex, miRNA-107 decrease was found in the motor cortex as well, which might be an indication of a generalized altered pattern in affected parts of AD brain [70].
Herbert et al. proved a statistically significant decrease of miRNA-29a/b-1 cluster, complementary to 3′UTR mBACE1 in sporadic AD patients. In agreement with data from literature [75–79], a two to five fold increase of BACE1 in the cortex of the same AD patients was proven, along with normal mRNA BACE1 levels as shown by qRTPCR. The modification of expression is not confined to a specific cortical area, as proven by comparison to cerebellum [72]. During normal fetal development miRNA-29a/b-1 and BACE1 are mutually exclusive, as it is normal for pared microRNA-mRNA. In HeLa both microRNAs were capable of repressing the expression of a luciferase reporter containing the mBACE13′UTR, but not of the control with mutated seed target. After overexpressing both microRNAs in HEK293 cells, the endogenous BACE1 and Aβ peptide synthesis were repressed. In contrast, the miRNA-29a/b-depletion led to Aβ accumulation [46].
Cogswell et al. quantified the expression of over 300 de microRNAs in hippocampus, frontal medial gyrus and cerebellum from AD patients in different stages of the disease. These data showed changes for certain microRNA species, correlating with evolution and localization of pathological lesions. Furthermore, they report microRNA detection in cerebrospinal fluid, with altered levels as compared to healthy controls. Some of the microRNAs reported by Cogswell’s group have been reported before and include miRNA-9 and miRNA-132, whereas others such as miRNA-29a and miRNA-29b showed contradictory results, being increased in frontal medial gyrus and miRNA-415 increases in hippocampus and frontal medial gyrus [80].
Alterations in microRNAs expression are not necessarily restricted to one pathogenic or metabolic chain. The upregulation of certain microRNA might not be the cause but the result of certain metabolic abnormality. For example, in AD brain, accumulation of Aβ 42 (the end result of amyloid abnormal processing) is a weak stimulant of toll receptors [81]. In some non-neuronal cell lines, Toll stimulation by LPS has led to miRNA-132 and miRNA-146b accumulation [82], whereas the depletion of miRNA-132 reported by Cogswell et al. in AD brain would correlate, under these circumstances, with an immune response deficit. Also known to be related to innate immune and inflammatory reactions, miRNA-146a was recently reported to be increased in the neocortex of AD patients versus age-matched control brains, increase that correlated positively and statistically significant with the stage of the disease [71]. Lukiw’s group observations were confirmed in AD transgenic animal models and recreated in vitro, in a neuronal-glial coculture, used to establish a direct relationship between miRNA-146a expression level and NF-kB activation. Interestingly, by in vitro manipulation in a neuronal cell culture model of neuroinflammation, the two were previously reported to be interdependent [66], and further inquiry on downstream effect of NF-kB induced miRNA-146a upregulation led to observations of decrease interleukin-1 receptor-associated kinase-1 (IRAK-1), but not IRAK-2 protein levels [67]. miRNA-146a profile was found to be also modified in other neurodegenerative disorders such as prion disease [73], in an epileptogenic animal model as well in hippocampal cells of temporal epilepsy patients [83] and in inflammatory neurodegeneration of viral encephalitis (e.g. herpes simplex encephalopathy) [84].
Recent evidence emerged of miRNA-29 overexpression in insulin resistance animal models that correlate well with reports of association between diabetes and sporadic AD. Memory impairment and AD have been associated with anomalies in insulin signaling pathway molecules, such as downregulation of insulin receptor IRS-1[85]. MiRNA-29b was correlated with repression of dihydrolipoylbranched chain acyltransferase (DBT), a subunit of a dehydrogenase that metabolizes ramified amino acids, an essential source of N for glutamate. Such enzymatic deficiency leads to neurodegeneration, closing the vicious circle between insulin resistance and AD.
Neurodegeneration also implies apoptosis, and there are several microRNAs involved in both caspase-dependent and caspase-independent apoptosis. A few apoptotic mechanisms in which miRNAs involvement was proven are: (i) histone-deacetylases suppression, (ii) p53 upregulation, (iii) Bcl2 family members regulation [86].Unsurprisingly, some of previously discussed microRNAs are reported to be related to apoptotic events (e.g. miRNA-128 and BCL2 upregulation [87]).
Concluding remarks
miRNAs biology is still an emerging science, with a lot of unfilled blanks in biogenesis and molecular mechanisms. The scientific world takes a great deal of interest in noncoding RNAs, in search for new pathogenic mechanims of rare, fatal or undertreated diseases and with hope for new molecular therapeutic targets. Maybe even more than other fields, neurodegeneration would greatly benefit from miRNA science in several pathologies with high social impact, such as AD, PD, ALS or HD.
References
Shabalina SA, Spiridonov NA (2004) The mammalian transcriptome and the function of non-coding DNA sequences. Genome Biol 5(4):105. doi:10.1186/gb-2004-5-4-105gb-2004-5-4-105
Condorelli G, Dimmeler S (2008) MicroRNAs: components of an integrated system controlling cardiac development, physiology, and disease pathogenesis. Cardiovasc Res 79(4):551–552. doi:10.1093/cvr/cvn189
Nelson PT, Keller JN (2007) RNA in brain disease: no longer just “The messenger in the middle”. J Neuropathol Exp Neurol 66(6):461–468. doi:10.1097/01.jnen.0000240474.27791.f3
Griffiths-Jones S (2004) The microRNA registry. Nucleic Acids Res 32(Database issue):D109–D111. doi:10.1093/nar/gkh023
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34(Database issue):D140–D144. doi:10.1093/nar/gkj112
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36(Database issue):D154–D158. doi:10.1093/nar/gkm952
Britten RJ, Davidson EH (1969) Gene regulation for higher cells: a theory. Science 165(891):349–357
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854
Wightman B, Ha I, Ruvkun G (1993) Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75(5):855–862
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A (2004) Identification of mammalian microRNA host genes and transcription units. Genome Res 14(10A):1902–1910. doi:10.1101/gr.2722704
Saini HK, Griffiths-Jones S, Enright AJ (2007) Genomic analysis of human microRNA transcripts. Proc Natl Acad Sci USA 104(45):17719–17724. doi:10.1073/pnas.0703890104
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M, Croce CM (2004) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101(9):2999–3004. doi:10.1073/pnas.0307323101
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ, Hammond SM (2005) A microRNA polycistron as a potential human oncogene. Nature 435(7043):828–833. doi:10.1038/nature03552
Lian J, Zhang X, Tian H, Liang N, Wang Y, Liang C, Li X, Sun F (2009) Altered microRNA expression in patients with non-obstructive azoospermia. Reprod Biol Endocrinol 7:13. doi:10.1186/1477-7827-7-13
Kim VN, Han J, Siomi MC (2009) Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10(2):126–139
Gregory RI, Yan K-p, Amuthan G, Chendrimada T, Doratotaj B, Cooch N, Shiekhattar R (2004) The microprocessor complex mediates the genesis of microRNAs. Nature 432(7014):235–240
Carthew RW, Sontheimer EJ (2009) Origins and mechanisms of miRNAs and siRNAs. Cell 136(4):642–655. doi:10.1016/j.cell.2009.01.035
Gregory RI, Shiekhattar R (2005) MicroRNA biogenesis and cancer. Cancer Res 65(9):3509–3512. doi:10.1158/0008-5472.CAN-05-0298
Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20(5):515–524. doi:10.1101/gad.1399806
Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R (2005) Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123(4):631–640. doi:10.1016/j.cell.2005.10.022
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20. doi:10.1016/j.cell.2004.12.035
Liu J (2008) Control of protein synthesis and mRNA degradation by microRNAs. Curr Opin Cell Biol 20(2):214–221. doi:10.1016/j.ceb.2008.01.006
Foldes-Papp Z, Konig K, Studier H, Buckle R, Breunig HG, Uchugonova A, Kostner GM (2009) Trafficking of mature miRNA-122 into the nucleus of live liver cells. Curr Pharm Biotechnol 10(6):569–578
Park CW, Zeng Y, Zhang X, Subramanian S, Steer CJ Mature microRNAs identified in highly purified nuclei from HCT116 colon cancer cells. RNA Biol 7(5)
Taft RJ, Simons C, Nahkuri S, Oey H, Korbie DJ, Mercer TR, Holst J, Ritchie W, Wong JJ, Rasko JE, Rokhsar DS, Degnan BM, Mattick JS (2010) Nuclear-localized tiny RNAs are associated with transcription initiation and splice sites in metazoans. Nat Struct Mol Biol 17(8):1030–1034. doi: 10.1038/nsmb.1841
Liu CG, Calin GA, Meloon B, Gamliel N, Sevignani C, Ferracin M, Dumitru CD, Shimizu M, Zupo S, Dono M, Alder H, Bullrich F, Negrini M, Croce CM (2004) An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues. Proc Natl Acad Sci USA 101(26):9740–9744. doi:10.1073/pnas.04032931010403293101
Sempere LF, Freemantle S, Pitha-Rowe I, Moss E, Dmitrovsky E, Ambros V (2004) Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol 5(3):R13. doi:10.1186/gb-2004-5-3-r13gb-2004-5-3-r13
Takada S, Berezikov E, Yamashita Y, Lagos-Quintana M, Kloosterman WP, Enomoto M, Hatanaka H, Fujiwara S, Watanabe H, Soda M, Choi YL, Plasterk RH, Cuppen E, Mano H (2006) Mouse microRNA profiles determined with a new and sensitive cloning method. Nucleic Acids Res 34(17):e115. doi:10.1093/nar/gkl653
Xu H, Wang X, Du Z, Li N (2006) Identification of microRNAs from different tissues of chicken embryo and adult chicken. FEBS Lett 580(15):3610–3616. doi:10.1016/j.febslet.2006.05.044
Ambros V (2004) The functions of animal microRNAs. Nature 431(7006):350–355. doi:10.1038/nature02871nature02871
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433(7027):769–773. doi:10.1038/nature03315
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM (2003) Bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113(1):25–36
Hipfner DR, Weigmann K, Cohen SM (2002) The bantam gene regulates Drosophila growth. Genetics 161(4):1527–1537
Xu P, Vernooy SY, Guo M, Hay BA (2003) The Drosophila microRNA Mir-14 suppresses cell death and is required for normal fat metabolism. Curr Biol 13(9):790–795
Chang S, Johnston RJ Jr, Frokjaer-Jensen C, Lockery S, Hobert O (2004) MicroRNAs act sequentially and asymmetrically to control chemosensory laterality in the nematode. Nature 430(7001):785–789. doi:10.1038/nature02752nature02752
Li X, Carthew RW (2005) A microRNA mediates EGF receptor signaling and promotes photoreceptor differentiation in the Drosophila eye. Cell 123(7):1267–1277. doi:10.1016/j.cell.2005.10.040
Chen CZ, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303(5654):83–86. doi:10.1126/science.1091903
Hatfield SD, Shcherbata HR, Fischer KA, Nakahara K, Carthew RW, Ruohola-Baker H (2005) Stem cell division is regulated by the microRNA pathway. Nature 435(7044):974–978. doi:10.1038/nature03816
Ivey KN, Muth A, Arnold J, King FW, Yeh RF, Fish JE, Hsiao EC, Schwartz RJ, Conklin BR, Bernstein HS, Srivastava D (2008) MicroRNA regulation of cell lineages in mouse and human embryonic stem cells. Cell Stem Cell 2(3):219–229. doi:10.1016/j.stem.2008.01.016
Kloosterman WP, Plasterk RH (2006) The diverse functions of microRNAs in animal development and disease. Dev Cell 11(4):441–450. doi:10.1016/j.devcel.2006.09.009
Blenkiron C, Miska EA (2007) miRNAs in cancer: approaches, aetiology, diagnostics and therapy. Hum Mol Genet 16(1):R106–R113. doi:10.1093/hmg/ddm056
Alvarez-Garcia I, Miska EA (2005) MicroRNA functions in animal development and human disease. Development 132(21):4653–4662. doi:10.1242/dev.02073
Giraldez AJ, Cinalli RM, Glasner ME, Enright AJ, Thomson JM, Baskerville S, Hammond SM, Bartel DP, Schier AF (2005) MicroRNAs regulate brain morphogenesis in zebrafish. Science 308(5723):833–838. doi:10.1126/science.1109020
Grishok A, Pasquinelli AE, Conte D, Li N, Parrish S, Ha I, Baillie DL, Fire A, Ruvkun G, Mello CC (2001) Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106(1):23–34
Bernstein E, Kim SY, Carmell MA, Murchison EP, Alcorn H, Li MZ, Mills AA, Elledge SJ, Anderson KV, Hannon GJ (2003) Dicer is essential for mouse development. Nat Genet 35(3):215–217. doi:10.1038/ng1253ng1253
Bushati N, Cohen SM (2008) MicroRNAs in neurodegeneration. Curr Opin Neurobiol 18(3):292–296. doi:10.1016/j.conb.2008.07.001
Barbato C, Giorgi C, Catalanotto C, Cogoni C (2008) Thinking about RNA? MicroRNAs in the brain. Mamm Genome 19(7–8):541–551. doi:10.1007/s00335-008-9129-6
Ashraf SI, Kunes S (2006) A trace of silence: memory and microRNA at the synapse. Curr Opin Neurobiol 16(5):535–539. doi:10.1016/j.conb.2006.08.007
Khudayberdiev S, Fiore R, Schratt G (2009) MicroRNA as modulators of neuronal responses. Commun Integr Biol 2(5):411–413
Tapia-Arancibia L, Aliaga E, Silhol M, Arancibia S (2008) New insights into brain BDNF function in normal aging and Alzheimer disease. Brain Res Rev 59(1):201–220. doi:10.1016/j.brainresrev.2008.07.007
Christensen M, Schratt GM (2009) MicroRNA involvement in developmental and functional aspects of the nervous system and in neurological diseases. Neurosci Lett 466(2):55–62. doi:10.1016/j.neulet.2009.04.043
Shibata M, Kurokawa D, Nakao H, Ohmura T, Aizawa S (2008) MicroRNA-9 modulates Cajal–Retzius cell differentiation by suppressing Foxg1 expression in mouse medial pallium. J Neurosci 28(41):10415–10421. doi:10.1523/JNEUROSCI.3219-08.2008
Leucht C, Stigloher C, Wizenmann A, Klafke R, Folchert A, Bally-Cuif L (2008) MicroRNA-9 directs late organizer activity of the midbrain–hindbrain boundary. Nat Neurosci 11(6):641–648. doi:10.1038/nn.2115
Cao X, Pfaff SL, Gage FH (2007) A functional study of miR-124 in the developing neural tube. Genes Dev 21(5):531–536. doi:10.1101/gad.1519207
Yu JY, Chung KH, Deo M, Thompson RC, Turner DL (2008) MicroRNA miR-124 regulates neurite outgrowth during neuronal differentiation. Exp Cell Res 314(14):2618–2633. doi:10.1016/j.yexcr.2008.06.002
Maller Schulman BR, Liang X, Stahlhut C, DelConte C, Stefani G, Slack FJ (2008) The let-7 microRNA target gene, Mlin41/Trim71 is required for mouse embryonic survival and neural tube closure. Cell Cycle 7(24):3935–3942
Wayman GA, Davare M, Ando H, Fortin D, Varlamova O, Cheng HY, Marks D, Obrietan K, Soderling TR, Goodman RH, Impey S (2008) An activity-regulated microRNA controls dendritic plasticity by down-regulating p250GAP. Proc Natl Acad Sci USA 105(26):9093–9098. doi:10.1073/pnas.0803072105
Hebert SS, De Strooper B (2009) Alterations of the microRNA network cause neurodegenerative disease. Trends Neurosci 32(4):199–206. doi:10.1016/j.tins.2008.12.003
Asikainen S, Rudgalvyte M, Heikkinen L, Louhiranta K, Lakso M, Wong G, Nass R (2010) Global microRNA expression profiling of Caenorhabditis elegans Parkinson’s disease models. J Mol Neurosci 41(1):210–218. doi: 10.1007/s12031-009-9325-1
Bak M, Silahtaroglu A, Moller M, Christensen M, Rath MF, Skryabin B, Tommerup N, Kauppinen S (2008) MicroRNA expression in the adult mouse central nervous system. RNA 14(3):432–444. doi:10.1261/rna.783108
Berezikov E, Thuemmler F, van Laake LW, Kondova I, Bontrop R, Cuppen E, Plasterk RH (2006) Diversity of microRNAs in human and chimpanzee brain. Nat Genet 38(12):1375–1377. doi:10.1038/ng1914
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E, Hannon G, Abeliovich A (2007) A microRNA feedback circuit in midbrain dopamine neurons. Science 317(5842):1220–1224. doi:10.1126/science.1140481
Wang G, van der Walt JM, Mayhew G, Li YJ, Zuchner S, Scott WK, Martin ER, Vance JM (2008) Variation in the miRNA-433 binding site of FGF20 confers risk for Parkinson disease by overexpression of alpha-synuclein. Am J Hum Genet 82(2):283–289. doi:10.1016/j.ajhg.2007.09.021
Johnson R, Buckley NJ (2009) Gene dysregulation in Huntington’s disease: REST, microRNAs and beyond. Neuromol Med 11(3):183–199. doi:10.1007/s12017-009-8063-4
Packer AN, Xing Y, Harper SQ, Jones L, Davidson BL (2008) The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington’s disease. J Neurosci 28(53):14341–14346. doi:10.1523/JNEUROSCI.2390-08.2008
Lukiw WJ, Zhao Y, Cui JG (2008) An NF-κB-sensitive Micro RNA-146a-mediated inflammatory circuit in Alzheimer disease and in stressed human brain cells. J Biol Chem 283(46):31315–31322. doi:10.1074/jbc.M805371200
Cui JG, Li YY, Zhao Y, Bhattacharjee S, Lukiw WJ (2010) Differential Regulation of interleukin-1 receptor-associated kinase-1 (IRAK-1) and IRAK-2 by MicroRNA-146a and NF-κB in stressed human astroglial cells and in Alzheimer disease. J Biol Chem 285(50):38951–38960. doi: 10.1074/jbc.M110.178848
Lukiw WJ (2007) Micro-RNA speciation in fetal, adult and Alzheimer’s disease hippocampus. Neuroreport 18(3):297–300. doi:10.1097/WNR.0b013e3280148e8b00001756-200702120-00020
Sethi P, Lukiw WJ (2009) Micro-RNA abundance and stability in human brain: specific alterations in Alzheimer’s disease temporal lobe neocortex. Neurosci Lett 459(2):100–104. doi:10.1016/j.neulet.2009.04.052
Wang WX, Rajeev BW, Stromberg AJ, Ren N, Tang G, Huang Q, Rigoutsos I, Nelson PT (2008) The expression of microRNA miR-107 decreases early in Alzheimer’s disease and may accelerate disease progression through regulation of beta-site amyloid precursor protein-cleaving enzyme 1. J Neurosci 28(5):1213–1223. doi:10.1523/JNEUROSCI.5065-07.2008
Li YY, Cui JG, Hill JM, Bhattacharjee S, Zhao Y, Lukiw WJ (2010) Increased expression of miRNA-146a in Alzheimer’s disease transgenic mouse models. Neurosci Lett 487(1):94–98. doi: 10.1016/j.neulet.2010.09.079
Hebert SS, Horre K, Nicolai L, Papadopoulou AS, Mandemakers W, Silahtaroglu AN, Kauppinen S, Delacourte A, De Strooper B (2008) Loss of microRNA cluster miR-29a/b-1 in sporadic Alzheimer’s disease correlates with increased BACE1/beta-secretase expression. Proc Natl Acad Sci USA 105(17):6415–6420. doi:10.1073/pnas.0710263105
Saba R, Goodman CD, Huzarewich RL, Robertson C, Booth SA (2008) A miRNA signature of prion induced neurodegeneration. PLoS One 3(11):e3652. doi:10.1371/journal.pone.0003652
Hou J, Wang P, Lin L, Liu X, Ma F, An H, Wang Z, Cao X (2009) MicroRNA-146a feedback inhibits RIG-I-dependent type I IFN production in macrophages by targeting TRAF6, IRAK1, and IRAK2. J Immunol 183(3):2150–2158. doi:10.4049/jimmunol.0900707
Fukumoto H, Cheung BS, Hyman BT, Irizarry MC (2002) Beta-secretase protein and activity are increased in the neocortex in Alzheimer disease. Arch Neurol 59(9):1381–1389
Sun A, Koelsch G, Tang J, Bing G (2002) Localization of beta-secretase memapsin 2 in the brain of Alzheimer’s patients and normal aged controls. Exp Neurol 175(1):10–22. doi:10.1006/exnr.2002.7875S0014488602978751
Holsinger RM, McLean CA, Beyreuther K, Masters CL, Evin G (2002) Increased expression of the amyloid precursor beta-secretase in Alzheimer’s disease. Ann Neurol 51(6):783–786. doi:10.1002/ana.10208
Yang LB, Lindholm K, Yan R, Citron M, Xia W, Yang XL, Beach T, Sue L, Wong P, Price D, Li R, Shen Y (2003) Elevated beta-secretase expression and enzymatic activity detected in sporadic Alzheimer disease. Nat Med 9(1):3–4. doi:10.1038/nm0103-3nm0103-3
Zhao J, Fu Y, Yasvoina M, Shao P, Hitt B, O’Connor T, Logan S, Maus E, Citron M, Berry R, Binder L, Vassar R (2007) Beta-site amyloid precursor protein cleaving enzyme 1 levels become elevated in neurons around amyloid plaques: implications for Alzheimer’s disease pathogenesis. J Neurosci 27(14):3639–3649. doi:10.1523/JNEUROSCI.4396-06.2007
Cogswell JP, Ward J, Taylor IA, Waters M, Shi Y, Cannon B, Kelnar K, Kemppainen J, Brown D, Chen C, Prinjha RK, Richardson JC, Saunders AM, Roses AD, Richards CA (2008) Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J Alzheimers Dis 14(1):27–41
Chen K, Iribarren P, Hu J, Chen J, Gong W, Cho EH, Lockett S, Dunlop NM, Wang JM (2006) Activation of toll-like receptor 2 on microglia promotes cell uptake of Alzheimer disease-associated amyloid beta peptide. J Biol Chem 281(6):3651–3659. doi:10.1074/jbc.M508125200
Takahashi Y, Satoh M, Minami Y, Tabuchi T, Itoh T, Nakamura M (2008) Expression of miR-146a/b is associated with the toll-like receptor 4 signal in coronary artery disease: effect of renin-angiotensin system blockade and statins on miRNA-146a/b and toll-like receptor 4 levels. Clin Sci Lond 119(9):395–405. doi: 10.1042/CS20100003
Aronica E, Fluiter K, Iyer A, Zurolo E, Vreijling J, van Vliet EA, Baayen JC, Gorter JA (2010) Expression pattern of miR-146a, an inflammation-associated microRNA, in experimental and human temporal lobe epilepsy. Eur J Neurosci 31(6):1100–1107. doi:10.1111/j.1460-9568.2010.07122.x
Hill JM, Zhao Y, Clement C, Neumann DM, Lukiw WJ (2009) HSV-1 infection of human brain cells induces miRNA-146a and Alzheimer-type inflammatory signaling. Neuroreport 20(16):1500–1505. doi:10.1097/WNR.0b013e3283329c05
Zhu X, Perry G, Smith MA (2005) Insulin signaling, diabetes mellitus and risk of Alzheimer disease. J Alzheimers Dis 7(1):81–84
Li P (2010) MicroRNAs in cardiac apoptosis. J Cardiovasc Transl Res 3(3):219–224. doi:10.1007/s12265-010-9175-9
Guidi M, Muinos-Gimeno M, Kagerbauer B, Marti E, Estivill X, Espinosa-Parrilla Y (2010) Overexpression of miR-128 specifically inhibits the truncated isoform of NTRK3 and upregulates BCL2 in SH-SY5Y neuroblastoma cells. BMC Mol Biol 11(1):95. doi:10.1186/1471-2199-11-95
Acknowledgments
This paper is supported by the Sectoral Operational Programme Human Resources Development (SOP HRD), financed from the European Social Fund and by the Romanian Government under the contract nu POSDRU/89/1.5/S/64109.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Enciu, AM., Popescu, B.O. & Gheorghisan-Galateanu, A. MicroRNAs in brain development and degeneration. Mol Biol Rep 39, 2243–2252 (2012). https://doi.org/10.1007/s11033-011-0973-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0973-1