Abstract
Soil salinity is widespread in rice-producing areas globally, restricting both vegetative growth and grain yield. Improving salt tolerance of rice is a promising approach to meet the increasing food demand. An extensive literature survey indicates that maintaining proper Na+/K+ ratio and reactive oxygen species (ROS) content is the key issues for rice adaption to salt stress. In this review, distinctive from the existing reviews, we mainly discuss recent progresses in identifying the components and pathways involved in the rice response to salt stress and the approaches that can be used for breeding and cultivating salt-tolerant rice, pointing out the potential phytohormonal regulation of the components and the homeostasis of Na+/K+ and ROS. Thus, this review attempts to provide a comprehensive overview of the recent research on rice adaption to salt stress, which may provide guidance for rice breeding to engineer better salt-tolerant rice varieties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With the increase of world population, more foods are needed to meet the demands. Due to the reduction of cultivated land and lack of fresh water, it is encouraged to use saline-alkali land to make up for the shortages. There are about 1 billion hectares of saline-alkali land in the world. Effective use and improvement of saline-alkali land will make full use of land resources. Rice (Oryza sativa L.) is an important monocotyledonous crop and a primary food for more than half of the world population. Its productivity is critically affected by various abiotic stresses, such as drought, salinity, cold, and heat (Almeida et al. 2016; Zhu 2016). As one of the major constraints in rice cultivation worldwide, salinity affects many aspects of rice growth and development (Zeng et al. 2002; Ganie et al. 2019). Thus, breeding and cultivating salt-tolerant rice varieties have become one of the most important approaches to increase grain yield and ameliorate saline-alkali soil.
The sensitivity of rice to salinity stress greatly depends on growth stages, organ types, and genotypes (Khan et al. 1997; Nam et al. 2015). Generally, seedling and reproductive stages of rice are considered to be the salt-susceptible stages, it becomes tolerant to salinity during active tillering and at maturity period. As the primary target site for perception of salt stress, roots are more susceptible to salt than other organs (Gupta and Huang 2014; Nam et al. 2015). Salt tolerance of indica rice is higher than japonica rice (Chen et al. 2004; Hussain et al. 2019). Excess salts adversely affect all the metabolic activities in rice, leading to substantial reduction in growth and yield (Yang and Guo 2018a, b).
Salinity stress can induce osmotic and ionic stresses in plants. Osmotic stress is rapidly sensed by the plants soon after exposure to salinity conditions, leading to water and solute deficit in plants. Ionic stress begins with the accumulation of Na+ and Cl− in the plant cell, and eventually resulting in premature leaf senescence (Munns and Tester 2008; Yang and Guo 2018a, b). Excess Na+ in the cytoplasm interferes with K+ function. K+ is important for the catalytic activities of enzymes in metabolic pathways (Fu and Luan 1998). Thus, the maintenance of a low intracellular Na+/K+ ratio is important for plants to survive under salinity stress (Shabala and Pottosin 2014; Munns et al. 2016). In addition to osmotic and ionic stresses, salinity stress causes accumulation of reactive oxygen species (ROS) in plant cells, which can severely damage cellular structures and macromolecules such as enzymes, DNA, and lipids (Wang et al. 2009; Genisel et al. 2015; Ahanger et al. 2017).
In this review, we mainly focus on the phytohormonal regulation of Na+/K+ and ROS homeostasis associated with salt tolerance in rice, with exploration of the problems in the current research, for understanding the molecular mechanism of salinity tolerance which may have implications for improvement of rice varieties.
Na+/K+ homeostasis is associated with salt tolerance in rice
Salt stress is commonly caused by high concentrations of Na+ and Cl− in soil (Ismail et al. 2014). Excess Na+ competes with K+ for uptake across the plasma membranes of plant cells, which is important for the catalytic activities of many enzymes (Fu and Luan 1998), eventually reducing plant growth and causing cellular injury and even death. Thus maintaining cellular Na+/K+ homeostasis is a crucial factor determining the plant’s survival ability during the response to salt stress (Yang and Guo 2018a, b).
One primary response in maintaining cellular ion homeostasis is by restricting the accumulation of toxic sodium (Na+) (Tester and Davenport 2003). Salt Overly Sensitive 1 (SOS1), which mediates Na+/H+ exchange at the plasma membrane and cellular Na+ extrusion, is responsible for salt stress signaling and Na+ tolerance (Martinez-Atienza et al. 2007; Ji et al. 2013). In Arabidopsis, SOS1 could function both in Na+ loading (under mild saline stress at 25 mM NaCl) and unloading (at high salinity, 100 mM NaCl) (Shi et al. 2000). In rice, lower expression of OsSOS1 in old leaves may decrease frequency of retrieving Na+ from old leaf cells (Wang et al. 2012). Overexpression of OsSOS1 in Arabidopsis increases salt tolerance (Martinez-Atienza et al. 2007). The high-affinity K+ channel (HKT) family is well-established plant Na+ and Na+/K+ transporters in controlling Na+ accumulation (Garciadeblas et al. 2003; Platten et al. 2006; Horie et al. 2007; Almeida et al. 2013). In rice, the HKT gene family is divided into two classes: class I comprises Na+-selective transporters, including OsHKT1;1, OsHKT1;3, OsHKT1;4, and OsHKT1;5, and class II comprises transporters permeable to both Na+ and K+, including OsHKT2;1, OsHKT2;3, and OsHKT2;4 (Platten et al. 2006; Hauser and Horie 2010). Mutation in OsHKT1;1 leads to increased accumulation of Na+ in the shoot and hypersensitivity to salt stress (Wang et al. 2015; Campbell et al. 2017). OsHKT1;4 plays an important role in restricting Na+ accumulation in aerial parts during salinity stress at the reproductive growth stage (Suzuki et al. 2016). OsHKT1;5 reduces the transport of Na+ to shoots and positively regulates salt tolerance (Ren et al. 2005). The evidence by analyzing transposon-insertion rice mutants disrupted in OsHKT2;1 indicates that OsHKT2;1 is a central transporter for nutritional Na+ uptake into K+-starved rice roots. But OsHKT2;1-mediated Na+ influx does not cause Na+ toxicity, as its transcription is down-regulated upon salt stress (Horie et al. 2007). These studies suggest that HKT-mediated Na+/K+ homeostasis contributes to salt tolerance in rice.
Accumulating studies showed that a number of genes regulate rice salt tolerance by regulating HKT genes, such as MYB-like transcription factor, OsMYBc, positively regulates salt tolerance by directly activating the expression of OsHKT1;5 and OsHKT2;1 (Wang et al. 2015). Ethylene insensitive3-like1/2 (OsEIL1/2), the core transcriptional regulators of the ethylene signaling pathway, negatively affect salt tolerance by directly activating the expression of OsHKT2;1 (Yang et al. 2015). Rice magnesium transporter OsMGT1 is required for conferring salt tolerance in rice through enhancing the transport activity of OsHKT1;5 (Chen et al. 2017). Mutation in rice reduced culm number1 (rcn1), encoding a G subfamily ABC transporter (OsABCG5), causes a high Na+/K+ ratio and salt-sensitive phenotype. Further analysis found that RCN1/OsABCG5 is essential for upregulation of OsHKT1;5 under salt stress (Matsuda et al. 2014). The above results suggest that the modulation of Na+/K+ homeostasis under salt stress may provide an effective way to improve salt tolerance in rice.
Enhancement of ROS scavenging improves salt tolerance in rice
As a by-product of the plant’s stress response, ROS such as superoxide radical (O2.−), hydroxyl radical (OH−) and hydrogen peroxide (H2O2) are able to cause oxidative damage to protein, DNA, and lipids (Apel and Hirt 2004; Miller et al. 2010). ROS were originally believed to merely represent toxic molecules, but they have been now recognized as signal molecules in many plant cellular processes (Xia et al. 2009; Tsukagoshi 2016; Lv et al. 2018). Due to the dual role of ROS in plants, a fine-tuned balance between ROS biosynthesis and scavenging is crucial for maintaining appropriate levels of ROS at different development stages and in different growing environments (Bose et al. 2014; Tsukagoshi 2016; Zhang et al. 2016a).
Plants harbor numerous ROS generating pathways. NADPH oxidase is the most extensively studied. The plant NADPH oxidases (NOXs), which are also called as respiratory burst oxidase homologs (Rbohs), are the most studied ROS-producing enzymes. Rbohs produce ROS through catalyzing O2 to O2•− (Torres and Dangl 2005). There are 9 Rboh genes in rice genomes, and each homolog has a specific role in a broad range of biological processes (Marino et al. 2012; Wang et al. 2013). Salt treatment induced the expression of OsRbohA and OsRbohI, but repressed that of OsRbohB, OsRbohC, OsRbohE, and OsNox6 (Wang et al. 2013), suggesting that diverse roles of OsRboh genes in response to salt stress. Moreover, NOX activity was significantly higher in salt-tolerant than salt-sensitive cultivars of rice (Kaur et al. 2016a; Saini et al. 2018). These studies suggest that OsRbohs are linked to salt stress adaptation mechanisms in rice.
In addition to controlling the production of ROS, plants have evolved two efficient pathways for ROS scavenging, namely, enzymatic and nonenzymatic antioxidant defense system (You and Chan 2015). In enzymatic system, O2•− first is converted into H2O2 by superoxide dismutase (SOD). Ascorbate peroxidase (APX), catalase (CAT), and glutathione peroxidase (GPX) then detoxify H2O2 (You and Chan 2015; Mittler 2017). In rice, overexpression of OsMn-SOD1 leads to less accumulation of mitochondrial O2•− under salt treatment (Li et al. 2013). Transgenic plants overexpressing OsCu/Zn-SOD exhibited higher germination of seeds and plant height than non-transgenic plants under salt stress (Guan et al. 2017). There are eight APX genes in rice, including two cytosolic APXs (OsAPX1 and OsAPX2), two peroxisomal APXs (OsAPX3 and OsAPX4), and four chloroplast APXs (OsAPX5, OsAPX6, OsAPX7, and OsAPX8) (Hong et al. 2007). The expression of OsAPX2, OsAPX7, and OsAPX8 was upregulated under salinity stress (Teixeira et al. 2006; Hong et al. 2007; Zhang et al. 2013). Mutant with loss-of-function of OsAPX2 showed reduction of APX activity and sensitivity to salt stress, whereas transgenic lines overexpressing OsAPX2 displayed the increase of APX activity and enhancement of salt tolerance (Zhang et al. 2013). All these studies suggest that increase of ROS-scavenging enzyme activity decreases oxidative stress damage and enhances rice tolerance to salt stress.
Nonenzymatic antioxidants, including glutathione (GSH), ascorbic acid (AsA), carotenoids, tocopherols, and flavonoids, are also crucial for ROS homeostasis in plants (Gill and Tuteja 2010). Exogenous application of GSH and AsA leads to reduced ROS accumulation and increased salt tolerance (Xu et al. 2017; Wang et al. 2018). Knockdown of the expression of genes involved in AsA synthesis increases the accumulation of ROS and decreases the salt resistance of rice (Qin et al. 2016a, b; Wang et al. 2018), suggesting that GSH and AsA play important roles in the fine control of ROS homeostasis to improve salt tolerance.
Recent analyses with mutational and transgenic plants revealed that multiple genes contributed to improve salt tolerance in rice by regulating genes involved in ROS biosynthesis and scavenging pathway. Calcium-dependent protein kinase12 (OsCPK12), encoding a calcium-dependent protein kinase (CDPK), positively regulates rice salt tolerance by reducing accumulation of ROS (Asano et al. 2012). The zinc-finger proteins (ZFP), such as OsZFP179, OsZFP182, OsZFP185, OsZFP213, OsZFP245, OsZFP252, and drought and salt tolerance (DST), are involved in regulating rice salt tolerance. Among them, OsZFP179, OsZFP182, OsZFP213, OsZFP245, and OsZFP252 positively regulate tolerance to salt by increasing ROS-scavenging ability (Huang et al. 2005; Xu et al. 2008; Sun et al. 2010; Huang et al. 2012; Zhang et al. 2018), whereas OsZFP185 and DST negatively regulate rice tolerance to salt through regulating ROS-scavenging gene transcriptions (Huang et al. 2009; Cui et al. 2015; Zhang et al. 2016b). All these studies suggest that enhancing ROS-scavenging ability can efficiently increase the salt tolerance of rice.
Phytohormonal regulation of rice in salt-triggered Na+/K+ homeostasis and ROS scavenging
Plant hormones regulate normal growth and mediate responses to abiotic stress (Kazan 2015; Van de Poel et al. 2015). Studies in rice have shown that several phytohormones are related to the salt tolerance of rice, such as auxin, ethylene, abscisic acid (ABA), and gibberellin (GA) (Xia et al. 2012; Shan et al. 2014; Tao et al. 2015; Sah et al. 2016). Among these phytohormones, the function of ethylene in salt tolerance of rice has been extensively studied (Tao et al. 2015; Zhang et al. 2016a).
In rice, ethylene treatment of etiolated seedlings exhibited double response, namely, promotion of coleoptile growth but inhibition of root elongation (Ma et al. 2013). Based on the ethylene double response, a series of ethylene-response mutants maohuzi (mhz) were identified (Ma et al. 2013). Among these mutants, rice ethylene signaling components MHZ6/ETHYLENE INSENSITIVE3-LIKE1 (OsEIL1) and MHZ7/ETHYLENE INSENSITIVE2 (OsEIN2) were isolated (Ma et al. 2013; Yang et al. 2015). Loss-of-function or suppression of OsEIN2, OsEIL1, or OsEIL2 results in improvement of salt tolerance, while overexpressing each of them leads to salt hypersensitivity at the seedling stage. Further investigations indicate that this negative regulation of OsEIL1 or OsEIL2 in salt tolerance is likely attributed in part to the direct regulation of OsHKT2;1 expression and Na+ uptake in roots (Yang et al. 2015). Recent studies in rice showed that salt treatment upregulates ethylene biosynthesis gene transcriptions and ethylene production, leading to the inhibition of primary root elongation and reduced salt tolerance. Moreover, the regulators involved in salt-induced ethylene biosynthesis are emerging, such as SALT INTOLERANCE1 (SIT1) and DNA-BINDING WITH ONE FINGER 15 (OsDOF15) (Li et al. 2014; Qin et al. 2019). SIT1, a lectin receptor-like kinase, positively regulates salt tolerance by activating MITOGEN-ACTIVATED PROTEIN KINASE3/6 (MPK3/6), which promotes ethylene and ROS overproduction (Li et al. 2014). OsDOF15 is a DOF-binding with one finger (DOF) transcription factor, negatively regulates ethylene biosynthesis by directly binding to the promoter of 1-aminocyclopropane-1-carboxylate synthase1 (OsACS1). Under salt treatment, the transcription of OsDOF15 was suppressed, resulting in activation of the ethylene biosynthesis genes and enhanced ethylene biosynthesis, thereby inhibiting root growth (Qin et al. 2019). These studies imply that salt treatment enhances ethylene biosynthesis in rice, which might promote Na+ uptake and ROS accumulation, thus leading to reduction of salt tolerance of rice.
In addition to directly regulating Na+/K+ uptake and ROS accumulation, ethylene also coordinates with other phytohormones to modulate plant response to salinity. Auxin coordinates many of the key processes in plant development and adaptive growth (Strader and Zhao 2016; Wang et al. 2019a, b). Comprehensive analysis and expression profiling of genes in rice under salt stress showed that many genes involved in auxin transport, auxin signaling, and auxin homeostasis pathway were regulated by salt stress (Jain and Khurana 2009; Chai and Subudhi 2016). Downregulation of two rice auxin receptor genes, transport inhibitor response1 (OsTIR1) and auxin signaling F-box2 (OsAFB2), via OsmiR393 overexpression, leads to reduced tolerance to salt in rice (Xia et al. 2012). Exogenous application of auxin analogue naphthalene acetic acid (NAA) induced ROS production, whereas inhibiting the auxin biosynthesis by aminoethoxyvinylglycine (AVG) suppressed ROS production (Du et al. 2012), suggesting that auxin contributes to rice salt tolerance by modulating ROS production. Interaction between ethylene and auxin was revealed by two ethylene-response mutants: rice ethylene-insensitive7 (rein7/yuc8) and mhz2/soil-surface rooting1 (sor1) (Qin et al. 2017; Chen et al. 2018). REIN7/YUC8 catalyzes the conversion of indole-3-pyruvic acid (IPA) to indole-3-acetic acid (IAA) in auxin biosynthesis, it is transcriptionally modulated by ethylene signaling component OsEIL1 (Qin et al. 2017). MHZ2/SOR1, a RING finger E3 ubiquitin ligase, regulates ethylene response in primary roots by interacting with OsIAA26, a typical Aux/IAA protein involved in the auxin signaling pathway (Chen et al. 2018). Collectively, these studies suggest that ethylene may regulate ROS accumulation by modulating auxin biosynthesis and signaling, subsequently affecting the salt tolerance of rice.
Abscisic acid (ABA) is the central regulator of abiotic stress resistance in plants (Finkelstein 2013; Sah et al. 2016). Salt stress causes an increase in the ABA accumulation, and exogenous ABA may alleviate the deleterious effects of salt stress (Chen et al. 2006; Park et al. 2008; Welsch et al. 2008). In rice, ethylene treatment induced the expressions of MHZ4 or MHZ5, which are involved in ABA biosynthesis, leading to increased accumulation of ABA in roots. Mutation of either MHZ4 or MHZ5 reduced ethylene sensitivity in root growth. Genetic analysis revealed that MHZ4 and MHZ5-dependent ABA pathways act downstream of ethylene receptors to positively regulate root response to ethylene (Ma et al. 2014; Yin et al. 2015). Given that ABA production and signaling are necessary for plant responses to salinity, MHZ4 and MHZ5 are anticipated to have some roles in plant responses to salinity. Thus, besides regulating seedling growth, MHZ4 and MHZ5 may also mediate the interaction between ethylene and ABA on controlling stress responses.
Gibberellins (GA) are plant hormones that govern many aspects of plant biology. Several studies have revealed that GA participate in the regulation of rice salt tolerance (Shan et al. 2014; Zhu et al. 2015). Slender rice1 (SLR1), the rice homolog of Arabidopsis DELLA genes that have been identified as GA signaling components, was highly induced by salt stress. Moreover, salt induction of SLR1 expression was dependent on OsMYB91, an R2R3-type MYB transcription factor in rice (Zhu et al. 2015). Overexpressing gibberellin2-oxidases5 (OsGA2ox5), a gene involved in the GA catabolic pathway, enhanced the resistant to high-salinity stress in rice (Shan et al. 2014), suggesting a negative role of GA in rice salt tolerance. Under submerging conditions, the ethylene signaling transcription factor OsEIL1 directly activates the transcription of semidwarf1 (SD1), thereby promoting GA biosynthesis (Kuroha et al. 2018), suggesting an ethylene-GA crosstalk in abiotic stress. It remains to be further investigated whether this interaction is involved in salinity stress response.
Several studies have shown that cytokinins (CKs) functionally control plant adaptation to environmental stresses. In Arabidopsis, the CK-deficient plants with reduced levels of various CKs exhibited enhanced salt tolerance (Nishiyama et al. 2011). However, exogenous application of CKs resulted in increased salinity tolerance in Solanum melongena (Wu et al. 2014). Knockdown cytokinin oxidase2 (OsCKX2) in rice, which encodes an enzyme that degrades CK, resulted in better vegetative growth, higher relative water content and photosynthetic efficiency than those of wild type under salt stress (Joshi et al. 2018). Loss-of-function type A response regulators (RRs), OsRR9 and OsRR10, which are negative regulators of CK signaling, exhibited higher salinity tolerance than wild-type rice seedlings (Wang et al. 2019a, b), suggesting that CKs positively regulate salinity tolerance in rice. Recent research in rice has shown that CK treatment increased ethylene level by upregulating the transcription of ethylene biosynthesis genes, leading to the inhibition of root growth (Zou et al. 2018), suggesting that CKs promote ethylene biosynthesis in roots, whether salt stress promotes ethylene biosynthesis in rice roots through CKs remains to be studied.
Brassinosteroids (BRs) are plant steroid hormones, which play essential roles in plant growth and developmental programs (Yang et al. 2011; Wei and Li 2016). Apart from their roles in the regulation of plant growth and development, BRs confer tolerance to a range of abiotic stresses (Krishna 2003; Divi et al. 2010). Under salinity conditions, BRs activate ethylene biosynthesis and signaling pathway, thereby improving the salt tolerance of tomato seedlings (Zhu et al. 2016). In rice, exogenous application of BRs enhances salt tolerance of rice (Sharma et al. 2013). Moreover, the expression levels of genes involved in BRs biosynthesis and signaling pathway in salt tolerance cultivars of rice are higher than those in salt-sensitive cultivars (Kaur et al. 2016b), suggesting that BRs play an positive role in rice salt tolerance; however, whether this process depends on ethylene remains to be studied.
The function of jasmonic acid (JA) in plant’s resilience to many environmental challenges has been well studied, and its role in salt tolerance has also been reported (Qiu et al. 2014; Zhao et al. 2014; Kazan 2015). Endogenous JA accumulations in roots of rice plants are subjected to salt stress, and exogenous JAs improved salt-stress tolerance in rice and wheat (Moons et al. 1997; Kang et al. 2005; Qiu et al. 2014). Rice mutants with defect in JA biosynthesis exhibit improved tolerance to salt stress (Hazman et al. 2015). Overexpression of the Cyt P450 family gene OsCYP94C2b, encoding a JA-catabolizing enzyme, shows decreased JA content along with improved performance on high concentrations of salt (Kurotani et al. 2015). Furthermore, constitutive overexpression of rice JASMONATE ZIM-domain (OsJAZ) genes leads to improved salt tolerance (Ye et al. 2009). All these studies suggest that JA play a vital role in the adaptation to salt stress. The interaction between JA and ethylene has been investigated in rice, ethylene signaling component, OsEIL2, directly binds to the promoter of JA biosynthesis gene GAOYAO1 (GY1) to suppress its promoter activity, thus leading to inhibited JA biosynthesis and promoted mesocotyl/coleoptile elongation (Xiong et al. 2017). Further investigation may focus on whether this interaction plays a role in salt stress.
Conclusions and perspectives
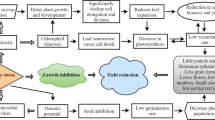
During the past few years, multiple studies were committed to elucidating the mechanism of salt tolerance in rice (Table 1), and the results show that maintaining Na+/K+ and ROS homeostasis is an effective way for rice to adapt to salt stress (Ganie et al. 2019). From above studies, a general conclusion could be made: on the one hand, salt stress induces SIT1 transcription, which in turn activates MPK3/6 to promote ethylene biosynthesis. On the other hand, salt stress inhibits OsDOF15 transcription, leading to increased ethylene biosynthesis. Ethylene overproduction promotes ABA, auxin, JA and GA biosynthesis, ultimately leading to enhanced Na+ uptake and ROS accumulation, thereby inhibiting growth and even causing plant death (Fig. 1).
The role of phytohormones in regulating salt tolerance in rice. Salt stress activates SIT1-MPK3/6 phosphorylation cascade and represses OsDOF15 transcription, leading to increased ethylene biosynthesis. Ethylene overproduction promotes ABA, auxin, JA and GA biosynthesis, ultimately leads to enhanced Na+ uptake and ROS accumulation, thereby exhibiting salt-sensitive phenotype. OsEIL1/2, the master transcriptional regulator of ethylene signaling in rice, directly regulate the expression of OsHKTs (a high affinity Na+ transporter), SD1 (GA biosynthesis gene), and OsYUC8 (auxin biosynthesis gene), and GY1 (JA biosynthesis gene). Auxin accumulation promotes SOR1-mediated degradation of OsIAA26, thus resulting in ROS accumulation. MHZ4 and MHZ5, which are involved in ABA biosynthesis, may mediate the interaction between ethylene and ABA in regulation of salinity response. CKs induce ethylene biosynthesis by upregulating the transcription of ethylene biosynthesis genes. Ethylene promotes ABA and auxin biosynthesis to inhibit root growth and inhibits JA biosynthesis to promote coleoptiles/mesocotyls growth. Under submergence, ethylene promotes internode elongation through increasing transcription of SD1 and GA production to escape flooding in deepwater rice. The solid lines indicate direct interactions, and the dashed lines indicate indirect interactions. The arrows indicate stimulatory effects, whereas the T sharp symbol indicates inhibitory effects
Plant hormones play important roles in regulating responses to a wide variety of internal and external stimuli (Kazan 2015; Sah et al. 2016). Engineering of hormone biosynthesis and signaling pathways can potentially offer new avenues to the improvement of abiotic stress tolerance in rice. However, changes in plant hormone biosynthesis and signaling can have undesirable consequences on rice growth and development (Yoshikawa et al. 2014; Yin et al. 2017). Thus, precise control of hormone productions and signaling may be critical for promotion of rice salinity tolerance. Several genes involved in hormone response have been reported to confer stress tolerance when overexpression in diverse species without adverse effects on plant development (Seo et al. 2010; Schmidt et al. 2013; Makhloufi et al. 2014), suggesting that focusing on downstream responses genes in hormone signaling pathway may be a good idea for genetic improvement of salt tolerance in rice. In addition, precise control hormone biosynthesis and signaling factor using specific promoters is another effective way to improve salt tolerance in rice.
Salt stress causes significant reductions in rice production worldwide, thus improving salt tolerance is a promising approach to meet the increasing food demand. Cultivar improvement through conventional breeding is feasible, but it takes a long time to minimize linkage drag through phenotypic screening (Iftekharuddaula et al. 2012; Hasan et al. 2015). Single nucleotide polymorphisms (SNPs) marker-assisted selection will greatly promote the molecular breeding process (Gimhani et al. 2016; Rana et al. 2019). Therefore, efforts should be made to capture useful quantitative trait loci (QTLs) associated with the salinity tolerance as possible genetic markers to introgress into elite rice varieties. Moreover, salt tolerance in rice is a cumulative effect of different salinity tolerance mechanisms governed by multiple genes (Horie et al. 2012), thus it remains to be further investigated how multiple genes are transferred at the same time with stable inheritance to offspring.
Wild rice with wide genetic diversity is considered a valuable source of genes for tolerance to salinity stress, which can be potentially used in rice breeding (Prusty et al. 2018; Quan et al. 2018; Yichie et al. 2018). Transcriptome analysis of salt stress responsiveness in the seedlings of Dongxiang wild rice showed that many genes involved in hormone biosynthesis or signaling pathway were upregulated or downregulated (Zhou et al. 2016), suggesting that hormone homeostasis is essential for salt-stress tolerance in rice. Further studies showed that wild rice confers high salt tolerance by modulating Na+/K+ uptake (Prusty et al. 2018; Yichie et al. 2018), but the relation between wild rice and the salt tolerant mechanism is largely unclear. Further studies should focus on cloning the salt tolerance gene of wild rice and elucidating its regulatory mechanism, which may be utilize in rice improvement. In addition to wild rice, tetraploid rice also exhibited resistance to salt stress (Tu et al. 2014), which inspires us that polyploid breeding will be a new way to improve salt tolerance in rice.
References
Ahanger MA, Tomar NS, Tittal M, Argal S, Agarwal RM (2017) Plant growth under water/salt stress: ROS production; antioxidants and significance of added potassium under such conditions. Physiol Mol Biol Plants 23:731–744. https://doi.org/10.1007/s12298-017-0462-7
Almeida P, Katschnig D, de Boer AH (2013) HKT transporters-state of the art. Int J Mol Sci 14:20359–20385. https://doi.org/10.3390/ijms141020359
Almeida DM, Almadanim MC, Lourenco T, Abreu IA, Saibo NJ, Oliveira MM (2016) Screening for abiotic stress tolerance in rice: salt, cold, and drought. Methods Mol Biol 1398:155–182. https://doi.org/10.1007/978-1-4939-3356-3_14
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399. https://doi.org/10.1146/annurev.arplant.55.031903.141701
Asano T, Hayashi N, Kobayashi M, Aoki N, Miyao A, Mitsuhara I, Ichikawa H, Komatsu S, Hirochika H, Kikuchi S, Ohsugi R (2012) A rice calcium-dependent protein kinase OsCPK12 oppositely modulates salt-stress tolerance and blast disease resistance. Plant J 69:26–36. https://doi.org/10.1111/j.1365-313X.2011.04766.x
Bose J, Rodrigo-Moreno A, Shabala S (2014) ROS homeostasis in halophytes in the context of salinity stress tolerance. J Exp Bot 65:1241–1257. https://doi.org/10.1093/jxb/ert430
Campbell MT, Bandillo N, Al Shiblawi FRA, Sharma S, Liu K, Du Q, Schmitz AJ, Zhang C, Very AA, Lorenz AJ, Walia H (2017) Allelic variants of OsHKT1;1 underlie the divergence between indica and japonica subspecies of rice (Oryza sativa) for root sodium content. PLoS Genet 13:e1006823. https://doi.org/10.1371/journal.pgen.1006823
Chai C, Subudhi PK (2016) Comprehensive analysis and expression profiling of the OsLAX and OsABCB auxin transporter gene families in rice (Oryza sativa) under phytohormone stimuli and abiotic stresses. Front Plant Sci 7:593. https://doi.org/10.3389/fpls.2016.00593
Chen ZD, Zhong WG, Yang J, Huang ZY (2004) Evaluation of salt tolerance of rice (Oryza sativa L.) germplasm. J Plant Genet Res 5:351–355. https://doi.org/10.13430/j.cnki.jpgr.2004.04.010
Chen CW, Yang YW, Lur HS, Tsai YG, Chang MC (2006) A novel function of abscisic acid in the regulation of rice (Oryza sativa L.) root growth and development. Plant Cell Physiol 47:1–13. https://doi.org/10.1093/pcp/pci216
Chen ZC, Yamaji N, Horie T, Che J, Li J, An G, Ma JF (2017) A magnesium transporter OsMGT1 plays a critical role in salt tolerance in rice. Plant Physiol 174:1837–1849. https://doi.org/10.1104/pp.17.00532
Chen H, Ma B, Zhou Y, He SJ, Tang SY, Lu X, Xie Q, Chen SY, Zhang JS (2018) E3 ubiquitin ligase SOR1 regulates ethylene response in rice root by modulating stability of Aux/IAA protein. Proc Natl Acad Sci U S A 115:4513–4518. https://doi.org/10.1073/pnas.1719387115
Cui LG, Shan JX, Shi M, Gao JP, Lin HX (2015) DCA1 acts as a transcriptional co-activator of DST and contributes to drought and salt tolerance in rice. PLoS Genet 11:e1005617. https://doi.org/10.1371/journal.pgen.1005617
Divi UK, Rahman T, Krishna P (2010) Brassinosteroid-mediated stress tolerance in Arabidopsis shows interactions with abscisic acid, ethylene and salicylic acid pathways. BMC Plant Biol 10:151. https://doi.org/10.1186/1471-2229-10-151
Du H, Wu N, Fu J, Wang S, Li X, Xiao J, Xiong L (2012) A GH3 family member, OsGH3-2, modulates auxin and abscisic acid levels and differentially affects drought and cold tolerance in rice. J Exp Bot 63:6467–6480. https://doi.org/10.1093/jxb/ers300
Finkelstein R (2013) Abscisic acid synthesis and response. Arabidopsis Book 11:e0166. https://doi.org/10.1199/tab.0166
Fu HH, Luan S (1998) AtKuP1: a dual-affinity K+ transporter from Arabidopsis. Plant Cell 10:63–73. https://doi.org/10.1105/tpc.10.1.63
Ganie SA, Molla KA, Henry RJ, Bhat KV, Mondal TK (2019) Advances in understanding salt tolerance in rice. Theor Appl Genet 132:851–870. https://doi.org/10.1007/s00122-019-03301-8
Garciadeblas B, Senn ME, Banuelos MA, Rodriguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801. https://doi.org/10.1046/j.1365-313X.2003.01764.x
Genisel M, Erdal S, Kizilkaya M (2015) The mitigating effect of cysteine on growth inhibition in salt-stressed barley seeds is related to its own reducing capacity rather than its effects on antioxidant system. Plant Growth Regul 75:187–197. https://doi.org/10.1007/s10725-014-9943-7
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930. https://doi.org/10.1016/j.plaphy.2010.08.016
Gimhani DR, Gregorio GB, Kottearachchi NS, Samarasinghe WL (2016) SNP-based discovery of salinity-tolerant QTLs in a bi-parental population of rice (Oryza sativa). Mol Gen Genomics 291:2081–2099. https://doi.org/10.1007/s00438-016-1241-9
Guan Q, Liao X, He M, Li X, Wang Z, Ma H, Yu S, Liu S (2017) Tolerance analysis of chloroplast OsCu/Zn-SOD overexpressing rice under NaCl and NaHCO3 stress. PLoS One 12:e0186052. https://doi.org/10.1371/journal.pone.0186052
Gupta B, Huang B (2014) Mechanism of salinity tolerance in plants: physiological, biochemical, and molecular characterization. Int J Genomics 2014:701596. https://doi.org/10.1155/2014/701596
Hasan MM, Rafii MY, Ismail MR, Mahmood M, Rahim HA, Alam MA, Ashkani S, Malek MA, Latif MA (2015) Marker-assisted backcrossing: a useful method for rice improvement. Biotechnol Biotechnol Equip 29:237–254. https://doi.org/10.1080/13102818.2014.995920
Hauser F, Horie T (2010) A conserved primary salt tolerance mechanism mediated by HKT transporters: a mechanism for sodium exclusion and maintenance of high K+/Na+ ratio in leaves during salinity stress. Plant Cell Environ 33:552–565. https://doi.org/10.1111/j.1365-3040.2009.02056.x
Hazman M, Hause B, Eiche E, Nick P, Riemann M (2015) Increased tolerance to salt stress in OPDA-deficient rice ALLENE OXIDE CYCLASE mutants is linked to an increased ROS-scavenging activity. J Exp Bot 66:3339–3352. https://doi.org/10.1093/jxb/erv142
Hong CY, Hsu YT, Tsai YC, Kao CH (2007) Expression of ASCORBATE PEROXIDASE 8 in roots of rice (Oryza sativa L.) seedlings in response to NaCl. J Exp Bot 58:3273–3283. https://doi.org/10.1093/jxb/erm174
Horie T, Costa A, Kim TH, Han MJ, Horie R, Leung HY, Miyao A, Hirochika H, An G, Schroeder JI (2007) Rice OsHKT2;1 transporter mediates large Na+ influx component into K+-starved roots for growth. EMBO J 26:3003–3014. https://doi.org/10.1038/sj.emboj.7601732
Horie T, Karahara I, Katsuhara M (2012) Salinity tolerance mechanisms in glycophytes: An overview with the central focus on rice plants. Rice (N Y) 5:11. https://doi.org/10.1186/1939-8433-5-11
Huang J, Wang JF, Zhang HS (2005) Rice ZFP15 gene encoding for a novel C2H2-type zinc finger protein lacking DLN box, is regulated by spike development but not by abiotic stresses. Mol Biol Rep 32:177–183. https://doi.org/10.1007/s11033-005-2338-0
Huang XY, Chao DY, Gao JP, Zhu MZ, Shi M, Lin HX (2009) A previously unknown zinc finger protein, DST, regulates drought and salt tolerance in rice via stomatal aperture control. Genes Dev 23:1805–1817. https://doi.org/10.1101/gad.1812409
Huang J, Sun SJ, Xu DQ, Lan HX, Sun H, Wang ZF, Bao YM, Wang JF, Tang HJ, Zhang HS (2012) A TFIIIA-type zinc finger protein confers multiple abiotic stress tolerances in transgenic rice (Oryza sativa L.). Plant Mol Biol 80:337–350. https://doi.org/10.1007/s11103-012-9955-5
Hussain S, Zhu CQ, Bai ZG, Huang J, Zhu LF, Cao XC, Nanda S, Hussain S, Riaz A, Liang QD, Wang LP, Li YF, Jin QY, Zhang JH (2019) iTRAQ-based protein profiling and biochemical analysis of two contrasting rice genotypes revealed their differential responses to salt stress. Int J Mol Sci 20. https://doi.org/10.3390/ijms20030547
Iftekharuddaula KM, Salam MA, Newaz MA, Ahmed HU, Collard BC, Septiningsih EM, Sanchez DL, Pamplona AM, Mackill DJ (2012) Comparison of phenotypic versus marker-assisted background selection for the SUB1 QTL during backcrossing in rice. Breed Sci 62:216–222. https://doi.org/10.1270/jsbbs.62.216
Ismail A, Takeda S, Nick P (2014) Life and death under salt stress: same players, different timing? J Exp Bot 65:2963–2979. https://doi.org/10.1093/jxb/eru159
Jain M, Khurana JP (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J 276:3148–3162. https://doi.org/10.1111/j.1742-4658.2009.07033.x
Ji HT, Pardo JM, Batelli G, Van Oosten MJ, Bressan RA, Li X (2013) The salt overly sensitive (SOS) pathway: established and emerging roles. Mol Plant 6:275–286. https://doi.org/10.1093/mp/sst017
Joshi R, Sahoo KK, Tripathi AK, Kumar R, Gupta BK, Pareek A, Singla-Pareek SL (2018) Knockdown of an inflorescence meristem-specific cytokinin oxidase-OsCKX2 in rice reduces yield penalty under salinity stress condition. Plant Cell Environ 41:936–946. https://doi.org/10.1111/pce.12947
Kang DJ, Seo YJ, Lee JD, Ishii R, Kim KU, Shin DH, Park SK, Jang SW, Lee IJ (2005) Jasmonic acid differentially affects growth, ion uptake and abscisic acid concentration in salt-tolerant and salt-sensitive rice cultivars. J Agron Crop Sci 191:273–282. https://doi.org/10.1111/j.1439-037X.2005.00153.x
Kaur N, Dhawan M, Sharma I, Pati PK (2016a) Interdependency of reactive oxygen species generating and scavenging system in salt sensitive and salt tolerant cultivars of rice. BMC Plant Biol 16:131. https://doi.org/10.1186/s12870-016-0824-2
Kaur N, Kirat K, Saini S, Sharma I, Gantet P, Pati PK (2016b) Reactive oxygen species generating system and brassinosteroids are linked to salt stress adaptation mechanisms in rice. Plant Signal Behav 11:e1247136. https://doi.org/10.1080/15592324.2016.1247136
Kazan K (2015) Diverse roles of jasmonates and ethylene in abiotic stress tolerance. Trends Plant Sci 20:219–229. https://doi.org/10.1016/j.tplants.2015.02.001
Khan MSA, Hamid A, Karim MA (1997) Effect of sodium chloride on germination and seedling characters of different types of rice (Oryza sativa L.). J Agron Crop Sci 179:163–169. https://doi.org/10.1111/j.1439-037X.1997.tb00512.x
Krishna P (2003) Brassinosteroid-mediated stress responses. J Plant Growth Regul 22:289–297. https://doi.org/10.1007/s00344-003-0058-z
Kuroha T, Nagai K, Gamuyao R, Wang DR, Furuta T, Nakamori M, Kitaoka T, Adachi K, Minami A, Mori Y, Mashiguchi K, Seto Y, Yamaguchi S, Kojima M, Sakakibara H, Wu J, Ebana K, Mitsuda N, Ohme-Takagi M, Yanagisawa S, Yamasaki M, Yokoyama R, Nishitani K, Mochizuki T, Tamiya G, McCouch SR, Ashikari M (2018) Ethylene-gibberellin signaling underlies adaptation of rice to periodic flooding. Science 361:181–186. https://doi.org/10.1126/science.aat1577
Kurotani K, Hayashi K, Hatanaka S, Toda Y, Ogawa D, Ichikawa H, Ishimaru Y, Tashita R, Suzuki T, Ueda M, Hattori T, Takeda S (2015) Elevated levels of CYP94 family gene expression alleviate the jasmonate response and enhance salt tolerance in rice. Plant Cell Physiol 56:779–789. https://doi.org/10.1093/pcp/pcv006
Li CR, Liang DD, Li J, Duan YB, Li H, Yang YC, Qin RY, Li L, Wei PC, Yang JB (2013) Unravelling mitochondrial retrograde regulation in the abiotic stress induction of rice ALTERNATIVE OXIDASE 1 genes. Plant Cell Environ 36:775–788. https://doi.org/10.1111/pce.12013
Li CH, Wang G, Zhao JL, Zhang LQ, Ai LF, Han YF, Sun DY, Zhang SW, Sun Y (2014) The receptor-like kinase SIT1 mediates salt sensitivity by activating MAPK3/6 and regulating ethylene homeostasis in rice. Plant Cell 26:2538–2553. https://doi.org/10.1105/tpc.114.125187
Lv B, Tian H, Zhang F, Liu J, Lu S, Bai M, Li C, Ding Z (2018) Brassinosteroids regulate root growth by controlling reactive oxygen species homeostasis and dual effect on ethylene synthesis in Arabidopsis. PLoS Genet 14:e1007144. https://doi.org/10.1371/journal.pgen.1007144
Ma B, He SJ, Duan KX, Yin CC, Chen H, Yang C, Xiong Q, Song QX, Lu X, Chen HW, Zhang WK, Lu TG, Chen SY, Zhang JS (2013) Identification of rice ethylene-response mutants and characterization of MHZ7/OsEIN2 in distinct ethylene response and yield trait regulation. Mol Plant 6:1830–1848. https://doi.org/10.1093/mp/sst087
Ma B, Yin CC, He SJ, Lu X, Zhang WK, Lu TG, Chen SY, Zhang JS (2014) Ethylene-induced inhibition of root growth requires abscisic acid function in rice (Oryza sativa L.) seedlings. PLoS Genet 10:e1004701. https://doi.org/10.1371/journal.pgen.1004701
Makhloufi E, Yousfi FE, Marande W, Mila I, Hanana M, Berges H, Mzid R, Bouzayen M (2014) Isolation and molecular characterization of ERF1, an ethylene response factor gene from durum wheat (Triticum turgidum L. subsp. durum), potentially involved in salt-stress responses. J Exp Bot 65:6359–6371. https://doi.org/10.1093/jxb/eru352
Marino D, Dunand C, Puppo A, Pauly N (2012) A burst of plant NADPH oxidases. Trends Plant Sci 17:9–15. https://doi.org/10.1016/j.tplants.2011.10.001
Martinez-Atienza J, Jiang XY, Garciadeblas B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012. https://doi.org/10.1104/pp.106.092635
Matsuda S, Nagasawa H, Yamashiro N, Yasuno N, Watanabe T, Kitazawa H, Takano S, Tokuji Y, Tani M, Takamure I, Kato K (2014) Rice RCN1/OsABCG5 mutation alters accumulation of essential and nonessential minerals and causes a high Na/K ratio, resulting in a salt-sensitive phenotype. Plant Sci 224:103–111. https://doi.org/10.1016/j.plantsci.2014.04.011
Miller G, Suzuki N, Ciftci-Yilmaz S, Mittler R (2010) Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ 33:453–467. https://doi.org/10.1111/j.1365-3040.2009.02041.x
Mittler R (2017) ROS are good. Trends Plant Sci 22:11–19. https://doi.org/10.1016/j.tplants.2016.08.002
Moons A, Prinsen E, Bauw G, Van Montagu M (1997) Antagonistic effects of abscisic acid and jasmonates on salt stress-inducible transcripts in rice roots. Plant Cell 9:2243–2259. https://doi.org/10.1105/tpc.9.12.2243
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Munns R, James RA, Gilliham M, Flowers TJ, Colmer TD (2016) Tissue tolerance: an essential but elusive trait for salt-tolerant crops. Funct Plant Biol 43:1103–1113. https://doi.org/10.1071/Fp16187
Nam MH, Bang E, Kwon TY, Kim Y, Kim EH, Cho K, Park WJ, Kim BG, Yoon IS (2015) Metabolite profiling of diverse rice germplasm and identification of conserved metabolic markers of rice roots in response to long-term mild salinity stress. Int J Mol Sci 16:21959–21974. https://doi.org/10.3390/ijms160921959
Nishiyama R, Watanabe Y, Fujita Y, Le DT, Kojima M, Werner T, Vankova R, Yamaguchi-Shinozaki K, Shinozaki K, Kakimoto T, Sakakibara H, Schmulling T, Tran LS (2011) Analysis of cytokinin mutants and regulation of cytokinin metabolic genes reveals important regulatory roles of cytokinins in drought, salt and abscisic acid responses, and abscisic acid biosynthesis. Plant Cell 23:2169–2183. https://doi.org/10.1105/tpc.111.087395
Park HY, Seok HY, Park BK, Kim SH, Goh CH, Lee BH, Lee CH, Moon YH (2008) Overexpression of Arabidopsis ZEP enhances tolerance to osmotic stress. Biochem Biophys Res Commun 375:80–85. https://doi.org/10.1016/j.bbrc.2008.07.128
Platten JD, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Horie T, Leigh RA, Lin HX, Luan S, Maser P, Pantoja O, Rodriguez-Navarro A, Schachtman DP, Schroeder JI, Sentenac H, Uozumi N, Very AA, Zhu JK, Dennis ES, Tester M (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11:372–374. https://doi.org/10.1016/j.tplants.2006.06.001
Prusty MR, Kim SR, Vinarao R, Entila F, Egdane J, Diaz MGQ, Jena KK (2018) Newly identified wild rice accessions conferring high salt tolerance might use a tissue tolerance mechanism in leaf. Front Plant Sci 9:417. https://doi.org/10.3389/fpls.2018.00417
Qin H, Deng Z, Zhang C, Wang Y, Wang J, Liu H, Zhang Z, Huang R, Zhang Z (2016a) Rice GDP-mannose pyrophosphorylase OsVTC1-1 and OsVTC1-3 play different roles in ascorbic acid synthesis. Plant Mol Biol 90:317–327. https://doi.org/10.1007/s11103-015-0420-0
Qin H, Wang Y, Wang J, Liu H, Zhao H, Deng Z, Zhang Z, Huang R, Zhang Z (2016b) Knocking down the expression of GMPase gene OsVTC1-1 decreases salt tolerance of rice at seedling and reproductive stages. PLoS One 11:e0168650. https://doi.org/10.1371/journal.pone.0168650
Qin H, Zhang Z, Wang J, Chen X, Wei P, Huang R (2017) The activation of OsEIL1 on YUC8 transcription and auxin biosynthesis is required for ethylene-inhibited root elongation in rice early seedling development. PLoS Genet 13:e1006955. https://doi.org/10.1371/journal.pgen.1006955
Qin H, Wang J, Chen X, Wang F, Peng P, Zhou Y, Miao Y, Zhang Y, Gao Y, Qi Y, Zhou J, Huang R (2019) Rice OsDOF15 contributes to ethylene-inhibited primary root elongation under salt stress. New Phytol. https://doi.org/10.1111/nph.15824
Qiu ZB, Guo JL, Zhu AJ, Zhang L, Zhang MM (2014) Exogenous jasmonic acid can enhance tolerance of wheat seedlings to salt stress. Ecotoxicol Environ Saf 104:202–208. https://doi.org/10.1016/j.ecoenv.2014.03.014
Quan RD, Wang J, Hui J, Bai HB, Lyu XL, Zhu YX, Zhang HW, Zhang ZJ, Li SH, Huang RF (2018) Improvement of salt tolerance using wild rice genes. Front Plant Sci 8:2269. https://doi.org/10.3389/fpls.2017.02269
Rana MM, Takamatsu T, Baslam M, Kaneko K, Itoh K, Harada N, Sugiyama T, Ohnishi T, Kinoshita T, Takagi H, Mitsui T (2019) Salt tolerance improvement in rice through efficient SNP marker-assisted selection coupled with speed-breeding. Int J Mol Sci 20. https://doi.org/10.3390/ijms20102585
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146. https://doi.org/10.1038/ng1643
Sah SK, Reddy KR, Li J (2016) Abscisic acid and abiotic stress tolerance in crop plants. Front Plant Sci 7:571. https://doi.org/10.3389/fpls.2016.00571
Saini S, Kaur N, Pati PK (2018) Reactive oxygen species dynamics in roots of salt sensitive and salt tolerant cultivars of rice. Anal Biochem 550:99–108. https://doi.org/10.1016/j.ab.2018.04.019
Schmidt R, Mieulet D, Hubberten HM, Obata T, Hoefgen R, Fernie AR, Fisahn J, San Segundo B, Guiderdoni E, Schippers JH, Mueller-Roeber B (2013) Salt-responsive ERF1 regulates reactive oxygen species-dependent signaling during the initial response to salt stress in rice. Plant Cell 25:2115–2131. https://doi.org/10.1105/tpc.113.113068
Seo YJ, Park JB, Cho YJ, Jung C, Seo HS, Park SK, Nahm BH, Song JT (2010) Overexpression of the ethylene-responsive factor gene BrERF4 from Brassica rapa increases tolerance to salt and drought in Arabidopsis plants. Mol Cell 30:271–277. https://doi.org/10.1007/s10059-010-0114-z
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279. https://doi.org/10.1111/ppl.12165
Shan C, Mei ZL, Duan JL, Chen HY, Feng HF, Cai WM (2014) OsGA2ox5, a gibberellin metabolism enzyme, is involved in plant growth, the root gravity response and salt stress. PLoS One 9:e87110. https://doi.org/10.1371/journal.pone.0087110
Sharma I, Ching E, Saini S, Bhardwaj R, Pati PK (2013) Exogenous application of brassinosteroid offers tolerance to salinity by altering stress responses in rice variety Pusa Basmati-1. Plant Physiol Biochem 69:17–26. https://doi.org/10.1016/j.plaphy.2013.04.013
Shi HZ, Ishitani M, Kim CS, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci U S A 97:6896–6901. https://doi.org/10.1073/pnas.120170197
Strader LC, Zhao Y (2016) Auxin perception and downstream events. Curr Opin Plant Biol 33:8–14. https://doi.org/10.1016/j.pbi.2016.04.004
Sun SJ, Guo SQ, Yang X, Bao YM, Tang HJ, Sun H, Huang J, Zhang HS (2010) Functional analysis of a novel Cys2/His2-type zinc finger protein involved in salt tolerance in rice. J Exp Bot 61:2807–2818. https://doi.org/10.1093/jxb/erq120
Suzuki K, Yamaji N, Costa A, Okuma E, Kobayashi NI, Kashiwagi T, Katsuhara M, Wang C, Tanoi K, Murata Y, Schroeder JI, Ma JF, Horie T (2016) OsHKT1;4-mediated Na+ transport in stems contributes to Na+ exclusion from leaf blades of rice at the reproductive growth stage upon salt stress. BMC Plant Biol 16:22. https://doi.org/10.1186/s12870-016-0709-4
Tao JJ, Chen HW, Ma B, Zhang WK, Chen SY, Zhang JS (2015) The role of ethylene in plants under salinity stress. Front Plant Sci 6:1059. https://doi.org/10.3389/fpls.2015.01059
Teixeira FK, Menezes-Benavente L, Galvao VC, Margis R, Margis-Pinheiro M (2006) Rice ascorbate peroxidase gene family encodes functionally diverse isoforms localized in different subcellular compartments. Planta 224:300–314. https://doi.org/10.1007/s00425-005-0214-8
Tester M, Davenport R (2003) Na+ tolerance and Na+ transport in higher plants. Ann Bot 91:503–527. https://doi.org/10.1093/aob/mcg058
Torres MA, Dangl JL (2005) Functions of the respiratory burst oxidase in biotic interactions, abiotic stress and development. Curr Opin Plant Biol 8:397–403. https://doi.org/10.1016/j.pbi.2005.05.014
Tsukagoshi H (2016) Control of root growth and development by reactive oxygen species. Curr Opin Plant Biol 29:57–63. https://doi.org/10.1016/j.pbi.2015.10.012
Tu Y, Jiang A, Gan L, Hossain M, Zhang J, Peng B, Xiong Y, Song Z, Cai D, Xu W, Zhang J, He Y (2014) Genome duplication improves rice root resistance to salt stress. Rice (N Y) 7:15–13. https://doi.org/10.1186/s12284-014-0015-4
Van de Poel B, Smet D, Van Der Straeten D (2015) Ethylene and hormonal cross talk in vegetative growth and development. Plant Physiol 169:61–72. https://doi.org/10.1104/pp.15.00724
Wang WB, Kim YH, Lee HS, Kim KY, Deng XP, Kwak SS (2009) Analysis of antioxidant enzyme activity during germination of alfalfa under salt and drought stresses. Plant Physiol Biochem 47:570–577. https://doi.org/10.1016/j.plaphy.2009.02.009
Wang H, Zhang M, Guo R, Shi D, Liu B, Lin X, Yang C (2012) Effects of salt stress on ion balance and nitrogen metabolism of old and young leaves in rice (Oryza sativa L.). BMC Plant Biol 12:194. https://doi.org/10.1186/1471-2229-12-194
Wang GF, Li WQ, Li WY, Wu GL, Zhou CY, Chen KM (2013) Characterization of rice NADPH oxidase genes and their expression under various environmental conditions. Int J Mol Sci 14:9440–9458. https://doi.org/10.3390/ijms14059440
Wang R, Jing W, Xiao L, Jin Y, Shen L, Zhang W (2015) The rice high-affinity potassium transporter1;1 is involved in salt tolerance and regulated by an MYB-type transcription factor. Plant Physiol 168:1076–1090. https://doi.org/10.1104/pp.15.00298
Wang Y, Zhao H, Qin H, Li Z, Liu H, Wang J, Zhang H, Quan R, Huang R, Zhang Z (2018) The synthesis of ascorbic acid in rice roots plays an important role in the salt tolerance of rice by scavenging ROS. Int J Mol Sci 19. https://doi.org/10.3390/ijms19113347
Wang WC, Lin TC, Kieber J, Tsai YC (2019a) Response regulator 9 and 10 negatively regulate salinity tolerance in rice. Plant Cell Physiol 29. https://doi.org/10.1093/pcp/pcz149
Wang PP, Shen LK, Guo JH, Jing W, Qu Y, Li WY, Bi RR, Xuan W, Zhang Q, Zhang WH (2019b) Phosphatidic acid directly regulates PINOID-dependent phosphorylation and activation of the PIN-FORMED2 auxin efflux transporter in response to salt stress. Plant Cell 31:250–271. https://doi.org/10.1105/tpc.18.00528
Wei Z, Li J (2016) Brassinosteroids regulate root growth, development, and symbiosis. Mol Plant 9:86–100. https://doi.org/10.1016/j.molp.2015.12.003
Welsch R, Wust F, Bar C, Al-Babili S, Beyer P (2008) A third phytoene synthase is devoted to abiotic stress-induced abscisic acid formation in rice and defines functional diversification of phytoene synthase genes. Plant Physiol 147:367–380. https://doi.org/10.1104/pp.108.117028
Wu X, He J, Chen J, Yang S, Zha D (2014) Alleviation of exogenous 6-benzyladenine on two genotypes of eggplant (Solanum melongena Mill.) growth under salt stress. Protoplasma 251:169–176. https://doi.org/10.1007/s00709-013-0535-6
Xia XJ, Wang YJ, Zhou YH, Tao Y, Mao WH, Shi K, Asami T, Chen Z, Yu JQ (2009) Reactive oxygen species are involved in brassinosteroid-induced stress tolerance in cucumber. Plant Physiol 150:801–814. https://doi.org/10.1104/pp.109.138230
Xia K, Wang R, Ou X, Fang Z, Tian C, Duan J, Wang Y, Zhang M (2012) OsTIR1 and OsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS One 7:e30039. https://doi.org/10.1371/journal.pone.0030039
Xiong Q, Ma B, Lu X, Huang YH, He SJ, Yang C, Yin CC, Zhao H, Zhou Y, Zhang WK, Wang WS, Li ZK, Chen SY, Zhang JS (2017) Ethylene-inhibited jasmonic acid biosynthesis promotes mesocotyl/coleoptile elongation of etiolated rice seedlings. Plant Cell 29:1053–1072. https://doi.org/10.1105/tpc.16.00981
Xu DQ, Huang J, Guo SQ, Yang X, Bao YM, Tang HJ, Zhang HS (2008) Overexpression of a TFIIIA-type zinc finger protein gene ZFP252 enhances drought and salt tolerance in rice (Oryza sativa L.). FEBS Lett 582:1037–1043. https://doi.org/10.1016/j.febslet.2008.02.052
Xu L, Zhao H, Ruan W, Deng M, Wang F, Peng J, Luo J, Chen Z, Yi K (2017) ABNORMAL INFLORESCENCE MERISTEM1 functions in salicylic acid biosynthesis to maintain proper reactive oxygen species levels for root MERISTEM activity in rice. Plant Cell 29:560–574. https://doi.org/10.1105/tpc.16.00665
Yang Y, Guo Y (2018a) Unraveling salt stress signaling in plants. J Integr Plant Biol 60:796–804. https://doi.org/10.1111/jipb.12689
Yang YQ, Guo Y (2018b) Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol 217:523–539. https://doi.org/10.1111/nph.14920
Yang CJ, Zhang C, Lu YN, Jin JQ, Wang XL (2011) The mechanisms of brassinosteroids' action: from signal transduction to plant development. Mol Plant 4:588–600. https://doi.org/10.1093/mp/ssr020
Yang C, Ma B, He SJ, Xiong Q, Duan KX, Yin CC, Chen H, Lu X, Chen SY, Zhang JS (2015) MAOHUZI6/ETHYLENE INSENSITIVE3-LIKE1 and ETHYLENE INSENSITIVE3-LIKE2 regulate ethylene response of roots and coleoptiles and negatively affect salt tolerance in rice. Plant Physiol 169:148–165. https://doi.org/10.1104/pp.15.00353
Ye H, Du H, Tang N, Li X, Xiong L (2009) Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Mol Biol 71:291–305. https://doi.org/10.1007/s11103-009-9524-8
Yichie Y, Brien C, Berger B, Roberts TH, Atwell BJ (2018) Salinity tolerance in Australian wild Oryza species varies widely and matches that observed in O. sativa. Rice (N Y) 11:66. https://doi.org/10.1186/s12284-018-0257-7
Yin CC, Ma B, Collinge DP, Pogson BJ, He SJ, Xiong Q, Duan KX, Chen H, Yang C, Lu X, Wang YQ, Zhang WK, Chu CC, Sun XH, Fang S, Chu JF, Lu TG, Chen SY, Zhang JS (2015) Ethylene responses in rice roots and coleoptiles are differentially regulated by a carotenoid isomerase-mediated abscisic acid pathway. Plant Cell 27:1061–1081. https://doi.org/10.1105/tpc.15.00080
Yin CC, Zhao H, Ma B, Chen SY, Zhang JS (2017) Diverse roles of ethylene in regulating agronomic traits in rice. Front Plant Sci 8:1676. https://doi.org/10.3389/fpls.2017.01676
Yoshikawa T, Ito M, Sumikura T, Nakayama A, Nishimura T, Kitano H, Yamaguchi I, Koshiba T, Hibara KI, Nagato Y, Itoh JI (2014) The rice FISH BONE gene encodes a tryptophan aminotransferase, which affects pleiotropic auxin-related processes. Plant J 78:927–936. https://doi.org/10.1111/tpj.12517
You J, Chan ZL (2015) ROS regulation during abiotic stress responses in crop plants. Front Plant Sci 6:1092. https://doi.org/10.3389/fpls.2015.01092
Zeng L, Shannon MC, Grieve CM (2002) Evaluation of salt tolerance in rice genotypes by multiple agronomic parameters. Euphytica 127:235–245. https://doi.org/10.1023/A:1020262932277
Zhang ZG, Zhang Q, Wu JX, Zheng X, Zheng S, Sun XH, Qiu QS, Lu TG (2013) Gene knockout study reveals that cytosolic ascorbate peroxidase 2 (OsAPX2) plays a critical role in growth and reproduction in rice under drought, salt and cold stresses. PLoS One 8:e57472. https://doi.org/10.1371/journal.pone.0057472
Zhang M, Smith JA, Harberd NP, Jiang C (2016a) The regulatory roles of ethylene and reactive oxygen species (ROS) in plant salt stress responses. Plant Mol Biol 91:651–659. https://doi.org/10.1007/s11103-016-0488-1
Zhang Y, Lan HX, Shao QL, Wang RQ, Chen H, Tang HJ, Zhang HS, Huang J (2016b) An A20/AN1-type zinc finger protein modulates gibberellins and abscisic acid contents and increases sensitivity to abiotic stress in rice (Oryza sativa). J Exp Bot 67:315–326. https://doi.org/10.1093/jxb/erv464
Zhang ZY, Liu HH, Sun C, Ma QB, Bu HY, Chong K, Xu YY (2018) A C2H2 zinc-finger protein OsZFP213 interacts with OsMAPK3 to enhance salt tolerance in rice. J Plant Physiol 229:100–110. https://doi.org/10.1016/j.jplph.2018.07.003
Zhao Y, Dong W, Zhang N, Ai X, Wang M, Huang Z, Xiao L, Xia G (2014) A wheat allene oxide cyclase gene enhances salinity tolerance via jasmonate signaling. Plant Physiol 164:1068–1076. https://doi.org/10.1104/pp.113.227595
Zhou Y, Yang P, Cui F, Zhang F, Luo X, Xie J (2016) Transcriptome analysis of salt stress responsiveness in the seedlings of Dongxiang wild rice (Oryza rufipogon Griff.). PLoS One 11:e0146242. https://doi.org/10.1371/journal.pone.0146242
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324. https://doi.org/10.1016/j.cell.2016.08.029
Zhu N, Cheng SF, Liu XY, Du H, Dai MQ, Zhou DX, Yang WJ, Zhao Y (2015) The R2R3-type MYB gene OsMYB91 has a function in coordinating plant growth and salt stress tolerance in rice. Plant Sci 236:146–156. https://doi.org/10.1016/j.plantsci.2015.03.023
Zhu T, Deng X, Zhou X, Zhu L, Zou L, Li P, Zhang D, Lin H (2016) Ethylene and hydrogen peroxide are involved in brassinosteroid-induced salt tolerance in tomato. Sci Rep 6:35392. https://doi.org/10.1038/srep35392
Zou X, Shao J, Wang Q, Chen P, Zhu Y, Yin C (2018) Supraoptimal cytokinin content inhibits rice seminal root growth by reducing root meristem size and cell length via increased ethylene content. Int J Mol Sci 19. https://doi.org/10.3390/ijms19124051
Funding
This work was funded by the Major Special Foundation of Transgenic Plants in China (grant number 2018ZX0800913B), the National Natural Science Foundation of China (grant numbers 31801445 and 31871551), and Innovation Project of Chinese Academy of Agricultural Sciences.
Author information
Authors and Affiliations
Contributions
All the authors discussed and created the review’s outline. Hua Qin wrote the manuscript. Rongfeng Huang edited the manuscript.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Rice Functional Genomics.
Rights and permissions
About this article
Cite this article
Qin, H., Huang, R. The phytohormonal regulation of Na+/K+ and reactive oxygen species homeostasis in rice salt response. Mol Breeding 40, 47 (2020). https://doi.org/10.1007/s11032-020-1100-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-020-1100-6