Summary
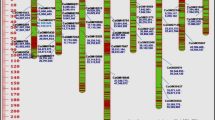
Microsatellites or simple sequence repeats (SSR) are the markers of choice for molecular genetic mapping and marker-assisted selection in many crop species. A microsatellite-based linkage map of cassava was drawn using SSR markers and a F2 population consisting of 268 individuals. The F2 population was derived from selfing the genotype K150, an early yielding genotype from an F1 progeny from a cross between two non-inbred elite cassava varieties, TMS 30572 and CM 2177-2 from IITA and CIAT respectively. A set of 472 SSR markers, previously developed from cassava genomic and cDNA libraries, were screened for polymorphism in K150 and its parents TMS 30572 and CM 2177-2. One hundred and twenty two polymorphic SSR markers were identified and utilized for linkage analysis. The map has 100 markers spanning 1236.7 cM, distributed on 22 linkage groups with an average marker distance of 17.92 cM. Marker density across the genome was uniform. This is the first SSR based linkage map of cassava and represents an important step towards quantitative trait loci mapping and genetic analysis of complex traits in M. esculenta species in national research program and other institutes with minimal laboratory facilities. SSR markers reduce the time and cost of mapping quantitative trait loci (QTL) controlling traits of agronomic interest, and are of potential use for marker-assisted selection (MAS).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Akkaya, M.S., A.A. Bhagwat & P.B. Cregan, 1992. Length polymorphisms of simple sequence repeat DNA in soybean. Genetics 132: 1131–1139.
Akkaya, M.S., R.C. Shoemaker, J.E. Specht, A.A. Bhagwat & P.B. Cregan, 1995. Integration of simple sequence repeat DNA markers into a soybean linkage map. Crop Sci 35: 1439–1445.
Allard, R.W., 1988: Genetic changes associated with the evolution of adaptedness in cultivated plants and their wild progenitors. J Hered 79: 225–238.
Awoleye, F., M. Van Duren, J. Dolezel & F.J. Novak, 1994. Nuclear DNA content and in vitro induced somatic polyploidization (Manihot esculenta Crantz) cassava breeding. Euphytica 76: 195–202.
Bennet, M.D., J.B. Smith & J.S. Heslop-Harrison, 1992. Nuclear DNA amounts in Angiosperms. Proc R Soc London Ser B 216: 179–-199.
Berloo, R. & P. Stam, 1998. Marker assisted selection in autogamous RIL populations: a simulation study. Theor Appl Genet 96: 147–154.
Best, R. & G. Henry, 1992. Cassava: towards the year 2000. In International network for cassava genetic resources. Report of the first meeting of the International Network for Cassava Genetics International Crop Network Series number 10. International plant Genetic resources institute (IPGRI). 10: 3–11.
Bryne, M., J.C. Murrell, J.V. Owen, A. Kriedemann, E.R. Williams & G.F. Moran, 1997. Identification and mode of action of quantitative trait loci affecting seedling height and leaf area in Eucalyptus nitens. Theor Appl Genet 94: 674–681.
Castiglioni, P., P. Ajmone-Marsan, R. Van Wijk & M. Motto, 1999. AFLP markers in amolecualr linkage map of maize: codominant scoring and linkage group distribution. Theor Appl Genet 99: 425–431.
Ceballos, H., C.A. Iglesias, J.C. Perez & A.G.O. Dixon, 2004. Cassava breeding: opportunities and challenges. Plant Molecular Biology 56: 503–516.
Churchill, G.A. & R.W. Doerge, 1994. Empirical threshold values for quantitative trait mapping. Genetics 138: 963–971.
Cock, J.H., 1985. Cassava:new potential for a neglected crop. Westview press, Boulder Colorado USA.
Dellarporta, S.L., J. Wood & J.R. Hicks, 1983. A plant DNA minipreparation: version II. Plant Mol Biol Rep 1: 19–21.
Dettori, M.T., R. Quarta & I. Verde, 2001. A peach linkage map integrating RFLPs, SSRs, RAPDs, and morphological markers. Genome 44: 783–-790.
Doege, R.W., 1993: Statistical methods for locating quantitative trait loci with molecular markers. PhD dissertation. North Carolina State University.
Eck van, H.J., J.M.E. Jacobs, P. Stam, J. Ton, W.J. Stiekema & E. Jacobsen, 1994. Multiple alleles for tuber shape in diploid potato detection by qualitative and quantitative genetic analysis using RFLPs. Genetics 137: 303–309.
El-Sharkawy, M.A. & J.H. Cock, 1990. Photosynthesis of cassava (Manihot esculenta). Expl Agric 26: 325–340.
Fregene, M., E. Okogbenin, C. Mba, F. Angel, M.C. Suarez, J. Guttierez, P. Chavarriaga, W. Roca, M. Bonierbale & J. Tohme, 2001. Genome mapping in cassava improvement: Challenges, achievements and opportunities. Euphytica 120:159–165.
Fregene, M., F. Angel, R. Gomez, F. Rodriguez, P. Chavariaga, W. Roca, J. Tohme & M. Bonierbale, 1997. A molecular genetic map of cassava. Theor Appl Genet 95: 431–441.
Grattapaglia, D., F.L.G. Bertolicci & R. Sederoff, 1995. Genetic mapping of QTLs controlling vegetative propagation in Eucalyptus grandis and E. urophylla using a pseudo-testcross mapping strategy and RAPD markers. Theor Appl Genet 90: 933–947
Grattapaglia, D. & R. Sederoff, 1994. Genetic linkage maps of Eucalyptus grandis and E. urophylla using a pseudo-testcross mapping strategy and RAPD markers. Genetics 137: 1121–1137.
Groover, A., M. Devey, T. Fiddler, J. Lee, R. Megraw, T. Mitchel-Olds, B. Sherman, S. Vujcic, C. Williams & D. Neale, 1994. Identification of quantitative trait loci influencing wood specific gravity in an outbred pedigree of loblolly pine. Genetics 138: 1293–1300.
Gruneberg, J.B., 1938. An analysis of the pleiotropic effects of a new lethal mutation in the rat (Mus norvegicus). Proc R Soc Lond B 125: 123–144.
Hamwieh, A., S.M. Udupa, W. Choumane, A. Sarker, F. Dreyer, C. Jung & M. Baum, 2005. A genetic linkage map of Lens sp. Based on microsatellite and AFLP markers and the localization of fusarium vascular kilt resistance. Theor Appl Gent 110: 669–677.
Hanley, S., J.H.A. Barrer, J.W. Van Ooijen, C. Aldam, S.L. Harris, I. Ahman, S. Larsson & A. Kart, 2002. A genetic linkage map of willow (Salix viminalis) based on two Lycopersicon esculentum x L. pennellii F2 populations. Theor Appl Genet 99: 254–271.
Jarret, R.L. & N. Bowen, 1994. Simple sequence repeats (SSRs) for sweet potato germplasm characterization. Plant Genet Res Newslet 100: 9–11.
Jorge, V., M. Fregene, C.M. Velez, M.C. Duque, J. Tohme & V. Verdier, 2001. QTL analysis of field resistance to Xanthomonas axonopodis pv manihotis in cassava Theor. Appl Genet 102: 564–571.
Jorge, V., M.A. Fregene, M.C. Duque, M.W. Bonierbale, J. Tohme & Verdier, 2000. Genetic mapping of resistance to bacterial blight disease in cassava (Manihot esculenta Crantz). Theor Appl Genet 101: 865–872.
Kawano, K., 1990. Harvest index and evolution of major food crop cultivars in the tropics. Euphytica 46: 195–202.
Kosambi, D.D., 1944. The estimation of map distances from recombination values. Ann Eugen 12: 172–175.
Kubisiak, T.L., C.D. Nelson, W.L. Nance & M. Stine (1995). RAPD linkage mapping in a longleaf pine x slash pine F1 family. Theor appl Genet 90:1119–1127.
Lander, E.S. & D. Botstein, 1989. Mapping Mendelian factors underlying quantitative traits by using RFLP linkage maps. Genetics 121: 185–199.
Lander, E.S., P. Green, J. Abraham, A. Barlow, M.J. Daly, S.E. Lincoln & L. Newburg, 1987. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1: 174–181.
Liebhard, R., L. Gianfrancesschi, B. Koller, C.D. Ryder, R. Tarchini, E. Van de Weg & C. Gessler, 2002. Development and characterization of 140 new microsatellites in apple (Malus x domestica Borkh). Mol Breed 10: 217–241.
Lin, S.Y., T. Sasa & M. Yano, 1998. Mapping quantitative trait loci controlling seed dormancy and heading date in rice, Oryza sativa, using backcross inbred lines. Theor Appl Genet 96: 997–1003.
Mann, C., 1997. Reseeding the green revolution. Science 277: 209 –220.
Mba, R.E.C., P. Stephenson, K. Edwards, S. Melzer, J. Mkumbira, U. Gullberg, K. Apel, M. Gale, J. Tohme & M. Fregene, 2001. Simple sequence repeat (SSR) markers survey of the cassava (Manihot esculenta Crantz) genome:towards an SSR-based molecular genetic map of cassava. Theor Appl Genet 102: 21–31.
Mogante, M., & A.M. Olivieri, 1993. PCR-amplified microsatellite markers in plant genetics. Plant J 3: 393–427.
Murranty, H., 1996. Power of tests for quantitative trait loci detection using full sib families in different schemes. Heredity 76:156–165.
Nweke, F.I., A.G.O. Dixon, R. Asiedu & S.A. Folayan, 1994. Cassava varietal needs of farmers and potential for production growth in Africa. COSCA working paper 10.
Okogbenin, E. & M. Fregene, 2002. Genetic and QTL mapping of early root bulking in an F1 mapping population of non-inbred parents in cassava (Manihot esculenta Crantz). Theor Appl Genet 106: 58–66.
Okogbenin, E., & M. Fregene, 2003. Genetic mapping of QTLs affecting productivity and plant architecture in a full-sib cross from non-inbred parents in cassava (Manihot esculenta Crantz). Theor Appl Genet 107: 1452–1462.
Olsen, K.M., & B.A. Schaal, 2001. Microsatellite variation in cassava (Manihot esculenta, Euphorbiacea) and its wild relatives: further evidence for a southern Amazonian origin of domestication. American Journal of Botany 88(1): 131–142.
Patterson, A.H., E.S. Lander, J.D. Hewitt, S. Peterson, S.E. Lincoln & S.D. Tanksley, 1988. Resolution of quantitative traits into Medelian factors, using a complete linkage map of restriction length polymorphisms. Nature 335: 721–726.
Patterson, A.H., S. Damon, J.D. Hewitt, D. Zamir, H.D. Rabinowitch, S.E. Lincoln, E.S. Lander & S.D. Tanksley, 1991. Mendelian factors underlying quantitative traits in tomato:comparison across species, generations and environments. Genetics 127: 181– 197.
Plaschke, J., M.W. Ganal, & M.S. Roder, 1995. Detection of genetic diversity in closely related bread wheat using microsatellite markers Theor. Appl Genet 91: 1001–1007.
Rector, B.G., J.N. All, W.A. Parott, & H.R. Boerma, 1998. Identification of molecular markers linked to quantitative trait loci for soybean resistance to corn earworm. Theor Appl Genet 96: 786–790.
Roa, A.C., P. Chavarriaga-Aguirre, M.C. Duque, M.M. Maya, M.W. Bonierbale, C. Iglesias & J. Tohme, 2000. Cross-species amplification of cassava (Manihot esculenta) (Euphorbiaceae) microsatellites: allelic polymorphism and degree of relationship. Am J Bot 87: 1647–1655.
Roder, M.S., J. Plaschke, S.U. Konig, A. Borner, M.E. Sorrels, S.D., Tanksley & M.W. Ganal, 1995. Abundance, variability and chromosomal location of microsatellites in wheat. Mol Gen Genet 246: 327–333.
Rongwen, J., M.S. Akkaya, A.A. Bhagwat, U. Lavi, & P.B. Cregan, 1995. The use of microsatellite DNA markers for soybean genotype identification. Theor Appl Genet 90: 43–48.
SAS Institute Inc, 1996. SAS/STAT software:changes and enhancement for release 6.12, Cary, NC: SAS Institute Inc. 158pp.
Senior, M.L., & M. Heun, 1993. Mapping maize microsatellites and polymerase chain reaction confirmation of the targeted repeats using a CT primer. Genome 36: 884–889.
Shapiro, S.S. & M.B. Wilk, 1965. An analysis of variance for normality (complete samples). Biometrika 52: 591–611.
Spickett, S.G. & J.M. Thoday, 1966: Regular responses to selection 3. Interaction between located polygenes. Genet Res 7: 96–121.
Titterington, D.M., A.F.M. Smith & U.E. Makov, 1985. Statitical analysis of finite mixture distributions. John Wiley and sons. N.Y.
Udupa, S.M., L.D. Robertson, F. Weigand, M. Baum & G. Kahl, 1999. Allelic variation at (TAA)n microsatellite loci in a world collection of chickpea (Cicer arietinum L.) germplasm. Mol Gen Genet 261: 354–363.
Winter P, T. Pfaff, S.M. Udupa, B. Huttel, P.C. Sharma, S. Sahi, R. Arreguin-Espinoza, F. Weigand, F.J. Muehlbauer & G. Kahl, 1999. Characterization and mapping of sequence-tagged microsatellite sites in the chickpea (Cicer arietinum L) genome. Mol Gen Genet 262: 90–101.
Xu, Y., L. Zhu, J. Xiao, N. Huang, & S.R. McCouch, 1997. Chromosomal regions associated with segregation distortion of molecular markers in F2, backcross, double-haploid, and recombinant inbred populations in rice (Oryza sativa L.). Mol Gen Gent 253: 535–545.
Yan, Z., A. Denneboom Hattendorf, O. Dolstra, T. Debener, P. Stam & P.B. Visse, 2005. Construction of an integrated map of rose with AFLP, SSR, PK, RGA, RFLP, SCAR and morphological markers. Theor Appl Genet 110: 766–777.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Okogbenin, E., Marin, J. & Fregene, M. An SSR-based molecular genetic map of cassava. Euphytica 147, 433–440 (2006). https://doi.org/10.1007/s10681-005-9042-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-005-9042-y