Abstract
Pm12, transferred from Aegilops speltoides (2n = 2x = 14, genome SS) to wheat, confers effective resistance to powdery mildew worldwide. By applying bulked segregant analysis in a BC3F2 segregating population consisting of 305 plants, 18 wheat genomic and EST-SSR markers linked to the resistance gene were identified. Pm12 was located in the 6SS portion of the T6BS-6SS.6SL translocation chromosome based on the physical bin positions of the genomic and EST-SSR markers in the Chinese Spring group six deletion stocks and their linkage relationship to the resistance gene. Twenty eight recombinants among 305 F2 plants indicated a low frequency of recombination between the alien chromosome segment and wheat chromosome 6B. Since recombination events occurred on both sides of Pm12, the materials generated provide opportunities for further reduction of alien chromatin by intercrossing selected individuals and using markers to select the required plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Powdery mildew, caused by Blumeria graminis f. sp. tritici, is one of the most important fungal diseases of wheat (Triticum aestivum L.) worldwide. Breeding for resistance to powdery mildew is the most economical and effective way to control the disease. However, evolution of the pathogen population often leads to virulence that enables the pathogen to overcome the host resistance, a situation made worse when single resistance genes are deployed in monocultures over wide areas. New and diversified sources of resistance to powdery mildew are needed to broaden the availability of resistant germplasm for wheat breeders. To date, 34 powdery mildew resistance gene loci have been characterized and assigned to specific chromosomes in common wheat (McIntosh et al. 2003; Hsam et al. 2003; Zhu et al. 2005; Miranda et al. 2006). Some of them were introduced from related species.
Diploid species with the S genome belong to the Aegilops section Sitopsis, and are part of the the secondary gene pool for common wheat improvement. The S genome is considered to be closely related to the B genome of T. aestivum. Ae. speltoides was proposed to be the B genome donor of both common and durum wheat (Huang et al. 2002; Kimber and Athwal 1972; Dvorak and Zhang 1990). It is the source of several disease resistance genes, including leaf rust resistance genes Lr28 (T4AS.4AL-7S#2S, McIntosh et al. 1982), Lr35 (Kerber and Dyck 1990), Lr36 (Dvorak and Knott 1990), Lr47 (Dubcovsky et al. 1998) and Lr51 (Dvorak 1977), stem rust resistance genes Sr32 (Friebe et al. 1996) and Sr39 (Kerber and Dyck 1990), and powdery mildew resistance genes Pm12 (Miller et al. 1988) and Pm32 (Hsam et al. 2003), that were transferred to wheat and are being used for wheat improvement.
Pm12 was first introgressed into hexaploid wheat cultivar Wembley in UK and a chromosome translocation involving group-6 chromosomes was identified (Miller et al. 1988). RFLP mapping indicated that the chromosome carrying Pm12 was a 6BS-6SS.6SL translocation (Jia et al. 1996), involving the entire long arm and most of the short arm of chromosome 6S. Pm12 was highly effective against powdery mildew isolates in China (Duan et al. 1998), Europe (Huang and Röder 2004) and the United States (Niewoehner and Leath 1998). However, due to the large amount of alien chromatin, lines with Pm12 appear to have yield depression and the gene has not contributed to commercial wheat cultivar production.
Molecular markers have been widely used to map powdery mildew resistance genes in wheat. So far, 18 of 34 wheat powdery mildew resistance genes have been mapped by RFLP, RAPD, AFLP and SSR approaches (Huang and Röder 2004; McIntosh et al. 2003, Hsam et al. 2003; Zhu et al. 2005, Miranda et al. 2006). Microsatellite markers, or simple sequence repeats (SSRs), have the advantages of being inexpensive, safe and easy to handle compared to RFLP markers. In the last decade, genetic (Röder et al. 1998b; Stephenson et al. 1998; Pestsova et al. 2000; Gupta et al. 2002; Song et al. 2002; Somers et al. 2004) and physical (Sourdille et al. 2004) maps of wheat microsatellites have been developed, and they greatly facilitate the molecular tagging of agronomically important genes. Several powdery mildew resistance genes, such as Pm5e (Huang et al. 2003), Pm24 (Huang et al. 2000), Pm27 (Järve et al. 2000), Pm30 (Liu et al. 2002), Pm33 (Zhu et al. 2005) and MlZec1 (Mohler et al. 2005) were mapped with microsatellite markers. Recently, a new kind of microsatellite marker, EST-SSR, derived from expressed sequence tags (ESTs) and more likely physically associated with the coding regions of the genome, provided additional PCR-based markers that could be used for genome mapping, and marker assisted selection (Cho et al. 2000; Scott et al. 2000; Eujayl et al. 2002, Yu et al. 2004).
In the present paper, we report (1) the genomic SSR and EST-SSR mapping of the powdery mildew resistance gene Pm12, and (2) the molecular identification of Pm12 introgression lines containing smaller segments of Ae. speltoides chromosome 6S.
Materials and methods
Plant materials
The Wembley line carrying Pm12, Line #31, was kindly provided by Dr. XY Duan, Chinese Academy of Agricultural Sciences (originally from Dr. MD Gale, JI Centre, Norwich, UK). Susceptible elite commercial common wheat lines 87-1, Jing 411, Nongda 101, Jin 190, and Jin 207 were used as parents in crosses with Line #31 and then used as recurrent parents for backcrossing to improve the agronomic traits. Five resistant plants of each cross were selected in each cycle to make further backcrosses with the recurrent parents. A BC3F2 population derived from Line #31/3*Jin 207 was chosen to construct a linkage map. A total of 305 BC3F2 plants and their progenies were used for powdery mildew evaluations and microsatellite analysis. Five resistant and 5 susceptible plants selected from the progenies of Line #31/8*Jing 411, Line #31/6*87-1, Line #31/3*Nongda 101 and Line #31/3*Jin 190 were used for marker validation.
Chinese Spring (CS) nullisomic-tetrasomics, ditelosomics and deletion lines of homoeologous group 6 (kindly provided by Drs. WJ Raupp and BS Gill, Wheat Genetics Resource Centre, Kansas State University, USA) were used for chromosomal arm assignments and physical mapping of the microsatellite markers.
Powdery mildew evaluations
The powdery mildew seedling reactions of the BC3F2 mapping population and BC3F3 progenies were determined as described by Liu et al. (2002). A local prevailing isolate of B. graminis f. sp. tritici E09 was used for inoculation. Reactions were scored on a 0, 0;, and 1 to 4 infection type (IT) scale, with 0 representing no visible symptoms, 0; representing necrotic flecks, and 1, 2, 3, 4 for highly resistant (necrosis with low sporulation), resistant (necrosis with medium sporulation), susceptible (no necrosis with medium to high sporulation), and highly susceptible (no necrosis with full sporulation) reactions, respectively. The reaction of each BC3F3 line was based on 20 seedlings and the reaction of each F2 plant was scored homozygous resistant (RR), heterozygous resistant (Rr) and homozygous susceptible (rr).
Genomic DNA isolation and microsatellite analysis
Genomic DNA was extracted from uninfected leaves by the cetyltrimethylammonium bromide method (Saghai-Maroof et al. 1984). Resistant and susceptible DNA pools were constructed by separately bulking equal amounts of DNA from 10 homozygous resistant (IT 0) and 10 homozygous susceptible (IT 4) plants.
Microsatellite markers from homoeologous group-6 chromosomes were used to screen the two DNA pools to find polymorphic markers. They were divided into two groups: genomic-SSRs, including GWM (Röder et al. 1998a, b), GDM (Pestsova et al. 2000), BARC (Song et al. 2002), CFD (Guyomarc’h et al. 2002), WMC (Gupta et al. 2002), and EST-SSRs, including PK (Gupta et al. 2003), KSUM (Yu et al. 2004) and CAU. The CAU primers were designed according to flanking sequences of microsatellite motifs in wheat ESTs from public EST databases in GenBank (Table 1). Each PCR reaction was performed in a total volume of 10 μl containing 10 mM Tris–HCl, pH 8.3, 50 mM KCl, 1.5 mM MgCl2, 0.2 mM of dNTP, 25 ng of each primer, 50–100 ng genomic DNA and 0.75 U Taq DNA polymerase. Amplifications were performed at 94°C for 3 min, followed by 45 cycles at 94°C for 1 min, 50–60°C for 1 min, and 72°C for 2 min, with a final extension at 72°C for 10 min. PCR products were separated on 8% non-denaturing polyacrylamide gels for the mapping population analysis and 5% denaturing polyacrylamide gels for chromosome arm assignments. Gels were silver stained and photographed.
Linkage analysis
Linkage between Pm12 and linked markers was determined using MAPMAKER (Lander et al. 1987), with an LOD threshold of 3.0. The Kosambi mapping function was used to calculate map distances (Kosambi 1944).
Physical mapping of the SSR and EST-SSR markers
Genomic SSR and EST-SSR markers linked to Pm12 were located using a set of Chinese Spring nullisomic-tetrasomics, ditelosomics and deletion lines. Each marker was mapped to a chromosome bin flanked by breakpoints of the largest deletion possessing the fragment and the smallest deletion lacking it after comparing the amplification patterns. A consensus deletion map of chromosome 6 was constructed by placing the breakpoints of 11 deletion lines on a hypothetical chromosome drawn to scale on the basis of the mean length of group-6 chromosomes. The deletion mapping data of each marker on the three homoeologous chromosomes was then combined to position the marker to the shortest chromosome interval.
Results
Genetic analysis of powdery mildew resistance gene Pm12
Line #31 and its derivatives remained highly resistant to race E09 (no symptoms or hypersensitive reaction was observed), after crossing and backcrossing with Chinese common wheat lines. The Line #31/3*Jin 207 BC3F2 population segregated 206 resistant (IT 0 or 0;) and 99 susceptible (IT 4) (χ2 3:1 = 9.27, P < 0.01) and the BC3F3 progenies segregated 54 homozygous resistant, 152 heterozygous resistant and 99 homozygous susceptible (χ2 1:2:1 = 15.92, P < 0.01). The significant deviations from 3:1 and 1:2:1 ratios reflected probable low male transmission of the 6B-6S alien translocation chromosome carrying Pm12.
Identification of genomic SSR and EST-SSR markers linked to the resistance gene
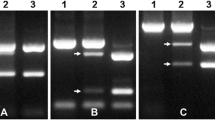
Of 182 genomic SSR and EST-SSR markers for homoeologous group-6 chromosomes, 18 SSR markers were polymorphic and linked to Pm12. These markers included: Xbarc198, Xbarc247, Xbarc1030, Xbarc1169, Xcau24, Xcau44, Xcau127, Xcau186, Xcau196, Xcfd13, Xcfd80, Xcfd190, Xgwm361, Xgwm570, Xgdm127, Xksum102, Xpk69 and Xwmc105 (Fig. 1). Four markers, Xbarc198, Xcfd80, Xcfd190, and Xgdm127, co-segregated with the resistance locus. Xcfd13 mapped 3.7 cM distal to Pm12, whereas the remaining markers were tightly linked to Pm12, revealing minimal recombination between wheat chromosome 6B and the Aegilops speltoides segment.
Physical mapping of the resistance gene Pm12 and its linked SSR markers
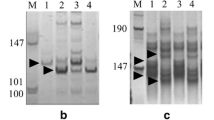
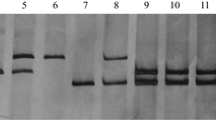
Among the genomic and EST SSR markers, Xcfd13, Xcfd190, Xbarc198 and Xwmc105 were physically mapped to the short arm; Xbarc1030 and Xgwm361 were assigned to the long arms of homoeologous group 6 chromosomes (Sourdille et al. 2004). Based on the amplification patterns in Chinese Spring and its nullisomic-tetrasomics, ditelosomics and deletion lines, the rest of the Pm12-linked genomic SSR and EST-SSR markers were mapped to chromosome 6B and physically assigned to specific bins. For example, Xcau127 amplified 3 DNA fragments in Chinese Spring and susceptible plants, but only 2 DNA fragments in Chinese Spring N6AT6B, N6BT6A, N6DT6A, Line #31 and resistant plants (Fig. 2). The absence of a 153 bp DNA fragment in Chinese Spring N6BT6A, Line #31 and resistant plants indicated that the Pm12 translocation involved chromosome 6B. Six microsatellite loci, Xcau44, Xcau127, Xcau186, Xcfd13, Xcfd190 and Xgwm570, were detected loci on all three homoeologous chromosomes 6A, 6B and 6D. Xcau196 and Xpk69 amplified loci on 6A and 6B, Xcau24 and Xbarc1169 detected loci on 6B and 6D, Xcfd80 detected loci on 6A and 6D. However, Xbarc1030 and Xgdm127 identified loci only on chromosome 6D and Xbarc198, Xbarc247, Xgwm361, Xksum102, and Xwmc105 detected loci only on chromosome 6B. Overall, of 18 polymorphic loci, 18, 16, 11 and 9 loci were located on 6S, 6B, 6A and 6D, respectively, indicating a relative closer relationship between 6S and 6B than between 6S and 6A or 6S and 6D. After combining the deletion mapping results from three homoeologous chromosomes, a consensus physical mapping of Pm12-linked microsatellite markers was constructed and each marker was assigned to the shortest possible chromosome interval (Fig. 3). Xcfd13, Xbarc1169 and Xksum102 were physically mapped to the distal telomeric region, along with Xcau127, Xbarc198, Xcfd190, Xgdm127 to the middle region, and Xwmc105 and Xcfd80 to the proximal region of the short arm. It was also shown that Xcau196, Xcau186, Xcau24, Xbarc247, Xcau44, Xgwm570 and Xpk69 were located on the distal telomeric region, along with Xbarc1030 and Xgwm361 to the middle part of the long arm.
Recombination between wheat 6B and Ae. speltoides 6S chromosomes
The telomeric physical locations of Xcfd13 and Xcau24 and their close genetic distances to Pm12 (3.7 cM and 1.7 cM, respectively) revealed severe recombination suppression between wheat chromosome 6B and Ae. spelotides chromosome 6S as expected for a large alien translocation. However, eight recombination breakpoints were observed between wheat chromosome 6B and Ae. speltoides chromosome 6S, resulting in 28 recombinants divided into eight classes. Three breakpoints close to markers Xcfd13, Xcau127 and Xwmc105 were located on the short arm while another 5 breakpoints were assigned to the long arm (Fig. 4). Among the recombination events, 18 occurred near Xcfd13; 4 close to Xcau186; and single recombinations were observed beside Xcau24, Xcau44, Xcau127, Xbarc1030, Xbarc1169, Xksum102, Xpk69 and Xwmc105 (Fig. 4). One heterozygous resistant plant, 1338-8, lacked the entire long arm of chromosome 6S. This plant was self-pollinated to obtain 4 homozygous resistant plants containing a much shorter segment of chromosome 6S with the aid of genomic and EST SSR markers.
Comparative organization of recombinants between chromosomes T6BS-6SS.6SL and 6B. The extent of 6B chromosome segments in Jin 207 (blank) and Line# 31 (gray) and 6S chromosome segments (black) in Line #31 and each recombinant is shown. Blank circles represent the centromeres. The breakpoint positions may lie anywhere between two adjacent SSR marker loci. The numbers on the left and right sides of chromosomes represent numbers and names of recombinants, respectively. The Pm12 gene lies between Xcau127 and Xwmc105
Genomic and EST SSR markers linked to Pm12 were validated using 5 resistant and 5 susceptible plants selected from crosses Line #31/8*Jing 411, Line #31/6*87-1, Line #31/3*Nongda 101 and Line #31/3*Jin 190. No recombinants were found between the marker loci and Pm12 among this set of materials. The lack of recombination was attributed to the small numbers of plants used in backcrossing and progeny testing.
Discussion
Genetic and physical mapping of powdery mildew resistance gene Pm12
Gene Pm12 was introduced into common wheat by crossing variety Wembley and Ae. speltoides and backcrossing 5 times with Wembley. The resistance gene was initially located on chromosome 6A by monosomic analysis (Miller et al. 1988). Using RFLPs, however, Pm12 was mapped to a 6BS-6SS.6SL translocation chromosome with the break-point proximal to Xpsr551-6B (Jia et al. 1996). SSR mapping confirmed that the translocation involved wheat chromosome 6B. Four SSR markers, Xbarc198, Xgdm127, Xcfd190, and Xcfd80, co-segregated with Pm12. However, the physical bins with these markers span most of the 6SS chromosome arm (76%). Xwmc105 was located on group 6 close to the centromere (Sourdille et al 2004). A recombination between Xwmc105 and Pm12 was identified in the mapping population. Hence the likely physical position of Pm12 is between Xwmc105 and Xcau127 on 6SS (Fig. 4).
Recombination between the wheat 6B and Ae. speltoides 6S chromosomes
It has been suggested that Ae. speltoides (SS genome) contributed the B genome to tetraploid and hexaploid wheats (Sarkar and Stebbins 1956; Riley et al. 1969); other researchers argued against Ae. speltoides as the likely donor (Kimber and Athwal 1972; Dvorak and Zhang 1990; Huang et al. 2002). In any case, no recombination was observed between wheat chromosome 6B and the 6S segment in Line #31 by Jia et al. (1996) and a very low frequency of recombination (5.4 cM) between distally located markers in group 6 chromosomes was detected in the present work (Fig. 1). Only 28 recombinants were identified in 305 plants (610 gametes); 18 involved Xcfd13 and this polymorphism probably represented a difference between Wembley (the original wheat parent) and Jin 207. Unfortunately Wembley was not included in the study. Because of their ordered groupings, it appears likely that the other 10 recombinants involved 6B-6S recombination events.
Pm12 is highly effective and confers powdery mildew resistance worldwide (Duan et al. 1998; Niewoehner and Leath 1998; Huang and Röder 2004). However, it has not been used in any cultivar because of poor agronomic traits probably associated with the alien segment. A smaller alien segment should be achieved by crossing recombinants 1336-7 and 1338-8 and selecting Pm12 and wheat alleles at Xcau127 and Xwmc105 as was done with rye and wheat recombinants by Lukaszewski (2000).
References
Cho YG, Ishii T, Temnykh S et al (2000) Diversity of microsatellite derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100:713–722
Duan XY, Sheng BQ, Zhou YL et al (1998) Monitoring of the virulence population of Erysiphe graminis f.sp. tritici. Acta Phytophylactica Sin 25:31–36
Dubcovsky J, Lukaszewski AJ, Echaide M et al (1998) Molecular characterization of two Triticum speltoides interstitial translocations carrying leaf rust and greenbug resistance genes. Crop Sci 38:1655–1660
Dvorak J (1977) Transfer of leaf rust resistance from Aegilops speltoides to Triticum aestivum. Can J Genet Cytol 19:133–141
Dvorak J, Knott DR (1990) Location of a Triticum speltoides chromosome segment conferring resistance to leaf rust in Triticum aestivum. Genome 33:892–897
Dvorak J, Zhang HB (1990) Variation in repeated nucleotide sequences sheds light on the phylogeny of the wheat B and G genomes. Proc Natl Acad Sci USA 87:9640–9644
Eujayl I, Sorrells ME, Baum M et al (2002) Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor Appl Genet 104:399–407
Friebe B, Jiang J, Raupp WJ et al (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87
Gupta PK, Balyan HS, Edwards KJ et al (2002) Genetic mapping of 66 new microsatellite (SSR) loci in bread wheat. Theor Appl Genet 105:413–422
Gupta PK, Rustgi S, Sharma S et al (2003) Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Mol Gen Genomics 270:315–323
Guyomarc′h H, Sourdille P, Charmet G et al (2002) Characterization of polymorphic microsatellite markers from Aegilops tauschii and transferability to the D genome of bread wheat. Theor Appl Genet 104:1164–1172
Hsam SLK, Lapochkina IF, Zeller FJ (2003) Chromosomal location of genes for resistance to powdery mildew in common wheat (Triticum aestivum L. em Thell.). 8. Gene Pm32 in a wheat-Aegilops speltoides translocation line. Euphytica 133:367–370
Huang XQ, Hsam SLK, Zeller FJ et al (2000) Molecular mapping of the wheat powdery mildew resistance gene Pm24 and marker validation for molecular breeding. Theor Appl Genet 101:401–414
Huang XQ, Röder MS (2004) Molecular mapping of powdery mildew resistance genes in wheat: a review. Euphytica 137:203–223
Huang SX, Sirikhachornkit A, Su XJ et al (2002) Genes encoding plastid acetyl-CoA carboxylase and 3-phosphoglycerate kinase of the Triticum/Aegilops complex and the evolutionary history of polyploid wheat. Proc Natl Acad Sci USA 99:8133–8138
Huang XQ, Wang LX, Xu MX et al (2003) Microsatellite mapping of the powdery mildew resistance gene Pm5e in common wheat (Triticum aestivum L.) Theor Appl Genet 106:858–865
Järve K, Peusha HO, Tsymbalova J et al (2000) Chromosomal location of a Triticum timopheevii-derived powdery mildew resistance gene transferred to common wheat. Genome 43:377–381
Jia JZ, Devos KM, Chao S et al (1996) RFLP-based maps of the homoeologous group-6 chromosomes of wheat and their application in the tagging of Pm12, a powdery mildew resistance gene transferred from Aegilops speltoides to wheat. Theor Appl Genet 92:559–565
Kerber ER, Dyck PL (1990) Transfer to hexaploid wheat of linked genes for adult-plant leaf rust and seedling stem rust resistance from an ampliploid of Aegilops speltoides × Triticum monococcum. Genome 33:530–537
Kimber G, Athwal RS (1972) A reassessment of the course of evolution of wheat. Proc Natl Acad Sci USA 69:912–915
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Lander ES, Green P, Abrahamson J et al (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Liu ZY, Sun QX, Ni ZF et al (2002) Molecular characterization of a novel powdery mildew resistance gene Pm30 in wheat originating from wild emmer. Euphytica 123:21–29
Lukaszewski AJ (2000) Manipulation of the 1RS.1BL translocation in wheat by induced homoeologous recombination. Crop Sci 40:216–225
McIntosh RA, Miller TE, Chapman V (1982) Cytogenetical studies in wheat XII. Lr28 for resistance to Puccinia recondita and Sr34 for resistance to P. graminis tritici. Z Pflanzenzuchtung 89:295–306
McIntosh RA, Yamazaki Y, Devos KM (2003) Catalogue of gene symbols for wheat. In: Pogna NE et al (eds) Proc 10th Int Wheat Genet Symp. Pasetum, Italy, vol 4. Istituto Sperimentale per la Cerealicoltura, Rome, Italy
Miller TE, Reader SM, Ainsworth CC et al (1988) The introduction of a major gene for resistance to powdery mildew of wheat, Erysiphe graminis f. sp. tritici, from Aegilops speltoides into wheat, Triticum aestivum. In: Jorna M, Shootmaker L (eds) Cereal breeding related to integrated cereal production: proceedings of the EUCARPIA conference. Pudoc, Wageningen, The Netherlands, pp 179–183
Miranda LM, Murphy JP, Marshall D et al (2006) Pm34: a new powdery mildew resistance gene transferred from Aegilops tauschii Coss. to common wheat (Triticum aestivum L.). Theor Appl Genet 113:1497–1504
Mohler V, Zeller FJ, Wenzel G et al (2005) Chromosomal location of genes for resistance to powdery mildew in common wheat (Triticum aestivum L. em Thell.). 9. Gene MlZec1 from the Triticum dicoccoides-derived wheat line Zecoi-1. Euphytica 142:161–167
Niewoehner AS, Leath S (1998) Virulence of Blumeria graminis f. sp. tritici on winter wheat in the eastern United States. Plant Dis 82:64–68
Pestsova E, Ganal MW, Röder MS (2000) Isolation and mapping of microsatellite markers specific for the D genome of bread wheat. Genome 43:689–697
Riley R, Unrau J, Chapman V (1969) Evidence on the origin of the B genome of wheat. J Heredity 49:90–98
Röder MS, Korzun V, Gill BS et al (1998a) The physical mapping of microsatellite markers in wheat. Genome 41:278–283
Röder MS, Korzun V, Wendehake K et al (1998b) A microsatellite map of wheat. Genetics 149:2007–2023
Saghai-Maroof MA, Soliman KM, Jorgensen RA et al (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal locations and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sarkar P, Stebbins GL (1956) Morphological evidence concerning the origin of the B genome in wheat. Am J Bot 43:297–304
Scott KD, Eggler P, Seaton G et al (2000) Analysis of SSRs derived from grape ESTs. Theor Appl Genet 100:723–726
Somers DJ, Isaac P, Edwards K (2004) A high density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Song QJ, Fickus EW, Cregan PB (2002) Characterization of trinucleotide SSR motifs in wheat. Theor Appl Genet 104:286–293
Sourdille P, Singh S, Cadalen T et al (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.) Funct Integr Genomics 4:12–25
Stephenson P, Bryan G, Kirby J et al (1998) Fifty new microsatellite loci for the wheat genetic map. Theor Appl Genet 97:946–949
Yu JK, Dake TM., Singh S et al (2004) Development and mapping of EST-derived simple sequence repeat markers for hexaploid wheat. Genome 47:805–818
Zhu ZD, Zhou RH, Kong XY et al (2005) Microsatellite markers linked to 2 powdery mildew resistance genes introgressed from Triticum carthlicum accession PS5 into common wheat. Genome 48:585–590
Acknowledgements
The authors are grateful to Dr. R. McIntosh for improving the manuscript. This work was financially supported by the National Fund for Distinguished Young Scholars (30425039), National Natural Science Foundation of China (30200174, 30571151), Fok Ying Tung Education Foundation (94021) and the State High Tech Programs (2006AA100102, 2006AA10Z1E9, 2006AA10Z1C4).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Wei Song and Hao Xie are contributed equally to this work.
Rights and permissions
About this article
Cite this article
Song, W., Xie, H., Liu, Q. et al. Molecular identification of Pm12-carrying introgression lines in wheat using genomic and EST-SSR markers. Euphytica 158, 95–102 (2007). https://doi.org/10.1007/s10681-007-9432-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9432-4