Abstract
High affinity ligands for a given target tend to share key molecular interactions with important anchoring amino acids and therefore often present quite conserved interaction patterns. This simple concept was formalized in a topological knowledge-based scoring function (GRIM) for selecting the most appropriate docking poses from previously X-rayed interaction patterns. GRIM first converts protein–ligand atomic coordinates (docking poses) into a simple 3D graph describing the corresponding interaction pattern. In a second step, proposed graphs are compared to that found from template structures in the Protein Data Bank. Last, all docking poses are rescored according to an empirical score (GRIMscore) accounting for overlap of maximum common subgraphs. Taking the opportunity of the public D3R Grand Challenge 2015, GRIM was used to rescore docking poses for 36 ligands (6 HSP90α inhibitors, 30 MAP4K4 inhibitors) prior to the release of the corresponding protein–ligand X-ray structures. When applied to the HSP90α dataset, for which many protein–ligand X-ray structures are already available, GRIM provided very high quality solutions (mean rmsd = 1.06 Å, n = 6) as top-ranked poses, and significantly outperformed a state-of-the-art scoring function. In the case of MAP4K4 inhibitors, for which preexisting 3D knowledge is scarce and chemical diversity is much larger, the accuracy of GRIM poses decays (mean rmsd = 3.18 Å, n = 30) although GRIM still outperforms an energy-based scoring function. GRIM rescoring appears to be quite robust with comparison to the other approaches competing for the same challenge (42 submissions for the HSP90 dataset, 27 for the MAP4K4 dataset) as it ranked 3rd and 2nd respectively, for the two investigated datasets. The rescoring method is quite simple to implement, independent on a docking engine, and applicable to any target for which at least one holo X-ray structure is available.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In absence of structural data on protein ligand complexes (X-ray diffraction, nuclear magnetic resonance spectroscopy, electron microscopy), molecular docking remains the computational method of choice to predict ligand binding modes [1]. Since the pioneering work of Kuntz et al. [2], over 100 different docking software have been reported [1, 3–6] that progressively addressed most of the issues related to this computational exercise: full ligand flexibility, accurate configurational sampling of the ligand in the protein binding site, partial protein flexibility, implicit or explicit solvation, prediction of relative or absolute binding (free) energies. Many benchmarking studies [7–11] comparing different algorithms across diverse datasets of protein–ligand X-ray structures, agree on the point that state-of-the-art docking algorithms are very efficient in predicting ligand poses: a relative position of a ligand with respect to a protein and a conformation of a protein-bound ligand. Unfortunately, these good solutions are hardly distinguishable from a much larger set of incorrect proposals (decoys) using any predicted energy criterion. Considering success in pose prediction as the ability to predict poses with root-mean square deviation (rmsd) from X-ray solution below 2 Å, most docking tools present in the best cases (self-docking) a success rate close to 70 % when considering the top-1 (best scored) solution [12]. Considering all possible solutions, this rate raises typically up to 85–90 % [12] thereby demonstrating that the best scored pose is not always the most reliable one. If handling multiple docking solutions is feasible albeit cumbersome for one particular ligand, this approach cannot be followed upon in silico screening a large compound library. Reasons accounting for repetitive failures in predicting either binding free energies or relative potency ranks [9, 12–14] are numerous: target flexibility, incorrect protonation/tautomeric states, incorrect treatment of many energy terms (solvation, entropy, metal chelation and weak non-covalent interactions).
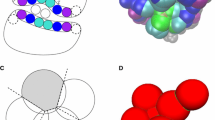
Three main approaches have been followed to rescue the inability of fast scoring functions to prioritize the best docking poses: (1) develop more sophisticated first-principle scoring functions, (2) use supervised machine learning (ML) algorithms to predict the likelihood of docking poses, (3) apply knowledge-based (chemical and topological) rules to filter out unreliable solutions. The first approach uses CPU-intensive energy calculations (e.g. MM-PBSA, MM-GBSA) to refine early docking results. Unfortunately, the benefit of this extra computational cost is controversial as it appears to be target-dependent and hardly predictable [15–17]. The second approach consists in training machine learning algorithms (e.g. support vector machines [18], random forest [19, 20]) with 3D protein–ligand structural descriptors in order to discriminate good from bad poses. If remarkable results in predicting binding affinities from protein–ligand X-ray structures have been recently published [20], such scoring functions have rarely been applied to prospective virtual screening campaigns and their true utility in virtual screening remains unknown. In any case, docking/ML combinations [21] must be regarded with great care due to the tendency of machine learning methods to be overtrained [22]. The third strategy, which is currently experiencing a revival, utilizes various knowledge-based approaches to rescore docking poses. The main idea is to use non-energetical topological criteria to address the quality of docking poses, notably by comparing docking solutions with protein–ligand complexes of known X-ray structure. Among knowledge-based approaches, we can clearly distinguish those methods aimed at constraining the docking algorithms towards expected poses (pharmacophore-constrained docking [23], shape-guided docking [24, 25], template matching [26]) from computational protocols that just restrain the analysis of docking poses to reward user-defined features. Both methods have proven useful in many examples for enhancing the quality of top-ranked poses as well as enriching virtual hit lists in true actives. Constrained docking may however be dangerous in forcing known inactive compounds to properly dock in a binding site. It is therefore common practice to conduct a totally free docking calculation and further apply simple cheminformatics descriptors (1D fingerprints [27], 3D similarity [28]) to enable the selection of docking solutions that look the most similar to experimentally-determined poses of known ligands. For example, we [29] and others [27, 30, 31] proposed several years ago, the concept of molecular interaction fingerprints (IFPs) [29] to post-process docking data and pick-up poses producing IFPs similar to that of known actives. Computing IFPs from docking poses is a robust and very efficient manner to predict ligand binding modes [32], propose reliable scaffold hops [33], and enrich virtual hits in true actives upon docking a compound library [34, 35]. The success of this post-processing approach relies on the fact that true ligands of a same target often share key interactions with key anchoring residues and thereby produce quite similar IFPs. A drawback of this method lies in the definition of a consensus binding site (fixed set of target residues) in order to generate fixed-sized and comparable interaction fingerprints. To overcome this limitation and extend the concept of interaction fingerprints to binding site-independent and coordinates frame-invariant fingerprints, we recently proposed to encode interaction patterns (sets of protein–ligand interactions) into either generic 1D fingerprints or 3D graphs [36]. Our GRIM algorithm for matching interaction pattern graphs has been described in details elsewhere [36], and here it will be just briefly summarized. Starting from 3D coordinates of a protein–ligand complex, molecular interactions (hydrophobic contacts, aromatic interactions, hydrogen bonds, ionic bonds, metal chelation) are first computed from a set of topological and chemical rules. Every detected interaction is then labelled by three interaction pseudoatoms (IPAs) located on (1) the ligand-interacting atom, (2) the protein-interacting atom and (3) the geometric barycenter of protein and ligand-interacting atoms (Fig. 1a, b). Starting from two sets of IPAs (reference, target), GRIM first creates a list of possible IPA matches. A pair is made if reference and target IPAs have the same label (same interaction type) and represent the same point of view (ligand, protein, barycenter). A product graph is created from the two reference and target graphs in which each successfully matched pair is consequently a vertex. A weight is added to each pair which is inversely proportional to the observed frequency among the 284,186 IPAs generated from the 9877 protein–ligand complexes of the sc-PDB dataset [37]. Assigned weights were as follows: hydrophobic IPA (0.299), aromatic IPA (0.990), h-bond acceptor (0.930), h-bond donor (0.834), negative ionizable (0.993), positive ionizable (0.966), metal complexation (0.985). An edge is observed between two vertices of the product graph after computing distances between the two reference IPAs and the two target IPAs. If the difference is below a given threshold [37], an edge is created. The largest cliques are then detected using the Bron–Kerbosch algorithm [38] with pivoting and pruning improvements [39]. Each IPA of the target is matched with the corresponding reference IPA using a quaternion-based characteristic polynomial [40]. It returns both the translation vector and the rotation matrices to match target and reference graphs as well as a Graph-alignment score (GRIMscore). As the graph is specific of a given protein–ligand interaction pattern, two sets of protein–ligand coordinates can therefore be easily compared by aligning the corresponding graphs (Fig. 1c). According to a previous benchmark, a GRIMscore value above 0.70 is indicative of a statically significant similarity of interaction patterns.
Principle of the graph alignment method for matching interaction patterns (GRIM). a The estrogen receptor β-WAY697 complex (PDB ID 1 × 76) is converted into a set of interaction pseudoatoms (IPAs) describing intermolecular interactions. For each interaction (e.g. hydrogen-bond between Arg346 of the protein and a phenolic oxygen atom of the ligand, displayed by a dashed black line), IPAs are placed on the ligand-interacting atom (l), the protein-interacting atom (p) and the geometric barycenter of both interacting atoms (c). IPAs are color-coded according the described interaction (gray, apolar and aromatic interactions; red, hydrogen bond (protein acceptor) and ionic bond (protein negatively charged); blue, hydrogen bond (protein donor) and ionic bond (protein positively charged). b Same procedure as above for a second complex between the same receptor and WAY-338. IPAs describing the same interaction with Arg346 are labelled L, C and P, respectively. IPAs are color-coded according the described interaction (light gray, apolar and aromatic interactions; light red, hydrogen bond (protein acceptor) and ionic bond (protein negatively charged), light blue, hydrogen bond (protein donor) and ionic bond (protein positively charged). c Graph-based alignment of the two sets of IPAs leading to an interaction-guided overlay of the two bound ligands. Please note that GRIM does not allow matching IPAs from different origin (e.g. l-type with p-types for example)
When applied to rescore docking poses generated by Surflex-Dock [41], GRIM rescoring significantly outperformed the Surflex-Dock scoring function in a retrospective virtual screening exercise aiming at recovering true actives from DUD-E decoys [42] for 10 targets of pharmaceutical interest [36]. We herewith present a prospective application of the GRIM graph matching method to the problem of docking pose selection by predicting, prior to the release of the corresponding X-ray coordinates (D3R Grand Challenge 2015) [43], the binding modes of 36 inhibitors bound to two different targets of pharmaceutical interest (HSP90α, MAP4K4). This manuscript will only highlight results obtained for Stage 1 (docking pose accuracy) of the D3R challenge.
Computational methods
HSP90α dataset
Four input protein structures (PDB ID: 2JJC, 2XDX, 4YKY, 4YKR) and 180 known HSP90α inhibitors (SMILES strings, Supplementary Table 1) were directly downloaded from the D3R Grand Challenge 2015 website [43] as a zipped archive file (279_data_303589.tar.gz).
HSP90α protein structures were prepared for docking as follows. First, existing hydrogen atoms were removed from the 4 input HSP90α structures and added back while optimizing both the protonation and tautomeric states using Protoss v.2.0 [44]. Two conserved waters mediating the interactions between protein and ligands (HOH2078 and HOH2166 for 2JJC; HOH2029 and HOH2054 for 2XDX; HOH6 and HOH233 for 4YKR; HOH5 and HOH198 for 4YKY) were kept, other water molecules were removed. Next, each protein structure was saved in 4 copies varying by the presence/absence of bound waters as follows: with both water molecules (wat2), with the first water molecule (wat1a), with the second water molecule (wat1b), without waters (dry). In total, 16 structures were therefore used as input for docking the 180 HSP90α inhibitors, which were provided by D3R Grand Challenge. Note, that binding mode should have been predicted only for 6 of those compounds.
Ligands from HSP90α crystal structures were checked manually (bond order, protonation and tautomeric states) and corrected whenever necessary. Protein and ligand structures were separately saved in MOL2 format using SYBYLX-2.1 [45]. In addition, 176 HSP90α-inhibitor complexes (Supplementary Table 2) were defined as templates for graph matching by searching the RCSB Protein Data Bank [46] for the P07900 UniProt [47] accession number and a known bound ligand. These 176 complexes were further processed as described above.
Starting from the provided input SMILES strings of 180 HSP90α inhibitors, hydrogen atoms were added and a 3D conformation was generated for every ligand using Corina v3.40 [48]. All Ligands were then saved in MOL2 format.
MAP4K4 dataset
Two input protein structures (PDB ID: 4OBO, 4U44) and 30 known MAP4K4 inhibitors (SMILES strings, Supplementary Table 3) were directly downloaded from the D3R Grand Challenge 2015 website [43] as a zipped archive file (280_data_473989.tar.gz). Furthermore, 6 additional MAP4K4-inhibitor complexes (Supplementary Table 4) were retrieved by searching the RCSB Protein Data Bank for the O95819 UniProt accession number and a known bound ligand. The 8 protein structures were prepared for docking using the protocol described for the HSP90α dataset. No bound water molecules were conserved in the present case.
Starting from the input SMILES strings of the 30 MAP4K4 inhibitors, hydrogen atoms were added and 3D conformations were generated using Corina v3.40 [48]. All Ligands were then saved in MOL2 format.
Docking
Ligands were docked to input protein structures using Surflex-Dock v.2745 [41]. For each protein input structure, a protomol was first generated using a list of binding site residues (including bound waters) for which at least one heavy atom was closer than 6.5 Å from at least one ligand heavy atom. The docking accuracy parameter set -pgeom was used. The -pgeom option starts each docking from 4 initial and different poses to ensure good search coverage, turns on ligand minimization prior to docking and after docking (in-pocket minimization), ensures that the returned poses are different from one another by at least 0.5 Å rmsd, and saves a total of 20 poses (ranked by Surflex-Dock energy score, from 000 to 019). In summary a total of 57,600 (180 × 16 × 20) and 4800 (30 × 8 × 20) poses were generated for the HSP90α, and MAP4K4 inhibitors respectively.
GRIM rescoring
Each Surflex-Dock pose was compared to the list of template complexes (176 for HSP90α inhibitors, 8 for MAP4K4 inhibitors). The interaction pattern of each docking pose was computed with IChem [36], aligned to that of the corresponding templates by graph-based alignment and ranked by GRIMscore. For every HSP90α ligand to dock, all poses were merged, regardless of the input protein structure and graph template used, and ranked by decreasing GRIMscore. For each MAP4K4 inhibitor, poses that do not exhibit at least one hydrogen-bond to the hinge region of the MAP4K4 kinase (residues E106, M107, C108) were discarded from further evaluation. Such poses were detected thanks to the protein–ligand interaction fingerprint generator [29] embedded in the IChem toolkit [36]. The 5 remaining poses with the highest GRIMscores (GRIM-1 to GRIM-5) were saved for every MAP4K4 ligand. For HSP90α inhibitors, a slightly different protocol was used to reflect the much higher number of templates and GRIM comparisons. To avoid retrieving too similar solutions, all poses were then clustered using an agglomerative method and a complete linkage clustering, starting from the highest GRIMscore (seed) and using a 2 Å rmsd threshold from the seed pose, until five different clusters were defined for each ligand. A representative pose (highest GRIMscore) for each of the 5 clusters was finally saved and ranked from 1 (GRIM-1) to 5 (GRIM-5) by decreasing GRIMscore.
Results and discussion
Predicting the binding mode of 6 HSP90α inhibitors
The first part of the challenge consisted in predicting the bound conformation of 180 HSP90α inhibitors from three chemical series (benzimidazolones, aminopyridines, benzophenones), given four reference input protein structures co-crystallized with at least one inhibitor of the above-cited three chemical series. A particular emphasis was put on six inhibitors (Fig. 2) whose protein-bound X-ray structures had to be released just at the closure of the first step (pose prediction accuracy) of the D3R Grand Challenge 2015. Since HSP90α inhibitors notoriously use conserved water molecules [49] to recognize the ATP-binding site, we decided to generate four sets of protein coordinates for each of the provided 4 input structures that just differ in the number of bound waters (none, one or two; see “Computational methods”). To use knowledge about inhibitor binding to the HSP90α target, we further retrieved 176 additional protein–ligand X-ray structures from the Protein Data Bank and ensured that all these inhibitors were occupying the same binding site that the 4 ligands co-crystallized with the input reference structures. Docking of all inhibitors to the 16 input structures was completely unrestrained (beside defining the common binding-site) and led to a total of 57,600 poses which were all compared to the 176 template structures using our GRIM interaction pattern matching method. To ascertain the generation of a few representative but diverse poses, we decided to cluster docking solutions using a 2 Å-rmsd threshold and provided up to 5 poses for each of 180 ligands (Table 1). Analyzing the rmsd of predicted poses to the true X-ray solution (released just after closure of the challenge) shows that our interaction pattern rescoring strategy achieves an outstanding accuracy since top-1 GRIM poses are predicted with a mean rmsd of 1.06 Å (Table 1). The top-1 GRIM pose of only two compounds (hsp90_44, hsp90_175) is predicted with a rmsd higher than 1 Å. The larger value of 2.47 Å (hsp90_44) is mainly due to pose differences occurring at the accessible pyridine-3-sulfonamide that does not strongly interact with the binding site; the position of the buried benzimidazolone core being nicely predicted with a rmsd of 0.53 Å (Fig. 3a). For compound hsp90_175, the main difference (rmsd = 1.67 Å) lies in the rotation of a single dihedral angle that drifts a phenol ring from its X-ray pose.
Predicted versus X-ray pose of two HSP90α inhibitors. Heteroatoms are colored in blue (nitrogen), red (oxygen), yellow (sulfur), and green (chlorine). The chemical structures of the two inhibitors are displayed below the binding poses. a Predicted binding mode of hsp90_44 (tan sticks) to HSP90α ATP-binding site (white surface). The pose has been selected by interaction pattern similarity to that of another HSP90α inhibitor co-crystallized with PDB entry 2ykr (plum sticks). The true X-ray pose is indicated by cyan sticks. b Predicted binding mode of hsp90_179 (green sticks) to HSP90α ATP-binding site (white surface). The pose has been selected by interaction pattern similarity to that of another HSP90α inhibitor co-crystallized with PDB entry 4fcq (plum sticks)
Since we intentionally clustered poses to avoid generating too many redundant answers, the quality of GRIM poses logically deteriorates when other solutions are considered (Table 1). Four 4 out of the 6 ligands, the GRIM-1 pose is by far the closest to the true X-ray structure which greatly facilitates the analysis of our rescoring. In all cases, the top solution selected by GRIM is better than that predicted by the native scoring function embedded in Surflex-Dock (mean rmsd = 1.91 Å; Table 1). As observed for almost all docking engines, the top-ranked pose is rarely the absolute best solution (the closest to the true X-ray pose). Among the set of 320 poses generated for each ligand, the lowest-rmsd pose is indeed very close (mean rmsd = 0.65 Å; Table 1) to the X-ray solution, thereby attesting the quality of Surflex-Dock as pose generator. For two out of the six ligands (hsp90_40, hsp90_164), the absolute best pose is ranked first by GRIM (Table 1). For two others (hsp90_73, hps90_179), the rmsd difference is so tiny (<0.3 Å) that these poses can be considered as almost equivalent.
We next looked for those cases where GRIM rescoring was able to rank at first position a near-native pose, and identified which protein input structures and which interaction pattern template had been used to select this particular pose (Table 2). In two cases (hsp90_40, hs90_175), the ligand to dock is a very close analog of the co-crystallized ligand from the input reference structure (2D similarity >0.90), it is therefore logical that the later protein structures and corresponding interaction patterns are used by GRIM to select the top pose. As a consequence, the interaction pattern of the predicted pose is very similar to that of the template (GRIMscore >0.85). However, the remaining ligands were posed by interaction pattern similarity to that of chemically different template ligands (2D similarity <0.60) thereby nicely illustrating the power of the knowledge-based rescoring method. A prototypical example is given by the 6-phenyl-1,3,5-triazine-2,4-diamine hsp90_179 (Table 2) whose correct pose (rmsd = 0.54 Å) has been deduced from that of a chemically unrelated 7H-pyrrolo[2,3-d]pyrimidine inhibitor (PDB entry 4fcq) that however exhibits a very similar binding mode (Fig. 3b) and interaction pattern (GRIMscore = 0.79).
Predicting the binding mode of 30 MAP4K4 inhibitors
The second challenge aims at predicting the bound conformation of MAP4K4 inhibitors, and is much more demanding than the previous one for many reasons: (1) the dataset to dock is larger (30 inhibitors; Fig. 4 and Supplementary Table 3) and much more chemically diverse (17 different chemotypes, 11 low molecular-weight fragments), (2) the number of templates (known MAP4K4-inhibitor X-ray structures) is lower with only 8 PDB complexes and three chemotypes (amino-quinazolines, amino-pyrrolotriazines, hydroxydihydropyridinone; Supplementary Table 4).
Although Surflex-Dock is able to propose at least one very reliable docking pose for 28 out of the 30 ligands (mean rmsd of the best possible pose = 0.94 Å; Table 3). The native Surflex-Dock scoring function and GRIMscore cannot detect near-native poses (<2 Å rmsd) as top-1 solution for 17 and 19 inhibitors, respectively. Rescoring by interaction pattern graph similarity (GRIM) provides overall better poses (mean rmsd of GRIM-1 pose = 3.18 Å) than Surflex-Dock (mean rmsd of Surflex-1 pose = 3.63 Å) but their accuracy remains lower than that observed for the previous HSP90α dataset. Despite their medium accuracy, it remains reassuring that the quality of the poses decreases with the GRIM rank (Table 3).
We next looked for the reasons explaining why it is so challenging to find near-native poses for MAP4K4 inhibitors. The first reason is that the MAP4K4 set of inhibitors contains a significant amount (11 out of 30 compounds) of low molecular weight fragments (MW < 250 and heavy atoms count <20). Out of these 11 fragments, only three of them (27 %) are well posed by GRIM (Fig. 5). Conversely, the success in predicting near-native poses for higher molecular weight ligands (heavy atom count ≥20) is significantly higher (8 out of 19, ratio = 42 %; Fig. 5).
Upon examining the GRIM docking poses of all 30 MAP4K4 inhibitors, we could identify three possible scenarios. The first one relates to 9 lead-like compounds (MAP01, MAP07, MAP08, MAP09, MAP11, MAP14, MAP18, MAP19, and MAP23) that were successfully docked (rmsd < 2.5 Å) and scored by GRIM (GRIMscore > 0.68) because the corresponding interaction patterns are quite similar to that from one of the six templates exhibiting either the same or a bioisosteric scaffold (e.g. MAP18 binding pose; Fig. 6). Importantly, polar interactions are those that contribute the most to the GRIMscore, thereby ensuring that both key hydrogen bonds to the kinase hinge region and overall shape of the bound inhibitors are shared between docked compounds and templates. The second scenario applies to the three fragments (MAP21, MAP27, MAP32) whose poses were also precisely recovered with GRIM. In all cases, the good pose was inferred by bioisosterism (same interaction pattern but different chemical structure) to a larger template ligand (Table 3). For example, the hydroxyphenyl-aminopyrimidine scaffold of MAP21 is perfectly docked (rmsd to X-ray structure = 0.51 Å) to MAP4K4 because of the high similarity of its interaction pattern graph to that observed in the 4rvt PDB template, although the latter ligand exhibits a chemically different but bioisosteric scaffold (Fig. 7). It is interesting to notice that GRIM did not select an interaction pattern graph generated by a much chemically closer template ligand (e.g. 4obo, 4obq) therefore demonstrating that our pose selection protocol is really biased by protein–ligand interactions and not dominated by simple ligand chemical neighborhood. Since Surflex-Dock was able to generate at least one reliable pose for all these ligands, the reason for GRIM failure to detect it (third scenario) usually lies in wrong graph alignments dominated by hydrophobic interactions. A prototypical example is illustrated with the incorrect pose of MAP06 (rmsd to X-ray pose = 5.19 Å) by analogy to that of the 4obp template (Fig. 8) where GRIM optimizes the shape overlap between the two interaction patterns without a single shared hydrogen-bond. The overlay of the GRIM pose to that of the 4obp template notably highlights a very good match of both pyridine rings which serve as pure hydrophobic anchors to the MAP4K4 binding site. In fact, MAP06 H-bonds to the kinase hinge by its pyridine nitrogen atom (Fig. 8). This interaction is indeed found in some poses which were not rewarded by GRIM because of a lower overall GRIMscore. Since fragments with a dominant hydrophobic character exhibit simpler interaction patterns, the risk of misaligning the corresponding graphs to that of larger templates is relatively high, therefore explaining many of the herein observed failures (Table 4).
Predicted versus X-ray pose of the MAP4K4 inhibitor MAP18. Heteroatoms are colored in blue (nitrogen), red (oxygen), and green (fluorine). A red arrow indicates the location of the hinge region (E106, M107, C108) of the kinase. The chemical structures of inhibitor and template are displayed below the binding poses. a Predicted (tan sticks) and X-ray poses (cyan sticks) of MAP18 bound to MAP4K4 (white ribbons). b The GRIM pose (tan sticks) has been selected by interaction pattern similarity to that of PDB template 4zk5 (plum sticks)
Predicted versus X-ray pose of the MAP4K4 inhibitor MAP21. Heteroatoms are colored in blue (nitrogen), and red (oxygen). A red arrow indicates the location of the hinge region (E106, M107, C108) of the kinase. The chemical structures of inhibitor and template are displayed below the binding poses. a Predicted (tan sticks) and X-ray poses (cyan sticks) of MAP21 bound to MAP4K4 (white ribbons). b The GRIM pose (tan sticks) has been selected by interaction pattern similarity to that of PDB template 4rvt (plum sticks)
Predicted versus X-ray pose of the MAP4K4 inhibitor MAP06. Heteroatoms are colored in blue (nitrogen), red (oxygen), and yellow (sulfur). A red arrow indicates the location of the hinge region (E106, M107, C108) of the kinase. The chemical structures of inhibitor and template are displayed below the binding poses. a Predicted (tan sticks) and X-ray poses (cyan sticks) of MAP06 bound to MAP4K4 (white ribbons). b The GRIM pose (tan sticks) has been selected by interaction pattern similarity to that of PDB template 4obp (plum sticks)
We recall here that all poses were prefiltered before GRIM scoring for hydrogen-bonding to the hinge region of the kinase. This filtering step improved the mean rmsd of the 30 MAP4K4 inhibitors from 3.62 to 3.18 Å. For 21 out of 30 inhibitors, the filter has no effect since exactly the same pose was selected by GRIM with or without filtering. For 4 inhibitors (MAP01, MAP004, MAP12, MAP32), the filter positively contributes to the selection of better poses. Notably, the mean rmsd of compound MAP01 could be decreased from 10.78 to 0.98 Å. Conversely, the filtering was detrimental in 5 cases, most of the time with a very marginal rmsd increase. All rmsd values with and without the hydrogen bond filter are described in Supplementary Table 5.
Comparative evaluation of GRIM rescoring
The release, by the D3R Grand challenge 2015 organizers, of results from all contributions permits a comparative evaluation of our pose selection method with respect to many others (Fig. 9). Two criteria have been retained to estimate the accuracy of every method. First, the mean rmsd of the best possible pose (lowest rmsd to the X-ray structure) was selected to reflect the overall quality of the posing algorithm. Second, the mean rmsd of the top-ranked pose illustrates the capacity of a scoring function to reward docking solutions that are very close to the true X-ray pose.
Comparative evaluation of GRIM (red triangles) with other approaches (gray dots) in predicting the binding mode of 36 inhibitors (6 HSP90α inhibitors, 30 MAP4K4 inhibitors) prior to the release of their protein-bound X-ray coordinates. Posing accuracy is evaluated by the rmsd of predicted poses to the X-ray solution. a Plotting the mean rmsd of the top-ranked pose versus the mean rmsd of the absolute best (lowest rmsd) pose for six HSP90α inhibitors; b Plotting the mean rmsd of the top-ranked pose versus the mean rmsd of the absolute best (lowest rmsd) pose for 30 MAP4K4 inhibitors
Among the 42 contributions to predict the binding pose of HSP90α inhibitors, GRIM is ranked 3rd when considering the average rmsd of the top-ranked pose (Fig. 9a). Seven methods deliver quite accurate answers with a mean rmsd of the top-ranked pose below 1.5 Å, one method being slightly better than GRIM (rmsd of 0.85 Å). Since contributions are anonymous, we are not aware, at the time this manuscript was written, of the corresponding method and its sophistication level.
The much more challenging MAP4K4 dataset drew less attention with 27 answers. The quality of the corresponding predictions is significantly lower than for the HSP90α dataset (Fig. 9b). Only 3 contributors predicted the pose of the 30 MAP4K4 inhibitors with an accuracy below 3.5 Å when considering the top-ranked pose. GRIM is one of these 3 methods being ranked second in this challenge (Fig. 9b). Looking at the accuracy of the best possible pose clearly highlights a docking problem since deviations to the X-ray pose remains between 2 and 3 Å for the best methods (Fig. 9b). Reasons for failures have already been discussed in the previous section of this manuscript and therefore do not only concern our docking engine (Surflex-Dock) but also all other dockers used in this competition.
We do not know whether it is the same method that slightly outperform GRIM in predicting the pose of both HSP90α and MAP4K4 inhibitors. Our interaction pattern-guided pose selection strategy is anyhow quite robust and accurate, with respect to competitor methods as it ranks 3rd and 2nd, respectively for the two sets of predictions. As to be expected, the quality of the results depends on the preexisting knowledge. When numerous and diverse interaction patterns are available for a particular target (e.g. HSP90α dataset), GRIM docking poses are very accurate. If less information is known (e.g. MAP4K4 dataset), the quality of the GRIM poses logically deteriorates but still remains better than that obtained without GRIM rescoring for the same set of poses. We have not investigated here the possibility to select more interaction pattern templates (e.g. from other protein kinases) for posing MAP4K4 inhibitors, as preliminary GRIM pairwise comparisons between the eight available MAP4K4-inhibitor complexes and 1548 protein kinase-inhibitor complexes (sc-PDB dataset) [37] were not particularly promising (GRIMscore <0.70). In some cases however, a protein family-based GRIM scoring strategy has been shown to be useful [36] and should not be forgotten. At this point, it should be recalled that selecting a near-native pose by GRIM matching does not mean that predicting the binding free energy of that pose with state-of-the art scoring functions will deliver good results. Indeed, we could not find any correlation (Pearson R = 0.19) between the Surflex-Dock score of GRIM-1 poses and experimental binding affinities of 180 HPS90α inhibitors (stage 2 of the challenge).
Conclusions
The herein presented GRIM method rescores docking poses by interaction pattern graph similarity to known protein–ligand X-ray structures. The methodology is both very simple and intuitive. Basically, the method automatizes the reasoning of a molecular modeler: Does this pose remind me the binding mode of known ligands for this protein or its close homologues?
Conceptually, it is different from many shape or template-matching docking methods [23–26] recently reported to outperform free docking in generating reliable poses. GRIM operates on freely generated docking poses but will just reward that poses which lead to interaction patterns similar to known ligands of the same or related target protein. In ca. 80–90 % of test cases, state-of-the-art docking engines propose a set of poses out of which at least one is close to the X-ray solution [12]. GRIM can therefore be used in addition to any of these dockers to prioritize the most relevant ones. The method is fast (20 ms/pose on average) and independent on the docking engine, however protein–ligand coordinates should be provided in a standard MOL2 format.
When applied to the a priori prediction of binding poses for 36 new inhibitors of two different targets, GRIM compares very favorably with competing methods as it ranked 3rd and 2nd, for HSP90α and MAP4K4 dataset respectively, in predicting near native poses. In most cases, the top-ranked pose as predicted by GRIM is the one that is the closest to the true solution. As any knowledge-based method, the accuracy of GRIM depends on existing experimental data. Depending on the target, the number and chemical diversity of co-crystallized ligands may vary quite significantly. GRIM rewards the pose with the closest interaction pattern to that seen in any other crystal of the same target, independently of how frequently this pose has already been obtained experimentally. The more chemically-diverse ligands co-crystallized with the target (or close homologues) are available, the higher the probability of the first GRIM pose being near native. The user should therefore be aware of the target-dependent applicability domain of the method, before using it blindly. The corresponding executable (IChem) is available for non-profit academic research at http://bioinfo-pharma.u-strasbg.fr/labwebsite/download.html.
Supporting information
List of 180 HSP90α inhibitors to dock, list of 176 PDB templates for HSP90α-inhibitor complexes, list of 30 MAP4K4 inhibitors to dock, list of 8 PDB templates for MAP4K4-inhibitor complexes, effect of hydrogen-bond filtering on the quality of GRIM top-ranked poses.
References
Chen YC (2015) Beware of docking! Trends Pharmacol Sci 36:78–95
Kuntz ID, Blaney JM, Oatley SJ, Langridge R, Ferrin TE (1982) A geometric approach to macromolecule–ligand interactions. J Mol Biol 161:269–288
Moitessier N, Englebienne P, Lee D, Lawandi J, Corbeil CR (2008) Towards the development of universal, fast and highly accurate docking/scoring methods: a long way to go. Br J Pharmacol 153(Suppl 1):S7–26
Yuriev E, Holien J, Ramsland PA (2015) Improvements, trends, and new ideas in molecular docking: 2012–2013 in review. J Mol Recognit 28:581–604
Sousa SF, Ribeiro AJ, Coimbra JT, Neves RP, Martins SA, Moorthy NS, Fernandes PA, Ramos MJ (2013) Protein–ligand docking in the new millennium—a retrospective of 10 years in the field. Curr Med Chem 20:2296–2314
Brooijmans N, Kuntz ID (2003) Molecular recognition and docking algorithms. Annu Rev Biophys Biomol Struct 32:335–373
Kellenberger E, Rodrigo J, Muller P, Rognan D (2004) Comparative evaluation of eight docking tools for docking and virtual screening accuracy. Proteins 57:225–242
Warren GL, Andrews CW, Capelli AM, Clarke B, LaLonde J, Lambert MH, Lindvall M, Nevins N, Semus SF, Senger S, Tedesco G, Wall ID, Woolven JM, Peishoff CE, Head MS (2006) A critical assessment of docking programs and scoring functions. J Med Chem 49:5912–5931
Smith RD, Damm-Ganamet KL, Dunbar JB Jr, Ahmed A, Chinnaswamy K, Delproposto JE, Kubish GM, Tinberg CE, Khare SD, Dou J, Doyle L, Stuckey JA, Baker D, Carlson HA (2016) CSAR benchmark exercise 2013: evaluation of results from a combined computational protein design, docking, and scoring/ranking challenge. J Chem Inf Model 56:1022–1031
Damm-Ganamet KL, Smith RD, Dunbar JB Jr, Stuckey JA, Carlson HA (2013) CSAR benchmark exercise 2011–2012: evaluation of results from docking and relative ranking of blinded congeneric series. J Chem Inf Model 53:1853–1870
Plewczynski D, Lazniewski M, Augustyniak R, Ginalski K (2011) Can we trust docking results? Evaluation of seven commonly used programs on PDBbind database. J Comput Chem 32:742–755
Li Y, Han L, Liu Z, Wang R (2014) Comparative assessment of scoring functions on an updated benchmark: 2. Evaluation methods and general results. J Chem Inf Model 54:1717–1736
Novikov FN, Zeifman AA, Stroganov OV, Stroylov VS, Kulkov V, Chilov GG (2011) CSAR scoring challenge reveals the need for new concepts in estimating protein–ligand binding affinity. J Chem Inf Model 51:2090–2096
Wang JC, Lin JH (2013) Scoring functions for prediction of protein–ligand interactions. Curr Pharm Des 19:2174–2182
Virtanen SI, Niinivehmas SP, Pentikainen OT (2015) Case-specific performance of MM-PBSA, MM-GBSA, and SIE in virtual screening. J Mol Graph Model 62:303–318
Kuhn B, Gerber P, Schulz-Gasch T, Stahl M (2005) Validation and use of the MM-PBSA approach for drug discovery. J Med Chem 48:4040–4048
Hou T, Wang J, Li Y, Wang W (2011) Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. J Chem Inf Model 51:69–82
Li L, Wang B, Meroueh SO (2011) Support vector regression scoring of receptor–ligand complexes for rank-ordering and virtual screening of chemical libraries. J Chem Inf Model 51:2132–2138
Zilian D, Sotriffer CA (2013) SFCscore(RF): a random forest-based scoring function for improved affinity prediction of protein–ligand complexes. J Chem Inf Model 53:1923–1933
Ballester PJ, Schreyer A, Blundell TL (2014) Does a more precise chemical description of protein–ligand complexes lead to more accurate prediction of binding affinity? J Chem Inf Model 54:944–955
Khamis MA, Gomaa W, Ahmed WF (2015) Machine learning in computational docking. Artif Intell Med 63:135–152
Gabel J, Desaphy J, Rognan D (2014) Beware of machine learning-based scoring functions-on the danger of developing black boxes. J Chem Inf Model 54:2807–2815
Hindle SA, Rarey M, Buning C, Lengauer T (2002) Flexible docking under pharmacophore type constraints. J Comput Aided Mol Des 16:129–149
Kelley BP, Brown SP, Warren GL, Muchmore SW (2015) POSIT: flexible shape-guided docking for pose prediction. J Chem Inf Model 55:1771–1780
Kumar A, Zhang KY (2016) Application of shape similarity in pose selection and virtual screening in CSARdock2014 exercise. J Chem Inf Model 56:965–973
Gao C, Thorsteinson N, Watson I, Wang J, Vieth M (2015) Knowledge-based strategy to improve ligand pose prediction accuracy for lead optimization. J Chem Inf Model 55:1460–1468
Deng Z, Chuaqui C, Singh J (2004) Structural interaction fingerprint (SIFt): a novel method for analyzing three-dimensional protein–ligand binding interactions. J Med Chem 47:337–344
Anighoro A, Bajorath J (2016) Three-dimensional similarity in molecular docking: prioritizing ligand poses on the basis of experimental binding modes. J Chem Inf Model 56:580–587
Marcou G, Rognan D (2007) Optimizing fragment and scaffold docking by use of molecular interaction fingerprints. J Chem Inf Model 47:195–207
Kelly MD, Mancera RL (2004) Expanded interaction fingerprint method for analyzing ligand binding modes in docking and structure-based drug design. J Chem Inf Comput Sci 44:1942–1951
Mpamhanga CP, Chen B, McLay IM, Willett P (2006) Knowledge-based interaction fingerprint scoring: a simple method for improving the effectiveness of fast scoring functions. J Chem Inf Model 46:686–698
Chalopin M, Tesse A, Martinez MC, Rognan D, Arnal JF, Andriantsitohaina R (2010) Estrogen receptor alpha as a key target of red wine polyphenols action on the endothelium. PLoS ONE 5:e8554
Venhorst J, Nunez S, Terpstra JW, Kruse CG (2008) Assessment of scaffold hopping efficiency by use of molecular interaction fingerprints. J Med Chem 51:3222–3229
de Graaf C, Rein C, Piwnica D, Giordanetto F, Rognan D (2011) Structure-based discovery of allosteric modulators of two related class B G-protein-coupled receptors. ChemMedChem 6:2159–2169
de Graaf C, Kooistra AJ, Vischer HF, Katritch V, Kuijer M, Shiroishi M, Iwata S, Shimamura T, Stevens RC, de Esch IJ, Leurs R (2011) Crystal structure-based virtual screening for fragment-like ligands of the human histamine H(1) receptor. J Med Chem 54:8195–8206
Desaphy J, Raimbaud E, Ducrot P, Rognan D (2013) Encoding protein–ligand interaction patterns in fingerprints and graphs. J Chem Inf Model 53:623–637
Desaphy J, Bret G, Rognan D, Kellenberger E (2015) sc-PDB: a 3D-database of ligandable binding sites—10 years on. Nucleic Acids Res 43:D399–D404
Bron C, Kerbosch J (1973) Algorithm 457: finding all cliques of an undirected graph. Commun ACM 16:575–577
Johnston HC (1976) Cliques of a graph—variations on the Bron–Kerbosch algorithm. Int J Parallel Prog 5:209–238
Theobald DL (2005) Rapid calculation of RMSDs using a quaternion-based characteristic polynomial. Acta Crystallogr A 61:478–480
Jain AN (2007) Surflex-Dock 2.1: robust performance from ligand energetic modeling, ring flexibility, and knowledge-based search. J Comput Aided Mol Des 21:281–306
Mysinger MM, Carchia M, Irwin JJ, Shoichet BK (2012) Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J Med Chem 55:6582–6594
Drug Design Data Resource. https://drugdesigndata.org/about/grand-challenge-2015
Bietz S, Urbaczek S, Schulz B, Rarey M (2014) Protoss: a holistic approach to predict tautomers and protonation states in protein–ligand complexes. J Cheminform 6:12
Tripos International, St. Louis, MO 63144–2319, USA
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
UniProt Consortium (2015) UniProt: a hub for protein information. Nucleic Acids Res 43:D204–D212
Molecular Networks GmbH, Erlangen, Germany
Kung PP, Sinnema PJ, Richardson P, Hickey MJ, Gajiwala KS, Wang F, Huang B, McClellan G, Wang J, Maegley K, Bergqvist S, Mehta PP, Kania R (2011) Design strategies to target crystallographic waters applied to the Hsp90 molecular chaperone. Bioorg Med Chem Lett 21:3557–3562
Acknowledgments
We thank the LABEX ANR-10-LABX-0034 Medalis for a post-doctoral fellowship to I.S. We also acknowledge the National Center for Scientific Research (CNRS, Institut de Chimie) and the Alsace Region for a doctoral fellowship to FDS. The High-performance Computing Center (University of Strasbourg, France) and the Calculation Center of the IN2P3 (CNRS, Villeurbanne, France) are acknowledged for allocation of computing time and excellent support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Slynko, I., Da Silva, F., Bret, G. et al. Docking pose selection by interaction pattern graph similarity: application to the D3R grand challenge 2015. J Comput Aided Mol Des 30, 669–683 (2016). https://doi.org/10.1007/s10822-016-9930-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-9930-3