Abstract
ICRISAT conserves a large collection of pearl millet [Pennisetum glaucum L. R. (Br.)] comprising of 21,392 accessions. This includes landraces, cultivars, genetic stocks, breeding lines, and wild relatives from 50 countries. However, only a small fraction of this huge collection has been exhaustively used in the pearl millet improvement program. The objective of our research was to develop a core collection of pearl millet to enhance utilization of genetic resources in improvement programs and simplify their management. For this purpose, accessions were initially stratified according to geographical distribution followed by hierarchical clustering on 11 quantitative traits using Ward’s method. This resulted in 25 distinct groups. Approximately 10% accessions were then randomly selected from each of these 25 distinct groups to form a core collection of 1,600 accessions. Different statistical methods like comparison of mean using Newman–Keuls test, variance using Levene’s test, frequency distributions using Chi-square test, and Wilcoxon’s rank-sum non-parametric test for the traits validated that the variation present in entire collection had been preserved in the core collection. The important phenotypic correlations among different traits that may be under the control of co-adapted gene complexes were also preserved in the core collection. The diversity represented in the core collection will therefore, be a guideline to breeders for a wider use of the pearl millet genetic resources available in the genebank.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pearl millet (Pennisetum glaucum (L) R. Br.) is the sixth most important cereal, primarily grown for grain production in the arid and semi-arid tropical areas of Africa and Asia (Khairwal et al. 1999). It has been used as a cereal crop for nearly 3,000 years in Africa and parts of Near East, and is grown in over 40 countries, predominantly in Asia and Africa. It is cultivated in 26 million ha in many countries of Africa namely Senegal, Mali, Burkina Faso, Niger, Nigeria, Chad, Sudan, and a few countries of Asia, particularly India. It is also grown in some parts of America and Australia, mainly as a forage and/or mulch component of minimum tillage-based cropping systems (FAOSTAT 2005). India is the largest producer of this crop, both in terms of area (9.1 m ha) and production (7.3 m t), with an average productivity of 780 kg ha−1 during the last 5 years. Nutritionally, pearl millet is a richer source of protein, calcium, phosphorus and iron in comparison to other important cereals like sorghum, maize, rice and wheat (Khairwal et al. 1999).
Being allogamous in nature, pearl millet accessions are highly heterogeneous reflecting high variability within and among the accessions. Protogyny and time lag between stigma emergence and anther dehiscence favor complete cross-pollination leading to greatest morphological diversity. The plant features include a diverse range of plant height, time to flowering, tillering, stem thickness, fodder and grain quality, high growth rate, and adaptability to varied agro-ecological environments. Therefore, this wide variability maintained by the landraces can be utilized for further improvement of the crop in enhancing the genetic potential for yield and also in alleviating the biotic and abiotic stresses.
International germplasm collections play a very important role in securing genetic diversity and promoting its use. Therefore, in last few decades, emphasis has been given for preserving crop germplasm that resulted in assemblage of large collections in national and international genebanks. Similarly, the germplasm collection at the International Crop Research Institute for the Semi-Arid Tropics (ICRISAT) has in excess of 21,000 accessions of pearl millet. This germplasm is the world’s most genetically diverse collection of pearl millet from 50 countries. The collection includes landraces collected from most of the pearl millet growing eco-systems and considerable number of wild relatives of the genus Pennisetum (750 accessions). However, the available diversity has not been adequately utilized in pearl millet improvement. The huge size of the germplasm collection has hindered the increased utilization of this diversity due to lack of proper evaluation data. Low use of germplasm has been also reported in other crops like wheat (Dalrymple 1986); spring barley (Vellve 1992); maize (Dowswell et al. 1996); groundnut (Jiang and Duan 1998); chickpea and pigeonpea (Shiv Kumar et al. 2004); and chickpea (Upadhyaya et al. 2006).
Like any other major crop, there are gaps in pearl millet collection and efforts are being made to collect or assemble new germplasm. In last few years, additional accessions have been acquired from six countries and at present ICRISAT gene bank hold 21,594 accessions (20,844 cultivated and 750 wild). The germplasm collections were acquired at different times and were characterized or evaluated as and when these became available. This resulted in characterization of the collection in batches of 1,000–2,000 accessions at a time for the morphological (qualitative and quantitative) traits. However, for practical plant breeding research, evaluation often requires replicated trials and agronomic traits often display genotype × environment (G × E) interaction. This necessitates reduction of entire collection to a manageable size that can be easily evaluated to generate good data and enhance utilization. Recognizing this, Frankel (1984) suggested the core collection approach for enhancing the management of a large germplasm collection, facilitating the use and study of the conserved germplasm. This implies that collection would be pruned to a manageable sample, called core collection, representing the rich genetic diversity of the crop with minimum redundancy within the sample. The core collection could further serve as a working collection and also as a guide for efficient utilization of the entire collection (Tohme et al. 1995; Brown 1989b). This core collection could be extensively evaluated, and the accessions that are not included in the core collection would be designated as reserve collection (Frankel 1984).
Frankel and Brown (1984) and Brown (1989a, b) suggested that a core collection could be established using information on the country of origin and morpho-agronomic characteristics of the accessions. Further, Brown (1989a) suggested that the issues to be considered while developing the core are the size, the sampling strategy, the grouping within the collection, and the number of accessions to be included in the core from each group. Using the sampling theory of selectively neutral alleles, Brown (1989a) therefore, suggested that about 10% sample size of the entire collection with an upper limit of 3,000 per species would effectively retain about 70% of the alleles in the sample. However, Crossa (1989) suggested a slightly different approach for cross-pollinated crops. They proposed the use of probability models and determined optimal sample sizes with 95% probability of including at least one copy of alleles with a given frequency. For example, if there are 50 loci with four alleles each, 156 individuals are required to retain at least one copy of alleles with 95% probability and with a frequency of 0.05. Although pearl millet is highly cross-pollinated crop, however, with lack of information available on number of loci and alleles per locus, this method could not be used in developing the core collection. Therefore, the hierarchical clustering suggested by Brown (1989b) was followed in which grouping starts with taxonomy (species, subspecies, and races) followed by grouping based on major geographical strata (country of origin, state), climate, or agro-ecological regions. The clustering within these broad geographical groups sort accessions into different clusters and the number of accessions can then be selected from each cluster depending on the strategy used. A good core therefore, would represent maximum genetic diversity with no genotypically redundant entries.
Ever since the concept on core collection is developed, a number of core collections have already been established for many crop species including perennial Glycine (Brown et al. 1987); peanut (Holbrook et al. 1993; Upadhyaya et al. 2003); perennial medicago species (Diwan et al. 1994; Basigulp et al. 1995); sorghum (Prasada Rao and Ramanath Rao 1995; Grenier et al. 2001); common bean (Tohme et al. 1995); okra (Mahajan et al. 1996); quinoa (Ortiz et al. 1998); Caribbean maize (Taba et al. 1998); alfalfa (Skinner et al. 1999); sweetpotato (Huaman et al. 1999); potato (Huaman et al. 2000); chickpea (Upadhyaya et al. 2001); Uruguayan maize (Malosetti and Abadie 2001) and pigeonpea (Reddy et al. 2005). The objective of the present study was to develop a pearl millet core collection using data on geographical origin and quantitative traits of 16,063 well-characterized cultivated accessions.
Materials and methods
Plant material
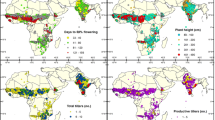
ICRISAT genebank holds 21,392 pearl millet accessions (20,642 cultivated and 750 wild) from 50 countries comprising of mostly landraces, breeding lines, and improved selections from landraces. These accessions are held in trust following the agreement signed in 1994 with the Food and Agriculture Organization of the United Nations (FAO). In the present study, 16,063 cultivated accessions (including landraces, breeding stocks, and advanced lines) from 25 countries that have data on quantitative traits were included. The accessions that do not have adequate characterization data were not included in the study. Of the 16,063 accessions, information on countries of origin was available for all the accessions. The characterization data consisted of data on 11 quantitative traits during rainy (June–October) and post-rainy (November–March) seasons. The data in two seasons were treated separately due to the large variation for the quantitative traits during these two seasons. These two seasons are characteristics of semi-arid tropical areas defined by the day-length and monsoons resulting in considerable difference in the expression of quantitative characters like days to 50% flowering, plant height, spike length, and spike thickness (Appa Rao et al. 1986). Data for all 11 quantitative traits were available in 13,321 accessions (Table 1). Data for days to 50% flowering, plant height, spike length, and spike thickness were available on approximately 15,775 accessions and approximately 15,850 accessions during post-rainy and rainy seasons, respectively (Table 1). Data on number of productive tillers and spike exertion were recorded only during rainy seasons owing to their better expression and on 1,000-grain weight during post-rainy seasons owing to quality factors. Data were available on 15,848, 15,393, and 16,059 accessions for number of productive tillers, spike exertion, and 1,000-grain weight, respectively.
Establishment of core collection
A flowchart of the methodology used in the establishment of core collection is schematically presented in Fig. 1. The entire pearl millet collection consisting of 16,063 accessions was first stratified into 25 groups based on their country of origin (Table 2). The data on 11 traits in each group was standardized using the range of each variable to eliminate scale differences (Milligan and Cooper 1985). A hierarchical cluster analysis was performed on the standardized quantitative data, using Ward’s minimum variance method (Ward 1963) with an R 2 (squared multiple correlation) of 0.70 for grouping the accessions. The Ward’s (1963) method was used as per the PROC-CLUSTER program in SAS (SAS Institute 1989). This method first computes a matrix based on Euclidean distances among group means and produces a dendrogram depicting successive fusion of individuals, and conclude at the stage in which all the individuals of the same group form a cluster (Romesburg 1984). The number of groups depends on the size of collection, the intended size of the core and the dissimilarity of the groups at the lowest level of sorting. Following the proportional sampling procedure, approximately, 10% of the accessions were randomly selected from each cluster (Brown 1989b) to be included in the core subset. The proportional allocation incorporates the alleles even with lower variance. At least one accession was included from those clusters that had less than 10 accessions.
Validation of the core selection method
Means of the entire and core collection were compared using Newman–Keuls procedure (Newman 1939; Keuls 1952) for the 11 traits. Levene’s homogeneity test for variances was used to compare the core and entire collection (Levene 1960). The frequency distributions observed for country of origin in the selected sample was compared to the expected frequency distribution from the entire collection using the chi-square test (Spagnoletti-Zeuli and Qualset 1993). The frequency deviations from the entire collection for these traits in the core sample were calculated. The deviations for country of origin were also compared by chi-square analysis. The Wilcoxon (1945) rank-sum non-parametric test was performed with the SAS NPAR1-WAY procedure (SAS Institute 1989), to determine if the core collection represented the entire germplasm collection for each of the 11 traits. The phenotypic correlation between different traits in core and entire collection were estimated separately to determine whether the associations which maybe under the same genetic control were conserved in the core. Phenotypic diversity was estimated by the Shannon–Weaver diversity index (SDI) (Shannon and Weaver 1949) as follows:
where, n = number of phenotypic classes for a trait, P i = proportion of the total number of entries in the ith class.
Results and discussion
The procedure used to establish the pearl millet core collection resulted in the selection of 1,600 accessions from the entire ICRISAT germplasm collection. The composition of the core collection reflected the predominance of accessions from India and North-West Africa, both representing dry semi-arid tropical ecology (Table 2). According to Harlan and de Wet (1971), the greatest morphological diversity in pearl millet occurred in the dry semi-arid tropical region of North-West Africa and the entire collection accounted for 3,801 (23.7%) accessions with 399 accessions (24.9%) in the core subset. India, which is considered as the secondary center of diversity for pearl millet, accounted for 5,373 accessions (33.45%) in the entire collection and 522 accessions (32.63%) in the core collection (Table 2). The germplasm from some of the countries like Yemen, Chad, and Sierra Leone appears to be under-represented in the collection held at ICRISAT genebank and consecutively in the core collection. Overall, the geographical distribution of the entire pearl millet landrace collection showed a disparity in the collection from different ecological zones. However, this represented a wide distribution of pearl millet growing areas over the world. The unequal representation of landraces from different countries may be the result of use of improved varieties instead of landraces according to farmer’s needs in the past. The poor representation of some countries could also be attributed to the difficulties in collection from the fields, the political situations in the prospected areas, the socio-economic conditions of the farmers, or even the countries with civil wars. The entire landrace collection, however, fits the evolutionary patterns, nearly representing the geographical distribution and ecological zones of the crop. Further, there was a wide range of variation captured in the entire collection for all the quantitative traits based on their country of origin.
Significant differences among means of the entire collection and core collection were recorded only for plant height (rainy) (Table 3). For four traits, variances of the entire collection and core subset were homogeneous. Of the remaining seven traits, variances in the core subset were higher, except number of productive tillers, indicating that the core captured greater variation. The range for most of the characters was retained in the core sample (Table 3). Between 85% and 100% of the range of the entire collection was included in the core for plant height (rainy and post-rainy), days to flowering (rainy and post-rainy), spike length (rainy and post-rainy) and spike thickness (rainy). For the remaining five traits, the range was between 73% and 83%. Thus, the selected core collection is representative of the entire collection.
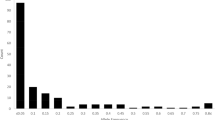
The analysis of deviation of frequency distribution indicated homogeneity among the entire collection and the core subset for country of origin. However, some countries were represented proportionately more (Nigeria, Namibia) and some less (India, Togo, and Zimbabwe) in the core sample. The percentages of accessions selected from different countries ranged from 0.38% for Yemen to 32.63% for India, in the core collection (Table 4). The proportional sampling strategy suggested by Brown (1989b), used for selecting the core sample, showed significant similarity in the frequency distribution for countries of origin. This was evident for most of the countries that were represented more or less in the entire and the core collection. For quantitative traits, the analysis of frequency distribution indicated homogeneity of distribution among the entire and core collection (data not shown). The Wilcoxon’s rank-sum test also indicated that all the characters except spike exertion (P = 0.011) have similar distribution in both core and entire collection (data not shown).
The Shannon–Weaver diversity indices calculated over all the 11 traits indicated similar pattern of diversity among entire collection and core collection (Table 5). No significant differences were observed in the diversity estimates for each of the traits in both entire as well as core collection. Significantly less diversity was observed for number of productive tillers in both the collections. A proper sampling strategy used in developing a core collection should consider the conservation of phenotypic associations arising from co-adapted gene complexes (Ortiz et al. 1998). The established core collection conserves the phenotypic correlations among the traits as observed for the entire collection (Table 6). This clearly indicates that most of the co-adapted gene complexes governing these traits were properly sampled. In the present study, strong associations were observed among some of the traits like days to 50% flowering (rainy) and plant height (rainy) (r = 0.550 in the entire collection and 0.529 in the core sample); spike length (rainy) and spike length (post-rainy) (r = 0.772 in the entire and 0.766 in the core collection); spike thickness (rainy) and spike thickness (post-rainy) (r = 0.585 in the entire collection and 0.594 in core sample) indicating that in future evaluation of germplasm fewer traits may be taken into consideration during rainy or post-rainy seasons, making it less laborious in regenerating the material. For other traits, correlations were low in both entire and core collection.
The core subset was developed considering 11 quantitative traits, which are influenced by G × E interaction, however, the data from two seasons were treated separately to nullify the effect of G × E interaction to some extent with an assumption that the effect of rainy and post-rainy seasons in different years were similar. Thus, the proportional sampling strategy used in the present study was found effective in establishing the core sample that represents the world variation of the entire pearl millet collection. There were no significant differences in the mean and range for the characters studied and frequency distributions were not affected due to proportional sampling. The core also retained overall phenotypic diversity present in the entire collection. Similar results were also obtained in other studies when proportional sampling strategy was adopted for selecting samples from each cluster to constitute the core collection (Erskine and Muehlbeur 1991; Schoen and Brown 1993; Bataillon et al. 1996; Ortiz et al. 1999). The established pearl millet core collection, therefore, can be used very efficiently as a starting point for further improvement programs, involving research on screening the germplasm collection for sources of desirable traits as well as photoperiod sensitivity, disease resistance, drought tolerance, and adaptation to saline or alkaline environments. For example, in diseases, like downy mildew (one of the most destructive diseases of pearl millet), the information on amount of variability present in the germplasm is very limited (only 4,727 cultivated accessions have been screened so far) and drought, a major abiotic stress, only 115 cultivated accessions have been screened, and it will take at least another few years to examine the entire germplasm collection for specific traits of interest. In such a case, the core collection would allow quick identification of new sources of alleles for resistance owing to the conservation of genetic variability in these accessions for most of the characters. The core subset will also help in tackling new constraints that may arise because of onset of new diseases and pests. The core being representative of the entire collection and seeds of the core subset being available, resistance sources for new constraints could be rapidly identified. Furthermore, additional sources of resistance can be found in the reserve collection by selectively examining the clusters from which the core accessions have been identified. The core will also provide a guideline to the curator while acquiring new accessions in the genebank collection and it should be revised periodically as and when additional accessions and information becomes available.
The list of pearl millet entries included in the core collection with details on country of origin, IP number, and the cluster composition are available on diskette from the corresponding author.
References
Appa Rao S, Mengesha MH, Sibale PK, Rajagopal Reddy C (1986) Collection and evaluation of pearl millet (Pennisetum) germplasm from Malawi. Eco Bot 40(1):27–37
Basigalup DH, Barnes DK, Stucker RE (1995) Development of a core collection for perennial Medicago plant introductions. Crop Sci 35:1163–1168
Bataillon TM, David JL, Schoen DJ (1996) Neutral genetic markers and conservation genetics: simulated germplasm collections. Genetics 144:409–417
Brown AHD, Grace JP, Speer SS (1987) Designation of a core collection of perennial Glycine. Soybean Genet Newslett 14:59–70
Brown AHD (1989a) Core collections: a practical approach to genetic resources management. Genome 31:818–824
Brown AHD (1989b) The case for core collections. In: Brown AHD, Frankel OH, Marshall DR, Williams JT (eds) The use of plant genetic resources. Cambridge University Press, Cambridge, UK, pp 136–156
Crossa J (1989) Methodologies for estimating the sample size required for genetic conservation of outbreeding crops. Theor Appl Genet 77:153–161
Dalrymple DG (1986) Development and spread of high-yielding wheat varieties in developing countries, 7th edn. US Agency for International Development, Washington DC
Diwan N, Bauchan GR, McIntosh MS (1994) A core collection for the United States annual Medicago germplasm collections. Crop Sci 34:279–285
Dowswell CR, Paliwal RL, Cantrell RL (1996) Maize in the third world. Westview Press, Boulder, USA
Erskine WA, Muehlbauer FJ (1991) Allozyme and morphological variability, outcrossing rate and core collectionformation in lentil germplasm. Theor Appl Genet 83:119–125
FAOSTAT (2005) http://apps.fao.org/
Frankel OH (1984) Genetic perspective of germplasm conservation. In: Arber W, Llimensee K, Peacock WJ, Starlinger P (eds) Genetic manipulations: impact on man and society. Cambridge University Press, Cambridge, England, pp 161–170
Frankel OH, Brown AHD (1984) Plant genetic resources today: a critical appraisal. In: Holden JHW, Williams JT (eds) Crop genetic resources: conservation and evaluation. George Allen and Unwin, London, UK, pp 249–257
Grenier C, Bramel-cox PJ, Hamon P (2001) Core collection of sorghum: I. Stratification based on eco-geographic data. Crop Sci 41:234–240
Harlan JR, de Wet JMJ (1971) Towards a relational classification of cultivated plants. Taxon 20(4):509–517
Holbrook CC, Anderson WF, Pittman RN (1993) Selection of a core collection from the U.S. germplasm collection of peanut. Crop Sci 33:859–861
Huaman Z, Aguilar C, Ortiz R (1999) Selecting a Peruvian sweetpotato core collection on the basis of morphological, eco-geographical, disease and pest reaction data. Theor Appl Genet 98:840–844
Huaman Z, Ortiz R, Zhang D, Rodriguez F (2000) Isozyme analysis of entire and core collections of Solanum tuberosum subsp. andigena potato cultivars. Crop Sci 40:273–276
Jiang HF, Duan NX (1998) Utilization of groundnut germplasm resources in breeding programme. Crop Genet Resour 2:24–25
Keuls M (1952) The use of the “Studentized range” in connection with an analysis of variance. Euphytica 1:112–122
Khairwal IS, Rai KN, Andrews DJ, Harinarayana G (1999) Pearl millet breeding. Oxford & IBH publishing Co. Pvt. Ltd., New Delhi, 506 pp
Levene H (1960) Robust tests for equality of variances. In: Olkin I (ed) Contributions to probability and statistics: essays in honor of Harold Hotelling. Stanford University Press, Stanford, pp 278–292
Mahajan RK, Bisht IS, Agarwal RC, Rana RS (1996) Studies on South Asian okra collection: methodology for establishing a representative core set using characterization data. Genet Res Crop Evol 43:249–255
Malosetti M, Abadie T (2001) Sampling strategy to develop a core collection of Uruguayan maize landraces based on morphological traits. Genet Res Crop Evol 48(4):381–390
Milligan GW, Cooper M (1985) An examination of procedures for determining the number of clusters in a data set. Psychometrika 50(2):159–179
Newman D (1939) The distribution of range in samples from a normal population expressed in terms of an independent estimate of standard deviation. Biometrika 31:20–30
Ortiz R, Ruia-Tapia EN, Mijica-Sanchez A (1998) Sampling strategy for a core collection of Peruvian Quinoa germplasm. Theor Appl Genet 96:475–483
Ortiz R, Madsen S, Ruiz-Tapia EN, Jacobsen SE, Mujica-Sanchez A, Christiansen JL, Stolen O (1999) Validating a core collection of Peruvian Quinoa germplasm. Genet Res Crop Evol 46:285–290
Prasada Rao KE, Ramanath Rao V (1995) The use of characterization data in developing a core collection of sorghum. In: Hodgkin T, Brown AHD, van Hintum MJL, Morales EAV (eds) Core collections of plant genetic resources. A Wiley-Sayee Publications, Chichester, UK, pp 109–116
Reddy LJ, Upadhyaya HD, Gowda CLL, Singh S (2005) Development of core collection in pigeonpea [Cajanus cajan (L.) Millspaugh] using geographic and qualitative descriptors. Genet Res Crop Evol 52:1049–1056
Romesburg HC (1984) Cluster analysis for researchers. Lifetime Learning Publ., Belmont, CA
SAS Institute (1989) SAS user’s guide: statistics. State University Press, Cary, NC, USA
Schoen DJ, Brown AHD (1993) Conservation of allelic richness in wild crop relatives is aided by assessment of genetic markers. Proc Natl Acad Sci USA 90:10623–10627
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana, USA
Shiv Kumar S, Gupta S, Chandra S, Singh BB (2004) How wide is genetic base of pulse crops? In: Ali M, Singh BB, Shiv Kumar S, Dhar V (eds) Pulses in new perspective. Proceedings of the National Symposium on Crop Diversification and Natural Resources Management, 20–22 December 2003. Indian Society of Pulses Research and Development, Indian Institute of Pulses Research, Kanpur, pp 211–221
Skinner DZ, Bauchan GR, Auricht G, Hughes S (1999) Developing core collections for a large annual Medicago collection. In: Johnson RC, Hodgkin T (eds), Core collections for today and tomorrow. International Plant Genetic Resources Institute, Rome, Italy
Spagnoletti-Zeuli PL, Qualset CO (1993) Evaluation of five strategies for obtaining a core subset from a large genetic resource collection of durum wheat. Theor Appl Genet 87:295–304
Taba S, Diaz J, Franco J, Crossa J (1998) Evaluation of Caribbean maize accessions to develop a core subset. Crop Sci 38:1378–1386
Tohme J, Jones P, Beebe S, Iwanga M (1995) The combined use of agro-ecological and characterization data to establish the CIAT Phaseolus vulgaris core collection. In: Hodgkin T, Brown AHD, van Hintum MJL, Morales EAV (eds) Core collections of plant genetic resources. A Wiley-Sayee Publications, Chichester, UK, pp 95–107
Upadhyaya HD, Bramel PJ, Singh S (2001) Development of a chickpea core subset using geographical distribution and quantitative traits. Crop Sci 41:206–210
Upadhyaya HD, Ortiz R, Bramel PJ, Singh S (2003) Development of a groundnut core collection using taxonomical, geographical, and morphological descriptors. Genet Resour Crop Evol 50:139–148
Upadhyaya HD, Furman BJ, Dwivedi SL, Udupa SM, Gowda CLL, Baum M, Crouch JH, Buhariwalla HK, Singh S (2006) Development of a composite collection for mining germplasm possessing allelic variation for beneficial traits in chickpea. Plant Genet Resour 4:13–19
Vellve R (1992) Saving the seed: genetic diversity and European agriculture. Earthscan Publications, London
Ward JH (1963) Hierarchical grouping to optimize an objective function. J Am Stat Assoc 58:236
Wilcoxon F (1945) Individual comparisons by ranking methods. Biom Bull 1:80–83
Acknowledgements
The lead author gratefully acknowledges ICRISAT for granting a research fellowship during her doctoral program and also for funding the project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bhattacharjee, R., Khairwal, I.S., Bramel, P.J. et al. Establishment of a pearl millet [Pennisetum glaucum (L.) R. Br.] core collection based on geographical distribution and quantitative traits. Euphytica 155, 35–45 (2007). https://doi.org/10.1007/s10681-006-9298-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-006-9298-x