Abstract
In recent years, watermelon (Citrullus lanatus) has been subjected to significant losses due to vine decline in Sicily (southern Italy). During a survey conducted in 2009, the predominant fungal species associated with root rot and vine decline were Rhizoctonia spp. The most isolates were characterized as binucleate Rhizoctonia AG-F through morphological observation, nuclear condition, anastomosis tests and sequence homology of rDNA-ITS. Occasionally, R. solani was found. The pathogenicity of binucleate Rhizoctonia and virulences of different isolates were tested in growth chamber on watermelon seedlings. All isolates were pathogenic on watermelon seedlings and showed statistically significant differences on the disease incidence and severity among them. To our knowledge, this is the first report worldwide of the occurrence of pathogenic binucleate Rhizoctonia responsible for root rot and associated with watermelon vine decline.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
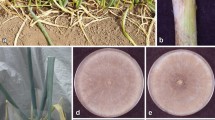
Watermelon [Citrullus lanatus (Thunb.) Matsum & Nakai] represents one of the most economically important vegetable crops in south-eastern Sicily covering approximately 2,000 ha. In recent years, this crop has been subjected to significant losses in yield and quality due to watermelon collapse (vine decline). Symptoms of the disease were similar to those described for other vine declines of melons (Miller et al. 1995; Zitter et al. 1996). On aerial parts the symptoms started with yellowing of the crown leaves followed by gradual collapse of the vines, generally just prior to harvest. Below-ground symptoms included necrosis on roots and moreover, in most cases the symptoms included a rot of secondary and feeder roots. Vine decline is a complex of diseases caused by a variety of fungal species belonging to different genera, even if one pathogen may predominate. Monosporascus cannonballus, Acremonium cucurbitacearum, Plectosporium tabacinum and Rhizopycnis vagum have been reported as the primary causal agents of this disease (Aegerter et al. 2000; García Jiménez et al. 2000; Martyn and Miller 1996). Root rot and vine decline have been previously observed in Italy in different cultivation areas of cucurbits. In addition to the prevalent fungi above mentioned, other species associated with the disease as Pyrenochaeta lycopersici, Fusarium spp., Macrophomina phaseolina, Rhizoctonia solani, Pythium spp., and Verticillium dahliae were recovered (Chilosi et al. 2008; Infantino et al. 2004).
A preliminary study conducted in some samples of collapsed watermelon plants collected during 2008 in south-eastern Sicily allowed us to ascertain a relevant association between Rhizoctonia spp. and a rot of secondary and feeder roots. The objectives of this paper were to characterize isolates of Rhizoctonia spp. recovered from root rot of watermelon in Sicily and associated to vine decline, to test their pathogenicity and to evaluate the virulence of different isolates. To identify the Rhizoctonia spp. morphological observation, nuclear condition, anastomosis tests and sequence homology of rDNA-ITS were done.
Surveys were conducted on watermelon grafted or not grafted on different rootstocks grown under tunnel and greenhouse in south-eastern Sicily (Table 1). Samples showing the vine decline incidence 10–100 % were collected in seven farms that had a history of vine decline. Isolations were performed from secondary and tertiary roots, tap roots and crowns. The infected tissues were washed under running water, surface disinfected in 1.5 % sodium hypochlorite for 1 min and placed on potato dextrose agar (PDA, Oxoid) amended with 100 ppm streptomycin sulphate. Hyphal tips or spores of representative fungal isolates were transferred to PDA for identification. The frequency of isolation of each fungus from field locations was recorded.
The most prevalent fungal species associated with vine decline and root rot were binucleate Rhizoctonia spp. and Fusarium spp. (Table 1). Another species frequently isolated was M. cannonballus. R. solani was detected only in one of the affected fields. The fungal isolates referable to Rhizoctonia spp. were identified on the basis of colony morphology and typical hyphal branching pattern. The numbers of nuclei within the hyphal cells were determined by staining with a 1 % Safranin O and 3 % KOH aqueous solution (Bandoni 1979). Most isolates of Rhizoctonia spp. were binucleate; the rest of the isolates were multinucleate and were characterized as R. solani AG-4 with anastomosis tests. Six representative binucleate isolates from different farms (DISTEF-AV, AC, AD, AL, AM, and AQ) and tester isolates of other binucleate Rhizoctonia AGs (AG A to AG U) were tested for anastomosis reaction on 2 % WA in Petri plates (Carling 1996). Hyphae of all six isolates fused with tester isolates of AG-F, but failed to fuse with hyphae of other AG tester isolates.
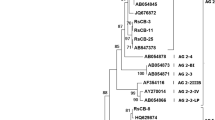
Mycelia for DNA extraction were cultured in a 9-cm Petri dish containing 10 ml of potato dextrose broth (PDB) at 25 °C. After 3 to 4 days of incubation, the mycelial mat was harvested by filtration and stored at −80 °C until use. Total genomic DNA was extracted as described by Hyakumachi et al. (1998). A polymerase chain reaction (PCR) with primers ITS 1 (50-TCCGTAGGTGAACCTGCGG-30) and ITS 4 (50-TCCTCCGCTTATTGATATGC-30) (White et al. 1990) was used to amplify rDNA-ITS regions of six isolates (Table 2). Amplification reactions were carried out using 50 ng of treated DNA. The PCR reaction mixture consisted of 200 μM each of dNTP, 50 pmol of oligonucleotide primers ITS 1 and ITS 4, 10 mM Tris–HCl at pH 8.3, 50 mM KCl, 1.5 mM MgCl2 and 1.25 units Taq DNA polymerase (Takara Bio Inc., Kyoto, Japan). The DNA Thermal Cycler (GeneAmp PCR system 2700; Applied Biosystems, Foster city, CA, USA) was programmed for one cycle of 5 min at 94 °C, followed by 25 cycles of 30 s at 94 °C, 30 s at 59 °C and 30 s at 72 °C, and one cycle of 7 min at 72 °C. The reaction mixture containing PCR products of rDNA-ITS region was mixed with 30 μl of PEG (30 % poly ethylene glycol 6000 containing 1.6 M NaCl), and centrifuged at 13,000 rpm at room temperature for 10 min. Amplified ITS products were precipitated, rinsed with 70 % ethanol and dissolved in 20 μl of distilled water. ITS region was sequenced using primers ITS 1, ITS 2 (50-GCTGCGTTC TTCATCGATGC-30), ITS3 (50-GCATCGATGAAGAAC GCAGC-30) (White et al. 1990) and ITS4 with a Ready Reaction Cycle Sequencing Kit (Applied Biosystems, Foster City, California, USA) and was analyzed with an ABI 3100 DNA sequencer (Applied Biosystems). The lengths of the rDNA-ITS region of the 6 isolates were approximately 620 bp based on their sequence data. Sequence similarity of rDNA-ITS within the isolates obtained from watermelon was 100 %. Sequence similarities between the isolates and BNR AG-F isolate (accession no. AB 219144) were 99 %.
Preliminarily, pathogenicity of two representative binucleate Rhizoctonia isolates was evaluated in the growth chamber on watermelon (cv. Crimson Sweet) not grafted or grafted on two rootstocks (Macis and Emphasis, Lagenaria siceraria hybrids). Twenty plants for each isolate were inoculated. The same number of plants served as controls. Two 6-mm-diameter mycelial plugs of 5 day-old colonies of each isolate, grown on PDA, were placed near the crown of plants. Plants were kept for 3 weeks at 25 °C and 95 % relative humidity with a 12-h fluorescent light/dark regimen. Each isolate caused root rot on seedlings and rootstocks, and occasionally necrotic lesion at the crown level. Binucleate Rhizoctonia was consistently reisolated from symptomatic tissue.
The variability of aggressiveness among isolates of binucleate Rhizoctonia AG-F used in molecular analysis was determined by comparing disease incidence (DI) and severity symptoms (SS) 4 weeks after inoculation. To this aim 42 to 48 seedlings cv. Crimson Sweet were selected and inoculated for each isolate. The same number of healthy seedlings served as controls. The inoculations were performed as above described. An empirical 0–5 rating scale was used for SS evaluation, where 0 = no symptoms, 1 = 1 to 25 % of infected root surface, 2 = 26 to 50 % of infected root surface, 3 = 51 to 75 % of infected root surface, 4 > 75 % of infected root surface, 5 = death of seedling. In the pathogenicity tests, the means of DI, SS index, and dry weight for each isolate were calculated averaging corresponding values determined for each replicate. Furthermore, analysis of variance (ANOVA; Statistica 7, Statsoft. Inc.) was performed for DI and weight data. The Bliss angular transformation was applied to DI data before statistical analysis. Mean values of DI were compared according to analysis of variance (ANOVA). The corresponding mean values were separated by least significant difference (LSD) test (P ≤ 0.01). Since an ordinal scale was adopted for disease severity assessment, SS data were analyzed according to the Kruskal-Wallis non parametric one-way analysis (Statistica 7, Statsoft. Inc.) followed by all possible pairwise comparison with the Mann–Whitney test. There was a significant effect of the pathogenicity on mean DI, dry weight and SS data among the Rhizoctonia AG-F isolates (Table 2).
In the present study we reported the first occurrence of binucleate Rhizoctonia spp. responsible for a watermelon root rot. Generally, these fungi are considered as biocontrol agents (Escanbde and Echandi 1991; Harris 2000) although isolates pathogenic to several plants were found (Manici and Bonora 2007; Nerey et al. 2010; Sneh et al. 1991).
For identifying AGs within binucleate Rhizoctonia, it is important to observe the differences in cultural morphology of their vegetative state, to observe the anastomosis reaction with the tester isolates and especially to analyze molecular profile data of the tester isolates. Numerous studies have demonstrated the usefulness of sequence analysis of the rDNA-ITS region for classifying and identifying intra-groups within Rhizoctonia sp. (Boysen et al. 1996; Gonzalez et al. 2001; Hsiang and Dean 2001; Johanson et al. 1998; Kuninaga et al. 1997; Salazar et al. 2000; Sharon et al. 2008). In this study, sequences of rDNA-ITS region contributed to the successful classification of BNR isolates associated to watermelon vine decline as BNR AG-F. These results were well matched with the anastomosis test.
On cucurbits, Rhizoctonia solani AG-4 was reported in Italy as agent of stem and fruit rot of cantaloupe melon (Corazza et al. 1992) and R. solani AG-7 was isolated from watermelon plants in Indiana (Baird and Carling 1994). To our knowledge, this is the first report of the occurrence of pathogenic binucleate Rhizoctonia associated with vine decline disease on cucurbits.
References
Aegerter, B. J., Gordon, T. R., & Davis, R. M. (2000). Occurrence and pathogenicity of fungi associated with melon root rot and vine decline in California. Plant Disease, 84, 224–230.
Baird, R. E., & Carling, D. E. (1994). First report of Rhizoctonia solani AG-7 in Indiana. Plant Disease, 79, 321.
Bandoni, R. J. (1979). Safranin O as a rapid nuclear stain for fungi. Mycologia, 71, 873–874.
Boysen, M., Borja, M., Delmont, C., Salazar, O., & Rubio, V. (1996). Identification at strain level Rhizoctonia solani AG 4 isolates by different sequence of asymmetric PCR products of the regions. Current Genetics, 29, 174–181.
Carling, D. E. (1996). Grouping in Rhizoctonia solani by hyphal anastomosis reaction. In B. Sneh, S. Jabaji-Hare, S. Neate, & G. Dijst (Eds.), Rhizoctonia species: Taxonomy, molecular, biology, ecology, pathology, and disease control (pp. 37–62). Dordrecht: Kluwer Academic Publishers.
Chilosi, G., Reda, R., Oleandri, M. P., Camele, I., Altieri, L., Montuschi, C., et al. (2008). Fungi associated with root rot and collapse of melon in Italy. EPPO Bulletin, 38, 147–154.
Corazza, L., Luongo, L., & Chilosi, G. (1992). Characterization of a strain of Rhizoctonia solani Kühn from melon in Italy. Phytopathologia Mediterranea, 31, 121–122.
Escanbde, A. R., & Echandi, E. (1991). Protection of potato from Rhizoctonia canker with binucleata Rhizoctonia fungi. Plant Pathology, 40, 197–202.
García Jiménez, J., Armengol, J., Sales, R., & Bruton, B. D. (2000). Fungal pathogens associated with melon collapse in Spain. EPPO Bulletin, 30, 169–173.
Gonzalez, D., Carling, D. E., Kuninaga, S., Vilgalys, R., & Cubeta, M. A. (2001). Ribosomal DNA systematic of Ceratobasidium and Thanatephorus with Rhizoctonia anamorphs. Mycologia, 93, 1138–1150.
Harris, A. R. (2000). Solid formulations of binucleate Rhizoctonia isolates suppress Rhizoctonia solani and Pythium ultimum in potting medium. Microbiological Research, 154, 333–337.
Hsiang, T., & Dean, J. D. (2001). DNA sequencing for anastomosis grouping of Rhizoctonia solani isolates from Poa annua. International Turfgrass Society Research Journal, 9, 674–678.
Hyakumachi, M., Mushica, T., Ogiso, Y., Toda, T., Kageyama, K., & Tsuge, T. (1998). Characterization of a new cultural type (LP) of Rhizoctonia solani AG 2–2 isolated from warm-season turfgrasses, and its genetic differentiation from other cultural types. Plant Patholology, 47, 1–9.
Infantino, A., Carlucci, A., Pucci, N., Ciuffreda, G., Montuschi, C., Lops, F., et al. (2004). Fungi causing root rots and collapse of cucurbits in Italy. Petria, 14, 77–89. In Italian.
Johanson, A., Turner, H. C., McKay, G. J., & Brown, A. E. (1998). A PCR-based method to distinguish fungi of the rice sheath-blight complex, Rhizoctonia solani, R. oryzae and R. oryzae-sativae. FEMS Microbiology Letters, 162, 289–294.
Kuninaga, S., Natsuaki, T., Takeuchi, T., & Yokosawa, K. (1997). Sequence variation of the rDNA ITS regions within and between anastomosis groups in Rhizoctonia solani. Current Genetics, 32, 237–243.
Manici, L. M., & Bonora, P. (2007). Molecular genetic variability of Italian binucleate Rhizoctonia spp. isolates from strawberry. European Journal of Plant Pathology, 118, 31–42.
Martyn, R. D., & Miller, M. E. (1996). Monosporascus root rot and vine decline: An emerging disease of melons worldwide. Plant Disease, 80, 716–725.
Miller, M. E., Martyn, R. D., Lovic, B. R., Bruton, B. D. (1995). An overview of vine decline diseases in melons. In: Proceedings Cucurbitaceae 94: Evaluation and enhancernent of Cucurbit Germplasm, (1994) pp. 31–35.
Nerey, Y., Pannecoucque, J., Hernandez, H. P., Diaz, M., Espinosa, R., De Vos, S., et al. (2010). Rhizoctonia spp. causing root and hypocotyl rot in Phaseolus vulgarisin Cuba. Journal of Phytopathology, 158, 236–243.
Salazar, O., Julian, M. C., Hyakumachi, M., & Rubio, V. (2000). Phylogenic grouping of cultural types of Rhizoctonia solani AG 2–2 based on specific rDNA-ITS sequences. Mycologia, 92, 505–509.
Sharon, M., Kuninaga, S., Hyakumachi, M., Naito, S., & Sneh, B. (2008). Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience, 49, 93–114.
Sneh, B., Bupee, L., & Ogoshi, A. (1991). Identification of Rhizoctonia species (p. 79). St. Paul: APS.
White, T. G., Burns, T., Lee, S., & Taylor, J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, & T. J. White (Eds.), PCR protocols: A guide to methods and applications (pp. 315–321). San Diego: Academic.
Zitter, T. A., Hopkins, D. L., & Thomas, C. E. (1996). Compendium of cucurbit diseases. St. Paul: APS.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aiello, D., Vitale, A., Hyakumachi, M. et al. Molecular characterization and pathogenicity of binucleate Rhizoctonia AG-F associated to the watermelon vine decline in Italy. Eur J Plant Pathol 134, 161–165 (2012). https://doi.org/10.1007/s10658-012-9973-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-012-9973-9