Abstract
A recent introduction of Puccinia striiformis f. sp. tritici (Pst) pathotype 134 E16A+ in Western Australia differed from existing pathotypes in its virulence for stripe rust resistance genes Yr8 and Yr9, and avirulence for Yr3, Yr4, Yr34 and some uncharacterized sources of resistance. The Australian wheat cultivar Rubric exhibited a very low seedling stripe rust response when tested against the Pst pathotype 134 E16A+. Genetic analysis of a Rubric/Avocet ‘S’ F3 population indicated monogenic inheritance of resistance. The resistance gene was tentatively designated YrRub. Bulked segregant analysis using multiplex-ready PCR technology placed YrRub distal to the microsatellite marker barc75 in chromosome 3BS. Follow up studies mapped SSR marker cfb3530 between YrRub (2.9 ± 1.3 cM) and barc75 (2.4 ± 1.2 cM). Genotypes Bolac, EMU ‘S’, Nesser, Hybrid 46 (Yr4) and Avalon (Yr4) amplified the YrRub-linked barc75 and cfb3530 alleles. The amplification of PCR products similar to that of Rubric in Yr4-carrying cultivars Hybrid 46 and Avalon suggested that YrRub was most likely to be Yr4. Due to unavailability of genotypes carrying Yr4 singly, test of allelism was not possible. Thirty-eight Australian wheat cultivars known to lack YrRub, an advanced breeding line WAWHT2046 (Yr34) and Vilmorin 23 (Yr3) amplified PCR products different from that of Rubric at the barc75 and cfb3530 marker loci. The amplification of non-Rubric alleles at the marker barc75 and cfb3530 loci among a set of diverse wheat genotypes demonstrated that these markers could be used for marker assisted selection of YrRub in combination with other molecularly tagged seedling and adult plant stripe rust resistance genes in wheat breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe rust, caused by Puccinia striiformis f. sp. tritici (Pst), was first reported in Australia in 1979 and resulted in an epidemic rendering several wheat cultivars susceptible (O’Brien et al. 1980; Wellings 2007). A committed breeding effort of deploying seedling and adult plant resistance gene combinations in eastern Australia resulted in improved levels of resistance in Australian cultivars (Bariana et al. 2001). No significant evolution in the stripe rust pathogen was detected for over a decade after the identification of Pst pathotype 110 E143A+ in 1986 (Wellings and McIntosh 1990). The Yr17 virulent pathotype 104 E137A− Yr17+ was detected in 1999 (Wellings 2007). This pathotype was avirulent on genotypes carrying stripe rust resistance genes Yr6, Yr7 and YrA and therefore a combination of these genes with Yr17 provided protection against predominant pathotypes in Australia (Bariana et al. 2004).
The wheat growing area in Western Australia is geographically separated from the eastern Australian wheat belt and remained free from stripe rust until the detection of Pst pathotype 134 E16A+ in 2002 (Wellings et al. 2003). This pathotype differed from eastern Australian Pst pathotypes in its virulence on seedling resistance genes Yr8 and Yr9 and its avirulence on Yr3, Yr4 and Yr34 and some other uncharacterised resistance sources. It also produced a significantly higher level of infection on many Australian cultivars previously reported to have moderate levels of adult plant resistance.
The Australian wheat cultivar Rubric (AUS33333), a selection from the CIMMYT genotype EMU ‘S’ (AUS17224), was released in 2002. It exhibited a high level of stripe rust resistance when tested under field conditions in 2003. Rubric showed susceptible seedling responses to the pre-2002 Pst pathotypes, whereas it showed a low seedling response against the Pst pathotype 134 E16A+. The seedling infection type (IT) of Rubric (IT;;C) was considerably lower than that expressed by WAWHT2046 (IT23C), which carries Yr34, another gene that confers seedling resistance to this pathotype. Genetic analysis was undertaken to study the mode of inheritance of seedling stripe rust resistance in cultivar Rubric. Bulked segregant analysis, using multiplex PCR technology, was performed to determine the genomic location of gene(s) involved.
Materials and methods
Generation of a segregating population
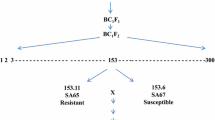
Cultivar Rubric was crossed with the susceptible line Avocet ‘S’. The F1 plants were grown during summer and F2 plants were space planted under rust free field conditions. A total of 89 F2 plants were successfully harvested and the F3 families were used for inheritance and genomic location studies.
Pathogen material
The Pst pathotypes used for seedling studies included: 110 E143A+ (444) and 134 E16A+ (572). The avirulence and virulence details of these pathotypes are described in Wellings (2007). The initial introduction of Pst pathotype, 104 E137A−, to Australia evolved to acquire virulence for Yr6 and YrA (108 E141A+), another single step mutation for virulence on Yr7 produced pathotype 110 E143A+ (Wellings 2007). The addition of A+ indicates virulence for the gene YrA. The post-2002 pathotype 134 E16A+ carries additional virulence for Yr8 and Yr9 when compared to the most virulent pre-2002 pathotype 110 E143A+. Pathotype 134 E16A+ also differs from 110 E143A+ for avirulence on Yr3, Yr4, Yr34 and some other uncharacterized sources of resistance.
Greenhouse screening
Twenty seeds of each F3 family were sown in 9 cm pots filled with a mixture of pinebark and sand. The parents Rubric and Avocet ‘S’ were included as controls. Ten grams of the water soluble fertiliser Aquasol® was dissolved in 10 l of tap water and applied to 100 pots. A single application of nitrogenous fertilizer urea was applied at the same rate as Aquasol to seven days old seedlings. Genotypes (with known resistance genes) Vilmorin 23 (Yr3), Hybrid 46 (Yr4), Avalon (Yr4) and WAWHT2046 (Yr6, Yr34) together-with Rubric and Avocet ‘S’ were sown in duplicate for evaluation against two pathotypes. Cultivars Bolac and Nesser showing response patterns similar to Rubric, were also included in this experiment. Seedlings were grown under rust free conditions in a temperature-controlled greenhouse maintained at 20°C before inoculation.
Twelve-day old Rubric/Avocet ‘S’-derived F2:3 seedlings (two leaf stage) were inoculated with urediniospores of the Pst pathotype 134 E16A+, whereas all other genotypes were inoculated with two pathotypes. Inoculated seedlings were incubated at 9–12°C for 24 h, under 100% relative humidity, and then moved to the greenhouse room maintained at 17 ± 2°C. Seedling responses were scored on a 0–4 scale as described in Bariana and McIntosh (1993). The F3 families were classified as non-segregating resistant (families showing uniform resistant response), segregating (families with both resistant and susceptible responses) and non-segregating susceptible (families with uniform susceptible response).
Molecular mapping
Genomic DNA was extracted from silica gel dried leaf tissue of each F3 family. Leaf sample for DNA isolation represented tissue from at least eight plants. Dried leaves were crushed using a stainless steel ball bearing in a Retsch MM300 Mixer Mill (Retsch, Germany) in 2 ml tubes. Seven hundred μl of pre-warmed (65°C) extraction buffer (50 mM Tris–HCl, 10 mM EDTA, 100 mM NaCl, 1% (w/v) SDS, 10 mM ß-mercaptoethanol, pH 8.0) was added to each tube. Samples were incubated at 65°C for 10 min and cooled at room temperature for 3 min. One hundred and fifty microliters of 3 M potassium acetate pH 5.2, was added and the tubes were shaken vigorously before cooling at −20°C for 20 min. The tubes were centrifuged at 13,000 rpm for 10 min and 700 μl of supernatant was transferred to fresh tubes. An equal volume of isopropanol was added and the tubes were inverted several times to mix supernatant with isopropanol. The tubes were then centrifuged at 10,000 rpm for 10 min to precipitate the DNA. The pellet was washed with 500 μl of 70% ethanol and air dried before re-suspension overnight in 200 μl of 10 mM Tris–HCl, pH 8.0. The tubes were centrifuged at 10,000 rpm for 5 min and 180 μl of supernatant containing the DNA was transferred to fresh tubes for storage at −20°C. The DNA samples were quantified using a Nanodrop ND-1000 Spectrophotometer (Nanodrop Technologies).
Bulk segregant analysis (BSA), as described by Michelmore et al. (1991), was used to determine the chromosomal location of the resistance gene(s) associated with low stripe rust response in the Rubric/Avocet ‘S’ cross. The resistant and susceptible bulks contained equal amounts of pooled DNA from nine resistant and nine susceptible F3 families, respectively. BSA was performed using a multiplex-ready PCR (Hayden et al. 2008) whole genome scan kit consisting of 488 published microsatellite (SSR) markers selected for high polymorphism in Australian wheat germplasm and genome coverage. Multiplex-ready PCR assays for BSA and genetic mapping were performed as described in Hayden et al. (2008). Electrophoresis and visualisation of the PCR products was performed on a GelScan2000 (Corbett Research) and ABI3730 DNA fragment analyser (Applied Biosystems). For analysis on the GelScan2000, the PCR products were mixed with an equal volume of gel loading dye (98% formamide, 10 mM EDTA, and 0.5% basic fuchsin as tracking dye), heated for 3 min at 95°C, chilled quickly on ice and separated on a 5% sequencing gel (Sambrook and Russell 2001). The 80 bp fluorescent DNA ladder was used to calculate allele sizes. For ABI3730 analysis, the procedure described by Hayden et al. (2008) was followed. SSR allele scoring was performed using GeneMapper v4.0 (Applied Biosystems). Four new primers mapped in the most distal deletion bin on 3BS (3BS3-0.87-1.00) from the physical map developed by Paux et al. (2008) were used to refine genomic location of the resistance gene. All these primers were distal to barc75.
Data analyses and genetic mapping
Chi-squared tests were used to determine the goodness-of-fit of observed segregation with expected genetic ratios. Recombination fractions were calculated with the MAP MANAGER version QTXb20 (Manly et al. 2001) using the Kosambi (1944) mapping function.
Results
Inheritance studies
Cultivar Rubric displayed a susceptible stripe rust response (IT3) when tested against pre-2002 Australian Pst pathotypes, and produced a resistant response (IT;;C) against the Pst pathotype 134 E16A+ (Table 1). The susceptible parent Avocet ‘S’ exhibited IT3+ against all pathotypes used. The Rubric/Avocet ‘S’ F3 families showed monogenic segregation (\( \chi^{2}_{1:2:1} = 0.06 \) non-significant at P = 0.05 and 1 df), when tested against the Pst pathotype 134 E16A+ (Table 2). The resistance gene conferring the low stripe rust response was tentatively designated YrRub.
Chromosome location and genetic mapping
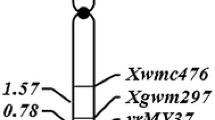
A whole genome scan kit consisting of 488 SSR markers was used to screen the parental lines and contrasting DNA bulks for loci associated with stripe rust response in the Rubric/Avocet ‘S’ cross. Nine markers revealed polymorphism between the parental lines and showed linkage with the resistant and susceptible DNA bulks. The entire F3 population was genotyped for the linked microsatellite markers. Four markers located on the short arm of chromosome 3B, namely barc75, gwm533, gwm493 and barc102 showed strong genetic association with resistance. Marker barc75 mapped closest to YrRub. In order to refine the location of YrRub in chromosome 3BS, four SSR markers (cpf3132, cfp140, cfb3530 and cfb3296) that mapped distal to barc75 were tested on parents and bulks. Marker cpf3132 was monomorphic, whereas markers cfp140, cfb3530 and cfb3296 were polymorphic and linked. Markers cfp140 and cfb3296 failed to amplify any product, when DNA from parent Avocet S was used, indicating the dominant inheritance of these markers. These markers were not mapped on the entire population as they were expected to be non-informative on 50% of heterozygous individuals of the F3 population. The co-dominant marker cfb3530 amplified 150 and 155 bp products, respectively, when DNA from Rubric and Avocet S were used. This marker was genotyped on the entire population. The marker cfb3530 mapped between YrRub and barc75 (Fig. 1). All markers conformed to the expected 1:2:1 segregation ratio. The LOD values for all of the linked markers were more than 30.0.
Genetic linkage map showing location of YrRub at the distal end of the chromosome 3BS (Kosambi map distances cM shown on the left side) and comparison of map from this study with that of the distal deletion bin 3BS3-0.87-1.00 of chromosome 3BS (Paux et al. 2008)
Tests on selected genotypes
Genotypes carrying known resistance genes Vilmorin 23 (Yr3), Hybrid 46 (Yr4) and WAWHT2046 (Yr6, Yr34) produced high infection types against the pre-2002 Pst pathotype 110 E143A+, whereas these genotypes produced infection types IT;1N, IT; and IT23C, respectively, against the Pst pathotype 134 E16A+ (Table 1). Cultivars Avalon, Bolac and Nesser produced infection types similar to that of Rubric. These genotypes and a set of current Australian wheat cultivars were genotyped using the SSR markers barc75 and cfb3530, and results are presented in Table 3. Rubric, Bolac, Nesser and EMU ‘S’ amplified the YrRub-linked barc75 132bp and cfb3530 150bp alleles. Stripe rust differential Hybrid 46 and the English wheat cultivar Avalon, both known to carry Yr4, amplified the YrRub-linked alleles at the barc75 and cfb3530 loci indicating that YrRub could be Yr4. Genotypes carrying Yr3, Yr34 and a set of current Australian wheat cultivars amplified PCR products different to that of Rubric (Table 3).
Discussion
Genetic analysis of stripe rust resistance in the Australian cultivar Rubric indicated the presence of a single gene tentatively designated as YrRub. Bulked segregant analysis indicated close genetic association of YrRub with the chromosome 3BS located marker barc75 (Fig. 1). Paux et al. (2008) published a physical map of chromosome 3BS and reported some additional markers that mapped distal to barc75 in the most distal deletion bin (3BS3-0.87-1.00). Marker gwm389 is the most distal marker (Fig. 1) and it segregated independently of YrRub. In contrast, marker cfb3530 mapped 2.9 ± 1.3 cM proximal to YrRub and 2.4 ± 1.2 cM distal to barc75 (Fig. 1). These results confirmed the location of YrRub in the most distal deletion bin of chromosome 3BS.
The possibility of YrRub being identical to one of the stripe rust resistance genes Yr3 and Yr4 and Yr34 that showed high and low responses against the pre- and post-2002 Australian Pst pathotypes, respectively, was considered. YrRub cannot be Yr34 as the latter was located in the chromosome arm 5AL (Bariana et al. 2006) and produces an infection type higher than that produced by YrRub (Table 1). Although location of Yr3 was listed in chromosome 5BL (Mclntosh et al. 1995), Chen et al. (1996) located it in chromosome 2B. Similarly Mclntosh et al. (1995) listed the location of Yr4 in chromosome 3B based on the 1987 Annual Report of the Plant Breeding Institute Cambridge, UK. Again Chen et al. (1996) located a gene thought to be Yr4 on chromosome 6B. These locations were never validated. Based on the amplification of the Rubric-specific PCR products at the cfb3530 and barc75 loci in genotypes known to carry Yr4 (Hybrid 46 and Avalon), we concluded that YrRub could be Yr4. Allelism test was not possible due to the absence of Yr4 carrying single gene stocks.
Chromosome 3BS carries durable stem rust resistance gene Sr2 (Hare and McIntosh 1979) and resistance to Fusarium head blight (Somers et al. 2003). The microsatellite marker gwm533 amplifies a 120 bp PCR product in Sr2 carrying genotypes (Spielmeyer et al. 2003). The absence of Sr2-linked physiological marker, pseudo black chaff, in Rubric and the amplification of a 143 bp allele at the gwm533 locus, suggested the absence of Sr2 in this cultivar. The amplification of non Sr2-linked alleles at marker loci (data not presented) stm559tcac, stm598tgag, stm560.3tgag and stm560tgag (Hayden et al. 2004, 2006) also indicated the absence of Sr2 in Rubric. YrRub may be linked closely in repulsion with Sr2. The location of the Sr2-linked marker gwm533 in the second deletion bin (Paux et al. 2008) and estimation of a genetic distance of approximately 10 cM between YrRub and gwm533 suggested that YrRub and Sr2 can be combined in a single genotype.
Validation of markers closely linked with commercially important traits such as disease resistance across diverse genetic backgrounds is essential to assure their robustness in marker assisted selection. Often, positive validation is performed on genotypes known to carry the target gene, however, negative validation is also necessary to show the absence of the gene-linked marker allele among commercial cultivars that are potential parents for marker assisted selection of the gene in question. A set of Australian cultivars and some additional genotypes were screened with markers barc75 and cfb3530. The barc75 132bp and cfb3530 150bp alleles were amplified in Rubric, Bolac, EMU S, Nesser and Yr4 carrying genotype Hybrid 46 and Avalon (Table 3). Australian wheat genotypes lacking YrRub and/or Yr4 carried allele(s) different from Rubric at the barc75 and cfb3530 loci (Table 3). These results demonstrated that barc75 and cfb3530 could be used for marker assisted selection of YrRub in breeding programs.
Although YrRub is not effective against the pre-2002 Pst pathotypes, it could still play an important role in gene combinations in Australia and elsewhere. It is effective against all five post-2002 pathotypes that are present in the eastern Australian wheatbelt (H·S. Bariana unpublished results). Redeployment of previously defeated rust resistance genes in gene combinations is not uncommon. Following the detection of virulence in stem rust pathogen for a widely deployed gene, Sr31, a previously defeated gene Sr36 did serve as a transient source of resistance against Ug99 in some cultivars. Virulence for Sr36 was detected in Australia in 1984 (Zwer et al. 1985), however, the current predominant pathotypes in Australia are avirulent on Sr36 (Park 2007). Similarly, the effectiveness of YrRub against post-2002 pathotypes supports the deployment of YrRub in strategically chosen gene combinations in new wheat cultivars.
This study clearly demonstrated the genomic location of YrRub in the short arm of chromosome 3BS. The presence of YrRub in CIMMYT (EMU S) and ICARDA (Nesser) germplasm was also demonstrated through marker genotyping. YrRub may have been transferred into CIMMYT and ICARDA germplasm through crosses involving winter wheats. Molecular markers linked with several stripe rust resistance genes (Yr10, Yr15, Yr24, Yr36 and Yr18) have been listed in Bariana et al. (2007). Although the use of combinations of APR genes for rust proofing the wheat industry globally will be desirable, strategic deployment of molecularly tagged seedling and adult plant resistance genes would ensure durability of stripe rust resistance in new wheat cultivars.
References
Bariana HS, McIntosh RA (1993) Cytogenetic studies in wheat XV Location of rust resistance genes in VPM1 and their genetic linkage with other resistance genes in chromosome 2A. Genome 36:476–482
Bariana HS, Cupitt CF, Warburton T (2001) Diversity of resistance to rust diseases in Australian wheats in 1999 and 2000 crop seasons. In: Eastwood R, Hollamby G, Rathjen A, Gororo N (eds) Proceedings of the 10th Assembly of Wheat Breeding Society of Australia. Mildura, pp 233–236
Bariana HS, Willey NJ, Venkata BP, Lehmenseik A, Standen GE, Lu M (2004) Breeding methodology to achieve durability for rust resistance in wheat. In: Black CK, Panozzo JF, Rebetzke GJ (eds) Proceedings of the 54th Australian Cereal Chemistry Conference and 11th Wheat Breeders Assembly. ACT, Canberra, pp 8–12
Bariana HS, Parry N, Barclay IR, Loughman R, McLean RJ, Shankar M, Wilson RE, Willey NJ, Francki M (2006) Identification and characterization of stripe rust resistance gene Yr34 in common wheat. Theor Appl Genet 112:1143–1148
Bariana HS, Brown GN, Bansal UK, Miah H, Standen GE, Lu M (2007) Breeding triple rust resistant wheat cultivars for Australia using conventional and marker-assisted selection technologies. Aust J Agric Res 58:576–587
Chen XM, Jones SS, Line RF (1996) Chromosomal location of genes for resistance to Puccinia striiformis in seven wheat cultivars with resistance genes at the Yr3 and Yr4 loci. Phytopathology 86:1228–1233
Hare RA, McIntosh RA (1979) Genetics and cytogenetics studies of durable adult-plant resistance in ‘Hope’ and related cultivars to wheat rusts. Z Pflanzenzücht 83:350–367
Hayden MJ, Kuchel H, Chalmers KJ (2004) Sequence tagged microsatellites for the Xgwm533 locus provide new diagnostic markers to select for the presence of stem rust resistance gene Sr2 in bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1641–1647
Hayden MJ, Stephenson P, Logojan AM, Khatkar D, Rogers C, Elsden J, Koebner RMD, Snape JW, Sharp PJ (2006) Development and genetic mapping of sequence-tagged microsatellites (STMs) in bread wheat (Triticum aestivum L.). Theor Appl Genet 113:1271–1281
Hayden MJ, Nguyen TM, Waterman A, McMichael GL, Chalmers KJ (2008) Application of multiplex-ready PCR for fluorescence-based SSR genotyping in barley and wheat. Mol Breeding 21:271–281
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Manly KF, Cudmore RH Jr, Meer JM (2001) Map Manager QTX, cross-platform software for genetic mapping. Mamm Genome 12:930–932
Mclntosh RA, Wellings CR, Park RF (1995) Wheat rusts: an atlas resistance genes. CSIRO Publishers, Australia
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions using segregation populations. Proc Natl Acad Sci USA 88:9828–9832
O’Brien L, Brown JS, Pascoe I (1980) Occurrence and distribution of wheat stripe rust in Victoria and susceptibility of commercial wheat cultivars. Australas Plant Pathol 9:14
Park RF (2007) Stem rust of wheat in Australia. Aust J Agric Res 58:558–566
Paux E, Sourdille P, Salse J, Saintenec C, Choulet F, Leroy P, Korol A, Michalak M, Kianian S, Spielmeyer W, Lagudah E, Somers D, Kilian A, Alaux M, Vautrin S, Berges H, Eversole K, Appels R, Safar J, Simkova H, Dolezel J, Bernard M, Feuillet C (2008) A physical map of the 1-gigabase bread wheat chromosome 3B. Science 322:101–104
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Somers DJ, Fedak G, Savard M (2003) Molecular mapping of novel genes controlling Fusarium head blight resistance and deoxynivalenol accumulation in spring wheat. Genome 46:555–564
Spielmeyer W, Sharp PJ, Lagudah ES (2003) Identification and validation of markers linked to broad spectrum stem rust resistance gene Sr2 in wheat (Triticum aestivum L.). Crop Sci 43:333–336
Wellings CR (2007) Puccinia striiformis in Australia: a review of the incursion, evolution and adaptation of stripe rust in the period 1979–2005. Aust J Agric Res 58:567–575
Wellings CR, McIntosh RA (1990) Puccinia striformis f. sp. tritici in Australasia: pathogenic changes during the first ten years. Plant Pathol 39:316–325
Wellings CR, Wright DG, Keiper F, Loughman R (2003) First detection of wheat stripe rust in Western Australia: evidence for a foreign incursion. Australas Plant Pathol 32:321–322
Zwer PK, Park RF, McIntosh RA (1985) Wheat stem rust in Australia 1969–1985. Aust J Agric Res 43:399–431
Acknowledgments
Financial support by the Grains Research Development and Corporation of Australia is highly acknowledged. We thank Profs. R. A. McIntosh and R. F. Park for reviewing the article.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bansal, U.K., Hayden, M.J., Gill, M.B. et al. Chromosomal location of an uncharacterised stripe rust resistance gene in wheat. Euphytica 171, 121–127 (2010). https://doi.org/10.1007/s10681-009-0007-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-009-0007-4