Abstract
Aphanizomenon flos-aquae, a freshwater cyanobacterium harvested from Upper Klamath Lake, Oregon, USA, is employed as a functional food material. Aphanizomenon flos-aquae can be fermented with Lactobacillus plantarum and Lactococcus lactis subsp. lactis. Here, to determine the dietary effects of A. flos-aquae (AFA) and fermented A. flos-aquae (F-AFA) on the mouse gut microbiome, high-sucrose diet containing no dietary fibre (NF), 5% AFA or 5% F-AFA was administered to ICR mice for 14 days. Compared with those in mice fed the NF, lower body weight gains and epididymal fat pad tissue weights were observed in mice fed the diet containing F-AFA. After feeding, the caecal microbiome was analysed by 16S rRNA (V4) gene amplicon sequencing using the Illumina MiSeq system. Sequences were clustered into operational taxonomic units, with a 97% identity cutoff, using the QIIME2 workflow script and SILVA database. The caecal microbiome alpha diversity was high in mice fed the F-AFA diet. The abundances of Muribaculum- and Alistipes inops-like bacteria belonging to the phylum Bacteroidetes and Ruminococcaceae UCG-014-like bacteria belonging to phylum Firmicutes were higher in the F-AFA group than in the AFA group. Isolation of typical F-AFA-susceptible gut indigenous bacteria and functional studies of the isolates in the presence of F-AFA are warranted to validate the current findings.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aphanizomenon flos-aquae (AFA) is a freshwater bloom-forming cyanobacterium that poses a threat to the marine environment and fishery industries worldwide (Cirés and Ballot 2016) due to its production of toxic compounds, such as paralytic shellfish toxins and respiratory toxic aphantoxins (Pereira et al. 2000; Zhang et al. 2016). However, ingestion of A. flos-aquae harvested from Upper Klamath Lake, Oregon, USA, and which is free of toxins (Carmichael et al. 2000) has demonstrated effects, such as human monocyte activation, antioxidant properties and hypolipidemia (Kushak et al. 2000; Pugh and Pasco 2001; Benedetti et al. 2004).
In vitro growth-promoting (prebiotic) activities of A. flos-aquae and its water extract were reported for Lactobacillus acidophilus (Campana et al. 2017). However, a Lactobacillus-promoting activity of AFA was not found in the colon of inbred C57BL/6J mice fed normal chow and the caecum of closed colony Institute of Cancer Research (ICR) mice fed a high-sucrose and low-dietary fibre diet (Rasmussen et al. 2006; Takei et al. 2019). In the case of ICR mice, the abundance of Muribaculum sp.- and Alistipes inops-like bacteria belonging to the phylum Bacteroidetes was increased, while that of Bifidobacterium pseudolongum-like bacteria was decreased after supplementation with AFA.
Humans and laboratory animals generally have hundreds to > 1000 species of bacteria at 11 to 12 log cells g−1 in their gut; the metabolic activities of the gut microbiome and host are known to affect each other significantly (Frazier and Chang 2020). An increase in the abundance of Muribaculum and Alistipes through dietary fibre intake has been reported (Chang et al. 2017; Wang et al. 2020). Although B. pseudolongum is regarded as a beneficial commensal bacterium in the murine gut and some other mammal commensal bacteria have beneficial properties for the host, such as hypolipidemic and gut protection activities (Mangin et al. 2018; Bo et al. 2020), B. pseudolongum is highly abundant in ICR mice fed a diet without insoluble dietary fibre (cellulose) (Kuda et al. 2017; Takei et al. 2019).
In our previous study in vitro antioxidant (O2− radical scavenging), anti-glycation (bovine serum albumen-fructose (BSA-Fru) model) and immune modulation activities of AFA were increased by fermentation with Lactobacillus plantarum and Lactococcus lactis subsp. lactis (Taniguchi et al. 2019). Fermented AFA (F-AFA) with such induced activities was separated into < 3 kDa and 30–100 kDa fractions. The active compounds were hypothesised to be lactate, oligopeptides and polypeptides. Based on these properties found in vitro, the effects of dietary supplementation with AFA and F-AFA on the gut microbiome and diet-related lifestyle diseases may differ. However, an in vivo study of F-AFA has not been conducted. Recently, researchers reported that the gut bacterial balance is rapidly altered by changes in the dietary composition before the appearance of lifestyle disease markers and/or symptoms (David et al. 2014; Williams et al. 2019).
In the present study, to determine the dietary effects of AFA and F-AFA on the gut microbiome, a high-sucrose diet containing no dietary fibre (NF), 5% AFA or 5% F-AFA was administered to ICR mice for 14 days, after which the caecal microbiome was analysed by 16S rRNA (V4) gene amplicon sequencing.
Materials and methods
Preparation of F-AFA
Lactobacillus plantarum AN7 (GenBank accession number: LC384876) isolated from a fermented fish product made in the Noto Peninsula, Ishikawa, Japan, and Lactococcus lactis subsp. lactis Oga-SU2 (LC208001) isolated from beach sand in the Oga Peninsula, Akita, Japan, were used as starter strains. The strains were precultured in 5 mL of de Man, Rogosa, Sharpe (MRS) broth (Oxoid, UK) at 37 °C for 24 h.
Freeze-dried powdered product of Aphanizomenon flos-aquae (AFA) harvested from Upper Klamath Lake was obtained from Dr’s Choice Co. (Tokyo, Japan). Aphanizomenon flos-aquae powder (100 g) was added to 900 mL of distilled water (DW), adjusted pH 7.0 with NaOH and autoclaved at 121 °C for 15 min. The precultured starter (5 mL) were inoculated into half (500 mL) of the suspended AFA and incubated at 37 °C for 7 days to produce F-AFA. The AFA and F-AFA suspensions were dried at 70 °C for 24 h for use in the animal experiments.
Lactic acid generation was confirmed using high-performance liquid chromatography (HPLC): column, ICSep ICE-ORH-801 (Tokyo Chemical Industry Co., Ltd., Japan); operating temperature, 35 °C; eluent, 0.005 mol L−1 of sulphuric acid (H2SO4); flow rate, 0.8 mL min−1. Eluted compounds were detected using a refractive index (RI) detector. Saccharides and organic acids in AFA and F-AFA were measured as previously reported (Shikano et al. 2019). Fourier-transform infrared (FT-IR) spectra of AFA and F-AFA were determined using an IR-attenuated total reflection (ATR) diamond attached Thermo Nicolet iS5 FT-IR spectrometer (Thermo Electron Corp., USA) and compared with those of cellulose and ethanol-washed cells of the starter strains (Kuda et al. 2013).
Animal care
Animal experiments were performed in accordance with the Fundamental Guidelines for Proper Conduct of Animal Experiments and Related Activities in Academic Research Institutions, under the jurisdiction of the Ministry of Education, Culture, Sports, Science and Technology, Japan. The study protocol was approved by the Animal Experiment Committee of the Tokyo University of Marine Science and Technology (Approval No. H31-5).
Eighteen 5-week-old male ICR mice purchased from Tokyo Laboratory Animal Science (Tokyo, Japan) were housed in metal wire cages (three mice per cage) at 22 ± 2 °C. The mice were acclimatised to the high-sucrose and low-dietary fibre diet and provided DW for drinking ad libitum. After 7 days, the mice were divided into three (NF, AFA and F-AFA) groups (n = 6, in each group) and fed a diet containing either no fibre (NF), 5% (w/w) AFA or 5% F-AFA for 14 days (Table 1). During the 8–10 feeding days, defecation frequency and faecal weight were measured. At the experimental endpoint, the mice were exsanguinated via the abdominal vein under anaesthesia with isoflurane (Fujifilm Wako Pure Chemical, Japan), and their liver, kidneys, spleen and epididymal fat pads were removed and weighed. After ligation with yarn, the caecum and colon were excised and placed on ice until microbial analysis was conducted.
Plasma triacylglyceride (TG), total cholesterol and glucose (Glu) levels were determined using commercial kits (Triglyceride E-Test Wako, Total Cholesterol E-Test Wako, Glucose CII-Test Wako, respectively; Fujifilm Wako Pure Chemical Corporation) according to the manufacturer’s instructions.
Bacterial cell count and isolation
The caecal contents were diluted with 99 volumes of phosphate-buffered saline (PBS; Nissui Pharmaceuticals, Japan), and the bacterial cells were counted by dielectrophoretic impedance measurement (DEPIM) (Hirota et al. 2014) using a bacteria counter (PHC Ltd., Japan).
A slight modification of the culture-dependent method described by Mitsuoka (2014) was used for viable cell counting. The caecal content was serially diluted with “dilution A” (KH2PO4, 4.5 g; Na2HPO4, 6 g; l-cysteine·HCl·H2O, 0.5 g; Tween 80, 0.5 g; agar, 0.75 g L−1). Aliquots of the diluted sample (0.03 mL) were plated separately on blood liver (BL) and Gifu Anaerobic Medium (GAM) agar (Nissui Pharmaceuticals Co., Ltd., Japan) plates containing 5% (v/v) defibrinated horse blood (Nippon Bio-Supply Center, Japan) and incubated at 37 °C for 48 h under anaerobic conditions using an AnaeroPack (Mitsubishi Gas Chemical, Japan). Detected colonies with typical morphologies were enumerated. Typical colonies were then isolated and further purified using the agar plates and conditions described above. The 16S rRNA genes of the isolates were amplified using polymerase chain reaction (PCR) primers 27F and 1492R, and amplicon sequencing was conducted by Macrogen Japan, Corp. (Kyoto, Japan). A homology search of the DNA Data Bank of Japan was performed using BLASTn (http://ddbj.nig.ac.jp/blast/blastn).
Analysis of the caecal microbiota using the MiSeq system
16S rDNA (V4) amplicon sequencing was conducted by Fasmac Co., Ltd. (Atsugi, Japan). DNA was extracted from the samples using the Mpure bacterial DNA extraction kit (MP Bio Japan, Japan). A DNA library was prepared using a two-step PCR (Sinclair et al. 2015). Then, the V4 region was amplified using a 23-cycle PCR with the following primers: forward, 515f 5′-ACACTCTTTCCCTACACGACGCTCTTCCGATCT-GTGCCAGCMGCCGCGGTAA-3′ and reverse, 806r 5′-GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT-GGACTACHVGGGTWTCTAAT-3′. Next, individual DNA fragments were tagged in an eight-cycle PCR with the following primers: forward, 5′-AATGATACGGCGACCACCGAGATCTACAC-[sequence for individual mouse]-ACACTCTTTCCCTACACCGACGC-3′ and reverse, 5′-CAAGCAGAAGACGGCATACGAGAT-[sequence for individual mouse]-CTGACTGGAGTTCAGACGTGTG-3′. DNA libraries were multiplexed and loaded onto an Illumina MiSeq system (Illumina, San Diego, CA, USA). Readings with a mismatched sequence at the start region were filtered using the FASTX Toolkit (http://hannonlab.cshl.edu/fastx_toolkit/); poor quality reads (below 20) and those shorter than 40 base pairs were omitted using Sickle (https://github.com/ucdavis-bioinformatics/sickle). Shortlisted reads were merged using the pair-end merge script FLASH (http://ccb.jhu.edu/software/FLASH/), and 240–260 base reads were selected. Chimeras in the selected reads were identified and omitted using the QIIME2 bioinformatics pipeline (https://qiime2.org/). Sequences were clustered into operational taxonomic units (OTUs), with a 97% identity cutoff, using the QIIME2 workflow script and SILVA database (https://www.arb-silva.de/).
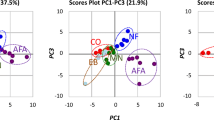
The alpha diversity (diversity within a microbiome) of the mouse gut microbiome was determined using the Shannon-Wiener (H′) and Simpson’s diversity (D) indices (Kim et al. 2017). Beta diversity (distance between groups based on differences in the OTUs present in each group) was assessed using unweighted and weighted UniFrac (Lozupone and Knight 2005) and expressed using principal coordinate analysis (PCoA) (Bunyavanich et al. 2016).
Statistical analysis
The body and organ weights and alpha diversity indices were expressed as the mean value ± standard error of the mean. Data were subjected to analysis of variance and Tukey’s and Dunnett’s post hoc tests, using a statistical software package (Excel Statistic Ver. 6, Japan). p < 0.05 was considered statistically significant.
Results
HPLC chromatogram and FT-IR spectra of AFA and F-AFA
The AFA suspension was fermented with L. plantarum AN7 and L. lactis subsp. lactis Oga-SU2 for 7 days, during which the pH decreased from approximately 6.3 to 5.2, and glucose and other saccharides were mainly converted to lactic acid (Fig. 1a). Lactate, already present in AFA, increased from 39 to 100 mg g−1 during fermentation. The saccharide and organic acid results concurred with those reported in a previous study (Taniguchi et al. 2019). A small quantity of acetate was also detected in F-AFA. The major peaks in the FT-IR spectra (Fig. 1b) were defined according to previous reports (Al-Qadiri et al. 2008). The carbohydrate absorbance (1200–1000 cm−1) of F-AFA was lower than that of AFA. The absorbance of F-AFA at 3000–2800 cm−1 (CH2, CH3), although not large, was higher than that of AFA, which might be due to the produced lactate.
Body, faecal, organ weights, plasma lipid and glucose levels
No abnormalities or adverse symptoms were observed in the mice fed NF, AFA, or F-AFA diets. Body weight gains were 32% lower in the F-AFA group and also tended to be lower in the AFA group than in the NF group (Table 2). On the other hand, faecal and caecal content weights and colon lengths were higher in mice fed AFA than in those fed F-AFA. A blue-green colour appeared in the faeces of mice fed AFA and F-AFA. The epididymal fat pad weights were 20% lower in the F-AFA group. The kidney weights were slightly higher in mice fed AFA. Though not significant, plasma TG levels tended to be lower in mice fed F-AFA rather than AFA. Plasma Glu levels tended to be lower in both the AFA and F-AFA groups than in the NF group.
Caecal viable and direct cell counts
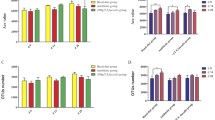
Three dominant typical colonies appeared on the BL agar plates (Table 3). Among them, colonies two and one were identified as Lactobacillus murinus and Lactobacillus johnsonii, respectively. The total colony count in the NF group was 8.5 log CFU g−1 and was lower in the AFA and F-AFA groups. Two dominant typical colonies were observed on the GAM blood agar plates and were identified as Bifidobacterium pseudolongum and Bacteroides intestinalis. These colony counts were also lower in the F-AFA group than in the NF and AFA groups. On the contrary, the direct total bacterial cell count in the caeca of tested mice was approximately 11 log cell g−1 and did not differ between diets (Table 4).
Caecal microbiota analysis by 16S rDNA (V4) amplicon sequencing
Alpha and beta diversity of the caecal microbiome
The total read number was found to be 45,000–52,000 base pairs upon 16S rDNA (V4) amplicon sequencing and was higher in mice fed F-AFA (Table 4) than in the mice fed NF. The number of OTUs (120–200) was 1.6 times higher in the AFA and F-AFA groups than in the NF group. The F-AFA group had higher alpha diversity, represented by the Shannon-Weaver H′ and Simpson’s D indices.
PCoA of the OTUs revealed that the gut microbiomes of the AFA and F-AFA groups differed from that of the NF group (Fig. 2) and that the gut microbiome clusters of the AFA and F-AFA groups overlapped.
Phylum level
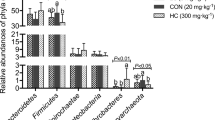
The caecal microbiome profiles at the phylum, family and genus levels are expressed by relative abundance, as shown in Fig. 3. The predominant phylum in mice fed NF was Firmicutes (65%) followed by Bacteroidetes (12%), Actinobacteria (19%) and Proteobacteria (2.5%) (Fig. 3a). The abundance of Bacteroidetes tended to be higher in the F-AFA group (23%) than in the NF group. Actinobacteria abundance was lower in the AFA and F-AFA groups. Verrucomicrobia was hardly detected in mice fed NF and tended to be higher in mice fed F-AFA (1.8%). Although present at low levels (0.22%), the abundance of Tenericutes was higher in the F-AFA group.
Composition of the caecal microbiome at the phylum (a), family (c) and genus (d) levels and ratio of caecal Firmicutes to caecal Bacteroidetes (F/B) (b) in mice fed a high-sucrose and low-dietary fibre diet containing no additional fibre (NF), 5% w/w dried Aphanizomenon flos-aquae (AFA) or 5% w/w fermented AFA (F-AFA). *Significant differences among the groups determined by Tukey’s method and/or Dunnett’s method. Superscript letters “a and b” means no significant difference between values with the same letters. Arrows indicate a significant difference from the NF group (p < 0.05)
Family, genus and OTU levels
The dominant family in the phylum Firmicutes in the NF group was Erysipelotrichaceae (44%), followed by Lactobacillaceae (7.4%), Clostridiaceae 1 (5.9%) and Lachnospiraceae (5.5%) (Fig. 2c). The abundance of Erysipelotrichaceae tended to be lower in the AFA (36%) and F-AFA (34%) groups. On the other hand, Ruminococcaceae and Lachnospiraceae abundance levels tended to higher in mice fed AFA and F-AFA. In the phylum Bacteroidetes in the NF group, families Bacteroidaceae (5.6%) and Muribaculaceae (3.8%) were dominant. Muribaculaceae abundance was 1.5 to 1.6 times higher in the AFA (8.3%) and F-AFA (8.9%) groups. Rikenerellaceae was hardly detected in the NF group and had an abundance of 3.8% in the F-AFA group. Bifidobacteriaceae was the predominant family of the phylum Actinobacteria, but the abundance of Eggerthellaceae, also belonging to Actinobacteria, was higher (0.9%) in mice fed F-AFA. Akkermansiaceae was the main family of the phylum Verrucomicrobia.
In Erysipelotrichaceae, the predominant family of phylum Firmicutes in the tested mice, genera Faecalibaculum (14–20%), Allobaculum (13–19%) and Turicibacter (2.5–4.4%) were dominant (Fig. 2d). In other families of the phylum Firmicutes, Lactobacillus (7.4–11%), Clostridium sensu_strict_1 (5.9–8.1%) and Lachnospiraceae g. were also dominant. Ruminococcaceae UCG-014 abundance was higher in mice fed F-AFA. Lachnoclostridium abundance was higher in the AFA and F-AFA groups. Alistipes was the predominant genus of the family Rikenellaceae in the F-AFA group.
Table 5 shows the read numbers of the 50 dominant OTUs. Several OTUs defined as Muribaculum intestinale (93–94%)- and Muribaculum sp. (99–100%)-like bacteria were higher in mice fed AFA and F-AFA. Clostridium fusiformis (100%)- and Parasutterella excrementihominis (94–100%)-like bacteria, although with smaller read numbers, were also higher in the AFA and F-AFA groups. Alistipes, dominant in the F-AFA group, was defined as A. inops-like bacteria. OTUs of Ruminococcaceae_UCG-014, Lactobacillus reuteri (100%)- and Clostridium aldenense/indolis (98%)-like bacteria were also higher in mice fed F-AFA.
Discussion
In the present study, to determine the dietary effects of AFA and F-AFA on the mouse gut microbiome, a high-sucrose diet containing no dietary fibre (NF), 5% AFA or 5% F-AFA was administered to ICR mice for 14 days. The caecal microbiomes were then analysed by 16S rRNA (V4) gene amplicon sequencing. As shown in Tables 2 and 4, compared with those in mice fed NF, body weight gain and epididymal fat pad weights were lower in mice fed F-AFA than in mice fed AFA, with higher alpha diversity of the caecal microbiome. The alpha diversity of the gut microbiome is positively correlated with dietary fibre intake and is responsible for its health functions (Payling et al. 2020); however, contrasting effects of probiotics on microbiome diversity have been reported (Preidis et al. 2013; Nagpal et al. 2018).
Although the hypolipidemic activity of AFA was previously reported (Kushak et al. 2000), the hypolipidemic and anti-obesity effects of F-AFA might be greater than those of AFA. In our previous study, the anti-glycation activity of F-AFA in the BSA-Fru model was higher than that of AFA (Taniguchi et al. 2019). L. plantarum AN7 has demonstrated in vitro cholesterol and bile acid lowering capabilities (Kuda et al. 2013; Kuda et al. 2016). These properties of fermented products and starter cells might be correlated with the anti-obesity effects of F-AFA.
There are many reports regarding the positive correlation between the gut Firmicutes to Bacteroidetes ratio (F/B) and diet-related lifestyle diseases, such as obesity and type 2 diabetes (Koliada et al. 2017; Zhao et al. 2019). In the present study, the F/B ratio was not significantly affected by diet, although it tended to be lower in mice fed AFA and F-AFA (Fig. 3b) than in mice fed NF. AFA was hypothesised to have a greater insoluble dietary fibre effect than F-AFA due to the colour and weight of faeces and colon length (Table 2). Cyanobacteria, such as AFA, have similar peptidoglycans and lipopolysaccharides to those of Gram-negative bacteria (Sharma et al. 2011), which may cause different effects from those induced by cellulose from general plant dietary fibre.
An increase in Muribaculum sp. was observed upon dietary supplementation with AFA and F-AFA (Table 5). Muribaculaceae (formerly, S24-7 family) is a dominant gut microbe in laboratory mice and has also been isolated from humans (Lagkouvardos et al. 2019). The abundance of Muribaculum in the mouse gut is reportedly lower when mice consume a high-fat diet and is increased by the intake of long-chain inulin and type 2-resistant starch, in addition to suppression of obesity and inflammation (Barouei et al. 2017; Hussain et al. 2019; Li et al. 2020). Gamma-aminobutyric acid-producing lactobacilli were also shown to increase gut Muribaculum abundance and reduce glycaemia (Ni et al. 2019; Patterson et al. 2019).
Alistipes sp. and Ruminococcaceae UCG-014 were increased by F-AFA (Table 5). Increasing the abundance of Alistipes sp. with insoluble dietary fibre intake was associated with the amelioration of obesity and type 2 diabetes (Lu et al. 2016; Chang et al. 2017). A. inops has been isolated from human faeces, and its major end products from saccharides are succinate and acetate (Shkoporov et al. 2015). Succinate is converted to propionate by other gut microbes (Reichardt et al. 2014). The family Ruminococcaceae, including the UCG-014 group, is regarded as a butyrate producer and exhibits a negative correlation with overweight and obesity (Gao et al. 2018); indeed, it was highly abundant in obesity model mice fed a high-fat diet (Zhang et al. 2019). Furthermore, the Ruminococcaceae UCG-014 abundance was much lower in ulcerative colitis model mice (Wang et al. 2018). There are many reports regarding gut propionate and butyrate functions, such as those indicating that they suppress blood cholesterol level and prevent colitis (Genda et al. 2018).
As mentioned above, although the typical dietary fibre effects were higher in the AFA group, such as increases in faecal and caecal content weights, body weight and epididymal fat tissue weight gains were lower in the F-AFA group. Furthermore, the plasma TG and Glu levels tended to be lower in mice fed F-AFA. These effects might be correlated with the increase in the abundances of dominant gut microbes, such as Muribaculum sp., A. inops and Ruminococcaceae UCG-014 and other minor microbes, although these could not be isolated with the standard culture methods employed in this study (Table 3).
Various species of caecal obligate anaerobes in ICR mice, such as B. pseudolongum, B. intestinalis, Bacteroides acidifaciens, Bacteroides vulgatus and [Clostridium] innocuum, could be detected and isolated by BL agar plating in our previous studies (Takei et al. 2019; Xia et al. 2019; Takei et al. 2020; Xia et al. 2020) but could not be detected in other studies (Yokota et al. 2018). Due to the direct cell count being a hundred times higher than the viable count, particularly in BL agar, severe obligate anaerobes might have been drastically reduced during the preparation and plating of caecal dilutions. Further study of the isolation of major obligate anaerobes is warranted to clarify the correlations among diet, gut microbiome and host. Caecal B. pseudolongum was previously detected in high abundance in ICR mice fed a diet with high sucrose and without insoluble dietary fibres, such as cellulose, and un-fermentable soluble dietary fibres, such as carrageenans (Kuda et al. 2017; Take et al. 2019).
Isolation of typical F-AFA-susceptible gut indigenous bacteria and functional studies of the isolates in the presence of F-AFA are warranted to validate the current findings. Additionally, to clarify the mechanisms of action of AFA and F-AFA on the gut microbiome and their applicability as functional food materials, further studies, particularly on the effects of longer AFA or F-AFA intake with respect to the sex, age and strain of laboratory animals and humans, are needed in the future.
Conclusion
To determine the dietary effects of AFA and F-AFA, a high-sucrose and low-dietary fibre diet containing no dietary fibre (NF), 5% AFA or 5% F-AFA was administrated to ICR mice for 14 days. Compared with those of mice fed NF, body weight and epididymal fat pad tissue weight gains were lower in mice fed AFA and even lower in mice fed F-AFA. The alpha diversity of the caecal microbiome was higher in mice fed F-AFA. The abundance levels of Muribaculum- and A. inops-like bacteria belonging to the phylum Bacteroidetes and Ruminococcaceae UCG-014-like bacteria belonging to the phylum Firmicutes were higher in the F-AFA group than in the AFA group. Isolation of typical F-AFA-susceptible gut indigenous bacteria and functional studies of the isolates in the presence of F-AFA are warranted to validate the current findings.
References
Al-Qadiri HM, Al-Holy AA, Cavinnato AG, Rasco BA (2008) Detection of sublethal thermal injury in Salmonella enterica serotype Typhimurium and Listeria monocytogenes using fourier transform infrared (FT-IR) spectroscopy. J Food Sci 73:M54–M61
Barouei J, Bendiks Z, Martinic A, Mishchuk D, Heeney D, Hsieh Y, Kieffer D, Zaragoza J, Martin R, Slupsky C, Marco M (2017) Microbiota, metabolome, and immune alterations in obese mice fed a high-fat diet containing type 2 resistant starch. Mol Nutr Food Res 61:20170084
Benedetti S, Benvenuti F, Pagliarani S, Francogli S, Scoglio S, Canestrati F (2004) Antioxidant properties of a novel phycocyanin extract from the blue-green alga Aphanizomenon flos-aquae. Life Sci 75:2353–2362
Bo T, Wen J, Zhao Y, Tian S, Zhang X, Wang D (2020) Bifidobacterium pseudolongum reduces triglycerides by modulating gut microbiota in mice fed high-fat food. J Steroid Biochem Mol Biol 198:105602
Bunyavanich S, Shen N, Grishin A, Wood R, Burks W, Dawson P (2016) Early-life gut microbiome composition and milk allergy resolution. J Allergy Clin Immunol 138:1122–1130
Campana R, Martinelli V, Scoglio S, Colombo E, Benedetti S, Baffone W (2017) Influence of Aphanizomenon flos-aquae and two of its extracts on growth ability and antimicrobial properties of Lactobacillus acidophilus DDS-1. LWT-Food Sci Technol 81:291–298
Carmichael WW, Drapeau C, Anderson DM (2000) Harvesting of Aphanizomenon flos-aquae Ralfs ex Born. & Flah. var. flos-aquae (Cyanobacteria) from Klamath Lake for human dietary use. J Appl Phycol 12:585–595
Chang S, Cui X, Guo M, Tian Y, Xu W, Huang K, Zhang Y (2017) Insoluble dietary fiber from pear pomace can prevent high-fat diet-induced obesity in rats mainly by improving the structure of the gut microbiota. J Microbiol Biotechnol 27:856–867
Cirés S, Ballot A (2016) A review of the phylogeny, ecology and toxin production of bloom-forming Aphanizomenon spp. and related species within the Nostocales (cyanobacteria). Harmful Algae 54:21–23
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, Ling AV, Devlin AS, Vama Y, Fishbach MA, Biddinger SB, Dutton RJ, Turnbaugh PJ (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505:559–563
Frazier K, Chang EB (2020) Intersection of the gut microbiome and circadian rhythms in metabolism. Trends Endocrinol Metab 31:25–36
Gao R, Zhu C, Li H, Yin M, Pan C, Huang L, Kong C, Wang X, Zhang Y, Qu S, Qin SH (2018) Dysbiosis signatures of gut microbiota along the sequence from healthy, young patients to those with overweight and obesity. Obesity 26:351–361
Genda T, Kondo T, Sugiura S, Hino S, Shimamoto S, Nakamura T, Ukita S, Morita T (2018) Bacterial fermentation of water-soluble cellulose acetate raises large-bowel acetate and propionate and decreases plasma cholesterol concentrations in rats. J Agric Food Chem 66:11909–11916
Hirota K, Inagaki R, Hamada R, Ishida K, Miyatake Y (2014) Evaluation of a rapid oral bacteria quantification system using dielectrophoresis and the impedance measurement. Biocontrol Sci 19:45–49
Hussain M, Bonilla-Rosso G, Chung CK, Bariswyl L, Rodriguez MP, Kim BS, Emgel P, Noti M (2019) High dietary fat intake induces a microbiota signature that promotes food allergy. J Clin Immunol 144:157–170
Kim B, Shin J, Guevarra RB, Lee JH, Kim DW, Seol K, Lee JH, Kim HB, Isaacson R (2017) Deciphering diversity indices for a better understanding of microbial communities. J Microbiol Biotechnol 27:2089–2093
Koliada A, Syzenko G, Moseiko V, Puchkov L, Budovska K, Perederiy V, Gavalko Y, Dorofeyev A, Romanenko M, Tkach S, Sineok L, Lushchak O, Vaiserman A (2017) Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol 17:120
Kuda T, Yazaki T, Ono M, Takahashi H, Kimura B (2013) In vitro cholesterol-lowering properties of Lactobacillus plantarum AN6 isolated from aji-narezushi. Lett Appl Microbiol 57:187–192
Kuda T, Masuko Y, Kawahara M, Kondo S, Nemoto M, Nakata T, Takahashi H, Kimura B (2016) Bile acid-lowering properties of Lactobacillus plantarum Sanriku–SU3 isolated from Japanese surfperch fish. Food Biosci 14:41–46
Kuda T, Yokota Y, Shikano A, Takei M, Takhashi H, Kimura B (2017) Dietary and lifestyle disease indices and caecal microbiota in high fat diet, dietary fibre free diet, or DSS induced IBD models in ICR mice. J Funct Foods 35:605–614
Kushak PI, Drapeau C, Van Cott EM, Winter HH (2000) Favorable effects of blue-green algae Aphanizomenon flos-aquae on rat plasma lipids. J Am Nutraceut Assoc 2:59–65
Lagkouvardos I, Lesker TR, Hitch TC, Gálvez EJC, Smit N, Neuhaus K, Wang J, Baines JF, Abt B, Stecher B, Overmann J, Strowig T, Clavel T (2019) Sequence and cultivation study of Muribaculaceae reveals novel species, host preference, and functional potential of this yet undescribed family. Microbiome 7:28
Li L, Wang Y, Zhu L, Liu Z, Ye C, Qin S (2020) Inulin with different degrees of polymerization protects against diet-induced endotoxemia and inflammation in association with gut microbiota regulation in mice. Sci Rep 10:978
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235
Lu Y, Fan C, Li P, Lu Y, Chang X, Qi K (2016) Short chain fatty acids prevent high-fat-diet-induced obesity in mice by regulating G protein coupled receptors and gut microbiota. Sci Rep 6:37589
Mangin I, Dossou-Yovo F, Lévêque C, Dessoy M, Sawoo O, Suau A, Pochart P (2018) Oral administration of viable Bifidobacterium pseudolongum strain Patronus modified colonic microbiota and increased mucus layer thickness in rat. FEMS Microbiol Ecol 94: fiy177
Mitsuoka T (2014) Establishment of intestinal bacteriology. Biosci Microbiota Food Health 33:99–116
Nagpal R, Wang S, Ahmadi S, Hayes J, Gagliano J, Subashchandrabose S, Kitzman DW, Becton T, Read R, Yadov H (2018) Human-origin probiotic cocktail increases short-chain fatty acid production via modulation of mice and human gut microbiome. Sci Rep 8:12649
Ni Y, Yang X, Zheng L, Wang Z, Wu L, Jiang J, Yang T, Ma L, Fu Z (2019) Lactobacillus and Bifidobacterium improves physiological function and cognitive ability in aged mice by the regulation of gut microbiota. Mol Nutr Food Res 63:1900603
Patterson E, Ryan PM, Wiley N, Carafa I, Sherwin E, Moloney G, Franciosi E, Mandal R, Wishart DS, Tuohy K, Ross RP, Cryan JF, Dinan TG, Stanton C (2019) Gamma-aminobutyric acid producing lactobacilli positively affect metabolism and depressive like behaviour in a mouse model of metabolic syndrome. Sci Rep 9:16323
Payling L, Fraser K, Loveday SM, Sims I, Roy N, McNabb W (2020) The effects of carbohydrate structure on the composition and functionality of the human gut microbiota. Trends Food Sci Technol 97:233–248
Pereira P, Onodera H, Andorinolo D, Franca S, Araújo F, Lagos N, Oshima Y (2000) Paralytic shellfish toxins in the freshwater cyanobacterium Aphanizomenon flos-aquae, isolated from Montargil reservoir. Portugal Toxicon 38:1689–1702
Pugh N, Pasco DS (2001) Characterization of human monocyte activation by a water soluble preparation of Aphanizomenon flos-aquae. Phytomedicine 8:445–453
Sharma NK, Tiwari SP, Tripathi K, Rai AK (2011) Sustainability and cyanobacteria (blue-green algae): facts and challenges. J Appl Phycol 23:105–1081
Shikano A, Kuda T, Shinayama J, Toyama A, Ishida Y, Takahashi H, Kimura B (2019) Effects of Lactobacillus plantarum Uruma-SU4 fermented green loofah on plasma lipid levels and gut microbiome of high-fat diet fed mice. Food Res Int 121:817–824
Shkoporov AN, Chaplin AV, Khokholova EK, Shcheribakova VA, Motuzova OV, Bozhenko VK, Kafarskaia LI, Efimov BA (2015) Alistipes inops sp. nov. and Coprobacter secundus sp. nov., isolated from human faeces. Int J Syst Evol Microbiol 65:4580–4588
Sinclair L, Osman OA, Bertilsson S, Eiler A (2015) Microbial community composition and diversity via 16S rRNA gene amplicons: evaluating the Illumina platform. PLoS One 10:e0116955
Takei M, Kuda T, Fukunaga M, Toyama A, Goto M, Takahashi H, Kimura B (2019) Effects of edible algae on caecal microbiomes of ICR mice fed a high-sucrose and low–dietary fibre diet. J Appl Phycol 31:3969–3978
Takei NM, Kuda T, Taniguchi M, Nakamura S, Takahashi H, Kimura B (2020) Detection and isolation of low molecular weight alginate- and laminaran susceptible gut indigenous bacteria from ICR mice. Carbohydr Polyomers 238:116205
Taniguchi M, Kuda T, Shibayama J, Sasaki T, Michihata T, Takahashi H, Kimura B (2019) In vitro antioxidant, anti-glycation and immunomodulation activities of fermented blue-green algae Aphanizomenon flos-aquae. Mol Biol Rep 46:1775–1786
Wang CS, Li WB, Wang HY, Ma YM, Zhao XH, Yang H, Qian JM, Li JN (2018) VSL#3 can prevent ulcerative colitis-associated carcinogenesis in mice. World J Gastroenterol 24:4254–4262
Williams BA, Mikkeisen D, Flanagan BM, Gidley MJ (2019) “Dietary fibre”: moving beyond the “soluble/insoluble” classification for monogastric nutrition, with an emphasis on humans and pigs. J Animal Sci Biotechnol 10:45
Xia Y, Kuda T, Toyama A, Goto M, Fukunaga M, Takahashi H, Kimura B (2019) Detection and isolation of bacteria affected by dietary cumin, coriander, turmeric, and red chili pepper in the caecum of ICR mice. J Funct Foods 61:103467
Xia Y, Fukunaga M, Kuda T, Goto M, Chiaraluce G, Hoshiba H, Takahashi H, Kimura B (2020) Detection and isolation of protein susceptible indigenous bacteria affected by dietary milk-casein, albumen and soy-protein in the caecum of ICR mice. J Biol Macromol 144:813–820
Yokota Y, Haraguchi Y, Shikano A, Kuda T, Takahashi H, Kimura B (2018) Induction of gut Lactobacillus reuteri in normal ICR mice by oral administration of L. plantarum AN1. J Food Biochem 42:e12589
Zhang L, Liu SY, Zhang J, Zhang JK, Hu C, Liu CX (2016) Respiratory toxicity of cyanobacterial aphantoxins from Aphanizomenon flos-aquae DC-1 in the zebrafish gill. Aquat Toxicol 176:106–115
Zhang C, Wu W, Li X, Xin X, Liu D (2019) Daily supplementation with fresh Angelica keiskei juice alleviates high-fat diet-induced obesity in mice by modulating gut microbiota composition. Mol Nutr Food Res 63:19000248
Zhao L, Lou H, Peng Y, Chen S, Zhang Y, Li X (2019) Comprehensive relationships between gut microbiome and faecal metabolome in individuals with type 2 diabetes and its complications. Endocrine 66:526–537
Acknowledgements
We would like to thank Editage (www.editage.com) for the English language editing.
Funding
This work was partially supported by the Towa Foundation for Food Science & Research, Tokyo, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Statements of animal rights
Animal experiments were performed in accordance with the Fundamental Guidelines for Proper Conduct of Animal Experiments and Related Activities in Academic Research Institutions, under the jurisdiction of the Ministry of Education, Culture, Sports, Science and Technology, Japan. The study protocol was approved by the Animal Experiment Committee of the Tokyo University of Marine Science and Technology (Approval No. H31-5).
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Taniguchi, M., Kuda, T., Takei, M. et al. Effects of fermented Aphanizomenon flos-aquae on the caecal microbiome of mice fed a high-sucrose and low-dietary fibre diet. J Appl Phycol 33, 397–407 (2021). https://doi.org/10.1007/s10811-020-02306-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-020-02306-x