Abstract
Low yields of crops, especially rice, are caused by climate change and environmental stress concerns such as drought, temperature fluctuations, and salinity in arid and semi-arid locations around the globe. Rice is one of the essential crops for human consumption and one of the most commonly farmed cereals on the planet earth, but its growth is severely retarded by excessive salt, which influences rice development and production, leading to economic loss. Salt stress induces osmotic stress and ionic toxicity in rice by altering the environment, leading to water deprivation and accumulation of toxic ions, thereby triggering specific physiological and molecular responses in the rice plants. Many factors may affect rice production and cereal quality via its interaction with salinity. This review focuses on some influential factors (photosynthesis, osmosis, micro and macronutrients, microbial flora, rice growth, development, and genes) that may reduce rice production in saline soils. The review also describes the responsive mechanism of rice to salinity and the genetic susceptibility of rice. In light of the challenges posed by the growing global population and limited agricultural land, it is imperative to consider the influential factors discussed in this review, along with genetic susceptibility to improve rice production in terms of quantity and quality under saline soil conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are subjected to a variety of environmental stresses like drought, salinity, heat, cold, flooding, ozone, UV radiation, heavy metals, etc., in a changing world. These factors have been found to affect plant growth, yield, and productivity, posing a threat to food security (Benevenuto et al. 2017; Raza et al. 2019). Salinity is a soil condition characterized by high amounts of soluble salt and is one of the harshest abiotic stresses among environmental conditions (Cramer et al. 2011; Hasanuzzaman et al. 2013a, b). Salinity directly or indirectly affects about 7% of the world's total land area and one-third of the world's irrigated land area (Hopmans et al. 2021). Rice can withstand a modest amount of saltwater without hurting its development or yield in most cases. Nevertheless, the extent of this tolerance varies considerably depending on the specific types and species of rice and the stage of their development (Hasanuzzaman et al. 2013a). Notably, indica rice exhibits a higher level of tolerance compared to japonica rice (Lee et al. 2003; Xu et al. 2020). Rice is classified as a salinity-susceptible cereal during its early developmental stage, limiting its production efficiency at maturity (Todaka et al. 2012). Consequently, understanding the different rice varieties and their responses to salinity at various growth stages is crucial for optimizing production and enhancing overall agricultural efficiency.
Excessive salt in rice plants induces ionic toxicity and osmotic stress (Rahman et al. 2017; Yan et al. 2020). Rice plants exhibit various morphological, physiological, and biochemical changes and symptoms when exposed to excessive salt, leading to reduced biomass production and grain yield (Riaz et al. 2019). Sodium ions harm plants and a higher concentration of Na+ in the root zone decreases K+ uptake due to their antagonistic effect (Xiong et al. 2002; Hussain et al. 2022a). K+ is essential for maintaining membrane potential, enzyme activities, and cell turgor; a reduction in K+ will reduce plant development (Xiong et al. 2002; Das et al. 2015). Aside from Na+, other anions/cations, like Cl– are poisonous to rice. Ionic stress causes chlorosis and necrosis, which can hasten senescence or delay development and growth (Bisht et al. 2019). The excess accumulation of Na+ impedes cellular metabolisms, such as protein synthesis and enzyme activity, thereby disturbing the process of photosynthesis (Horie et al. 2012). Sodium accumulation in shoots is connected to rice plant survival under salinity and hence maintaining a lower cytosolic Na+, which is has been found to an important strategy for salt tolerance in glycophytes (Horie et al. 2012). Regulation of water transport becomes a critical adaptive mechanism for rice plants under osmotic stress, as a sufficient amount of water is essential for cell growth and physiological activities like photosynthesis and metabolism. Furthermore, it will close the stomatal pores, reducing evaporation and water transport (Horie et al. 2012). Many factors can influence the rice plant's developmental and growth process. The effect can be direct or indirect, occurring during developmental stage, at maturity, or at the productive stage, etc. Here, some important factors have been discussed to understand its potentiality and the mechanism to deal with it through plant physiology or genetic backup.
Influential factors and salinity
Effect on plant development, metabolism and microbial activities
Plant development and metabolism are affected by increased soil salt levels, resulting in lower productivity or severe symptoms, which may lead to death (Hameed et al. 2021; Xiao and Zhou 2023). Salts in soil water have been found to restrict development by interfering with water intake, inhibiting K+ uptake in plant cells, and resulting in an ion imbalance (Parihar et al. 2015). Plants deal with the negative effects of salt stress in various methods, including the selective exclusion of Na+, faster signaling networks, ion uptake regulation, and the development of osmotic-adjustable solutes (Reddy et al. 2017; Ponce et al. 2021). Plants collect organic or compatible solutes (trehalose, glycerol, proline, glycine betaine, sucrose, and glucose) to adjust for osmotic stress (Zivcak et al. 2016). Many studies have demonstrated that when plants are deprived of water and exposed to osmotic and salt stress, they acquire a variety of appropriate solutes (Gull et al. 2019; Gupta and Huang 2014; DeGomez 2019). Glycine betaine and proline are the most important solutes in osmotic adjustment linked to salt tolerance, free amino acids, and carbohydrates (Hajlaoui et al. 2010).
The presence of plenty of microorganisms in arid and semi-arid soils plays a crucial role in enhancing soil fertility, and promoting aeration, thereby influencing the growth of cultivated plants (Soussi et al. 2016). However, the deleterious impacts of soil salinity on microbial processes in agricultural land are well documented, with salinity primarily affecting microbial activity indicators such as respiration and enzymatic activity (Rath et al. 2015). Consequently, there is a decline in microbial biomass (Sardinha et al. 2003), which affects the enzymatic functions of microorganisms. To mitigate this effect and enhance microbial population activity in high saline or sodic soils, one such method is the incorporation of compost, which has shown the potential to reduce salinity-induced toxicity (Lakhdar et al. 2009) (Fig. 1).
Organic amendment for saline soils
Soil is a mixture of many chemical ingredients which may be present inside the soil; and the plant's requirement for these ingredients depends on their potential for development and growth (DeGomez 2019). The significant reasons for agricultural production losses in irrigated areas are soil salinity produced using water with a medium salt concentration in solution and excessive fertilizer application (Pla-Sentís 2021). Aside from erosion and salinization, have caused annual loss of $27.3 billion, in irrigation land, affecting approximately 20% of the worlds lands.causing (Mustafa 2007; Qadir et al. 2014).
Organic matter (OM) has been linked to enhanced soil quality due to radicular growth and the generation of root exudates like organic acids, which regulate soil pH and reduces the negative impacts of soil salt concentration; hence boosting the accessibility of nutrients (DeLuca and DeLuca 1997; Ernst et al. 2004). Under saline circumstances, organic and biofertilizers have been demonstrated to boost rice output (Zayed et al. 2013). The treatments of compost horticultural remains mixtures considerably boosted the field cation exchange capacity "CEC" (Wang et al. 2014; Asaye et al. 2022). The addition of compost mixture raises nitrogen (N), phosphorous (P), and potassium (K) levels, which may be related to organic additions and the number of accessible macronutrients (Table 1).
Most saline soils exhibit notable N, P, and K deficiencies, impacting plant growth and development (Shereen et al. 2005). The accessible proportion of potassium may rise due to increased cation exchange capacity (CEC) associated with compost. In saline soils, compost mineralization can enhance the proportion of available potassium to plants in at least two ways: firstly, it increases the (CEC) and the contribution of cations held in the compost's negative charges, which gradually becomes available over time (Bhattacharyya et al. 2007; Dhanorkar et al. 1994). Secondly, during compost mineralization, the decomposition process leads to the release of potassium and other vital nutrients into the soil through microbial breakdown, thus enhancing their accessibility to plants (Bhattacharyya et al. 2007; Dhamodharan et al. 2015). These combined processes create a more favorable environment for plant growth in saline soils.
Salinity in the soil has been shown to have a deleterious effect on soil fertility (Mohanavelu et al. 2021). Rice plants grown in saline environments face a nutritional imbalance in the absorption and transport of imperative plant elements, i.e., N, Ca, K, Mg, and Zn, resulting in decreased growth and production (Zayed et al. 2013). Therefore, it contends that the usage of compost has alleviated this discrepancy, at least in part (Litardo et al. 2022). Litardo et al. (2022) investigated the effects of different treatments, including leonardite, gypsum, compost, sugarcane cake, pig manure, and control applications, to discover that compost application was usually linked to the utmost accumulation of integral macronutrients in rice grains, although no significant variations were observed in the leaves.
Ion imbalances in salt-stressed plants
Salinity stress can lead to ion imbalance and toxicity in plants due to intense competition between Na+ and Cl– ions and other needed ions such as K+, Ca2+, and NO3– (Assaha et al. 2017). This stress induces a nutritional imbalance in plants, characterized by reduced levels of N, P, K, Ca, and Mg and increased levels of ratios (Jouyban 2012; Razzaque et al. 2011). Ionic imbalance occurs when excessive Na+ and Cl– build up in plant cells and tissues, preventing the uptake of other nutrients. According to Lee et al. (2009), high salt stress alters the concentrations of Na+, Ca2+, K+, and Mg2+ in the root and shoot of rice plants. When plants were exposed to high salt levels, their boron (B) and silicon (Si) availability was reduced (Wimmer et al. 2001; Currie et al. 2007). High NaCl concentrations increased cadmium (Cd) toxicity and lowered zinc (Zn) availability in various cereal crops, including rice (Amanullah et al. 2016).
Effects of salt stress on rice growth and development
Salinity affects both the developmental and growth stages of crop plants. This effect depends on the salt concentration in the soil, type of salt constituents, and type of crop (rice) cultivar, as some rice cultivars are comparatively more susceptible than others. The salinity may affect different effects at different climatic conditions and plant developmental stages.
Effect of salt stress on rice plant growth
For sufficient rice production, plant growth is the primary factor influencing it. The growth may be at the root of getting a sufficient amount of nutrients/solutes and getting an adequate amount of nutrients/solutes. If the growth of the root is affected, the plant will be unable to intake an adequate amount of food, and thus, the life span and survival capability of the plant will decrease. Retardation of the upper parts, like stems and leaves, directly affects plant production. Under saline conditions, the leaves may be unable to do activities normally like photosynthesis, transpiration, etc. (Negrão et al. 2017). Rice crop is more susceptible to salinity stress in the seedling stage than during the tillering stage (Grattan et al. 2002). The root interacts directly with the soil environment's biotic and abiotic features and the rest of the plant (De-Smet et al. 2012). Salinity can affect cell division and cell elongation in rice plants, which causes a lessening in the growth of roots, leaves, and the whole crop (Munns 2002). In root growth, organelles, nuclei stability, and normal epidermal cell frequency, tetraploid rice (HN2026-4x and Nipponbare-4x) beat diploid rice (Tu et al. 2014a, b). This type of rice has lowered the Na+ in the roots of the rice plant. This is due to the exposure of a protective gap between pericycle and cortical cells in the rice (Tetraploid), which raises root confrontation against salinity by enhancing the transport of H+ ions on the surface and decreasing the entry of Na+ in rice roots (Tu et al. 2014a, b; Reddy et al. 2017).
At the early seedling stage, enhanced salt stress increased rice leaf mortality in all rice cultivars (Shereen et al. 2005). It may cause leaf mortality and reduce the leaf area, lowering the plant's photosynthetic rate (Amirjani 2011). The process of metabolism inside leaf senescence is affected by salt stress. It can also negatively affect the cells of transpiring leaves, causing them to grow more slowly. Higher concentrations of salt in soil, especially affect older leaves, thus leading to the death of the leaves, which is necessary for the plant's survival (Munns et al. 2006).
Salt stress and rice grain yield
Paniculate sterility is a severe problem in rice grain production under salt stress (Hussain et al. 2017). Due to the salinity effect's genetic processes and food deficits, panicle sterility is caused in many rice cultivars, particularly during the pollination and fertilization stages (Hassanuzzaman et al. 2009; Pariha et al. 2015). Multiple studies have indicated that salt stress after fertilization causes panicle sterility, decreasing grain setting, pollen-bearing capability, stigmatic surface, or both (Abdullah et al. 2001; Arif et al. 2020). Lower grain yield during salt stress is due to a lack of glucose transition to vegetative and spikelet development (Liu et al. 2022a, b). Many yield factors, including the number of tillers plant−1, spikelets panicle−1, and the % sterile florets, have negative linear relationships with increased salt stress (Zeng and Shannon 2000; Huang et al. 2020).
Furthermore, salinity stress significantly reduces soluble sugar translocation to superior and inferior spikelets and suppresses starch synthase activity during grain development, contributing to poorer rice grain yield (Abdullah et al. 2001; Arif et al. 2020); Chen et al. 2019). Salinity stress negatively impacts various aspects of rice growth, including reduced spikelet number, panicle length, number of tillers per plant, florets number, and 1000-grain weight (Aref et al. 2012). It is necessary to comprehend the phenomenon of salt stress concerning rice yield reduction (Hussain et al. 2017).
Effect of salt stress on rice physiological characteristics
The physiological effects of salt stress on rice plants include reduced PAR (photosynthetically active radiation), Pn (net photosynthesis), Gs (stomatal conductance), Tr (transpiration rate), pigment degradation, and RWC (relative water content) (Thompson et al. 2006). Rice plant water usage efficiency (WUE) is considerably affected by salinity stress (Ramezani et al. 2012). "WUE" is related to biomass and seed production in a beneficial way. All of these factors/parameters growth has a negative pleiotropic effect on rice development and physiological characteristics (Amanullah et al. 2016) and can also induce aberrant rice development and growth at both biochemical and molecular levels (Nishimura et al. 2011).
Effects of salinity on photosynthesis
Photosynthesis is the main physiological factor, a critical metabolic process that provides essential ingredients for plant development and survival, and has been affected by salinity (Chaves et al. 2009; Ponce et al. 2021). Under salt stress, the photosystem II (PSII) complex exhibits a decreased photosynthetic efficiency in rice plants. According to Suriyan (2009), the amount of chlorophyll and carotenoids in rice leaves was dramatically reduced after salinity stress treatments. Reduction in the ratio of quantum yield of the PSII and K+ /Na+ is related to high salt stress (García Morales et al. 2012).
Maintaining a net photosynthetic rate (Pn) under stress is linked to salt tolerance (Liu et al. 2011). Nounjan et al. (2020) investigated the photosynthetic effect of salinity on rice cultivars, finding that KDML105 had a significantly lower rate of photosynthesis than CSSLs. In all genotypes studied, a drop in Pn was closely associated with a decrease in Gs. This demonstrates that Pn is predominantly regulated by stomata closure, restraining the delivery of CO2 to the Calvin cycle. Apart from stomatal variables, other parameters, such as the ratio of Ci/Ca in response to Gs, have decreased Pn during salt stress (Dadkhah 2011). Under stress, Ci/Ca at upper levels in KDML105 and CSSL8-103 proposed that these genotypes have more photosynthetic non-stomatal limitations than the other genotypes (Liu et al. 2011). Under salt stress, non-stomatal restriction factors may slow photosynthesis by lowering mesophyll conductance (Wang et al. 2018) and modifying ATP generation, resulting in ribulose bisphosphate (RuBP) shortage (Lawlor et al. 2009) and reduced activity of essential CO2 fixation enzymes. Liu et al. (2011) reported that Pokkali had the greatest Pn value, and salt stress did not affect the PSII in Pokkali and KDML105. It has been discovered that the stability of both photochemical machinery (PSII and PSI) is linked to the speedy recovery of the photosynthetic rate (Yi et al. 2016). Photosynthesis and energy absorption require chlorophyll pigments. Hence, the long-term stability of PSII could help improve the process of photosynthesis when the plant recovers from salt after stress, resulting in plant growth.
Salinity stress and chlorophyll
Chlorophyll pigments are essential for the process of photosynthesis. High salinity has been found to activate some enzymes promoting the breakdown of chlorophyll; as a result, the concentration of chlorophyll is decreased and influences the process of photosynthesis (Kordrostami et al. 2017). According to Nounjan et al. (2020), when subjected to salinity treatment, two rice cultivars (CSSL8-103 and KDML105) exhibited a reduction in chlorophyll levels as measured by SPAD values. In contrast, Pokkali, DH103, and CSSL8-106 plants did not show any change in chlorophyll levels under the same conditions. The decrease in chlorophyll ratio (a:b) due to salt stress is associated with reduced SPAD values and total chlorophyll contents, as demonstrated in the study by Mahlooji et al. (2018). Moreover, the chlorophyll concentration decreased in both KDML105 and CSSL8-103 varieties. Salinity has also been found to disintegrate thylakoids, resulting in reducing chlorophyll (a:b ratio) (Shu et al. 2012) due to accumulation and toxicity caused by Na+ (Yeo and Flowers 1983; Blum 2018). The effect of salinity has been found in the chlorophyll contents in various rice varieties (Lutts et al. 1996; Tatar et al. 2010).
Osmotic regulation in salt-stressed plants
Maintaining a low osmotic potential in plant cells due to a high accumulation of osmolytes is known as osmoregulation or osmotic adjustment (Turner 2018). Under abiotic stress, plant protection and survival depend on distinguishing between compatible and non-compatible solutes. It depends on the physiology of the plant or its genotypes, as Nounjan et al. (2020) reported two different cultivars of rice with different susceptibility after its treatment with salt stress. Rice type Pokkali was found to have the lowest osmotic potential than KDML105, which has the highest osmotic potential. On the other hand, both DH103 and CSSLs cultivars have moderate osmotic potential (Nounjan et al. 2020). This is due to the ability of the cultivar to accumulate sugar or other solutes. Salt-resistant genotypes in many plant species have accumulated much sugar in reaction to salt stress (Hajlaoui et al. 2010). When salt stress was eliminated from all rice lines/cultivars, osmotic potential increased, and sugar content decreased, indicating that plants may recover to normal growth conditions (Nounjan et al. 2020).
Intracellular compartmentalization
One of the most important considerations in preventing plant salt damage is limiting excessive Na+ transfer to the shoots (Nishimura et al. 2011). Plants have the ability to move hazardous elements to older leaves, as well as leaf sheaths, to save the young developing tissues. Long-term salt transfer into transpiring leaves leads to extremely high Cl– and Na+ applications, as well as the leaves eventually peris h (Amanullah et al. 2016). The speed at which leaves die plays a critical role in plant's survival (Amanullah et al. 2016). There may be enough photosynthesizing leaves for the plant to generate flowers and seeds if the rate of new leaf development continues to outstrip the rate of old leaf demise. The plant may not be able to replicate seeds if the death rate exceeds the rate of growth (Li et al. 2011). The plants ability to isolate ions in older leaves and structural tissues may have a big impact on plant longevity, especially in the case of rice cultivars.
When exposed to salinity, old leaves collect significantly more Na+, Cl–, and NO3– compared to young leaves (Nath et al. 2016). While up-regulation of OsHKT1; 1, OsHAK10, as well as OsHAK16 in response to salt enhances Na+ accumulation in old leaves, augmented OsNHX1 expression causes Na+ compartmentalization in old leaves (Ren et al. 2005; Nath et al. 2016). In salt-tolerant varieties have a significantly lesser salt concentration in the panicle compared to husks and rachis, with grains having the lowest concentration (Parida and Das 2005; Isayenkov 2019).
Approaches to improving rice salt tolerant
Apart from conventional methods, new techniques are now used to overcome the stress and enhance the plant's production. With the increase in the human population, food scarcity is a significant problem. So, scientists are searching for susceptible cultivars which may be able to survive in saline soils and produce sufficient grain and fodder for livestock or other industries.
The plant breeding
The plant breeding approach is one of the approaches that may offer assistance in managing the circumstance. The culmination strategy is to identify germplasm lines with salt-tolerant traits that can help transmit resilience qualities into rice cultivars. A salt-tolerant genotype should be able to show resistance in the presence of Na+ concentrations (lower) while sustaining the concentration of K+. Marker-assisted selection (MAS) uses molecular markers associated with tolerance traits that have been incorporated to speed up traditional breeding (MAS). The genetic variation may lead to discovering economical molecular markers and novel alleles of interest, potentially yielding salt-tolerant cultivars. Seven (CR1009, ADT45, Pusa 44, MTU1010, Gayatri, PR114 and Sarjoo 52) major regionally adapted rice cultivars have been transplanted with Saltol, a significant QTL for salinity susceptibility (Singh et al. 2016). About 376 SNP were found in Saltol QTL for salt tolerance (Rahman et al. 2016). Rahman et al. 2016 reported about seven landraces that can accumulate lower Na+ and higher levels of K+ in leaves leading to a reduction in the ratio of Na+/K+.
These rice landraces offer an additional reservoir of salt tolerance genes, contributing to diverse sources of genetic salt tolerance and producing varieties with functional alleles for different tolerance traits. It is the need of the day to investigate candidate genes and potential mutation/allelic variation for higher production and salt tolerance (Das et al. 2014).
The system biology
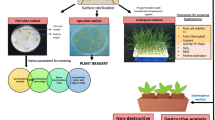
Several methodologies have been under consideration to better understand and deconstruct the rice resistance mechanism against salinity. Some of these strategies, like phenomics, transcriptomics, proteomics, and metabolic research, is required, which will aid in producing salt-tolerant germplasms. In fundamental research, many phenotyping technologies have been used, including noninvasive ones that use visible or fluorescent imaging. Visible chlorophyll fluorescence imaging, red–green–blue imaging, hyperspectral imaging, and Thermo imaging are high-throughput phenotyping platforms (Humplík et al. 2015). Image analysis was used to distinguish between two types of salinity stresses (ion-dependent and independent), one of the main tools for the physiological and genetic understanding of rice salt tolerance/resistance pathways (Hairmansis et al. 2014; Hussain et al. 2022b). Two rice varieties (Fatmawati and IR64) have been assessed for salt tolerance with the help of a non-destructive image-based phenotyping procedure, and the rice response to salinity could be readily discriminated from the control.
The knowledge of the function of candidate genes of interest, the germplasm for salt tolerance, and plant physiology research can assist in bridging the gap amid genotype and phenotype (Sexcion et al. 2006). Transpiration through stoma is the most prevalent transpiration type in plants. Even when transpiration is limited, salt tolerance can be achieved by increasing water intake capacity at night. Plant phenomics can help build diverse tolerance mechanisms into breeding lines by revealing the genetic basis for salt tolerance mechanisms (Hairmansis et al. 2014). The study of complete transcriptome changes in various biological situations is known as transcriptomics (Thompson et al. 2006). Microarrays detect the expression of all genes expressed in a single trial, thus recognized as the gold standard for analyzing genome-wide transcriptional response (Zhou et al. 2007). Using the microarray approach, rice salinity inducible transcripts were found (Kumari et al. 2009), for example, (Dongxiang wild rice) with the help of the Illumina HiSeq 2000 platform (Zhou et al. 2016; Zhou et al. 2007). Pandit et al. (2010) reported using two strategies (bulked segregant analysis and genome-wide microarray) to explore putative salt tolerance genes utilizing differential transcriptome analysis.
The study of protein molecules both at the structural and functional levels is known as proteomics. The most basic and straightforward technique for explaining the function of a gene is to understand the protein structure and function (Lee et al. 2009). OsRMC is a root-secreted protein that is reported to be strongly responsive under salt stress (Zhang et al. 2009). Mishra et al. (2016) studied metabolic pathways related to salinity tolerance by investigating CSR27, salt-sensitive MI48, along with their extremely tolerant and sensitive RIL progenies. Variations were observed in the production of different antioxidant enzymes, amino acids, and proteins across various rice genotypes, and their genes are co-located in QTLs mapped with the bi-parental population method, implying its role in salt tolerance. Another approach is selective nuclear magnetic resonance-based metabolomics was used to investigate salt sensitive and salt-tolerant variations (Nam et al. 2015). With the help of this technique, it was explored that the shift in carbohydrate and aliphatic areas is the most important factor in determining salt tolerance and salt sensitivity. Nam et al. (2015) established a relationship between metabolite changes and rice genotypes' salt tolerance and growth potential. Salt tolerance QTLs have been discovered on chromosome 1 in rice, notably Saltol (OsHKT1; 5) from Nona Bokra and SKC1 (OsHKT1; 5) from Pokkali. Salton is hypothesized to play a function in maintaining Na+/K+ equilibrium in salinity stress (Platten et al. 2013). Genetic polymorphism techniques like single nucleotide polymorphism (SNP) and simple sequence repeat (SSR) were used to find salt-tolerant rice variations (Dhar et al. 2011). Along with salt tolerance, some traits were also discovered through a transgenic approach, like the yield/production discovered by the golden gate SNP array (Kurokawa et al. 2016). Tu et al. (2014a, b) proposed that the duplication of the genome has increased root salt tolerance and the transport of proton to the root's surface, reducing the Na+ accumulation.
The transgenic method
In this type of approach, recombinant DNA technology (ies) is utilized to introduce plants with desirable and potentially novel features. In transgenic plants, the overexpression of OsPP1a has increased OsNAC6, SnRK1A, and OsNAC5, which showed better resistance to high salt treatment (Liao et al. 2016). Overexpression of OsNHX1 in rice has also been associated with enhanced biomass output and germination and altered Na+ and K+ accumulations in the shoots and roots (Ishak et al. 2020; Chen et al. 2007). OsTPS1 Overexpression has been found to be related to trehalose-6-phosphate synthase in rice plants and is linked to higher compatible solute accumulation. This mechanism helps improve plant life and photosynthetic efficiency, thus increasing plant growth and reducing its wilting (Li et al. 2011). Transgenic rice plants are more susceptible to salinity and a higher amount of Na+. Another gene (PDH45) has been similarly capable of resistance against salinity (Nath et al. 2016). These findings indicate that PtCYP714A3 has a vital role in how rice shoots react to salt's harmful effects. This discovery provides a groundwork for creating genetically engineered crops that withstand high salt levels (Wang et al. 2016). Developing stress-tolerant rice varieties is crucial to managing salt stress and ensuring high-yield rice production. All the international and national research organizations, like IRRT and NRSP in Asia, are mandated to enhance plant production.
The contribution of candidate genes to salt tolerance
The candidate gene approach is another landmark for producing salt-tolerant plants with high production capability for both grain and fodder. The accumulation of Na+ and Cl– ions in the cytoplasm at Na+ and Cl– ions in the cytoplasm at a higher amount is physiologically harmful to plants (Wang et al. 2016). Rice contains a conserved SOS pathway, and OsSOS1, a Na+/H+ antiporter, lowers the cell's total Na+ level (Kumari et al. 2013). Yasmine et al. (2015) converted the rice variant BRRI than 28 into a salt-tolerant variety using the OsSOS1 gene to a salt-tolerant variety capable of resisting salinity levels of 150–200 mmol/L. In the context of salinity, Diédhiou and team (2008) found that SAPK4 plays a crucial role in maintaining ion balance, growth, and development. Overexpression of OsHAKs and OsAKT1 in alkali-stressed shoots may assist in maintaining potassium nutrition by allowing K+ to be released from roots to shoots or taken up by roots (Yang et al. 2014). However, at the higher concentration, the Na+/K+ ratio externally has a connection between AKT1 expression and root Na+, suggesting that AKT1 may be involved in Na+ absorption. OsHAK5 overexpression transgenic rice plants developed faster than wild-type seedlings in the 100 mmol/L NaCl (Yang et al. 2014). The integration of high-affinity K+ transporter genes from wild type into modern high-yielding could offer a solution for boosting rice production in salt-affected regions. Miyamoto et al. (2015) noted that enhancing OsHKT2:1 overexpression led to increased shoot Na+ accumulation in low K+ conditions and hypothesized that OsHKT2:1 expression plays a pivotal role in determining rice varieties' potential for Na+ accumulation.
Increases in the number of vacuolar Na+/H+ antiporters (also known as vacuolar H+ pyrophosphatases), glycine betaine, osmoprotectants such as proline, and enzymes involved in the ROS detoxification will yield diverse impacts on rice salt tolerance. For example, HKT1;5 and HKT2;3 haplotypes H5 and H1 are linked to high salt tolerance of about 150 mmol/L NaCl (Ernst et al. 2004). OsHKT1;1 was found to control the amount of salt in phloem sap and protect leaf blades against toxicity (Wang et al. 2015; Chen et al. 2013). OsHKT1;4 removed Na+ from the xylem to prevent Na+ in the leaf blades (Cotsaftis et al. 2012). Additionally, PDH45 is employed to engineer transgenic rice that can survive salt chloride concentrations of 200 mmol/L (Nath et al. 2016). The overexpression of the SIDP361 gene both at the heading and seedling stages of transgenic rice plants was highly tolerant to the salt stress of about 200 mmol/L NaCl (Li et al. 2016). Rice OsNHX1 overexpression improves salt tolerance by promoting sodium compartmentalization into vacuoles (Chen et al. 2007). The transgenic approach was also found to alter or influence the physiology of rice plant-like photosynthesis. In the presence of salt stress of about 50–150 mmol/L, the transgenic plant (Nipponbare) with an introgressed gene (OsNHX1) and a large leaf area have greater photosynthetic rates as compared to the wild variety of rice (Xiang et al. 2018; Chen et al. 2021).
Rice transgenic plants fed salt-containing media had a notably enhanced shoot and root, as well as greater fresh weight per plant compared to wild-type plants. The Over-expression of the OsCIPK15 gene in transgenic rice has increased salt tolerance, growth, and production compared to wild rice, where the gene is absent or down-regulated (Xiang et al. 2007). The stress resistance profiles of nine salt-tolerant rice types and transgenic rice lines having intrinsically expressed genes, presumably implicated with salt tolerance, were evaluated by examining their growth and viability under conditions of hyperosmotic, temperature, ionic, and salt stress. The amplification of the CYP94C2b gene promotes jasmonate deactivation (Kurotani et al. 2015). Organ and product specificity can be ensured in the candidate gene approach to get the desired results/products. In salt-tolerant lines, root-specific genes (such as Os11g34460 Per-ARNT-Sim motif, Os02g02170, and OsTIP2; 1 aquaporin) were found to have a higher affinity for the nitrate transporter gene, which can be paired with other genes for further research (Cotsaftis et al. 2011).
Salt stress did not affect GSA, Os08g41990, or Os02g07230 expression in Pokkali, DH103, or CSSL8-106, although KDML105 and CSSL8-103 showed a more drastic down-regulation. An aminotransferase enzyme generated by the GSA gene transforms glutamate 1-semialdehyde (GSA) to -aminolevulinic acid (ALA) (Dalal and Tripathy 2012). Os08g41990 is involved in the biosynthesis of tetrapyrroles and porphyrins, which is a crucial step in the chlorophyll manufacturing process (Nounjan et al. 2020), whereas Os02g07230 (porphobilinogen deaminase) polymerizes porphobilinogen molecules to produce 1-hydroxymethylbillane, a linear tetrapyrrole Os02g07230 (Nounjan et 2018). KDML105 and CSSL8-103 show lower GSA, Os08g41990, and Os02g07230 expression than Pokkali, DH103, and CSSL8-106, indicating that these genotypes are more vulnerable to salt stress. According to Dalal and Tripathy (Dalal and Tripathy 2012), demonstrated a connection between reduced chlorophyll production in stressed rice seedlings and the downregulation of preliminary intermediates of chlorophyll biosynthesis, such as GSA and ALA. As a result, the expression changes in Os08g41990 led to a moderate reduction in chlorophyll in PK, DH103, and CSSL8-106, but a dramatic decrease in KDML105 and CSSL8-103. Tetrapyrrole has been shown to activate the ROS detoxifying mechanism and boost drought stress signaling (Nagahatenna et al. 2015), and plants require porphyrin formation to continue photosynthesis under drought stress (Tanaka and Tanaka 2007). These findings led to the conclusion that higher levels of Os08g41990 are linked with improved salt stress tolerance. The gene's stability under salt stress could be linked to the retention of chlorophyll, ultimately preservating photosynthetic efficiency. Under stress conditions, the gene Os02g07230 can also affect photosynthetic pigment synthesis (Cornah et al. 2002). In contrast to CSSL8-103, where these two genes are highly expressed and encode enzymes for earlier steps in the chlorophyll biosynthesis pathway, the Os02g07230 gene, which is responsible for the first step in porphyrin biosynthesis, exhibits a marked reduction in expression (Tanaka and Tanaka 2007). Interestingly, chlorophyll production genes were downregulated during stress, whereas the gene responsible for chlorophyll breakdown were upregulated (Liu et al. 2018; Gan et al. 2022), and the reverse was observed after recovery. The amount of chlorophyll was quantified using the complex activities of chlorophyll production and degradation genes, alongside the flow of intermediatary compounds within the process (Dalal and Tripathy 2012). As a result, more research is needed into the mechanisms that impact chlorophyll metabolic pathways during and after salt stress.
Conclusion and future perspectives
This review focuses on some influential factors (photosynthesis, osmosis, micro and macronutrients, microbial flora, growth, rice development, and genes) that play pivotal roles in the development and growth of rice within saline soil. The review also describes rice’s significant impact on the microbial flora present in the soil, as it can profoundly influence the rice plant's health and productivity. Moreover, the role of genes in shaping the plant's response to salinity is thoroughly explored, uncovering the genetic susceptibility that underlies the rice plant's adaptive strategies. The influential factors and genetic susceptibility needed in the rice breeding program offer tremendous potential. It uncovers valuable coping mechanisms that enable the rice plant to withstand and thrive under salinity-induced stress. These approaches are projected to augment both the quantitative and the qualitative production of rice in the future. This comprehensive approach is incredibly beneficial in enhancing agriculture endeavors toward a more sustainable and prosperous future, crucially meeting the escalating demands for sustenance posed by an ever-growing global population.
References
Abdullah Z, Khan MA, Flowers T (2001) Causes of sterility in seed set of rice under salinity stress. J Agro Crop Sci 187:25–32
Amanullah I, Inamullah X (2016) Dry matter partitioning and harvest index differ in rice genotypes with variable rates of phosphorus and zinc nutrition. Rice Sci 23:78–87
Amirjani MR (2011) Effect of salinity stress on growth, sugar content, pigments and enzyme activity of rice. Int J Bot 7:73–81
Aref F, Rad HE (2012) Physiological characterization of rice under salinity stress during vegetative and reproductive stages. Ind J Sci Technol 5:2578–2586
Arif Y, Singh P, Siddiqui H, Bajguz A, Hayat S (2020) Salinity induced physiological and biochemical changes in plants: an omic approach towards salt stress tolerance. Plant Physiol Bioch 156:64–77
Asaye Z, Kim DG, Yimer F, Prost K, Obsa O, Tadesse M, Gebrehiwot M, Brüggemann N (2022) Effects of combined application of compost and mineral fertilizer on soil carbon and nutrient content, yield, and agronomic nitrogen use efficiency in maize-potato cropping systems in southern ethiopia. Land 11:784
Assaha DV, Ueda A, Saneoka H, Al-Yahyai R, Yaish MW (2017) The role of Na+ and K+ transporters in salt stress adaptation in glycophytes. Front Physiol 8:509
Benevenuto RF, Agapito-Tenfen SZ, Vilperte V, Wikmark OG, van Rensburg NRO (2017) Molecular responses of genetically modified maize to abiotic stresses as determined through proteomic and metabolomic analyses. PLoS ONE 12:e0173069
Bhattacharyya P, Chakrabarti K, Chakraborty A, Nayak D, Tripathy S, Powell M (2007) Municipal waste compost as an alternative to cattle manure for supplying potassium to lowland rice. Chemosphere 66:1789–1793
Bisht N, Tiwari S, Singh PC, Niranjan A, Chauhan PS (2019) A multifaceted rhizobacterium Paenibacillus lentimorbus alleviates nutrient deficiency-induced stress in Cicer arietinum L. Microbiol Res 223:110–119
Blum A (2018) Plant breeding for stress environments. CRC Press
Chaves MM, Flexas J, Pinheiro C (2009) Photosynthesis under drought and salt stress: regulation mechanisms from whole plant to cell. Ann Bot 103:551–560
Chen M, Chen QJ, Niu XG, Zhang R, Lin HQ, Xu CY, Wang GY, Chen J (2007) Expression of OsNHX1 gene in maize confers salt tolerance and promotes plant growth in the field. Plant Soil Enviro 53:490
Chen T, Xu Y, Wang J, Wang Z, Yang J, Zhang J (2013) Polyamines and ethylene interact in rice grains in response to soil drying during grain filling. J Exper Bot 64:2523–2538
Chen L, Deng Y, Zhu H, Hu Y, Jiang Z, Tang S, Wang S, Ding Y (2019) The initiation of inferior grain filling is affected by sugar translocation efficiency in large panicle rice. Rice 2:75
Chen T, Shabala S, Niu Y, Chen ZH, Shabala L, Meinke H, Venkataraman G, Pareek A, Xu J, Zhou M (2021) Molecular mechanisms of salinity tolerance in rice. The Crop J 9:506–520
Cornah JE, Terry MJ, Smith AG (2002) Green or red: what stops the traffic in the tetrapyrrole pathway? Trends Plant Sci 8:224–230
Cotsaftis O, Plett D, Johnson AA, Walia H, Wilson C, Ismail AM, Close TJ, Tester M, Baumann U (2011) Root-specific transcript profiling of contrasting rice genotypes in response to salinity stress. Mol Plant 4:25–41
Cotsaftis O, Plett D, Shirley N, Tester M, Hrmova M (2012) A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporters and alternative splicing. PLoS ONE 7:e39865
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:1–14
Currie HA, Perry CC (2007) Silica in plants: biological, biochemical and chemical studies. Ann Bot 100:1383–1389
Dadkhah A (2011) Effect of salinity on growth and leaf photosynthesis of two sugar beet (Beta vulgaris L.) cultivars. J Agricul Sci Technol 13:1001–1012
Dalal VK, Tripathy BC (2012) Modulation of chlorophyll biosynthesis by water stress in rice seedlings during chloroplast biogenesis. Plant Cell Environ 35:1685–1703
Das P, Nutan KK, Singla-Pareek SL (2015) Pareek A (2012) Understanding salinity responses and adopting ‘omics-based’approaches to generate salinity tolerant cultivars of rice. Front Plant Sci 6:712
Das P, Mishra M, Lakra N, Singla-Pareek S, Pareek A (2014) Mutation breeding: a powerful approach for obtaining abiotic stress tolerant crops and upgrading food security for human nutrition. In: Mutagenesis: exploring novel genes and pathways: Wageningen Academic Publishers 615–621.
DeGomez, T (2019). Basic Soil Components. In Climate, forests and woodlands. university of arizona, peter kolb, montana state university, and sabrina kleinman, university of arizona: Climate, Forests and Woodlands Community of Practice.
DeLuca T, DeLuca D (1997) Composting for feedlot manure management and soil quality. J Prod Agri 10:235–241
De-Smet I, White PJ, Bengough AG, Dupuy L, Parizot B, Casimiro I, Heidstra R, Laskowski M, Lepetit M, Bennett M (2012) Analyzing lateral root development: how to move forward. Plant Cell 24:15–20
Dhamodharan K, Ravi KS, Balachandar D (2015) Changes in nutrient status and enzymatic activities in soil amended with composted urban green waste. Int J Recycl 4:101–108
Dhanorkar B, Borkar D, Puranik R, Joshi R (1994) Forms of soil potassium as influenced by long-term application of FYM and NPK in Vertisol. J Potassium R 10:42–48
Dhar R, Sägesser R, Weikert C, Yuan J, Wagner A (2011) Adaptation of Saccharomyces cerevisiae to saline stress through laboratory evolution. J Evol Boil 24:1135–1153
Diédhiou CJ, Popova OV, Dietz K-J, Golldack D (2008) The SNF1-type serine-threonine protein kinase SAPK4 regulates stress-responsive gene expression in rice. BMC Plant Boil 8:1–13
Ernst WH, Peter D, Herman JP (2004) Chapter 3 vegetation, organic matter and soil quality developments in soil science. Develop Soil Sci 29:41–98
García Morales S, Trejo-Téllez LI, Gómez Merino FC, Caldana C, Espinosa-Victoria D, Herrera Cabrera BE (2012) Growth, photosynthetic activity, and potassium and sodium concentration in rice plants under salt stress. Acta Sci Agron 34:317–324
Grattan SR, Zeng L, Shannon MC, Roberts SR (2002) Rice is more sensitive to salinity than previously thought. California Agri 56:189–198
Gull A, Lone AA, Wani NUI (2019) Biotic and abiotic stresses in plants. Abiotic Biot Stress Plants 1–19.
Gupta BH, B, (2014) Mechanism of salinity tolerance in plants: physiological, biochemical, and molecular characterization. Int J Genom. https://doi.org/10.1155/2014/701596
Hairmansis A, Berger B, Tester M, Roy SJ (2014) Image-based phenotyping for non-destructive screening of different salinity tolerance traits in rice. Rice 7:1–10
Hajlaoui H, El Ayeb N, Garrec JP, Denden M (2010) Differential effects of salt stress on osmotic adjustment and solutes allocation on the basis of root and leaf tissue senescence of two silage maize (Zea mays L.) varieties. Ind Crops Prod 31:122–130
Hameed A, Ahmed MZ, Hussain T, Aziz I, Ahmad N, Gul B, Nielsen BL (2021) Effects of salinity stress on chloroplast structure and function. Cells 7:2023
Hasanuzzaman M, Nahar K, Fujita M (2013 a) Plant response to salt stress and role of exogenous protectants to mitigate salt-induced damages. In: Ecophysiology and responses of plants under salt stress: Springer, New York, NY 25–87. https://doi.org/10.1007/978-1-4614-4747-4_2
Hasanuzzaman M, Nahar K, Fujita M, Ahmad P, Chandna R, Prasad M (2013 b) Enhancing plant productivity under salt stress: relevance of poly-omics. Salt stress in plants. Springer, New York, NY 113–156. https://doi.org/10.1007/978-1-4614-6108-1_6
Hopmans JW, Qureshi A, Kisekka R, Munns S, Grattan P, Rengasamy A, Ben-Gal S, Assouline M, Javaux M, Minhas P (2021) Critical knowledge gaps and research priorities in global soil salinity. Adv Agron 169:1–191
Horie T, Karahara I, Katsuhara M (2012) Salinity tolerance mechanisms in glycophytes: an overview with the central focus on rice plants. Rice 5:1–18
Huang M, Shan S, Cao J, Fang S, Tian A, Liu Y, Zou Y (2020) Primary-tiller panicle number is critical to achieving high grain yields in machine-transplanted hybrid rice. Sci Rep 10:2811
Humplík JF, Lazár D, Husičková A, Spíchal L (2015) Automated phenotyping of plant shoots using imaging methods for analysis of plant stress responses–a review. Plant Methods 11:1–10
Hussain S, Zhang R, Liu S, Li R, Zhou Y, Chen Y, Hou H, Dai Q (2022b) Transcriptome-wide analysis revealed the potential of the high-affinity potassium transporter (HKT) gene family in rice salinity tolerance via ion homeostasis. Bioengineering 9:410
Hussain S, Zhang JH, Zhong C, Zhu LF, Cao XC, Yu SM, Bohr JA, Hu JJ, Jin QY (2017) Effects of salt stress on rice growth, development characteristics, and the regulating ways: a review. J Integ Agri 16:2357–2374
Hussain S, Zhang R, Liu S, Wang Y, Ahmad I, Chen Y, Hou H, Dai Q (2022a) Salt stress affects the growth and yield of wheat (Triticum aestivum L.) by altering the antioxidant machinery and expression of hormones and stress-specific genes. Phyton 92:3
Isayenkov SV (2019) Genetic sources for the development of salt tolerance in crops. Plant Growth Regul 89:1–17
Ishak NK, Sulaiman Z, Tennakoon KU (2015) Comparative study on growth performance of transgenic (Over-expressed OsNHX1) and wild-type nipponbare under different salinity regimes. Rice Sci 22:275–282
Jouyban Z (2012) The effects of salt stress on plant growth. Tech J Eng Appl Sci 2:7–10
Kordrostami M, Rabiei B, Hassani Kumleh H (2017) Biochemical, physiological and molecular evaluation of rice cultivars differing in salt tolerance at the seedling stage. Physiol Mol Biol Plants 23:529–544
Kumar K, Kumar M, Kim SR, Ryu H, Cho YG (2013) Insights into genomics of salt stress response in rice. Rice 6:1–15
Kumari S, Panjabi nee Sabharwal V, Kushwaha HR, Sopory SK, Singla-Pareek SL, Pareek A (2009) Transcriptome map for seedling stage specific salinity stress response indicates a specific set of genes as candidate for saline tolerance in Oryza sativa L. Funct Integr Genom 9:109–123
Kurokawa Y, Noda T, Yamagata Y, Angeles-Shim R, Sunohara H, Uehara K, Furuta A, Negai K, Jena KK, Yasui H, Yoshimura A, Ashikari M, Doi K (2016) Construction of a versatile SNP array for pyramiding useful genes of rice. Plant Sci 242:131–139
Kurotani K, Yamanaka K, Toda Y, Ogawa D, Tanaka M, Kozawa H, Nakamura H, Hakata M, Ichikawa H, Hattori T, Takeda S (2015) Stress tolerance profiling of a collection of extant salt-tolerant rice varieties and transgenic plants overexpressing abiotic stress tolerance genes. Plant Cell Physiol 56:1867–1876
Lakhdar A, Rabhi M, Ghnaya T, Montemurro F, Jedidi N, Abdelly C (2009) Effectiveness of compost use in salt-affected soil. J Hazard Mater 171:29–37
Ben-Ari G (2012) Plant biotechnology and agriculture, marker-assisted selection in plant breeding. Academic Press 163–184. https://doi.org/10.1016/B978-0-12-381466-1.00011-0
Lawlor DW, Tezara W (2009) Causes of decreased photosynthetic rate and metabolic capacity in water-deficient leaf cells: a critical evaluation of mechanisms and integration of processes. Ann Bot 103:561–579
Lee KS, Choi WY, Ko JC, Kim TS, Gregorio GB (2003) Salinity tolerance of japonica and indica rice (Oryza sativa L.) at the seedling stage. Planta 216:1043–1046
Lee DG, Ahsan N, Lee SH, Lee JJ, Bahk JD, Kang KY, Lee BY (2009) Chilling stress-induced proteomic changes in rice roots. J Plant Physiol 166:1–11
Li HW, Zang BS, Deng XW, Wang XP (2011) Overexpression of the trehalose-6-phosphate synthase gene OsTPS1 enhances abiotic stress tolerance in rice. Planta 234:1007–1018
Li M, Guo L, Guo C, Wang L, Chen L (2016) Over-expression of a DUF1644 protein gene, SIDP361, enhances tolerance to salt stress in transgenic rice. J Plant Biol 59:62–73
Liao YD, Lin KH, Chen CC, Chiang CM (2016) Oryza sativa protein phosphatase 1a (OsPP1a) involved in salt stress tolerance in transgenic rice. Mol Breeding 36:1–19
Litardo RCM, Bendezú SJG, Zenteno MDC, Pérez-Almeida IB, Parismoreno LL, García EDL (2022) Effect of mineral and organic amendments on rice growth and yield in saline soils. J Saudi Soci Agri Sci 21:29–37
Liu Y, Du H, Wang K, Huang B, Wang Z (2011) Differential photosynthetic responses to salinity stress between two perennial grass species contrasting in salinity tolerance. Hort Sci 46:311–316
Liu X, Li L, Li M, Su L, Lian S, Zhang B, Li X, Ge K, Li L (2018) AhGLK1 affects chlorophyll biosynthesis and photosynthesis in peanut leaves during recovery from drought. Sci Rep 8:1–11
Liu C, Mao B, Yuan D, Chu C, Duan M (2022a) Salt tolerance in rice: physiological responses and molecular mechanisms. Crop J 1:13–25
Liu C, Mao D, Yuan C, Chu M, Duan D (2022b) Salt tolerance in rice: physiological responses and molecular mechanisms. Crop J 10:13–25
Lutts S, Kinet J, Bouharmont J (1996) NaCl-induced senescence in leaves of rice (Oryza sativa L.) cultivars differing in salinity resistance. Ann Bot 78:389–398
Mahlooji M, Seyed Sharifi R, Razmjoo J, Sabzalian M, Sedghi M (2018) Effect of salt stress on photosynthesis and physiological parameters of three contrasting barley genotypes. Photosynthetica 56:549–556
Mishra S, Singh B, Panda K, Singh BP, Singh N, Misra P, Rai V, Singh NK (2016) Association of SNP haplotypes of HKT family genes with salt tolerance in indian wild rice germplasm. Rice 9:1–13
Miyamoto T, Ochiai K, Nonoue Y, Matsubara K, Yano M, Matoh T (2015) Expression level of the sodium transporter gene OsHKT2; 1 determines sodium accumulation of rice cultivars under potassium-deficient conditions. J Soil Sci Plant Nutr 61:481–492
Mohanavelu A, Naganna SR, Al-Ansari N (2021) Irrigation induced salinity and sodicity hazards on soil and groundwater: an overview of its causes. Impacts Mitig Strat Agri 11:983
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250
Munns R, James RA, Läuchli A (2006) Approaches to increasing the salt tolerance of wheat and other cereals. J Exper B 57:1025–1043
Mustafa MA (2007) Desertification processes. UNESCO chair of desertification/university of khartoum, Khartoum, Sudan
Nagahatenna DS, Langridge P, Whitford R (2015) Tetrapyrrole-based drought stress signalling. Plant Biotechnol J 13:447–459
Nam MH, Bang E, Kwon TY, Kim Y, Kim EH, Cho K, Park WJ, Kim BG, Yoon IS (2015) Metabolite profiling of diverse rice germplasm and identification of conserved metabolic markers of rice roots in response to long-term mild salinity stress. Int J Mol Sci 16:21959–21974
Nath M, Yadav S, Sahoo RK, Passricha N, Tuteja R, Tuteja N (2016) PDH45 transgenic rice maintain cell viability through lower accumulation of Na+, ROS and calcium homeostasis in roots under salinity stress. J Plant Physiol 191:1–11
Negrão S, Schmöckel S, Tester M (2017) Evaluating physiological responses of plants to salinity stress. Ann Bot 119:1–11
Nishimura T, Cha-Um S, Takagaki M, Ohyama K, Kirdmanee C (2011) Survival percentage, photosynthetic abilities and growth characters of two indica rice (Oryza sativa L. spp. indica) cultivars in response to iso-osmotic stress. Spanish J Agri R 9:262–270
Nounjan N, Chansongkrow P, Charoensawan V, Siangliw JL, Toojinda T, Chadchawan S, Theerakulpisut P (2018) High performance of photosynthesis and osmotic adjustment are associated with salt tolerance ability in rice carrying drought tolerance QTL: physiological and co-expression network analysis. Front Plant Sci 9:1135
Nounjan N, Mahakham W, Siangliw JL, Toojinda T, Theerakulpisut P (2020) Chlorophyll retention and high photosynthetic performance contribute to salinity tolerance in rice carrying drought tolerance quantitative trait loci (QTLs). Agriculture 10:620
Pandit A, Rai V, Bal S, Sinha S, Kumar V, Chauhan M (2010) Combining QTL mapping and transcriptome profiling of bulked RILs for identification of functional polymorphism for salt tolerance genes in rice (Oryza sativa L.). Mol Genet Genom 284:121–136
Parida AK, Das AB (2005) Salt tolerance and salinity effects on plants: a review. Ecot Environ Safety 6:324–349
Parihar P, Singh S, Singh R, Singh VP, Prasad SM (2015) Effect of salinity stress on plants and its tolerance strategies: a review. Environ Sci Pollut Res Int 22:4056–4075
Pla-Sentís I (2021) Prediction of soil salinization and sodification processes as affected by groundwater under different climate and management conditions. In: intensified land and water use: Springer; 25–42.
Platten JD, Egdane JA, Ismail AM (2013) Salinity tolerance, Na+ exclusion and allele mining of HKT1; 5 in Oryza sativa and O. glaberrima: many sources, many genes, one mechanism? BMC Plant Biol 13:1–16
Ponce KS, Guo L, Leng Y, Meng L, Ye G 2021 Advances in sensing, response and regulation mechanism of salt tolerance in rice. Int J Mol Sci 22:2254
Qadir M, Quillérou E, Nangia V, Murtaza G, Singh M, Thomas RJ, Noble AD (2014) Economics of salt‐induced land degradation and restoration. In: Natural resources forum: Wiley Online Library 282–295
Rahman MA, Thomson MJ, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail AM (2016) Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization. Ann Bot 117:1083–1097
Rahman A, Nahar K, Al Mahmud J, Hasanuzzaman M, Hossain MS, Fujita M (2017) Salt stress tolerance in rice: emerging role of exogenous phytoprotectants. Adv Int Rice Res 9:139–174
Ramezani M, Seghatoleslami M, Mousavi G, Sayyari-Zahan M (2012) Effect of salinity and foliar application of iron and zinc on yield and water use efficiency of ajowan (Carum copticum). Int J Agri Crop Sci 4:421–426
Rath KM, Rousk J (2015) Salt effects on the soil microbial decomposer community and their role in organic carbon cycling: a review. Soil Biol Bioch 81:108–123
Raza A, Razzaq A, Mehmood SS, Zou X, Zhang X, Lv Y, Xu J (2019) Impact of climate change on crops adaptation and strategies to tackle its outcome: a review. Plants 8:34
Razzaque MA, Talukder NM, Islam MT (2011) Dutta RK (2011) Salinity effect on mineral nutrient distribution along roots and shoots of rice (Oryza sativa L.) genotypes differing in salt tolerance. Arch Agron Soil Sci 57:33–45
Reddy INBL, Kim BK, Yoon IS, Kim KH, Kwon TR (2017) Salt tolerance in rice: focus on mechanisms and approaches. Rice Sci 24:123–144
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146
Riaz M, Arif MS, Ashraf MA, Mahmood R, Yasmeen T, Shakoor MB, Shahzad SM, Ali M, Saleem I, Arif M (2019) A comprehensive review on rice responses and tolerance to salt stress. Advances in rice research for abiotic stress tolerance, 133–158.
Sardinha M, Müller T, Schmeisky H, Joergensen RG (2003) Microbial performance in soils along a salinity gradient under acidic conditions. Appl Soil Ecol 23:237–244
Sexcion FS, Egdane JA, Ismail AM, Dionisio-Sese ML (2006) Morpho-physiological traits associated with tolerance of salinity during seedling stage in rice (Oryza sativa L.). Phili J Crop Sci 34:27–37
Shereen A, Mumtaz S, Raza S, Khan M, Solangi S (2005) Salinity effects on seedling growth and yield components of different inbred rice lines. Pak J Bot 37:131–139
Shu S, Guo SR, Sun J, Yuan LY (2012) Effects of salt stress on the structure and function of the photosynthetic apparatus in cucumis sativus and its protection by exogenous putrescine. Physiol Plantarum 146:285–296
Singh R, Singh Y, Xalaxo S, Verulkar S, Yadav N, Singh S, Singh AK (2016) From QTL to variety-harnessing the benefits of QTLs for drought, flood and salt tolerance in mega rice varieties of india through a multi-institutional network. Plant Sci 242:278–287
Soussi AR, Ferjani R, Marasco A, Guesmi H, Cherif E, Rolli F, Mapelli H, Ouzari I, Hadda D, Daffonchio A, Cherif A (2016) Plant-associated microbiomes in arid lands: diversity, ecology and biotechnological potential. Plant Soil 405:357–370
Suriyan CU, Supaibulwattana K, Kirdmanee C (2009) Comparative effects of salt stress and extreme pH stress combined on glycinebetaine accumulation, photosynthetic abilities and growth characters of two rice genotypes. Rice Sci 16:274–282
Tanaka R, Tanaka A (2007) Tetrapyrrole biosynthesis in higher plants. Annu Rev Plant Biol 58:321–346
Tatar Ö, Brueck H, Gevrek MN, Asch F (2010) Physiological responses of two turkish rice (Oryza sativa L.) varieties to salinity. Turk J Agric for 34:451–459
Thompson GA, Goggin FL (2006) Transcriptomics and functional genomics of plant defence induction by phloem-feeding insects. J Exper Bot 57:755–766
Todaka D, Nakashima K, Shinozaki K, Yamaguchi-Shinozaki K (2012) Toward understanding transcriptional regulatory networks in abiotic stress responses and tolerance in rice. Rice 5:1–9
Tu Y, Jiang A, Gan L, Hossain M, Zhang J, Peng B, Xiong Y, Song Z, Cai D, Xu W, Zhang J, He J (2014a) Genome duplication improves rice root resistance to salt stress. Rice 7:1–13
Tu YA, Jiang L, Gan M, Hossain J, Zhang B, Peng Y, Xiong Z, Song D, Cai W, Xu Zhang J (2014b) Genome duplication improves rice root resistance to salt stress. Rice 7:1–13
Turner NC (2018) Turgor maintenance by osmotic adjustment: 40 years of progress. J Exp Bot 69:3223–3233
Wang L, Sun X, Li S, Zhang T, Zhang W, Zhai P (2014) Application of organic amendments to a coastal saline soil in north china: effects on soil physical and chemical properties and tree growth. PLoS ONE 9:e89185
Wang Z, Xu Y, Chen T, Zhang H, Yang J, Zhang J (2015) Abscisic acid and the key enzymes and genes in sucrose-to-starch conversion in rice spikelets in response to soil drying during grain filling. Planta 241:1091–1107
Wang W-S, Zhao X-Q, Li M, Huang L-Y, Xu J-L, Zhang F, Cui YR, Fu BY, Li ZK (2016) Complex molecular mechanisms underlying seedling salt tolerance in rice revealed by comparative transcriptome and metabolomic profiling. J Experi Bot 67:405–419
Wang X, Wang W, Huang J, Peng S, Xiong D (2018) Diffusional conductance to CO2 is the key limitation to photosynthesis in salt-stressed leaves of rice (Oryza sativa L.). Physiol Plantarum 163:45–58
Wimmer M, Muehling K, Läuchli A, Brown P, Goldbach H (2001) Interaction of salinity and boron toxicity in wheat (Triticum aestivum L) In: Plant nutrition: Springer Dordrecht 92:426–427
Xiang J, Chen X, Hu W, Xiang Y, Yan M, Wang J (2018) Overexpressing heat-shock protein OsHSP50. 2 improves drought tolerance in rice. Plant Cell Rep 37:1585–1595
Xiang Y, Huang Y, Xiong L (2007) Characterization of stress-responsive CIPK genes in rice for stress tolerance improvement. Plant Physiol 144:1416–1428
Xiao F, Zhou H (2023) Plant salt response: perception, signaling, and tolerance. Front Plant Sci 13:1053699
Xiong L, Zhu JK (2002) Molecular and genetic aspects of plant responses to osmotic stress. Plant Cell Environ 25:131–139
Xu Y, Zhang L, Ou S, Wang R, Wang Y, Chu C, Yao S (2020) Natural variations of SLG1 confer high-temperature tolerance in indica rice. Nat Commun 11:5441
Yan G, Fan X, Peng M, Yin C, Xiao Z, Liang Y (2020) Silicon improves rice salinity resistance by alleviating ionic toxicity and osmotic constraint in an organ-specific pattern. Front Plant Sci 11:260
Yang T, Zhang S, Hu Y, Wu F, Hu Q, Chen G, Cai J, Wu T, Moran N, Yu L, Xu G (2014) The role of a potassium transporter OsHAK5 in potassium acquisition and transport from roots to shoots in rice at low potassium supply levels. Plant Physiol 166:945–959
Yasmin F, Biswas S, Jewel GNA, Elias SM, Seraj ZI (2015) Constitutive overexpression of the plasma membrane Na+/H+ antiporter for conferring salinity tolerance in rice. Plant Tissue Cult Biotechnol 25:257–272
Yeo A, Flowers T (1983) Varietal differences in the toxicity of sodium ions in rice leaves. Physiol Plant 59:189–195
Yi X-P, Zhang Y-L, Yao H-S, Luo H-H, Gou L, Chow WS, Zhang WF (2016) Rapid recovery of photosynthetic rate following soil water deficit and re-watering in cotton plants (Gossypium herbaceum L.) is related to the stability of the photosystems. J Plant Physiol 194:23–34
Zayed B, Elkhoby W, Salem A, Ceesay M, Uphoff N (2013) Effect of integrated nitrogen fertilizer on rice productivity and soil fertility under saline soil conditions. J Plant Bio Res 2:14–24
Zeng L, Shannon MC (2000) Effects of salinity on grain yield and yield components of rice at different seeding densities. Agron J 92:418–423
Zivcak M, Brestic M, Sytar O (2016) Osmotic adjustment and plant adaptation to drought stress. In Drought stress. Physiol Biochem 1: 105–143
Zhang L, Tian LH, Zhao JF, Song Y, Zhang CJ, Guo Y (2009) Identification of an apoplastic protein involved in the initial phase of salt stress response in rice root by two-dimensional electrophoresis. Plant Physiol 149:916–928
Zhou J, Wang X, Jiao Y, Qin Y, Liu X, He K, Chen C, Ma L, Wang J, Xiong L, Zhang Q, Fan L, Deng XW (2007) Global genome expression analysis of rice in response to drought and high-salinity stresses in shoot, flag leaf, and panicle. Plant Mol Biol 63:591–608
Zhou Y, Yang P, Cui F, Zhang F, Luo X, Xie J (2016) Transcriptome analysis of salt stress responsiveness in the seedlings of Dongxiang wild rice (Oryza rufipogon Griff.). PLoS ONE 11:e0146242
Acknowledgements
The authors gratified the National Natural Science Foundation of China (32101817), Jiangsu Agriculture Science and Technology Innovation Fund (JASTIF, Grant No. CX(21)3111), and Natural Science Foundation of the Jiangsu Higher Education Institutions (21KJD210001) for their financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Ahmad Mohammad Alqudah.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hussain, S., Zhang, R., Chen, Y. et al. An overview on salt-induced physiological changes, molecular mechanism of salinity tolerance and application strategies for its management in rice. CEREAL RESEARCH COMMUNICATIONS (2024). https://doi.org/10.1007/s42976-023-00487-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42976-023-00487-y