Abstract
Numerous stripe rust resistance genes have been identified from wheat, and new virulent races of Puccinia striiformis f. sp. tritici have also emerged in recent years. Deployment of diverse combinations of resistance genes is an efficient way to combat virulent evolution of strip rust pathogen. In this study, publically available molecular markers were used to identify the distribution of 36 Yr genes in 672 wheat accessions. The effectiveness of Yr genes individually and in combinations was also evaluated in field conditions. The result showed effective resistance of some recently applied genes, such as Yr15 and Yr65. It also showed the lost efficacy of some once widely used genes, such as Yr9 and Yr10. Moreover, significant additive effects were observed in some gene combinations, such as Yr9 + Yr18 and Yr30 + Yr46. Proper deploying of Yr genes and utilizing the positive interactions will be helpful for durable resistance breeding in wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe rust, caused by Puccinia striiformis f. sp. tritici (Pst), is a serious fungal disease for wheat, especially in hypothermal and moist environments (Chen et al. 2014). To control this disease genetically, numerous Yr (yellow rust) genes have been found (McIntosh et al. 2016). The formally designated Yr genes were up to Yr76 (Dracatos et al. 2016; Xiang et al. 2016). Yr15, Yr24/Yr26, Yr35, Yr36, Yr53, Yr64 and Yr65 were derived from tetraploid wheat, Yr8 from Aegilops. comosa (Niu et al. 2004), Yr9 from Secale cereale (Mago et al. 2005), Yr17 from Ae. ventricosa (Jia et al. 2011), Yr28 and Yr48 from Ae. tauschii (Singh et al. 2000; Lowe et al. 2011), Yr37 from Ae. kotschyi (Heyns et al. 2011), Yr38 from Ae. sharonensis (Marais et al. 2010), Yr40 from Ae. geniculata (Kuraparthy et al. 2009), Yr42 from Ae. neglecta (Marais et al. 2009), Yr50 from Thinopyrum intermedium (Liu et al. 2013), Yr70 from Ae. umbellulata (Bansal et al. 2016), and others mainly from hexaploid landraces (McIntosh et al. 2016).
To date, Yr10, Yr18, Yr36 and Yr46 have been cloned (Liu et al. 2014; Krattinger et al. 2009; Fu et al. 2009; Moore et al. 2015). Yr10 encodes a NBS–LRR protein; Yr18 encodes an ATP-binding cassette (ABC) transporter; Yr36 encodes a wheat Kinase-START (WKS) protein; Yr46 encodes a hexose transporter. Fine mapping of Yr9, Yr15 and Yr26 was also conducted (Mago et al. 2005; Zhang et al. 2013; Abdollahi Mandoulakani et al. 2015). Moreover, a lot of functional genes involved in the wheat-stripe rust responses have been identified, such as TaHLRG, TaMDHAR and TaADF7 (Liu et al. 2008a, b; Feng et al. 2014; Fu et al. 2014). Concerning coevolution of plant and pathogen, the effectiveness of single gene to the resistance is limited and short-term. Gene pyramiding will be an effective way to improve plant durable resistance (Ellis et al. 2014).
When proposing pyramiding strategies, distribution of Yr genes needs to be identified in wheat germplasms and breeding lines. Marker-assisted detection (MAD) is the most commonly used method to identify the presence of Yr genes (Goutam et al. 2015). Since most markers are linkage markers instead of gene markers, the validity of them remains to be assessed. The presence of Yr5, Yr9, Yr10, Yr15, Yr17, Yr18, Yr26 or Yr36 has been identified in a limited number of wheat cultivars and breeding lines (Tabassum et al. 2010; Yuan et al. 2012; Zeng et al. 2014). There are a great number of Pst resistance genes, especially newly reported ones (McIntosh et al. 2016), having not been detected in diverse germplasms and breeding lines. Moreover, the effectiveness of most Yr genes against the newly emerged Pst races, such as PST-V26 (Tian et al. 2016), is not very clear. The function of resistance gene also depends on the genetic background (Ellis et al. 2014). So, it is meaningful to evaluate the effectiveness of Yr genes in diverse germplasms not just in the near-isogenic lines (NILs).
Many researches have indicated that additive effects exist extensively among Yr genes. For example, complex additive interactions were observed by Yang et al. (2013). Yr31 suppressed the additive effect of Yr30 and a 3D locus, but not of Yr18 in Mexican. The 3D and 5BL loci were generally not additive with each other, but were additive when combined with other loci in China. Additive effects were also observed between Yr58 and Yr46. The recombinant inbred lines (RILs) carrying both genes showed a lower IT than those carrying Yr58 or Yr46 individually (Chhetri et al. 2016). Studies also showed that rust resistance can be enhanced by combining all stage resistance (ASR) or seedling resistance (SR) genes with adult plant resistance (APR) or slow rusting resistance (SLR) genes (Chen et al. 2013; Ellis et al. 2014). It is necessary to evaluate the interactions between different Yr genes pyramided together.
The Sichuan Province in China is an important overwintering area for stripe rust races. The prevalent Pst races are PST-CYR32 and PST-CYR33 (Zhou et al. 2014a, b). New virulent races also emerged frequently, such as PST-G22 (Xiang et al. 2013). If stripe rust could not be controlled in the Chengdu Plain, it would put a threat on the wheat production of the middle and lower reaches of the Yangtze River. To control epidemics of stripe rust, the evaluation for the efficacy of Yr genes in wheat germplasms and breeding lines should be timely assayed.
Materials and methods
Plant materials
A total of 672 wheat accessions (Table S1) were collected, including 17 Yr gene NILs of Avocet “S” (AvS), 21 cultivars and 147 breeding lines of “Chuanyu” wheats, 170 landraces and 140 cultivars in China, 148 accessions from all over the world, and 29 synthetic wheats. Among these materials, the NILs were received from the Sichuan Academy of Agricultural Sciences (SAAS), the 200 China core collections and 84 foreign germplasms were obtained from the Chinese Academy of Agricultural Sciences (CAAS, http://www.cgris.net/), and 23 accessions containing specific Yr genes were acquired from USDA-ARS (http://www.ars-grin.gov/).
Field testing
All 672 wheat accessions were evaluated for stripe rust reaction at Shuangliu, Shifang and Jitian in Sichuan Province in 2013, 2014 and 2015. Twenty seeds of each accession were planted in a row, and susceptible wheat strains “Minxian169” and “Chuanyu12” were inserted after every 9 rows. Mixed Pst spores of races PST-CYR32, PST-CYR33, PST-SU11, PST-Hybrid46 and PST-G22 (provided by SAAS), were suspended in 0.05% Tween 20 and were sprayed on wheat seedling leaves at trefoil stage. The infection types (ITs) were recorded at the adult plant stages (twice for each environment at 150 and 164 days after seeding) by a modified method according to the standard classification system from 0 to 4 (McIntosh et al. 1995).
Molecular markers
Two closely linked (usually flanking) markers of each Yr gene were chosen to identify its presence/absence in the wheat accessions, except a few genes for which only one closely linked marker was reported. As shown in Table 1, a total of 77 markers (37 SSR markers, 15 STS markers and 25 EST or gene based markers) for 39 stripe rust resistance genes were employed in this study. The primer sequences were synthesized by GENEWIZ Biotech (China).
DNA extraction
A total of 100 mg fresh leaf tissue was collected from each accession, frozen in liquid nitrogen, and ground to powder with a high-throughput tissue grinder. Then, 2× hexadecyltrimethylammonium bromide (CTAB) extraction buffer containing 1.4 M NaCl, 100 mM Tris–HCl (pH 8.0), 2% CTAB and 20 mM EDTA was added to extract the genomic DNA according to Riede and Anderson (1996). Finally, the DNA was dissolved in 100 μl TE buffer with 10 mM Tris–HCl (pH 8.0), 1 mM EDTA and 20 μg/ml RNase, and was incubated for 1 h at 37 °C before storing at −20 °C.
PCR and electrophoresis
Polymerase chain reaction (PCR) was performed using a Master Cycler Pro PCR System (Eppendorf, Germany). A 10 μl PCR mixture consisted of 100 ng template DNA, 5 μl 2× Es Taq MasterMix (CWBIO Biotech, China), 0.3 μl 10 μM forward primer and 0.3 μl 10 μM reverse primer. Amplifications were programmed with 5 min of denaturation at 94 °C; 35 cycles of 30 s at 94 °C, 30 s at 40–60 °C (depending on the primer pair) and 30 s at 72 °C; and 72 °C for 10 min followed by a 4 °C hold step. PCR products were separated by 1.5% agarose gel, 8% non-denaturing polyacrylamide gel or 6% denaturing polyacrylamide gel, and were stained with ethidium bromide or silver nitrate (An et al. 2009), respectively. For the primer pairs STS-7/8, STS-9/10 and URIC/LN2, the PCR products were digested with DpnII (New England Biolabs, USA) according to Chen et al. (2003) before electrophoresis.
Phenotyping and genotyping
The immune plant with no visible symptoms was scored as IT 0, the highly resistant plant with little necrotic flecks and no sporulation was scored as IT 1, the moderately resistant plant with a few necrotic flecks and trace sporulation was scored as IT 2, the moderately susceptible plant with necrotic blotches and moderate sporulation was scored as IT 3, the highly susceptible plant with chlorotic stripes and abundant sporulation was scored as IT 4. The average ITs from the scores of all environments was used to represent the resistant/susceptible (R/S) phenotype of a wheat accession. Few wheat accessions in some environments were not scored because of their absence, but it did not affect the scores collected from other environments. While analyzing genotyping results, the wheat accession was counted for carrying a specific Yr gene only when both flanking markers were presented, except for a few genes where one closely linked marker has been reported.
Data analysis
All data of phenotypes and genotypes were recorded in Microsoft office excel 2010 for statistical analysis. To compare the effective of each gene in diverse genetic backgrounds, the data was divided into two groups (presence of a gene and absence of a gene). To evaluate the effective of different combinations of two resistance genes, the data was divided into four groups (group one, where both selected genes were present; group two and group three, where only one of the selected genes was present; group four, where none of the two genes was present). One-way analysis of variance (ANOVA) was conducted to evaluate the variance and significance among these groups.
Results
Detection of stripe rust resistance genes in 672 wheat accessions
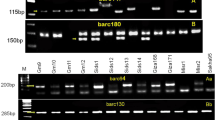
The accuracy of MAD is affected by the distance between the target gene and the linkage markers. So, closely linked markers are desired. For example, a recent report indicated that YrSP was different from Yr5/Yr7 (Feng et al. 2015). Both of them were located on the 2BL chromosome, and were hard to be distinguished. Seven published markers were screened in CS, Avocet and Yr-NILs of Yr5, Yr7 and YrSP (Fig. 1). S19M93 and STS7/8 can be used to identify Yr5; gwm526 and barc349 can be used to identify Yr7; dp269 and WC5/WC6 can be used to identify YrSP. Moreover, some markers were easy to be distinguished, such as STS (S19M93 for Yr5) and SCAR (SC385 for Yr17), and were helpful for high-throughput detection. By using the NILs or donors of the corresponding gene as the positive control, the detection of 36 Yr genes in 672 wheat accessions was accomplished (Fig. S1).
Electrophoretogram of seven PCR based markers for the Yr5/Yr7–YrSP locus. An 8% non-denaturing polyacrylamide gel was used to separate the PCR products. A molecular weight standard of base pairs was listed in the right. Right arrow indicates the polymorphism bands of each molecular marker among the five wheat accessions
Distribution of 36 Pst resistance genes in different taxon groups
Totally, Yr10, Yr17 and Yr18 were identified positively in numerous accessions, while Yr8, Yr36, Yr61 and YrZH84 were detected positively only in a few accessions. As shown in Table 2, Yr5, Yr9, Yr15, Yr17, Yr30, Yr48, Yr65, Yr67, YrSP and TaHLRG showed high frequency in “Chuanyu” breeding lines; Yr10, Yr18, Yr33, Yr51, Yr59 and Yr62 showed high frequency in Chinese landraces; Yr5, Yr9, Yr29, Yr30, Yr39, Yr41, Yr48, Yr49, Yr57 and Yr59 showed high frequency in Chinese modern cultivars; Yr4, Yr5, Yr7, Yr9, Yr29, Yr30, Yr33, Yr41, Yr52, Yr57, Yr60 and YrSP showed high frequency in introduced foreign germplasms. These results revealed broad diversity of resistance genes in different wheat taxon groups. Moreover, the accessions carrying multiple Yr genes identified in this study might be useful as parental lines for diversifying Pst resistance sources in wheat breeding.
Stripe rust reactions in field testing
By analyzing the IT scores collected from three places and three years, environmental variation was non-significant (p = 0.87), but significant variances were observed among the average ITs of different wheat accessions from all environments (p < 0.01). The overview for the percentages of each Pst infection phenotype and the average ITs of whole wheat accessions and four taxon groups are shown in Fig. S2. The ratio of resistant to susceptible phenotype in the total wheat accessions is near to 1:1, and the average IT of the total wheat accessions is between 2.5 and 3.
Relationship between the number of pyramided genes and Pst resistance
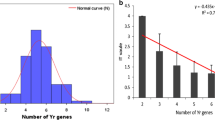
To obtain a realistic relationship between Pst resistance and the pyramided gene number, the results of MAD and field testing were combined for analysis. As shown in Fig. 2, a significant positive correlation (R2 > 0.8, p < 0.01) between the number of Yr genes and the Pst resistance was observed in the 672 wheat accessions. This high correlation suggested that pyramiding of Yr genes will be an effective way to improve stripe rust resistance in wheat breeding. Moreover, the accessions carrying multiple Yr genes might be useful as intermediate materials for introducing Pst resistance sources into commercial varieties.
The relationship of stripe rust resistance and pyramided gene number. The trend line was added based on the sum percentages of MR and HR, a or the average ITs, b of each pyramided gene number. HS highly susceptible phenotype, MS moderately susceptible phenotype, MR moderately resistant phenotype, HR highly resistant phenotype. Each bar represents the mean value of the stripe rust IT (0, 1, 2, 3, 4) and standard deviation (SD; n ≥ 17) in b
Contribution of Yr genes individually to Pst resistance
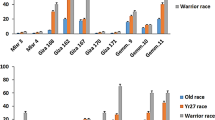
The contribution of each Yr gene to stripe rust resistance was evaluated in diverse genetic backgrounds. As shown in Fig. 3, the wheat accessions carrying Yr15, Yr17, Yr18, Yr65 and Yr67 had significantly lower ITs than those without the corresponding gene, whilst the situation for Yr29 was opposite. The other genes did not meet a significant level to stripe rust resistance in this study. Moreover, the contribution of eleven genes to Pst resistance in the four taxon groups was shown in Fig. S3. These results revealed some Yr genes contributed effective resistance to the current Pst races. The other genes that did not show significance but still had lower ITs, such as Yr46 and Yr60, might contribute to partial resistance in pyramids, which will be analyzed in the following section.
Contribution of each gene to Pst resistance. Each bar represents the mean value of the stripe rust IT (0, 1, 2, 3, 4) and SD (n ≥ 15). One-way ANOVA was used to determine the significance level between the presence and absence groups of each Yr gene. Significance at *p < 0.05. Significance at **p < 0.01
Comparing different combinations of two genes with respect to Pst resistance
A total of 183 combinations of each two genes were suitable for the statistical analysis, and partial results of them were shown in Fig. 4. 30.23% combinations showed a similar average ITs with those carrying either one gene, 27.13% combinations showed a higher average ITs than those either one gene present, and 42.64% combinations showed a lower average ITs than those either one gene present. The results revealed that some combinations of two genes, one conferring significant resistance and another conferring non-significant resistance (Fig. 3), such as Yr17 + Yr26 and Yr9 + Yr18, effectively improved the Pst resistance in the field trails. The results also showed that some combinations of two genes conferring non-significant resistance, such as Yr30 + Yr46, effectively improved the Pst resistance when pyramided together. But in some other combinations, there was a reduction of resistance compared with those carrying either of the two genes, such as Yr48 + Yr67. These results revealed the additive effects or epistatic effects between the resistance genes when pyramided.
Contribution of different combinations of two Yr genes to stripe rust resistance. The difference value (D value) of the ITs between the combination group and the higher or lower group (indicated by  ) are shown on the column to represent the resistance improvement (negative D value) or reduction (positive D value). The results of a absolute D value >15% are shown here. Each bar represents the mean value of the stripe rust IT (0, 1, 2, 3, 4) and SD (n ≥ 10). One-way ANOVA was used to determine the significance between the combination group and the higher or lower group of each pair of Yr genes. Significance at *p < 0.05, significance at **p < 0.01
) are shown on the column to represent the resistance improvement (negative D value) or reduction (positive D value). The results of a absolute D value >15% are shown here. Each bar represents the mean value of the stripe rust IT (0, 1, 2, 3, 4) and SD (n ≥ 10). One-way ANOVA was used to determine the significance between the combination group and the higher or lower group of each pair of Yr genes. Significance at *p < 0.05, significance at **p < 0.01
Discussion
Although numerous Yr genes have been reported, Yr11, Yr12, Yr13, Yr14, Yr19, Yr20, Yr22, Yr23 and Yr25 have not been mapped, Yr27, Yr31, Yr32, Yr37, Yr38, Yr40, Yr42, Yr45, Yr50, Yr54, Yr56, Yr63, Yr66, Yr70, Yr73 and Yr74 have no appropriate markers, Yr2, Yr3, Yr21, Yr34, Yr43, Yr47, Yr55 and other temporarily named Yr genes have no closely linked markers. So, 36 Yr genes were finally screened and used to evaluate their contribution to the current Pst resistance in this study.
When conducting marker-assisted selection (MAS) in breeding, validity and convenience of a molecular marker should be considered. Here, PCR based markers were employed (Table 1) for identification of the corresponding gene (Fig. S1). There are still some markers used to map a gene but not suitable for detecting the gene in diverse genetic backgrounds, such as Owm45F3R3 for Yr51 (Randhawa et al. 2014). It might be because the polymorphism of a marker showed in the mapping population could not be distinguished in some wheat germplasms. Some RGAP markers also have this problem, such as the RGAP markers for Yr44 (Sui et al. 2009), Yr45 (Li et al. 2011) and Yr59 (Zhou et al. 2014a, b). So, it is hard to apply these markers directly for MAD, even with the donor lines. Moreover, the brightness of the target band of some markers was too weak to be distinguished from polymorphic bands even with a touchdown PCR program, such as stm673acag for Yr1 (Bansal et al. 2009) and gwm508 for Yr35 (Dadkhodaie et al. 2011).
To enhance the reliability of molecular markers, much improvement has been made from polymorphic markers to specific markers, to gene specific markers, and to functional markers. There were many types of marker conversion having been conducted, such as amplified fragment length polymorphism (AFLP) to SCAR (SC-OPD11, Niu et al. 2004), SSR to SCAR (gwm415 to SC-372, Jia et al. 2011), SSR to STM (gwm533 to stm559tgag, Hayden et al. 2004), EST to SSR (bu099658, Hasancebi et al. 2014), DArT to STS (sun104, Randhawa et al. 2014) and RGAP to STS (wgp5467 to STS5467, Zhou et al. 2014a, b). Still, fine mapping of some Yr genes is required for valid detection, such as distinguishing the closely linked loci Yr5/Yr7-YrSP (Fig. 1). Moreover, sequence specific marker could indicate the presence of the cloned Yr10, Yr18 and Yr36 in the genomic DNA accurately (Fig. S1). But, whether it could be used to indicate the functional type still needs to be considered. Because there are many haplotypes existed in the hexaploid wheat, such as the resistant (Lr67res) and susceptible types (Lr67sus) of Yr46 (Moore et al. 2015). So, function specific markers were developed to indicate the functional mutant. For example, the marker THR1 developed in the non-coding region was used to identify the association of TaHLRG and stripe rust resistance (Liu et al. 2008a, b). Two markers cib-Yr28M1 and cib-Yr28M2 developed in the gene coding region were used to distinguish the resistance associated Yr28 (unpublished). To utilize these cloned Pst resistance-associated genes, such as TaMDHAR (Feng et al. 2014) and TaADF7 (Fu et al. 2014), more functional markers are required.
Through MAD, the distribution of 36 resistance genes in 672 wheat accessions was illustrated. The result showed consistence with previous studies. Yr9, Yr10, Yr17 and Yr18 were the mostly identified genes, while Yr8 and Yr36 were identified in a few accessions (Tabassum et al. 2010; Yuan et al. 2012; Zeng et al. 2014). Many “Chuanyu” cultivars and advanced breeding lines were released in Sichuan and other provinces of China. They showed a good adaptability to the local Pst races (Fig. S2). This might be related to the introduction of new Yr genes (such as Yr15, Yr65 and Yr67) and the reduction of previously widely applied Yr genes (such as Yr9, Yr10 and Yr24/Yr26). However, there are two prominent problems. One problem is the over-use of a few Yr genes, such as Yr17 presenting in 74.40% of “Chuanyu” breeding lines (Table 2). This will lead to diversity reduction of resistance genes in breeding population and is unfavourable for breeding durable resistance varieties. The other problem is that the durable genes have not been widely deployed in “Chuanyu” wheat. The Chinese landraces maintaining rust resistance chronically (Fig. S2) showed a high frequency of Yr18 to 59.41% (Table 2). Many reports revealed the importance of durable resistance genes in combining with other Yr genes (Krattinger et al. 2009; Yang et al. 2011). The enhancement effect was also confirmed in this study, such as Yr9 + Yr18 (Fig. 4). Therefore, through MAD, the overuse of a few Yr genes can be avoided, and durable genes with low frequency can be introduced intentionally. Additionally, the newly reported heterogenous genes, such as Yr37 (Heyns et al. 2011), Yr40 (Kuraparthy et al. 2009) and Yr50 (Liu et al. 2013), should also be applied properly. To perform resistance breeding for a long period, extensive resistance resources are required to broaden the genetic basis of breeding materials.
As prevalent Pst races were variant with environments, it is necessary to understand the effective of Yr genes under a specific condition. Here, a case study was performed in Sichuan province, and we hope that it would attract enough attention to the breeders and researchers in the world-wide. Some Yr genes identified in this study showed significant effects under current conditions, such as Yr15 and Yr65, while some once massively used Yr genes did not confer significant resistance, such as Yr9 (Fig. 3). These results were basically consistent with the rust testing results of some available NILs. Only the NILs of Yr5, Yr15 and Yr18 showed resistance, others (such as the NILs of Yr9 and Yr26) were susceptible to Pst in this study. But variances existed in the results of Yr17. Yr17 contributed significantly to the current Pst resistance in this study (Fig. 3), but Yr17-NIL was susceptible. Further analysis showed that the contribution significance of Yr17 varied with different taxon groups, non-significant in “Chuanyu” breeding lines but significant in Chinese modern cultivars and introduced foreign germplasms (Fig. S3). Other genes, such as Yr67, also showed similar results. The function of one gene depends on the genetic background, so it is meaningful to evaluate the contribution of Yr genes to stripe rust resistance in diverse genetic backgrounds rather than in their NILs. MAD combined with rust testing results could be used to select the most effective genes against current Pst races when breeding high resistant varieties.
The genes conferring significant resistance will be applied widely in breeding, such as Yr15, Yr17 and Yr65 (Table 2; Fig. 3). But other genes showing non-significant effectiveness under current conditions still need to be utilized in gene pyramiding. Because the results identified in this study (Fig. 2) and the presence of multi-QTLs in resistant wheat (Lowe et al. 2011; Rosewarne et al. 2013; Yang et al. 2013) suggested that gene pyramiding could improve the durability of rust resistance. Additive effects and epistatic effects exist extensively in Yr gene pyramids (Fig. 4). And 42.64% of the combinations of two Yr genes improved Pst resistance than those either one present. Although showing non-significance, these combinations with substantially lower ITs (negative D value, Fig. 4) than those carrying either one gene still need to be considered. Moreover, some durable resistance genes conferring partial resistance to Pst showed enhancing effect to race-specific genes in this study, such as Yr17, Yr18, Yr30 and Yr46 (Fig. 4). Taking advantage of the positive interactions and avoiding the negative interactions should be carefully considered in resistance gene pyramiding.
References
Abdollahi Mandoulakani B, Yaniv E, Kalendar R, Raats D, Bariana HS, Bihamta MR, Schulman AH (2015) Development of IRAP- and REMAP-derived SCAR markers for marker-assisted selection of the stripe rust resistance gene Yr15 derived from wild emmer wheat. Theor Appl Genet 12:211–219
Agenbag GM, Pretorius ZA, Boyd LA, Bender CM, Prins R (2012) Identification of adult plant resistance to stripe rust in the wheat cultivar Cappelle-Desprez. Theor Appl Genet 125:109–120
An Z, Xie L, Cheng H, Zhou Y, Zhang Q, He X, Huang H (2009) A silver staining procedure for nucleic acids in polyacrylamide gels without fixation and pretreatment. Analyt Biochem 391:77–79
Bansal UK, Hayden MJ, Keller B, Wellings CR, Park RF, Bariana HS (2009) Relationship between wheat rust resistance genes Yr1 and Sr48 and a microsatellite marker. Plant Pathol 58:1039–1043
Bansal UK, Hayden MJ, Gill MB, Bariana HS (2010) Chromosomal location of an uncharacterised stripe rust resistance gene in wheat. Euphytica 171:121–127
Bansal M, Kaur S, Dhaliwal HS, Bains NS, Bariana HS, Chhuneja P, Bansal UK (2016) Mapping of Aegilops umbellulata-derived leaf rust and stripe rust resistance loci in wheat. Plant Pathol. doi:10.1111/ppa.12549
Chen XM, Soriac MA, Yanb G, Sunb J, Dubcovsky J (2003) Development of sequence tagged site and cleaved amplified polymorphic sequence markers for wheat stripe rust resistance gene Yr5. Crop Sci 43:2058–2064
Chen XM, Coram T, Huang X, Wang M, Dolezal A (2013) Understanding molecular mechanisms of durable and non-durable resistance to stripe rust in wheat using a transcriptomics approach. Curr Genomics 14:111–126
Chen W, Wellings C, Chen X, Kang Z, Liu T (2014) Wheat stripe (yellow) rust caused by Puccinia striiformis f. sp. tritici. Mol Plant Pathol 15:433–446
Cheng P, Xu LS, Wang MN, See DR, Chen XM (2014) Molecular mapping of genes Yr64 and Yr65 for stripe rust resistance in hexaploid derivatives of durum wheat accessions PI 331260 and PI 480016. Theor Appl Genet 127:2267–2277
Chhetri M, Bariana H, Kandiah P, Bansal U (2016) Yr58: a new stripe rust resistance gene and its interaction with Yr46 for enhanced resistance. Phytopathology 106:1530–1534
Dadkhodaie NA, Karaoglou H, Wellings CR, Park RF (2011) Mapping genes Lr53 and Yr35 on the short arm of chromosome 6B of common wheat with microsatellite markers and studies of their association with Lr36. Theor Appl Genet 122:479–487
Dracatos PM, Zhang P, Robert F, Park RF, McIntosh RA, Wellings CR (2016) Complementary resistance genes in wheat selection ‘Avocet R’ confer resistance to stripe rust. Theor Appl Genet 129:65–76
Ellis JG, Lagudah ES, Spielmeyer W, Dodds PN (2014) The past, present and future of breeding rust resistant wheat. Front Plant Sci 5:641
Feng H, Wang X, Zhang Q, Fu Y, Feng C, Wang B, Huang L, Kang Z (2014) Monodehydroascorbate reductase gene, regulated by the wheat PN-2013 miRNA, contributes to adult wheat plant resistance to stripe rust through ROS metabolism. Biochim Biophys Acta 1839:1–12
Feng JY, Wang MN, Chen XM, See DR, Zheng YL, Chao SM, Wan AM (2015) Molecular mapping of YrSP and its relationship with other genes for stripe rust resistance in wheat chromosome 2BL. Phytopathology 105:1206–1213
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L, Chen X, Sela H, Fahima T, Dubcovsky J (2009) A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360
Fu Y, Duan X, Tang C, Li X, Voegele RT, Wang X, Wei G, Kang Z (2014) TaADF7, an actin-depolymerizing factor, contributes to wheat resistance against Puccinia striiformis f. sp. tritici. Plant J 78:16–30
Goutam U, Kukreja S, Yadav R, Salaria N, Thakur K, Goyal AK (2015) Recent trends and perspectives of molecular markers against fungal diseases in wheat. Front Microbiol 6:861
Hasancebi S, Mert Z, Ertugrul F, Akan K, Aydin Y, Senturk Akfirat F, Altinkut Uncuoglu A (2014) An EST-SSR marker, bu099658, and its potential use in breeding for yellow rust resistance in wheat. Czech J Genet Plant Breed 50:11–18
Hayden MJ, Kuchel H, Chalmers KJ (2004) Sequence tagged microsatellites for the Xgwm533 locus provide new diagnostic markers to select for the presence of stem rust resistance gene Sr2 in bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1641–1647
Herrera-Foessel SA, Singh RP, Lillemo M, Huerta-Espino J, Bhavani S, Singh S, Lan C, Calvo-Salazar V, Lagudah ES (2014) Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor Appl Genet 127:781–789
Heyns I, Pretorius Z, Marais F (2011) Derivation and characterization of recombinants of the Lr54/Yr37 translocation in common wheat. Open Plant Sci J 5:1–8
Jia JQ, Li GR, Liu C, Lei MP, Yang ZJ (2011) Characterization of wheat yellow rust resistance gene Yr17 using EST-SSR and rice syntenic region. Cereal Res Commun 39:88–99
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Kuraparthy V, Sood S, See DR, Gill BS (2009) Development of a PCR assay and marker-assisted transfer of leaf rust and stripe rust resistance genes Lr57 and Yr40 into hard red winter wheats. Crop Sci 49:120–126
Li Y, Niu YC (2007) Identification of molecular markers for wheat stripe rust resistance gene Yr6. Acta Agric Boreali Sin 22:189–192
Li Y, Niu YC, Chen XM (2009) Mapping a stripe rust resistance gene YrC591 in wheat variety C591 with SSR and AFLP markers. Theor Appl Genet 118:339–346
Li Q, Chen XM, Wang MN, Jing JX (2011) Yr45, a new wheat gene for stripe rust resistance on the long arm of chromosome 3D. Theor Appl Genet 122:189–197
Lin F, Chen XM (2007) Genetics and molecular mapping of genes for race-specific all-stage resistance and non-race-specific high-temperature adult-plant resistance to stripe rust in spring wheat cultivar Alpowa. Theor Appl Genet 114:1277–1287
Liu C, Yang ZJ, Li GR, Zeng ZX, Zhang Y, Zhou JP, Liu ZH, Ren ZL (2008a) Isolation of a new repetitive DNA sequence from Secale africanum enables targeting of Secale chromatin in wheat background. Euphytica 159:249–258
Liu D, Xia X, He Z, Xu S (2008b) A novel homeobox-like gene associated with reaction to stripe rust and powdery mildew in common wheat. Phytopathology 98:1291–1296
Liu J, Chang Z, Zhang X, Yang Z, Li X, Jia JQ, Zhan H, Guo H, Wang J (2013) Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor Appl Genet 126:265–274
Liu W, Frick M, Huel R, Nykiforuk CL, Wang X, Gaudet DA, Eudes F, Conner RL, Kuzyk A, Chen Q, Kang Z, Laroche A (2014) Stripe rust resistance gene Yr10 encodes an evolutionary-conserved and unique CC–NBS–LRR sequence in wheat. Mol Plant 7:1740–1755
Lowe I, Jankuloski L, Chao S, Chen X, See D, Dubcovsky J (2011) Mapping and validation of QTL which confer partial resistance to broadly virulent post-2000 North American races of stripe rust in hexaploid wheat. Theor Appl Genet 123:143–157
Lu Y, Wang M, Chen X, See D, Chao S, Jing J (2014) Mapping of Yr62 and a small-effect QTL for high-temperature adult-plant resistance to stripe rust in spring wheat PI 192252. Theor Appl Genet 127:1449–1459
Luo PG, Hu XY, Ren ZL, Zhang HY, Shu K, Yang ZJ (2008) Allelic analysis of stripe rust resistance genes on wheat chromosome 2BS. Genome 51:922–927
Mago R, Miah H, Lawrence GJ, Wellings CR, Spielmeyer W, Bariana HS, McIntosh RA, Pryor AJ, Ellis JG (2005) High-resolution mapping and mutation analysis separate the rust resistance genes Sr31, Lr26 and Yr9 on the short arm of rye chromosome 1. Theor Appl Genet 112:41–50
Marais GF, Marais AS, McCallum B, Pretorius ZA (2009) Transfer of leaf rust and stripe rust resistance genes Lr62 and Yr42 from Aegilops neglecta to common wheat. Crop Sci 49:871–879
Marais GF, Badenhorst PE, Eksteen A, Pretorius ZA (2010) Reduction of Aegilops sharonensis chromatin associated with resistance genes Lr56 and Yr38 in wheat. Euphytica 171:15–22
McIntosh RA, Wellings CR, Park RF (1995) Wheat rust: an atlas of resistance genes. CSIRO, Sydney
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Appels R, Xia XC (2016) Catalogue of gene symbols for wheat. http://www.shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp. Accessed Dec 2016
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S, Kong X, Spielmeyer W, Talbot M, Bariana H, Patrick JW, Dodds P, Singh R, Lagudah E (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498
Murphy LR, Santra D, Kidwell K, Yan G, Chen X, Campbell KG (2009) Linkage maps of wheat stripe rust resistance genes Yr5 and Yr15 for use in marker-assisted selection. Crop Sci 49:1786–1790
Niu YC, Li SM, Li WF, Wu LR, Xu SC (2004) Molecular markers assisted selection for the stripe rust (Puccinia striiformis tritici) resistance genes Yr8 and Yr10 in wheat breeding. In: Plant protection towards the 21st century-proceedings of the XVth International Plant Protection Congress, Beijing, China
Randhawa M, Bansal U, Valárik M, Klocová B, Doležel J, Bariana H (2014) Molecular mapping of stripe rust resistance gene Yr51 in chromosome 4AL of wheat. Theor Appl Genet 127:317–324
Randhawa M, Bariana H, Mago R, Bansal U (2015) Mapping of a new stripe rust resistance locus Yr57 on chromosome 3BS of wheat. Mol Breeding 35:65
Ren RS, Wang MN, Chen XM, Zhang ZJ (2012) Characterization and molecular mapping of Yr52 for high-temperature adult-plant resistance to stripe rust in spring wheat germplasm PI 183527. Theor Appl Genet 125:847–857
Riede CR, Anderson JA (1996) Linkage of RFLP markers to an aluminum tolerance gene in wheat. Crop Sci 36:905–909
Rosewarne GM, Singh RP, Huerta-Espino J, William HM, Bouchet S, Cloutier S, McFadden H, Lagudah ES (2006) Leaf tip necrosis, molecular markers and beta1-proteasome subunits associated with the slow rusting resistance genes Lr46/Yr29. Theor Appl Genet 112:500–508
Rosewarne GM, Herrera-Foessel SA, Singh RP, Huerta-Espino J, Lan CX, He ZH (2013) Quantitative trait loci of stripe rust resistance in wheat. Theor Appl Genet 126:2427–2449
Singh RP, Nelson JC, Sorrells ME (2000) Mapping Yr28 and other genes for resistance to stripe rust in wheat. Crop Sci 40:1148–1155
Smith PH, Hadfiled J, Hart NJ, Koebner RMD, Boyd LA (2007) STS markers for the wheat yellow rust resistance gene Yr5 suggest a NBS-LRR-type resistance gene cluster. Genome 50:259–265
Sui XX, Wang MN, Chen XM (2009) Molecular mapping of a stripe rust resistance gene in spring wheat cultivar Zak. Phytopathology 99:1209–1215
Tabassum S, Ashraf M, Chen XM (2010) Evaluation of Pakistan wheat germplasms for stripe rust resistance using molecular markers. Sci China Life Sci 53:1123–1134
Tian Y, Zhan G, Chen X, Tungruentragoon A, Lu X, Zhao J, Huang L, Kang Z (2016) Virulence and simple sequence repeat marker segregation in a Puccinia striiformis f. sp. tritici population produced by selfing a Chinese isolate on Berberis shensiana. Phytopathology 106:185–191
Xiang YJ, Zhang ZY, Shen L, Kuang WJ, Ji HL, Ni JY, Chen WQ, Peng YL (2013) Virulence dynamics of Puccinia striiformis f. sp. tritici in Sichuan province from 2005 to 2010. Southwest China J Agric Sci 26:1858–1863
Xiang C, Feng J, Wang M, Chen X, See DR, Wan A, Wang T (2016) Molecular mapping of stripe rust resistance gene Yr76 in winter club wheat cultivar Tyee. Phytopathology 106:1186–1193
Xu LS, Wang MN, Cheng P, Kang ZS, Hulbert SH, Chen XM (2013) Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor Appl Genet 126:523–533
Yang EN, Zou YC, Yang WY, Tang YL, He ZH, Singh RP (2011) Breeding adult plant resistance to stripe rust in spring bread wheat germplasm adapted to sichuan province of China. Czech J Genet Plant Breed 47:S165–S168
Yang EN, Rosewarne GM, Herrera-Foessel SA, Huerta-Espino J, Tang ZX, Sun CF, Ren ZL, Singh RP (2013) QTL analysis of the spring wheat “Chapio” identifies stable stripe rust resistance despite inter-continental genotype × environment interactions. Theor Appl Genet 126:1721–1732
Yao ZJ, Lin RM, Xu SC, Li ZF, Wan AM, Ma ZY (2006) The molecular tagging of the yellow rust resistance gene Yr7 in wheat transferred from differential host Lee using microsatellite markers. Sci Agric Sin 39:1146–1152
Yin GH, Wang JW, Wen WE, He ZH, Li ZF, Wang H, Xia XC (2009) Mapping of wheat stripe rust resistance gene YrZH84 with RGAP markers and its application. Acta Agron Sin 35:1274–1281
Yuan CL, Jiang H, Wang HG, Li K, Tang H, Li XB, Fu DL (2012) Distribution, frequency and variation of stripe rust resistance loci Yr10, Lr34/Yr18 and Yr36 in Chinese wheat cultivars. J Genet Genom 39:587–592
Zahravi M, Bariana HS, Shariflou MR, Balakrishna PV, Banks PM, Ghannadha MR (2003) Bulk segregant analysis of stripe rust resitance in wheat (Triticum aestivum) using microsatellite markers. In: Proceedings 10th international wheat genetics symposium, Instituto Sperimentale per Cerealcoltura, Rome (Pogna NE, Romano M, Pogna EA, Galterio, et al.), pp 861–863
Zeng QD, Han DJ, Wang QL, Yuan FP, Wu JH, Zhang L, Wang XJ, Huang LL, Chen XM, Kang ZS (2014) Stripe rust resistance and genes in Chinese wheat cultivars and breeding lines. Euphytica 196:271–284
Zhang P, McIntosh RA, Hoxha S, Dong C (2009) Wheat stripe rust resistance genes Yr5 and Yr7 are allelic. Theor Appl Genet 120:25–29
Zhang X, Han D, Zeng Q, Duan Y, Yuan F, Shi J, Wang Q, Wu J, Huang L, Kang Z (2013) Fine mapping of wheat stripe rust resistance gene Yr26 based on collinearity of wheat with Brachypodium distachyon and rice. PLoS One 8:e57885
Zhou XL, Han DJ, Chen XM, Gou HL, Guo SJ, Rong L, Wang QL, Huang LL, Kang ZS (2014a) Characterization and molecular mapping of stripe rust resistance gene Yr61 in winter wheat cultivar Pindong 34. Theor Appl Genet 127:2349–2358
Zhou XL, Wang MN, Chen XM, Lu Y, Kang ZS, Jing JX (2014b) Identification of Yr59 conferring high-temperature adult-plant resistance to stripe rust in wheat germplasm PI 178759. Theor Appl Genet 127:935–945
Acknowledgements
This project was partially funded by the Special Fund for Strategic Pilot Technology of the Chinese Academy of Sciences (Class A, No. XDA08030106) and the National Natural Science Foundation of China (General Program, No. 31371608).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
Electrophoretogram of the primers for different resistance genes. The corresponding NILs (Table S1) of Yr4, Yr5, Yr6, Yr7, Yr8, Yr9, Yr10, Yr15, Yr17, Yr18, Yr26, Yr29 and YrSP, and the related wheat donors that were reported to carry Yr16, Yr28, Yr30, Yr36, Yr39, Yr41, Yr48, Yr52, Yr53, Yr59, Yr62, Yr64, Yr65, Yr67 and YrZH84 were used as the positive control and are shown at right near the marker. Although there is no control accession for Yr33, Yr46, Yr49, Yr51, Yr57, Yr60, Yr61 and TaHLRG, the postulated bands were easy to be distinguished according to previously reported base pairs of the PCR product. The names of the positively identified wheat accessions of each gene are highlighted in bold (PDF 521 kb)
Figure S2
The overview of stripe rust resistance in all wheat accessions. The percentages of each Pst infection phenotype (a) and averages ITs (b) in total wheat accessions and the four taxon groups (“Chuanyu” breeding lines, Chinese landraces, Chinese modern cultivars, and introduced foreign germplasms) were shown upon the column. HS highly susceptible phenotype; MS moderately susceptible phenotype; MR moderately resistant phenotype; HR highly resistant phenotype. Each bar represents the mean value of stripe rust ITs (0, 1, 2, 3, 4) and SD (n ≥ 140) in b (PDF 94 kb)
Figure S3
The contributions of Yr genes to Pst resistance in different taxon groups. The other genes having insufficient sample size were not analyzed. Each bar represents the mean value of stripe rust ITs (0, 1, 2, 3, 4) and SD (n ≥ 10). One way ANOVA was used to reveal the significant level between the presence and absence groups of each Yr gene. * indicates the significance of p < 0.05. ** indicates the significance of p < 0.01 (PDF 278 kb)
Table S1
All wheat accessions used in this paper. The total 672 wheat accessions were subdivided into 6 groups. Group 0: 17 NILs of corresponded Yr gene; Group 1: 168 “Chuanyu” cultivars and breeding lines; Group 2: 170 Chinese landraces; Group 3: 140 Chinese modern cultivars; Group 4: 148 Introduced foreign germplasms; Group 5: 29 synthetic wheats (PDF 62 kb)
Rights and permissions
About this article
Cite this article
Zheng, S., Li, Y., Lu, L. et al. Evaluating the contribution of Yr genes to stripe rust resistance breeding through marker-assisted detection in wheat. Euphytica 213, 50 (2017). https://doi.org/10.1007/s10681-016-1828-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-016-1828-6