Abstract
Landraces are considered as storehouses of valuable genetic diversity. Understanding the structure of this diversity and identification of distinct clusters with complementary traits is an important goal for a sustained and successful pearl millet breeding program. In this study a collection of 214 pearl millet accessions from different geographical regions of Sudan, and 11 accessions from West Africa, were assessed for their genetic diversity using 30 simple sequence repeat markers (SSRs) and evaluated for 15 agro-morphological traits. The diversity of the studied pearl millet landraces was demonstrated by a large number (400) of detected SSR alleles. The average polymorphic information content (PIC), gene diversity and observed heterozygosity of the 30 SSRs were 0.77, 0.82 and 0.72 respectively. A total of seven phylogenic groups with variable sizes were identified. Low correlation was observed between the agro-morphological matrix and the genetic matrix (r = 0.20). The average PIC values obtained across the seven linkage groups varied significantly. Weak genetic differentiations were observed among the geographical regions, suggesting a high seed and pollen-mediated gene flow among the regions. Analysis of molecular variance revealed that the variation of pearl millet accessions within the regions was much higher than among the regions. The large divergence observed among the landraces of this study is promising for developing new cultivars and for development of heterotic groups which can be used to develop population and hybrid varieties with higher degrees of heterozygosity and therefore hybrid vigor and stability, as well as to intensify yield production in the harsh production environments of Sudan.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pearl millet [(Pennisetum glaucum (L.) R. Br., [syn. Cenchrus americanus (L.) Morrone]) (2n = 2x = 14)] (Brunken 1977; Stapf and Hubbard 1934) is one of the most important staple food and fodder crop of millions of the poorest people living on the most marginal agricultural lands of sub-Saharan Africa and Asia. It is the preferred crop and main source of dietary energy and protein (Abdalla et al. 1998) for the majority of western Sudanese people. Harlan et al. (1975) and Oumar et al. (2008) suggested a defused belt stretching from Sudan to Senegal as the center of origin for pearl millet, the region in which the greatest genetic diversity is expected. The crop is mostly grown under harsh environmental conditions on infertile soils of low water-holding capacity, where other cereal crops generally fail (Manning et al. 2011). Because of the expected increase of these harsh conditions, the future suitability of pearl millet cultivation is expected to expand to additional geographic regions under various climate scenarios (Lane and Jarvis 2007; Waithaka et al. 2013).
Pearl millet exhibits a wide range of morphological and agronomical variability for several traits such as plant height, days to flowering, panicle length and width, grain yield, tolerance to biotic and abiotic stresses as well as nutritional value (Bhattacharjee et al. 2007; Haussmann et al. 2006; Stich et al. 2010; Bashir et al. 2014). Although there is a great expected genetic variability among Sudanese pearl millet landraces, the Sudanese pearl millet breeding program, founded in the early 1970s exploited a narrow gene pool to develop the currently used improved cultivars. Landraces possess valuable genetic variability and are often well adapted to the local soil type and climatic conditions (Khairwal et al. 2007). Assessing the extent of genetic variation among landrace accessions is essential to understand the pattern of diversity and evolutionary relationship among those accessions, to develop a sampling strategy for genetic resources in a more systematic fashion for subsequent conservation, and to efficiently use the diversity in crop improvement efforts (Thiyagu et al. 2011; Wu et al. 2007).
Genetic diversity assessment can be useful to identify contrasting parental materials to enhance heterozygosity or to optimize the genetic heterogeneity in a hybrid population and hence enhance yield stability in variable and changing climates (Haussmann et al. 2007, 2012). Development of high yielding and stable varieties requires the inclusion of diverse and adapted germplasm in breeding programs as sources of desirable genes and/or gene complexes.
Assessment of genetic diversity based on morphological characteristics might not provide an accurate classification of the genetic divergence among the genetic materials, due to the restricted number of morphological traits evaluated, environmental influences and development-specific trait expression. On the other hand, molecular markers have been proven to be a powerful tool for assessing the genetic variation and relatedness among cultivars of various crops (Hu et al. 2009; Wu et al. 2007). PCR-compatible markers based on co-dominant simple sequence repeats (SSRs) are often considered as one of the most appropriate and reliable markers in applied breeding programs. SSRs markers have been widely employed to assess the extent of genetic diversity among various crop species, such as sorghum (Ramu et al. 2013), maize (Reif et al. 2005), wheat (Fufa et al. 2005), barley (Naeem et al. 2011), cotton (Wu et al. 2007) and pearl millet (Stich et al. 2010; Budak et al. 2003; Kapila et al. 2008; Mariac et al. 2006; Nepolean et al. 2012). To date no attempts have been made to study the genetic diversity of Sudanese pearl millet using molecular markers, thus providing a basis for a more efficient use of genetic diversity in the narrow pearl millet breeding programs of Sudan.
Therefore, the objectives of this study were to
-
1.
determine the level of genetic diversity of Sudanese pearl millet germplasm,
-
2.
assess the regional distribution of genetic diversity within Sudan,
-
3.
define potential distinct groups among the studied pearl millet landraces; and
-
4.
examine the relationship between molecular and morphological diversity.
Materials and methods
Plant materials
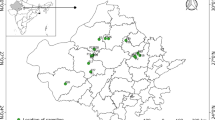
A set of 225 pearl millet accessions including 214 accessions of pearl millet landraces collected from different ecological zones of Western Sudan, and preserved at the Genetic Resource Unit at Agricultural Research Corporation (ARC), Sudan, and 11 landraces or improved cultivars from West Africa (WA, in our case encompassing Mali, Niger, Mauritania and Togo), provided by the International Crops Research Institute for the Semi-Arid Tropics (ICRISAT), Niger, were used in the present study (Table S1). The collection regions were classified to five different geographical regions, these are: North Darfur (ND), North Kordofan (NK), South Kordofan (SK), West Darfur (WD) and WA, and different sample sizes were collected from each (Bashir et al. 2014). Morphological diversity assessment (15 traits) was carried out under field conditions at three distinct locations (Wad Medani, El Fasher and El Obeid) in Sudan during the 2010 rainy season, where, the field experiments were arranged in 15 × 15 lattice designs with three replications. For more detailed information regarding the field trials please see Bashir et al. (2014). For DNA extraction seeds of each accession were sown in a tray in the greenhouse of University of Hohenheim in April 2010 for the genotyping study.
Genomic DNA extraction
From seedlings of each accession, leaf samples of 15 individuals at three leaves stage were collected separately and lyophilized. Genomic DNA was extracted from a bulk leaf tissues of 15 individual plants per accession using a modified CTAB protocol (Mace et al. 2003). The extracted DNA was evaluated for its quality using agarose gel electrophoresis, and spectrophotometry for DNA quantification. The final working DNA samples were maintained at a uniform concentration of 20 ng/µL.
Genotyping by SSR markers
The 225 pearl millet accessions were genotyped with a set of 30 SSR markers selected from a set of several previously used SSRs in pearl millet diversity (Kapila et al. 2008; Oumar et al. 2008) and bi-parental mapping studies (Qi et al. 2004; Yadav et al. 2004). They provide a reasonable coverage with at least three markers for each linkage group. The SSRs were grouped into multiplex sets of three, where forward primers were labeled with 6-FAM, HEX and TET fluorescent dyes (Biomers GmbH, Germany). For amplification conditions a touchdown PCR program was followed consisting of an initial denaturation cycle of 94 °C for 5 min; followed by 5 cycles at 94 °C for 30 s, 60 °C (−1 °C per cycle) for 30 s, elongation at 72 °C for 30 s; 39 cycles of 94 °C for 30 s, 52 °C for 30 s, elongation at 72 °C for 30 s; and a final 20 min elongation at 72 °C. 3 µl of each PCR product was multiplexed with 0.05 µl MegaBACE ET400-R Size Standards and 4.95 µl deionised H2O, centrifuged and denatured at 95 °C for 2 min, cooled in ice and then size-fractioned using capillary electrophoresis on a MegaBACE DNA analysis system (Amersham Biosciences, Sweden). In each of the 96-well PCR plates, two negative PCR controls were included. The MegaBACE Fragment Profiler v1.2 was used for fragment sizing and allele calling and then manually checked.
Analysis of diversity
The degree of polymorphism of each SSR locus was estimated based on the polymorphic information content (PIC), following Botstein et al. (1980):
where k is number of alleles, p i and p j are the frequency of the ith and the jth alleles respectively. Furthermore, we estimated for each locus the following descriptors of genetic diversity: total number of alleles (At), number of rare alleles (Ar, freq. <5 %), allelic richness (Arich) and private alleles (Apr). The observed heterozygosity (H0) and the expected heterozygosity (He) or gene diversity were calculated according to Nei (1987). To explore the genetic differentiation among the groups, F statistics (F ST ) were estimated according to Weir and Cockerham (1984). Pair-wise F ST was calculated according to (Nei 1973) using the Adegenet 1.4-0 R package (Jombart 2014). A Wilcoxon test was used to determine the significance differences between the groups for the different estimates.
Analysis of molecular variance and population structure
The pattern and degree of molecular genetic variation between and within the groups (geographical regions) was performed by analysis of molecular variance (AMOVA) according to Excoffier et al. (1992) using the R package pegas (Paradis 2010), with 1,000 permutations.
The genetic dissimilarity matrix based on Euclidean distance was calculated using Adegenet 1.4-0 R package (Jombart 2014) to identify the genetic relatedness among the collected pearl millet accessions. The illustration of the relationship structure among the five collection regions were obtained by principal component analysis (PCA), while an unrooted bootstrapped Neighbor Joining (NJ) tree was constructed to perform the individual relationship using ape R package (Paradis et al. 2004). Both PCA and NJ were performed based on the Euclidean distance matrix. A typology was also performed to display the diversity among the regions using an interclass PCA (Dolédec and Chessel 1987) implemented in Adegent 1.4-0 R package (Jombart 2014).
The software STRUCTURE 2.3.4 (Pritchard et al. 2000) was used to detect the population structure and assign the pearl millet accessions into subgroups. To identify the optimal value of K, the analysis was carried out by setting the burn-in Markov Chain Monte Carlo (MCMC) replication to 10,000 and data were collected over 100,000 MCMC replications in each run. With a model allowing for admixture and correlated allele frequencies (Falush et al. 2003, Evanno et al. 2005), which improves clustering for closely related populations, five independent runs were performed setting the number of population (K) from 2 to 10. The basis of this kind of clustering method is the allocation of individual genotypes to K clusters in such a way that Hardy–Weinberg equilibrium and linkage equilibrium are valid within clusters, whereas these kinds of equilibrium are absent between clusters (Vetriventhan et al. 2014). The final K value was determined based on the rate of change in mean log probability of the data [LnP(D)] between successive K, stability of grouping pattern across five runs and germplasm information of the examined material.
Molecular and morphological diversity relationship
Correlations between the Euclidean distance matrix computed from standardized morphological data, i.e. from adjusted entry means for 15 agro-morphological traits across three evaluation sites in Sudan (Bashir et al. 2014) and the distance matrix for the molecular marker were examined by performing a Mantel test (Mantel 1967) (1,000 permutation replicates, mantel.rtest function) using ADE4 package implemented in R Software 3.03 (R Development Core Team 2011).
Results
Overall genetic diversity
The 225 accessions differed significantly (P ≤ 0.01) for most agro-morphological traits (Bashir et al. 2014). Based on the 30 SSR loci distributed across the seven pearl millet chromosome pairs, a total of 400 alleles were detected among the 225 pearl millet accessions using bulked DNA samples. The number of alleles per locus ranged from 6 (Xpsmp2059) to 27 (Xpsmp2263) alleles with an average of 13.3 per locus, showing that the accessions exhibited a high level of genetic diversity (Table 1). All 30 SSR loci assessed were highly polymorphic, showing PIC values ranging from 0.40 (Xpsmp2059) to 0.91 (Xpsmp2255) with an average of 0.74. Gene diversity (He) varied from 0.41 (Xpsmp2059) to 0.92 (Xpsmp2255) with an average of 0.77. An average observed heterozygousity (H0) of 0.81 was obtained, with values ranging from 0.43 (Xpsmp2059) to 0.99 (Xpsmp2072). The number of rare alleles observed ranged from 1 (Xpsmp2059I, Xpsmp2202, Xpsmp2263 and Xpsmp2267) to 8 (Xpsmp2001 and Xpsmp2066), with an average of 3.5 (Table 1). The 30 SSR markers indicated significantly different average PIC values among the seven linkage groups with LG1 having the lowest value (PIC = 0.62) and LG7 showing the highest PIC (PIC = 0.75) (data not shown).
Regional genetic diversity
There were only slight differences for the different genetic diversity measures among the pearl millet accessions collected in the different geographic regions (Table 2). Among the 400 alleles detected, when correcting for the sample size for the collection regions in Sudan, the highest number of alleles (27 %) was observed among North Darfur accessions, while the lowest number (21 %), was detected among accessions of North Kordofan (Wilcoxon test, significant P ≤ 0.05). The average number of alleles per marker among the geographical regions ranged from 6.4 (WA) to 10.6 (North Darfur) (Table 2). No significant differences were observed among the regions for allele richness (Wilcoxon test, P value = 0.06), gene diversity (P = 0.06) and observed heterozygosity (P = 0.06). WA material showed a higher private allele richness compared to the Sudanese regions (p ≤ 0.05). Further, there was generally a marginally greater observed heterozygosity than expected heterozygosity (gene diversity) across all regions.
AMOVA and genetic structure of the Sudanese pearl millets
AMOVA revealed that most of the variation was observed within geographical regions and not among regions, as shown in sixfold larger sums of squares of the within region level (Table S1). The small portion of variation observed among the geographical regions was mostly attributed to differences between the materials from Sudan and those from WA. High average genetic similarity was observed within the regions when comparing the different regions according to their estimated genetic similarity matrix (Table 3). Pairwise F ST comparisons among the different geographical regions showed that F ST values varied from 0.004 to 0.037, with the highest values obtained between the four Sudanese regions and WA (Table 3).
Even though it can be biologically sound, the assumption of a fixed number of populations is arbitrary. The structure analysis revealed that the number of populations can be assumed to be three (K = 3) based on the rate of change in LnP(D) (Fig. S2). The analysis also revealed that there was an increase of likelihood at K = 5, however, the results are similar to K = 3. The graphical bar plot generated by STRUCTURE showed clear admixture among the identified sub-groups (Fig. S3).
Principal component and phylogeny analyses
The first two principal components explained only 10.54 and 8.55 % of the total genetic variation, respectively (Fig. 1). No clear structure pattern was observed among the accessions of the different regions. Accessions from West Darfur were located in the upper part of the PCA biplot. A typology displaying the diversity among the regions using an interclass PCA (Dolédec and Chessel 1987), showed that the different Sudanese regions were genetically very similar (Fig. 2). The WA accessions were slightly different, but showed still a strong intermixture with the Sudanese accessions. A neighbor joining tree based on the genetic distances of the 225 pearl millet accessions revealed a similar picture with no clear separation of geographical regions and a probable high level of admixture (Fig. 3; Fig. S3). The correlation between the Euclidean distance matrix for molecular marker data and similarity matrix for 15 agro-morphological traits (Bashir et al. 2014) as revealed by the Mantel matrix correspondence test was low (r = 0.20, P < 0.05).
Discussion
Genetic diversity of pearl millet in Sudan
Efficient use of locally adapted diversity is a major requirement to enhance selection gains in pearl millet breeding in Sudan. To our knowledge, this is a first study describing the diversity of a large number of Sudanese pearl millet landraces using microsatellite markers. Characterization of germplasm collections can be done using morphological descriptors and/or molecular markers. Morphological descriptors are reliable and relatively simple and easy to study (Beyene et al. 2005), but also show some limitations compared to molecular markers, such as limited polymorphism and environmental influence on the phenotype which affects the evaluation process (Ferreira 2006). In the case of very large collections, the phenotypic approach has relatively high costs and includes the challenges of large-scale experimental trials (Korir et al. 2013; Ferreira 2006). Although there were significant variations observed among the studied pearl millet accessions for most of the agro-morphological traits assessed, the variation for some traits such as flowering time, plant height and panicle length was relatively limited compared to global or other West African pearl millet collections (Bashir et al. 2014). Despite this limitation, agro-morphological traits are useful for preliminary evaluation because they relate to functional diversity and enable selection of germplasm with contrasting, complementary traits. The 30 SSR markers used in this study enable conclusions about the overall gene diversity, polymorphism and number of alleles observed in the studied Sudanese pearl millet accessions, but do not relate explicitly to functional diversity and specific traits. Ultimately, the combined use of agro-morphological and molecular marker data is required to enable a comprehensive diversity assessment and informed diversity-based decision making in germplasm selection.
The average gene diversity (H e = 0.77) observed in this study among the 225 accessions was higher than previously reported for other pearl millet studies (Nepolean et al. 2012 (H e = 0.62), Stich et al. 2010 (H e = 0.74) and Mariac et al. 2006 (H e = 0.49)). These differences might be due to the fact that the first two studies used inbreds, while we used landraces, which generally show a higher diversity and heterozygosity level, whereas Mariac et al. (2006) considered a single plant per accession. Furthermore, the mentioned studies used DNA derived from individual plants, whereas we used DNA from bulk samples and the di-nucleotide repeat SSR markers we used tend to be more polymorphic.
In agreement with the higher gene diversity observed in this study, the mean observed heterozygozity and the average number of alleles per marker were considerably higher than previously reported in several studies (Mariac et al. 2006; Kapila et al. 2008; Nepolean et al. 2012; Oumar et al. 2008). This points to a broad genetic base of pearl millet landraces in Sudan which then can be considered as a promising source of variation for improving pearl millet of Sudan, by widening the genetic base of the current breeding material using the available locally adapted diversity in a strategic manner. Moreover, through a larger genetic diversity plant breeders can reach a higher response to selection with the same selection intensity. Higher genetic diversity is favorable for genetic marker development, construction of segregating populations, functional gene cloning and association mapping and provides enriched gene resources for gene mining in the grass family (Wang et al. 2012).
Regional genetic diversity
The AMOVA revealed that most of the differentiation was present within regions and not among regions. These findings were confirmed by the high gene diversity, observed heterozygosity and average allelic number per locus observed within the regions. Moreover, the low level of differentiation observed by the PCA and the neighbor-joining tree confirmed the previous observation. The lack of regional differentiation can be attributed to the biological characteristic of pearl millet as highly allogamous crop with out-crossing rates exceeding 85 % (Stich et al. 2010; Vom Brocke et al. 2003). The high out-crossing rate, as indicated by the high level of heterozygosity, increases the admixture level among the regions; this was clearly revealed by the STRUCTURE analysis. Within the geographical zones sampled in this study, the environmental conditions are highly variable both spatially and temporally, resulting in high genetic variation among the pearl millet landraces. Moreover, the overlapping between the early and late flowering landraces within the growing regions decreases the differentiation between the regions. Consistent with this interpretation, the AMOVA analysis and pairwise F ST comparisons also confirmed absence of significant variation among the different geographical regions of Sudan, indicating that most of the accessions of the different regions share high levels of allelic exchange (Table 3). The lacking differentiation among regions was probably not only due to the influence of a high rate of outcrossing and concomitant high rate of pollen-mediated gene flow, but might be also due to a high frequency of seed exchange of cultivated landraces among the farmers across the Sudanese regions. The slight differentiation of the WA accessions compared to the Sudanese accessions strengthens this argument, because little or no recent seed exchange among farmers from WA and Sudan can be expected to have occurred. Previous studies assessing the diversity of pearl millet showed similar results, pointing to a high within population diversity most likely caused by pollen-mediated gene flow and/or by seed-mediated gene flow (Mariac et al. 2006; Vom Brocke et al. 2003). Moreover, farmers in pearl millet growing regions of Sudan generally grow several landraces in one field during the growing season, in order to increase the buffering capacity and prevent complete crop losses due to diseases, pests or abiotic stresses. Similar observations were reported for WA by Stich et al. (2010), who attributed this to farmers’ strategies to overcoming the famine prone period at the beginning of the rainy season. By cultivating early maturing landraces farmers can achieve grain yields at a time period when food shortages might occur. The late maturing landraces possess generally a higher yield potential, especially in years without any terminal droughts, hence they serve as the major grain producers. The high similarity among regions revealed by molecular marker analysis can be also attributed to the high inter-annual climate variability within the agro-ecological zones, which leads to divergent selection pressures in different years and prevents detectable genetic differentiation between the landraces (Stich et al. 2010).
As revealed by molecular marker analysis, the patterns of genetic diversity among the geographical regions were overlapping. However, the number of the sub-groups shown by the neighbor joining tree (Fig. 3) in the present study was higher than previously reported on pearl millet in several studies (Stich et al. 2010; Tostain 1994). Since the effectiveness of clustering is influenced by the number of markers, sample size and the differentiation among the populations (Rosenberg et al. 2005), we suggest that the higher rate of clustering in our study is due to the larger number of pearl millet accessions evaluated and due to the more informative SSRs we used. Moreover, we could explain more of the genetic variation based on the first two PCAs compared to WA material (Stich et al. 2010). These findings show that the pearl millet accessions examined possess a high level of genetic diversity and are suitable for improving pearl millet breeding in Sudan.
Implications of the diversity observed in pearl millet breeding
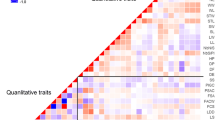
The dendrogram constructed using agro-morphological traits of the same pearl millet accessions resulted in the establishment of seven distinct clusters of accessions with complementary traits (Bashir et al. 2014). These results suggest the existence of variation among the studied materials for both quantitative and qualitative traits. Similarly, clustering patterns based on SSR markers in this study revealed a clear distinction among the genotypes and formed seven distinct sub-groups. The differentiation among the studied pearl millet germplasm based on the molecular markers as presented by PCA (Fig. 1), explained less variation (19 %) than did the variation observed in PCA based on agro-morphological traits (46 %, Bashir et al. 2014). Despite the similarities of the clustering patterns (seven groups with both agro-morphological and marker data) mentioned above, the correlation observed between the agro-morphological and SSR markers based on their distance matrices was very low. This could be explained by the fact that molecular SSR markers are mostly neutral markers, cover a larger proportion of the genome and are less influenced by the environment, while morphological traits are controlled by a relatively small number of specific gene loci (Bruschi et al. 2003; Zhang et al. 2010). Nevertheless, some genotypes appeared together in the same group with both agro-morphological and molecular diversity assessment methods, reflecting both genetic and morphological similarity between these genotypes. For example, G210 (NDF_08019), G213 (NDF_08023), G145 (HSD_6607) and G146 (HSD_6616) grouped together in both morphological diversity (group VII) and genetic diversity analysis (group IV) and were clearly distinct from G100 (HSD_3525), G108 (HSD_3533), G182 (HSD_7226), G223 and (GB8735xMoro) and G224 (B9_Tabi) who equally grouped together in both analysis types (groups III and VII in morphological and genetic diversity analyses, respectively). Such genetically and morphologically distinct genotype groups could serve as initial reference in the development of the heterotic groups. Overall, the results obtained from agro-morphological characterization (Bashir et al. 2014) and molecular markers, indicated clearly substantial differentiation among the studied pearl millet accessions, and therefore provide a good basis for further pearl millet breeding in Sudan.
Based on genetic diversity assessments it is possible to initially assign genotypes to specific heterotic pools in a hybrid breeding program (Xie et al. 2014). Heterotic pools for crops such as maize, sorghum and rye have been developed over a long period of time, are therefore genetically distinct from each other, hence genotype assignment to one group using molecular markers is a rather straight forward procedure (Ben-Israel et al. 2012; Fischer et al. 2010; Reif et al. 2005). In pearl millet in Sudan, there are mostly no clear heterotic patterns documented, hence there is the necessity to define initial heterotic pools for an optimal hybrid breeding program. Therefore, crosses among members of different clusters in this study and analysis of combining ability patterns and hybrid performance might be promising for establishing different heterotic pools.
The heterotic deviation is expected to increase with an increase in genetic distance between the parents; i.e. the expected level of heterosis of crosses between related lines is smaller compared to the crosses between unrelated lines. However, this increase holds only up to a certain optimum level of genetic distance, and subsequently (in the case of very wide crosses) the hybrid performance starts to decrease (Melchinger and Gumber 1998), because of increasingly reduced genome integrity and negative epistatic effects.
Estimated correlations between marker-based diversity measures and the level of heterosis in inter-pool crosses are not always very tight (Zhang et al. 1995; Xiao et al. 1996; Lanza et al. 1997; Melchinger 1999; Benchimol et al. 2000; Jordan et al. 2003; Zhao et al. 2010; Xie et al. 2014). This can be related to the fact that most of the markers used in calculation of the genetic distance are neutral markers and are not linked to yield, its components or other functions that are important for heterosis and hybrid vigor (Jordan et al. 2003). In hybrid rice, Zhang et al. (1995) found that the use of a subset of markers linked to yield predict hybrid performance better than a large number of random markers. Similar results were reported by Jordan et al. (2003) for sorghum. Jordan et al. (2003) concluded that a model combining phenotypic trait data and parental diversity on particular linkage groups relevant to performance traits is required to predict hybrid yield and appropriately select hybrid parents. The same applies to the definition of heterotic groups. However the formation of initial heterotic groups (as required for Sudanese pearl millets) based on genetic diversity as revealed by molecular markers reveals a practical value, because genetic diversity between the hybrid parents is a major prerequisite to enhance heterozygosity and therefore the potential hybrid vigor, hence increasing opportunities to develop performing hybrids. Also Ferreira et al. (1995) stated that heterosis and the combining ability of parents depend directly on the genetic diversity between them, and that the chance of finding promising combinations is better when more divergent materials are used. The initial heterotic groups based on genetic diversity must then be validated using combining ability studies.
Conclusions
The marker-based analysis implemented in this study clearly identified substantial genetic diversity existing among Sudanese pearl millet accessions. This indicates that these assessed materials are useful sources for future pearl millet improvement programs. Crosses involving parental materials with larger genetic distance or belonging to different sub-groups and/or with introduced pearl millet genotypes from West Africa would be useful to identify and validate initial heterotic groups, which could be used both in pearl millet population and hybrid breeding to improve adaptation, yield stability as well as to intensify yield in the harsh production environments of Sudan. The weak genetic differentiation observed between geographical regions point to the importance of pollen-mediated gene flow, divergent selection pressures due to high spatial and temporal environmental variability and human factors in shaping the genetic structure of pearl millet landraces.
References
Abdalla AA, El Tinay AH, Mohamed BE, Abdalla AH (1998) Effect of traditional processes on phytate and mineral content of pearl millet. Food Chem 63:79–84. doi:10.1016/S0308-8146(97)00194-5
Bashir EMA, Ali Abdelbagi M, Ali Adam M, Melchinger AE, Parzies HK, Haussmann BIG (2014) Characterization of Sudanese pearl millet germplasm for agro-morphological traits and grain nutritional values. Plant Genet Resour 12:35–47. doi:10.1017/S1479262113000233
Benchimol LL, Souza CL Jr, Garcia AAF, Kono PMS, Mangolim CA, Barbosa AMM, Coelho ASG, Souza AP (2000) Genetic diversity in tropical maize inbred lines: heterotic group assignment and hybrid performance determined by RFLP marker. Plant Breed 119(6):491–496. doi:10.1046/j.1439-0523.2000.00539.x
Ben-Israel I, Kilian B, Nida H, Fridman E (2012) Heterotic trait locus (HTL) mapping identifies intra-locus interactions that underlie reproductive hybrid vigor in Sorghum bicolor. PLoS One 7:e38993. doi:10.1371/journal.pone.0038993
Beyene Y, Botha AM, Myburg AA (2005) A comparative study of molecular and morphological methods of describing genetic relationships in traditional Ethiopian highland maize. Afr J Biotechnol 4(7):586–595
Bhattacharjee R, Khairwal IS, Bramel PJ, Reddy KN (2007) Establishment of a pearl millet [Pennisetum glaucum (L.) R. Br.] core collection based on geographical distribution and quantitative traits. Euphytica 155:35–45. doi:10.1007/s10681-006-9298-x
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Brunken JN (1977) A systematic study of Pennisetum sect. Pennisetum (Gramineae). Am J Bot 64(2):161–176. doi:10.2307/2442104
Bruschi P, Vendramin GG, Bussotti F, Grossoni P (2003) Morphological and molecular diversity among Italian populations of Quercus petraea (Fagaceae). Ann Bot 91:707–716. doi:10.1093/aob/mcg075
Budak H, Pedraza F, Cregan PB, Baenziger PS, Dweikat I (2003) Development and utilization of SSRs to estimate the degree of genetic relationships in a collection of pearl millet germplasm. Crop Sci 43:2284–2290. doi:10.2135/cropsci2003.2284
Dolédec S, Chessel D (1987) Rythmes saisonniers et composantes stationnelles en milieu aquatique. I. description d’un plan d’observation complet par projection de variables. Acta Oecol Oecol Gen 8:403–426
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Ferreira ME (2006) Molecular analysis of gene banks for sustainable conservation and increased use of crop genetic resources. In: Ruane J, Sonnino A (eds) The role of biotechnology in exploring and protecting agricultural genetic resources. FAO, Rome, pp 121–127
Ferreira DF, Oliveira AC, Santos MXD, Ramalho MAP (1995) Methods to evaluate the genetic maize divergence and its relation with the diallelic crossings. Pesq Agropec Bras, Brasília 30:1189–1194
Fischer S, Melchinger AE, Korzun V, Wilde P, Schmiedchen B, Möhring J, Piepho H-P, Dhillon BS, Würschum T, Reif JC (2010) Molecular marker assisted broadening of the Central European heterotic groups in rye with Eastern European germplasm. Theor Appl Genet 120:291–299. doi:10.1007/s00122-009-1124-0
Fufa H, Baenziger PS, Beecher BS, Dweikat I, Graybosch RA, Eskridge KM (2005) Comparison of phenotypic and molecular marker-based classifications of hard red winter wheat cultivars. Euphytica 145:133–146. doi:10.1007/s10681-005-0626-3
Harlan JR, de Wet JMJ, Stemler ABL (1975) Plant domestication and indigenous African Agriculture. In: Harlan JR, de Wet JMJ, Stemler ABL (eds) Origin of African domestication, Mouton Publishers, The Hague, The Netherlands, pp 3–20
Haussmann BIG, Boubacar A, Boureima SS, Vigouroux Y (2006) Multiplication and preliminary characterization of West and Central African pearl millet landraces. Int Sorghum Millets Newsl 47:110–112
Haussmann BIG, Boureima SS, Kassari IA, Moumouni KH, Boubacar A (2007) Mechanisms of adaptation to climate variability in West African pearl millet landraces—a preliminary assessment. J SAT Agric Res 3. http://oar.icrisat.org/2624/1/Mechanisms_of_adaptation.pdf
Haussmann BIG, Fred Rattunde H, Weltzien-Rattunde E, Traoré PSC, vom Brocke K, Parzies HK (2012) Breeding strategies for adaptation of pearl millet and sorghum to climate variability and change in West Africa. J Agron Crop Sci 198:327–339. doi:10.1111/j.1439-037X.2012.00526.x
Hu X, Wang J, Lu P, Zhang H (2009) Assessment of genetic diversity in broomcorn millet (Panicum miliaceum L.) using SSR markers. J Genet Genomics 36:491–500
Jombart T (2014) An introduction to adegenet 1.4-0. http://adegenet.r-forge.r-project.org/files/tutorial-basics.pdf
Jordan DR, Tao Y, Godwin ID, Henzell RG, Cooper M, McIntyre CL (2003) Prediction of hybrid performance in grain sorghum using RFLP markers. Theor Appl Genet 106:559–567. doi:10.1007/s00122-002-1144-5
Kapila RK, Yadav RS, Plaha P, Rai KN, Yadav OP, Hash CT, Howarth CJ (2008) Genetic diversity among pearl millet maintainers using microsatellite markers. Plant Breed 127:33–37. doi:10.1111/j.1439-0523.2007.01433.x
Khairwal IS, Yadav SK, Rai KN, Upadhyaya HD, Kachhawa D, Nirwan B, Bhattacharjee R, Rajpurohit BS, Dangaria CJ, Srikant S (2007) Evaluation and identification of promising pearl millet germplasm for grain and fodder traits [WWW document]. J SAT Agric Res. http://oar.icrisat.org/2628/. Accessed 26 July 2012
Korir NK, Han J, Shangguan L, Wang C, Kayesh E, Zhang Y, Fang J (2013) Plant variety and cultivar identification: advances and prospects. Crit Rev Biotechnol 33:111–125. doi:10.3109/07388551.2012.675314
Lane A, Jarvis A (2007) Changes in climate will modify the geography of crop suitability: agricultural biodiversity can help with adaptation. J SAT Agric Res 4:1–12. http://www.icrisat.org/journal/SpecialProject/sp2.pdf
Lanza LLB, Souza CL Jr, Ottoboni LMM, Vieira MLC, de Souza AP (1997) Genetic distance of inbred lines and prediction of maize single-cross performance using RAPD markers. Theor Appl Genet 94:1023–1030. doi:10.1007/s001220050510
Mace ES, Buhariwalla KK, Buhariwalla HK, Crouch JH (2003) A high-throughput DNA extraction protocol for tropical molecular breeding programs. Plant Mol Biol Rep 21:459–460. doi:10.1007/BF02772596
Manning K, Pelling R, Higham T, Schwenniger J-L, Fuller DQ (2011) 4500-year old domesticated pearl millet (Pennisetum glaucum) from the Tilemsi Valley, Mali: new insights into an alternative cereal domestication pathway. J Archaeol Sci 38:312–322. doi:10.1016/j.jas.2010.09.007
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Mariac C, Robert T, Allinne C, Remigereau M, Luxereau A, Tidjani M, Seyni O, Bezancon G, Pham J, Sarr A (2006) Genetic diversity and gene flow among pearl millet crop/weed complex: a case study. Theor Appl Genet 113:1003–1014. doi:10.1007/s00122-006-0360-9
Melchinger AE (1999) Genetic diversity and heterosis. In: Coors JG, Pandey S (eds) The genetics and exploitation of heterosis in crops. ASA, CSSA, and SSSA, Madison, WI, pp 99–118
Melchinger AE, Gumber RK (1998) Overview of heterosis and heterotic groups in agronomic crops. In: Larnkey KR, Staub JE (ed) Concepts and breeding of heterosis in crop plants. CSSA Special Publication 25, pp 29–44
Naeem R, Dahleen L, Mirza B (2011) Genetic differentiation and geographical relationship of Asian barley landraces using SSRs. Genet Mol Biol 34:268–273. doi:10.1590/S1415-47572011005000014
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci (USA) 70:3321–3323
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York, NY
Nepolean T, Gupta SK, Dwivedi SL, Bhattacharjee R, Rai KN, Hash CT (2012) Genetic diversity in maintainer and restorer lines of pearl millet. Crop Sci 52:2555–2563. doi:10.2135/cropsci2011.11.0597
Oumar I, Mariac C, Pham J, Vigouroux Y (2008) Phylogeny and origin of pearl millet (Pennisetum glaucum [L.] R. Br.) as revealed by microsatellite loci. Theor Appl Genet 117:489–497
Paradis E (2010) pegas: an R package for population genetics with an integrated–modular approach. Bioinformatics 26:419–420. doi:10.1093/bioinformatics/btp696
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20:289–290
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qi X, Pittaway TS, Lindup S, Liu H, Waterman E, Padi FK, Hash CT, Zhu J, Gale MD, Devos KM (2004) An integrated genetic map and a new set of simple sequence repeat markers for pearl millet, [Pennisetum glaucum (L.) R. Br.]. Theor Appl Genet 109:1485–1493
Ramu P, Billot C, Rami J-F, Senthilvel S, Upadhyaya HD, Reddy LA, Hash CT (2013) Assessment of genetic diversity in the sorghum reference set using EST-SSR markers. Theor Appl Genet 126:2051–2064. doi:10.1007/s00122-013-2117-6
R Development Core Team (2011) R: A language and environment for statistical computing. the R Foundation for Statistical Computing: Vienna, Austria. http://www.R-project.org/
Reif JC, Hamrit S, Heckenberger M, Schipprack W, Peter Maurer H, Bohn M, Melchinger AE (2005) Genetic structure and diversity of European flint maize populations determined with SSR analyses of individuals and bulks. Theor Appl Genet 111:906–913
Rosenberg NA, Mahajan S, Ramachandran S, Zhao C, Pritchard JK, Feldman MW (2005) Clines, clusters, and the effect of study design on the inference of human population structure. PLoS Genet 1:e70. doi:10.1371/journal.pgen.0010070
Stapf O, Hubbard CE (1934) Pennisetum. In: D. Prain (Ed.) The flora of tropical Africa. Reeve & Co. Ltd. Ashfnrd. Kent. 9:954–1070
Stich B, Haussmann BIG, Raj P, Sankalp B, Hash CT, Melchinger AE, Parzies HK (2010) Patterns of molecular and phenotypic diversity in pearl millet [Pennisetum glaucum (L.) R. Br.] from West and Central Africa and their relation to geographical and environmental parameters. BMC Plant Biol 10:1–10
Thiyagu K, Narayanan Manikanda Boopathi N, Nadarajan AG, Selvakumar P, Magadum S, Ravikesavan R (2011) Sampling and exploitation of genetic variation exist in locally adapted accessions using phenotypic and molecular markers for genetic improvement of cotton. Geneconserve 10:129–153
Tostain S (1994) Isozymic classification of pearl millet (Pennisetum glaucum, Poaceae) landraces from Niger (West Africa). Plant Syst Evol 193:81–93
Vetriventhan M, Upadhyaya HD, Anandakumar CR, Senthilvel S, Varshney RK, Parzies HK (2014) Population structure and linkage disequilibrium of ICRISAT foxtail millet (Setaria italica (L.) P. Beauv.) core collection. Euphytica 196:423–435. doi:10.1007/s10681-013-1044-6
Vom Brocke K, Christinck A, Weltzien RE, Presterl T, Geiger HH (2003) Farmers’ seed systems and management practices determine pearl millet genetic diversity patterns in semiarid regions of India. Crop Sci 43:1680–1689
Waithaka M, Nelson GC, Thomas TS, Kyotalimye M (2013) East African agriculture and climate change: a comprehensive analysis. International Food Policy Research Institute (IFPRI), Washington DC, USA. doi:10.2499/9780896292055
Wang C, Jia G, Zhi H, Niu Z, Chai Y, Li W, Wang Y, Li H, Lu P, Zhao B, Diao X (2012) Genetic diversity and population structure of chinese foxtail millet [Setaria italica (L.) Beauv.] landraces. G3 Genes Genomes. Genetics 2:769–777. doi:10.1534/g3.112.002907
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wu Y-X, Daud MK, Chen L, Zhu S-J (2007) Phylogenetic diversity and relationship among Gossypium germplasm using SSRs markers. Plant Syst Evol 268:199–208. doi:10.1007/s00606-007-0565-7
Xiao J, Li J, Yuan L, Mc Couch SR, Tanksley SD (1996) Genetic diversity and its relationship to hybrid performance and heterosis in rice as revealed by PCR-based markers. Theor Appl Genet 92:637–643. doi:10.1007/BF00226083
Xie F, He Z, Esguerra MQ, Qiu F, Ramanathan V (2014) Determination of heterotic groups for tropical Indica hybrid rice germplasm. Theor Appl Genet 127:407–417. doi:10.1007/s00122-013-2227-1
Yadav RS, Hash CT, Bidinger FR, Devos KM, Howarth CJ (2004) Genomic regions associated with grain yield and aspects of post-flowering drought tolerance in pearl millet across stress environments and tester background. Euphytica 136:265–277. doi:10.1023/B:EUPH.0000032711.34599.3a
Zhang QF, Gao YJ, Yang SH, Saghai Maroof MA, Li JX (1995) Molecular divergence and hybrid performance in rice. Mol Breed 1:133–142
Zhang X, Zhang Y, Yan R, Han J, Fuzeng-Hong F, Cao K (2010) Genetic variation of white clover (Trifolium repens L.) collections from China detected by morphological traits. RAPD and SSR. Afr J Biotechnol 9:3032–3041
Zhao K, Wright M, Kimball J, Eizenga G, McClung A, Kovach M, Tyagi W, Ali ML, Tung C-W, Reynolds A, Bustamante CD, McCouch SR (2010) Genomic diversity and introgression in O. sativa reveal the impact of domestication and breeding on the rice genome. PLoS One 5:e10780
Acknowledgments
We acknowledge financial support for this research by the Global Crop Diversity Trust (project GS10016), the German Academic Exchange Service (DAAD, PhD scholarship to E.M.A. Bashir), and the McKnight Foundation Collaborative Crop Research Program (Support to B.I.G. Haussmann). The authors are indebted to the Genetic Resource Unit, ARC and ICRISAT, Niger for providing the genetic materials. The ultimate support from C. Tom Hash throughout the research steps is truly appreciated. We thank Sabine Böger for technical assistance in the laboratory. The statistical analysis advice from Tobias Schrag and Fabian Freund and valuable comments on the manuscript by Willmar Leiser are highly appreciated.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bashir, E.M.A., Ali, A.M., Ali, A.M. et al. Genetic diversity of Sudanese pearl millet (Pennisetum glaucum (L.) R. Br.) landraces as revealed by SSR markers, and relationship between genetic and agro-morphological diversity. Genet Resour Crop Evol 62, 579–591 (2015). https://doi.org/10.1007/s10722-014-0183-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-014-0183-5