Abstract
During surveys conducted in 2010–2012 Rhizoctonia symptoms were observed on 30 ornamental species in different nurseries located in eastern Sicily (Southern Italy). Eighty-eight isolates of Rhizoctonia spp. were obtained from symptomatic leaves, roots and stems. Fifty-six of the isolates were binucleate and 32 were multinucleate Rhizoctonia. Characterisation of anastomosis groups (AGs) was performed using morphological characteristics and sequence analysis of the internal transcribed spacer of ribosomal DNA ( rDNA-ITS) region. Most isolates collected were Rhizoctonia solani AG-4 HG-I (35.2% of all isolates) and one isolate was AG-2-2 IIIB. The binucleate isolates belonged to AG-R (27.3%), AG-A (21.6%), AG-G (12.5%), AG-V (1.1%) and AG-Fb (1.1%). The pathogenicity of 38 representative isolates collected from each host was tested on seedlings or cuttings grown in a growth chamber. All R. solani AG-4 HG-I isolates, most of the binucleate AG-R, AG-A and AG-G and AG-V were pathogenic and reproduced symptoms identical to that observed in nurseries, while binucleate AG-Fb and R. solani AG-2-2 IIIB isolates were nonpathogenic. This is the first report of the occurrence of Rhizoctonia species on some ornamental plants and the first report of binucleate Rhizoctonia AG-R and AG-V in Europe.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rhizoctonia species are soilborne pathogens causing root and foliar diseases on a wide range of agronomic crops, turfgrasses, ornamental plants, fruit and forest trees worldwide (Sneh et al. 1991; Couch 1995).
Rhizoctonia spp. are classified into three groups based on the differences in nuclei number per cell (Ogoshi 1975; Ogoshi 1987; Ogoshi 1996): multinucleate Rhizoctonia (MNR) (teleomorphs: Thanatephorus and Waitea), binucleate Rhizoctonia (BNR) (teleomorphs: Ceratobasidium and Tulasnella) and uninucleate Rhizoctonia (UNR) (teleomorph: Ceratobasidium). Each group is composed by different anastomosis groups (AGs). Moreover, several AGs of R. solani and binucleate Rhizoctonia are subdivided into subgroups that differ for biochemical, genetic and pathogenic characteristics (Sneh et al. 1991, 1998; Priyatmojo et al. 2001; Naito 2004; Aoyagi et al. 1998; Sharon et al. 2006).
Binucleate Rhizoctonia spp. have been divided into 21 AGs designated AG-A to AG-U (Ogoshi et al. 1983; Sneh et al. 1991; Hyakumachi et al. 2005). Of the BNR AGs, AG-J and AG-N are excluded, the representative isolates of AG-M are lost, and AG-T and AG-U reported as new BNR AGs by Hyakumachi et al. (2005) were subsequently confirmed to belong to AG-A and AG-P, respectively (Sharon et al. 2008). Moreover, AG-V and AG-W were recently reported in China (Yang 2013; Yang et al. 2015). BNR were weakly virulent, not-pathogenic or even mycorrhizal or biocontrol agents (Harris et al. 1994; Andersen and Rasmussen 1996; Sneh 1998; Hwang and Benson 2002). However, several studies reported BNR as pathogenic on economically important agricultural and horticultural crops (Priyatmojo et al. 2001; Kuramae et al. 2007; Aiello et al. 2012; Muzhinji et al. 2015; Yang et al. 2015).
The multinucleate species are represented by R. solani Kühn (teleomorph: Thanatephorus cucumeris (A.B. Frank) Donk, Waitea circinata var. oryzae (anamorph = Rhizoctonia oryzae Ryker & Gooch) and W. circinata var. zeae (anamorph = Rhizoctonia zeae Voorhees) (Leiner and Carling 1994) and W. circinata var. circinata (Toda et al. 2005). R. solani is composed of 14 anastomosis groups: AG-1 to AG-10, AG-BI (Sneh et al. 1991), AG-11 (Carling et al. 1994), AG-12 (Carling et al. 1999), and AG-13 (Carling et al. 2002) and three subgroups HG-I, HG-II (Kuninaga and Yokosawa 1984) and HG-III (Stevens Johnk and Jones 2001). R. solani is the most widespread species, with a host range that includes over 500 plant species (Farr et al. 1995). Hyphal anastomosis was not used because it does not always provide accurate results and currently, rDNA internal transcribed spacer region (ITS) sequence analysis is the most accurate method to determine AGs and subgroups of Rhizoctonia spp. and establish phylogenetic relationships among these (Hyakumachi et al. 2005; Sharon et al. 2008).
Rhizoctonia diseases of ornamental plants can occur and in some cases may cause severe epidemics. A wide range of disease symptoms have been recorded, including root and stem rot, leaf spot, seedlings damping-off and foliar web blight on economic important ornamental species such as Azalea spp., Begonia spp., Petunia × hybrida, Rosa spp., Vinca minor and Pittosporum tobira worldwide (Chase 1991; Benson and Cartwright 1996; Hyakumachi et al. 2005; Rinehart et al. 2007). In South Italy, binucleate Rhizoctonia and R. solani are widespread in nurseries and cause extensive damage to young ornamental plants (Aiello et al. 2008a, b, 2009a, b; Polizzi et al. 2009a, b, c, 2010a, b, c, 2011a, b). Prevention is the first strategy to control Rhizoctonia diseases and in particular the adoption of control measures to reduce primary inoculum (use of healthy plants and sterilized pots and potting media, and the removal of infected plants). Chemical management is the approach most often used to control Rhizoctonia diseases. A wide range of chemicals has been reported as effective and selective against R. solani and binucleate Rhizoctonia-like fungi (Jager et al. 1991; Haralson et al. 2013). However, different Rhizoctonia species and AGs possess a different sensitivity towards fungicides (Kataria et al. 1991; Ueyama et al. 1990; Kataria and Gisi 1996; Benson and Cartwright 1996; Csinos and Stephenson 1999; Virgen-Calleros et al. 2000). Therefore, an accurate diagnosis through the determination of Rhizoctonia AGs present in a particular area or in an ornamental nursery is important for the selection of effective disease management strategies and for understanding the distribution and spread of pathogen.
Considering the importance of Rhizoctonia diseases and high economic losses caused by these fungi, surveys were conducted over a 3-year period, in commercial ornamental nurseries located in Catania province, eastern Sicily, Italy. The aims of the present study were to identify the AGs and subgroups of Rhizoctonia spp. obtained from ornamentals using morphological characteristics and ITS sequence; and evaluate the pathogenicity of representative Rhizoctonia isolates on the ornamental hosts from which they were isolated.
Materials and methods
Field surveys, sample collection and fungal isolation
Surveys were conducted during 2010–2012 in 10 nurseries located in eastern Sicily. The disease incidence was recorded for each host species based on the number of symptomatic plants on the total of those producted. Approximately, 20 plants per species per nursery showing Rhizoctonia-like symptoms were randomly collected for analysis. Small sections (0.2–0.5 cm long) from the edge of symptomatic tissues were surface disinfected with 1.5% sodium hypochlorite for 1 min, rinsed once in sterile distilled water (SDW), dried on sterile absorbent paper and placed on potato dextrose agar (PDA, Oxoid) plates amended with 100 ppm streptomycin sulphate (Sigma-Aldrich). Plates were incubated at 25 ± 1 °C under continuous dark conditions. Following 48 to 72 h of incubation, hyphae from the margin of colonies with features characteristic of Rhizoctonia spp. were placed on PDA plates. After 5 days, single-hyphal or tip were selected and transferred into PDA plates for monomycelic cultures.
A total of 88 isolates were obtained and used for morphological and molecular characterisation. Stock cultureswere stored in tubes on PDA covered with mineral oil.
Morphological characteristics and nuclear conditions
The Rhizoctonia isolates were identified morphologically by examining the hyphal branching after 3–4 days of growth on PDA. To distinguish binucleate Rhizoctonia isolates from multinucleate R. solani isolates, the number of nuclei per hyphal cell was determined. Agar disks (5 mm in diameter) containing mycelium from 2- to 3-day-old cultures growing on 2% water agar (WA, Oxoid) were placed on sterile, glass slides in a moist chamber at 25 °C for 2 to 3 days in the dark. Nuclei were stained with one drop of each 1% safranin O and 3% KOH solution (Bandoni 1979). From each isolate, number of nuclei of 20 cells in hyphae at merging two solutions was determined microscopically at × 400 magnification using an Olympus BX61 microscope.
Molecular characterisation
Genomic DNA from each of the 88 isolates was extracted using a conventional method (Izumitsu et al. 2012). Briefly, small piece of mycelia of each isolate was added to 100 μL TE buffer in a 1.5-mL tube. After the sequential treatments by microwaving twice for 1 min, cooling for 10 min, and centrifuging for 5 min, the supernatant was used as the template DNA for PCR. PCR amplification of the rDNA, including regions of ITS1, 5.8S rDNA and ITS2 for each isolate was performed with the primer set of ITS1-F and ITS4-B (Gardes and Bruns 1993). Amplification was performed in 10 μL reaction mixture containing 2 μL template DNA, 1 μL PCR buffer, 0.5 μL dNTP (2.5 mM), 0.15 μL of each primer (20 μM) and 0.05 μL (5 units/μL) Taq DNA polymerase (TaKaRa Bio Inc. Japan). Amplification was performed with a thermal cycler (Verity, Life Technologies Applied Biosystems, USA) with the following program: an initial denaturation at 96 °C for 2.5 min; 40 cycles consisting of denaturation at 96 °C for 30 s, annealing at 55 °C for 30 s and extension at 72 °C for 30 s; a final extension at 72 °C for 5.5 min. A 3 μL aliquot of PCR product was separated by electrophoresis on 1.5% (w/v) agarose gel, stained with ethidium bromide and visualized by UV transilluminator. A 5 μL aliquot of remaining each PCR product was cleaned up for sequencing by the addition of 2 μL ExoSAP-IT (Affymetrix, USA) according to the manufacturer’s instructions. One μL of template solution was sequenced using BigDye terminator cycle sequencing kit v. 3.1 (Life Technologies Applied Biosystems, USA) with the same primers used for the PCR amplification. Electrophoretic separation of sequencing products was performed on the Genetic Analyzer 3130xl (Life Technologies Applied Biosystems, USA). The nucleotide sequences generated by the sequencing of each isolate in both directions by primer ITS1-F and ITS4-B were edited and assembled using the combination of Sequencher 5.0.1 (Gene Codes Corp., USA) and Indelligent (Dmitriev and Rakitov 2008) with manual adjustment. Sequences from all isolates were compared with those in the GenBank nucleotide database provided by the National Center for Biotechnology Information (www.ncbi.nlm.nih.gov) using BLAST algorithm to determine sequence identity and find the closest match based on maximal percent identity. Sequences derived in this study were lodged at GenBank.
Phylogenetic analysis
Sequences of 37 isolates of Rhizoctonia spp., selected as representative isolates from different hosts, were used for phylogenetic analysis to confirm the AGs and subgroups recovered. Separate phylogenetic trees were constructed for BNR isolates and R. solani isolates. Additional reference sequences of 11 isolates from known AGs worldwide were retrieved from GenBank database and included together with the binucleate Rhizoctonia isolates obtained in this work. For R. solani tree, 15 reference sequences from the same database were added to our isolates (Table 1). The nucleotide sequences generated by the sequencing were aligned using Clustal W algorithm in MEGA v. 6 (Tamura et al. 2013) and the alignment was corrected manually where necessary. The evolutionary history was inferred using the Neighbor-Joining method (Saitou and Nei 1987). The bootstrap (BP) consensus tree inferred from 1000 replicates was considered to represent the evolutionary history of the taxa analyzed (Felsenstein 1985). The evolutionary distances were computed using the Kimura 2-parameter method (Kimura 1980) and are in the units of the number of base substitutions per site. The tree was drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The analysis involved 34 nucleotide sequences of BNR and 30 nucleotide sequences of R. solani. All positions containing gaps and missing data were eliminated (complete deletion option). The tree with BNR was rooted with an isolate of R. orizae (accession number: KC590575) as out-group. The tree with R. solani was rooted with an isolate of Athelia rolfsii (accession number: AY684917).
Pathogenicity
Pathogenicity tests were performed with 38 representative isolates on potted, healthy, seedlings or cuttings of all symptomatic species recovered (Table 2, except Carissa macrocarpa) grown in a growth chamber for 4 months. Each experiment was conducted twice and similar results were obtained in both tests. For each experiment three replicates per isolate were used with 20–50 plants per replicate. All plants were inoculated at the base of each stem plants with two 6 mm in diameter mycelial plugs obtained from cultures grown on PDA plates for 5 days at 25 ± 1 °C in the dark. Uninoculated plants served as a control, for all the hosts. After inoculation plants were covered with a plastic bag for 48 h and maintained at 25 ± 1 °C and 95% relative humidity (RH) under a 12-h fluorescent light/dark regimen. All plants were irrigated 2–3 times per week and examined weekly for disease symptoms. Disease incidence (DI) was assigned to each host species and isolate by determining the percentage of seedlings or cuttings with symptoms of Rhizoctonia disease after 7 days to 4 months from pathogen inoculation.
Results
Field surveys, sample collection and fungal isolation
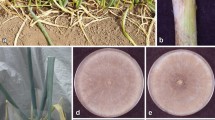
Symptoms referable to Rhizoctonia spp. were detected over a 3-year period in 10 nurseries investigated and on 30 different ornamental species (Figs. 1 and 2). Rhizoctonia diseases were observed in the period from March to November during propagation stage on unrooted and rooted cuttings (1–4 months old) in greenhouse and on established plants (1–4 years old) in open field. Diseases incidence varied, approximately, from 10 to 50%, according to the host species (Table 2). The symptoms observed consisted of crown and root rot, stem rot, damping-off and web blight (Table 2). Early in the disease development, crown and stem rot was characterized by water-soaked lesions at the soil line that turned light reddish brown to dark brown and expanded to girdle the stem and internal brown discolouration of cortical tissues; root rot sometimes occurs in association with these symptoms. The infected roots become dark brown or black and were partially or completely destroyed. As a consequence of root and stem rot, basal leaves initially turned chlorotic and gradually became necrotic and sometimes infected plants wilted and died. Damping-off consisted of the decay of the stem at soil level, causing it to fall over because it has not yet thickened supporting tissue. Symptoms of either web blight or aerial blight included interveinal and marginal irregular necrotic lesions that progress to total leaf necrosis and leaf-drop. During nursery production, especially under the hot humid conditions (22–30 °C and 85–95% RH), the web-like brown mycelium of the pathogen covered portions of the infected plants and resulted in brown patch disease. The web blight symptom was associated with crown and root rot. Unrooted cuttings and cuttings during rooting stage were susceptible to infection.
Morphological characteristics and nuclear conditions
All isolates showed typical features of Rhizoctonia spp. including branching at right angles with constriction at the base of hyphae and septum near the point of origin. Fifty-six of the isolates recovered were binucleate, the other 32 isolates were multinucleate.
Molecular characterisation and phylogenetic analysis
Analysis of the rDNA-ITS region using the BLAST algorithm (against Genbank database) revealed that 31 isolates belonged to R. solani AG-4-HGI, 1 to AG-2-2 IIIB, 24 to binucleate AG-R, 19 to AG-A, 11 to AG-G, AG-Fb and AG-V. The similarity range of 79 isolates was from 98% to 100% while nine isolates showed an sequence identity lower (from 85% to 97%) (Table 3). Highest sequence similarities (98–100%) were observed among our R. solani AG-4, AG-2-2 IIIB and binucleate Rhizoctonia AG-Fb, AG-V and AG-G isolates and representative isolates from Genbank. Binucleate AG-R and AG-A isolates showed an sequence similarity with isolates from Genbank lower (from 93% to 99% and 85% to 99%, respectively).
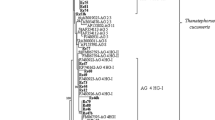
The optimal tree for R. solani with the sum of branch length = 0.56051825 was shown in Fig. 3. The percentages of replicate trees in which the associated taxa clustered together in the bootstrap test were shown next to the branches (BP analysis; Fig. 3). There were a total of 342 positions in the final dataset. R. solani AG-4 HG-I and AG-2-2 IIIB isolates clustered with representative isolates from Genbank with BP values of 95% and 98%, respectively (Fig. 3).
The optimal tree for BNR with the sum of branch length = 0.49361651 was shown in Fig. 4. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test were shown next to the branches (BP analysis; Fig. 4). There were a total of 308 positions in the final dataset. All BNR isolates formed clades together with representative isolates with bootstrap supports of 98% (AG-A and AG-Fb), 97% (AG-R), 96% (AG-V) and 94% (AG-G) (Fig. 4).
Pathogenicity
Of thirty-eight isolates, thirty Rhizoctonia isolates tested were pathogenic to the different original hosts inoculated and produced symptoms identical or similar to those observed on diseases plants in the nurseries (Table 3). Eight isolates were not pathogenic. The DI (%) caused by Rhizoctonia species on different hosts ranging from 75 to 100% after 7 days to 4 months (Table 3).
All R. solani AG-4 HG-I isolates were pathogenic and caused 100% of DI on Trachycarpus fortunei, citrus species, Chamaerops humilis, Osteospermum sp., Tabebuia impetiginosa and Streptosolen jamesonii, Arbutus unedo, Lagunaria patersonii, and Bignonia sp., whereas lower DI on Murraya paniculata and Thevetia peruviana (80% and 75%, respectively).
Binucleate AG-A isolates caused high DI (from 80% to 100%) on Dodonaea viscosa, Cistus salvifolius, Quercus ilex, Thryptomene saxicola, Passiflora mollissima, Carissa grandiflora, and Catharanthus roseus but were non pathogenic on Viburnum tinus and Grevillea sp. The binucleate AG-V isolate (61 LNO) was pathogenic to original host (Laurus nobilis) and caused 100% of DI. Binucleate AG-R isolates didn’t cause disease on C. grandiflora but were pathogenic to Butia capitata, C. salvifolius, P. mollissima and Polygala myrtifolia causing DI from 90% to 100%. Binucleate AG-G isolates were pathogenic and caused DI from 85% to 100% on Pittosporum tobira, V. tinus, Phillyrea angustifolia, Quercus ilex and C. roseus but not were pathogenic to B. capitata and Phormium spp.
On some host species two AGs were recognized and both were pathogenic (AG-A and AG-R on C. salvifolius and P. mollissima, AG-A and AG-G on C. roseus and Q. ilex) whereas on other species only one AG recovered was pathogenic (AG-A on C. grandiflora, AG-R on B. capitata and AG-G on V. tinus).
None AGs recognized from Phormium spp. (AG-Fb, AG-2-2 IIIB and AG-G) was pathogenic on this species. The pathogens were re-isolated from the artificially inoculated plants and identified as previously described, completing Koch’s postulates. No symptoms were observed on control plants.
Discussion
In this study, 88 Rhizoctonia isolates were recovered from 30 ornamental species in eastern Sicily (Southern Italy) over a 3-year period, and their AGs and subgroups determined. Fifty-six isolates were binucleate Rhizoctonia and 32 were multinucleate and were identified as R. solani. Rhizoctonia symptoms were observed in 10 ornamental nurseries and included damping-off, crown, root and stem rot and web blight thought damping-off was mainly associated with R. solani AG-4 while crown and stem rot was caused primarily from BNR. Infections were observed in the period from March to November during young growing stages in greenhouse (i.e., seeding, rooting and pot transplanting) and during summer (June–July) in the containerised field-grown species, probably, favored by optimal conditions for growth of the Rhizoctonia species.
Based on results of characterisation and of phylogenetic analysis, the most prevalent anastomosis group was R. solani AG-4 subgroup HG-I (35.2%) followed by binucleate Rhizoctonia AG-R (27.3%), AG-A (21.6%) and AG-G (12.5%), while only one isolate of AG-2-2 IIIB (R. solani), AG-Fb and AG-V was recognized.
In the pathogenicity tests, R. solani AG-4 isolates, the majority of binucleate AG-R, AG-A and AG-G isolates and AG-V isolate were pathogenic to original host, fulfilling Koch’s postulates. None AGs recognized from Phormium spp. (AG-Fb, AG-2-2 IIIB and AG-G) and the AG from Grevillea sp. (AG-A) were pathogenic on these species that seem not be susceptible to Rhizoctonia disease. Our study revealed a diversity in the composition of Rhizoctonia populations recovered from ornamental nurseries (binucleate AG-R, AG-A, AG-G, AG-V and AG-Fb; R. solani AG-4 and AG-2-2 IIIB). Moreover, in some cases, different binucleate AGs were recovered from the same host and both were pathogenic (AG-A and AG-R on C. salvifolius and P. mollissima, AG-A and AG-G on C. roseus and Q. ilex). In other cases, only one AG among those recovered appeared pathogenic to original host (AG-A on C. grandiflora, AG-R on B. capitata and AG-G on V. tinus. These results are consistent with reports that show that the host range of individual AG differs (Ogoshi 1996). Various studies have documented that certain hosts are susceptible to specific AGs and not others (Ohkura et al. 2009). These results could be also explained with the presence of some non-pathogenic or hypovirulent binucleate Rhizoctonia isolates. Several studies reported the presence of these isolates that colonize plant roots and can have an antagonistic activity against pathogenic Rhizoctonia isolates (Harris et al. 1994; Hwang and Benson 2002; Khan et al. 2005) or considered as mycorrhizal species (Andersen and Rasmussen 1996).
R. solani is widespread in ornamental nurseries in Italy (Aiello et al. 2008a, b, 2009a, b; Polizzi et al. 2009b, 2010a, c, 2011a, b; Garibaldi et al. 2003, 2006, 2009a, b) where represents a very limiting factor for ornamental plants cultivated in Sicily (Southern Italy). The pathogen has a very broad host range worldwide (Farr et al. 1995) and among these several ornamental species have been reported in the literature (Chase 1991; Sneh et al. 1991; Priyatmojo et al. 2001). However, no reports have been published of diseases caused by Rhizoctonia spp. on Citrus volkameriana, citrange, T. fortunei, A. unedo, T. peruviana and Bignonia sp.
BNR AG-A and AG-G are the two most common groups associated with root rot on strawberry in the world (Martin 2000) and widespread in strawberry-growing areas in Northern Italy (Manici and Bonora 2007). Recently, have been also reported on ornamentals in Southern Italy (Polizzi et al. 2009a, c, 2010b) but have not been reported diseases caused by Rhizoctonia AG-A on Carissa spp., C. roseus, Eugenia sp., Q. ilex, and Rhizoctonia AG-G on P. angustifolia, P. tobira and C. roseus.
R. solani was reported in Florida on C. grandiflora, Carissa spp. and Eugenia sp. (Alfieri et al. 1972, 1984), and R. solani and Rhizoctonia sp. were reported on Quercus spp. (Collado et al. 1996; Mulenko et al. 2008). Chase (1991) reported a binucleate Rhizoctonia sp. causing aerial blight and root rot of P. tobira while Alfieri et al. (1984) reported R. solani in Florida. Binucleate Rhizoctonia sp. and R. solani has been reported on C. roseus in United States (Alfieri et al. 1984; Chase 1991; Holcomb and Carling 2002) and R. solani AG-1 IB in Italy (Garibaldi et al. 2006).
AG-R, the binucleate AG most frequently found in this survey, has only been reported in the USA, Australia, Brasil and China (Burpee et al. 1980; Sumner 1985; Yang et al. 2006, Rinehart et al. 2007), and was not previously reported to be present in Italy (Europe), as well as AG-V recently reported only in China (Yang 2013). This study is the first report of Rhizoctonia disease on C. salvifolius, B. capitata, P. myrtifolia caused by AG-R and on L. nobilis caused by AG-V. In addition, this is also the first report of binucleate Rhizoctonia AG-R and AG-V in Europe.
The results of these surveys showed that, overall, BNR were the most prevalent Rhizoctonia species isolated from ornamental nurseries in Sicily. The higher frequency of binucleate Rhizoctonia spp. recovered could be explained with the cropping history of the study site as reported by other authors (Gill et al. 2000, 2001, 2004; Schroeder and Paulitz 2008). However, data from a European survey shows that Rhizoctonia spp. can be expected to be present in the soil irrespective of the previous crop in the field (Goll et al. 2014).
The high incidence of disease observed in several ornamental nurseries in the last years and the frequency and distribution of AGs will therefore depend on other factors such as climatic conditions and farming practice (Virgen-Calleros et al. 2000). The environmental conditions during propagation of plants in the greenhouse heated and irrigated overhead could provides ideal condition for Rhizoctonia disease development (warm temperatures and high humidity), such as the use of non-disinfected substrate. The containerised plants production could have a role in promoting infections because the ornamental plants are frequently stressed, remain containerised throughout production, and several wounds could be incurred during transplanting. Moreover, the predominant potting component in field-grown nurseries in most of the eastern Sicily is autochtone volcanic soil mixed with peat and perlite or vermiculite to assure plant stability and proper development. In eastern Sicily, potato-growing area are widespread near to the ornamental nurseries, and different reports show the high susceptibility of potato to binucleate Rhizoctonia and R. solani (Muzhinji et al. 2015). The use of non-disinfected soil taken from these infected area or the use of soil recycled could represent a possible source of pathogen inoculum and increase infection risks by Rhizoctonia. Prevention, thus, is the first strategy to control Rhizoctonia diseases but an accurate diagnosis through the determination of Rhizoctonia AGs present in a particular area or in an ornamental nursery is important for selection of effective disease management strategies.
References
Aiello, D., Castello, I., Vitale, A., Lahoz, E., Nicoletti, R., & Polizzi, G. (2008a). First report of damping-off on African daisy caused by Rhizoctonia solani AG-4 in Italy. Plant Disease, 92, 1367.
Aiello, D., Parlavecchio, G., Vitale, A., Lahoz, E., Nicoletti, R., & Polizzi, G. (2008b). First report of damping-off caused by Rhizoctonia solani AG-4 on Lagunaria patersonii in Italy. Plant Disease, 92, 836.
Aiello, D., Caiazzo, R., Carella, A., Lahoz, E., Nicoletti, R., Vitale, A., et al. (2009a). Characterisation of Rhizoctonia species isolated from ornamental nurseries. Journal of Plant Pathology, 91(S4), 45.
Aiello, D., Vitale, A., Lahoz, E., Nicoletti, R., & Polizzi, G. (2009b). First report of crown and root rot caused by Rhizoctonia solani AG-4 on orange jessamine in Italy. Plant Disease, 93, 204.
Aiello, D., Vitale, A., Hyakumachi, M., & Polizzi, G. (2012). Molecular characterisation and pathogenicity of binucleate Rhizoctonia AG-F associated to the watermelon vine decline in Italy. European Journal of Plant Pathology, 134, 161–165.
Alfieri Jr., S. A., Seymour, C. P., & Denmark, J. C. (1972). Aerial blight of Carissa grandiflora caused by Rhizoctonia solani. The Plant Disease Reporter, 56, 511–514.
Alfieri Jr., S. A., Langdon, K. R., Wehlburg, C., & Kimbrough, J. W. (1984). Index of plant diseases in Florida (revised). Florida: Division of Plant Industry, Florida Department of Agriculture and Consumer Services.
Andersen, T. F., & Rasmussen, H. N. (1996). The mycorrhizal species of Rhizoctonia. In B. Sneh, S. Jabaji-Hare, S. Neate, & G. Dijst (Eds.), Rhizoctonia species: taxonomy, molecular biology, ecology, pathology and disease control (pp. 379–390). The Netherlands: Kluwer Academic Publishers.
Aoyagi, T., Kageyama, K., & Hyakumachi, M. (1998). Characterisation and survival of Rhizoctonia solani AG2-2 LP associated with large patch disease of zoysia grass. Plant Disease, 82, 857–863.
Bandoni, R. J. (1979). Safranin O as a rapid stain for fungi. Mycologia, 71, 873–874.
Benson, D. M., & Cartwright, D. K. (1996). Ornamental diseases incited by Rhizoctonia spp. In S. B. Sneh, S. Jabaji-Hare, G. Neate, & Dijst (Eds.), Rhizoctonia species: taxonomy, molecular, biology, ecology, pathology, and disease control (pp. 303–314). The Netherlands: Kluwer Academic Publishers.
Burpee, L. L., Sanders, P. L., Cole Jr., H., & Sherwood, R. T. (1980). Pathogenicity of Ceratobasidium cornigerum and related fungi representing five anastomosis groups. Phytopathology, 70, 843–846.
Carling, D. E., Rothrock, C. S., Macnish, G. C., Sweetingham, M. W., Brainard, K. A., & Winters, S. W. (1994). Characterisation of anastomosis group-11 (AG-11) of Rhizoctonia solani. Phytopathology, 84, 1387–1393.
Carling, D. E., Pope, E. J., Brainard, K. A., & Carter, D. A. (1999). Characterisation of mycorrhizal isolates of Rhizoctonia solani from an orchid, including AG-12, a new anastomosis group. Phytopathology, 89, 942–946.
Carling, D. E., Baird, R. E., Gitaitis, R. D., Brainard, K. A., & Kuninaga, S. (2002). Characterisation of AG-13, a newly reported anastomosis group of Rhizoctonia solani. Phytopathology, 92, 893–899.
Chase, A. R. (1991). Characterisation of Rhizoctonia species isolated from ornamentals in Florida. Plant Disease, 75, 234–238.
Collado, J., Platas, G., & Pelaez, F. (1996). Fungal endophytes in leaves, twigs, and bark of Quercus ilex from Central Spain. Nova Hedwigia, 63, 347–360.
Couch, H. B. (1995). Diseases of turfgrasses (3rd ed.). Malabar: Krieger Publishing Company.
Csinos, A. S., & Stephenson, M. G. (1999). Evaluation of fungicides and tobacco cultivar resistance to Rhizoctonia solani incited target spot, damping off and sore shin. Crop Protection, 18, 373–377.
Dmitriev, D., & Rakitov, R. (2008). Decoding of superimposed traces produced by direct sequencing of heterozygous indels. PLoS Computational Biolology, 4(7), e1000113.
Fang, X., Finnegan, P. M. & Barbetti M. J. (2013). Wide variation in virulence and genetic diversity of Binucleate Rhizoctonia isolates associated with root rot of strawberry in Western Australia. Plos One, 8, number e55877.
Farr, D. F., Bills, G. F., Chamuris, G. P., & Rossman, A. Y. (1995). Fungi on plants and plant products in the United States. Saint Paul: APS Press.
Felsenstein, J. (1985). Confidence limits on phylogenies: an approach using the bootstrap. Evolution, 39, 783–791.
Gardes, M., & Bruns, T. D. (1993). ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Molecular Ecology, 2, 113–118.
Garibaldi, A., Minuto, A., Bertetti, D., Nicoletti, R., & Gullino, M. L. (2003). First report of web blight on yellow-sage (Lantana camara) caused by Rhizoctonia solani in Europe. Plant Disease, 87, 875.
Garibaldi, A., Bertetti, D., & Gullino, M. L. (2006). First report of leaf blight caused by Rhizoctonia solani AG 1B on Madagascar periwinkle (Catharanthus roseus) in Italy. Plant Disease, 90, 1361.
Garibaldi, A., Bertetti, D., & Gullino, M. L. (2009a). First report of leaf blight on Hosta fortune caused by Rhizoctonia solani AG 4 in Italy. Plant Disease, 93, 432.
Garibaldi, A., Gilardi, G., Bertetti, D., & Gullino, M. L. (2009b). First report of leaf blight on Fan columbine (Aquilegia flabellata) caused by Rhizoctonia solani AG 4 in Italy. Plant Disease, 93, 433.
Gill, J. S., Sivasithamparam, K., & Smettem, K. R. J. (2000). Soil types with different texture affects development of Rhizoctonia root rot of wheat seedlings. Plant and Soil, 221, 113–120.
Gill, J. S., Sivasithamparam, K., & Smettem, K. R. J. (2001). Effect of soil moisture at different temperatures on Rhizoctonia root rot of wheat seedlings. Plant and Soil, 231, 91–96.
Gill, J. S., Hunt, S., Sivasithamparam, K., & Smettem, K. R. J. (2004). Root growth altered by compaction of a sandy loam soil affects severity of Rhizoctonia root rot of wheat seedlings. Australian Journal of Experimental Agriculture, 44, 595.
Goll, M. B., Schade-Schutze, A., Swart, G., Oostendorp, O., Schott, J. J., Jaser, B., & Felsenstein, F. G. (2014). Survey on the prevalence of Rhizoctonia spp. in European soils and determination of the baseline sensitivity towards sedaxane. Plant Pathology, 63, 148–154.
González, D., Carling, D. E., Kuninaga, S., Vilgalys, R., & Cubeta, M. A. (2001). Ribosomal DNA systematics of Ceratobasidium and Thanatephorus with Rhizoctonia anamorphs. Mycologia, 93, 1138–1150.
Haralson, J. C., Brannen, P. M., NeSmith, D. S., & Scherm, H. (2013). Chemical control of Cylindrocladium and Rhizoctonia root rots in blueberry propagation. Crop Protection, 44, 1–5.
Harris, A. R., Schisler, D. A., Neate, S. M., & Ryder, M. H. (1994). Suppression of damping-off caused by Rhizoctonia solani, and growth promotion, in bedding plants by binucleate Rhizoctonia spp. Soil Biology and Biochemistry, 26, 263–268.
Holcomb, G. E., & Carling, D. E. (2002). First report of web blight caused by Rhizoctonia solani on Catharanthus roseus in Louisiana. Plant Disease, 86, 1272.
Hua, G. K. H., Bertier, L., Soltaninejad, S., & Hofte, M. (2014). Cropping Systems and Cultural Practices Determine the Rhizoctonia Anastomosis Groups Associated with Brassica spp. in Vietnam. PLoS ONE, 9, number e111750. doi:10.1371/journal.pone.0111750
Hwang, J., & Benson, D. M. (2002). Biocontrol of Rhizoctonia stem and root rot of poinsettia with Burkholderia cepacia and binucleate Rhizoctonia. Plant Disease, 86, 47–53.
Hyakumachi, M., Priyatmojo, A., Kubota, M., & Fukui, H. (2005). New anastomosis groups, AG-T and AG-U of binucleate Rhizoctonia spp. causing root and stem rot of cut- flower and miniature roses. Phytopathology, 95, 784–792.
Izumitsu, K., Hatoh, K., Sumita, T., Kitade, Y., Morita, A., & Tanaka, C. (2012). Rapid and simple preparation of mushroom DNA directly from colonies and fruiting bodies for PCR. Mycoscience, 53, 396–401.
Jager, G., Velvis, H., Lamers, J. G., Mulder, A., & Roosjen, J. (1991). Control of Rhizoctonia solani in potato by biological, chemical and integrated measures. Potato Research, 34, 269–284.
Kataria, H. R., Hugelshofer, U., & Gisi, U. (1991). Sensitivity of Rhizoctonia species to different fungicides. Plant Pathology, 40, 203–211.
Kataria, H. R. & Gisi, U. (1996). Chemical control of Rhizoctonia species. In: SnehB, Jabaji-Hare S, Neate S, Dijst G, eds. Rhizoctonia species: taxonomy, molecular biology, ecology, pathology and disease control. Dordrecht, the Netherlands: Kluwer Academic Publishers. p 537–547.
Khan, F. U., Nelson, B. D., & Helms, T. C. (2005). Greenhouse evaluation of binucleate Rhizoctonia for control of R. solani in soybean. Plant Disease, 89, 373–379.
Kimura, M. (1980). A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16, 111–120.
Kuninaga, S., & Yokosawa, R. (1984). DNA base sequence homology in Rhizoctonia solani Kuhn. IV. Genetic relatedness within AG-4. Annals of the Phytopathological Society of Japan, 50, 322–330.
Kuramae, E. E., Buzeto, A. L., Nakatani, A. K., & Souza, N. L. (2007). rDNA-based characterisation of a new binucleate Rhizoctonia spp. causing root rot on kale in Brazil. European Journal of Plant Pathology, 119, 469–475.
Leiner, R. H., & Carling, D. E. (1994). Characterization of Waitea circinata (Rhizoctonia) isolated from agricultural soils in Alaska. Plant Disease, 78, 385–388.
Manici, L., & Bonora, P. (2007). Molecular genetic variability of Italian binucleate Rhizoctonia spp. isolates from strawberry. European Journal of Plant Pathology, 118, 31–42.
Martin, F. N. (2000). Rhizoctonia spp. recovered from strawberry roots in central coastal California. Phytopathology, 90, 345–353.
Mulenko, W., Majewski, T., & Ruszkiewicz-Michalska, M. (2008). A preliminary checklist of micromycetes in Poland. In W. Mułenko, T. Majewski, & M. Ruszkiewicz-Michalska (Eds.), A preliminary checklist of micromycetes in Poland (p. 752). Polish Academy of Sciences: Institute of Botany.
Muzhinji, N., Truter, M., Woodhall, J. W., & Van der Waals, J. E. (2015). Anastomosis groups and pathogenicity of Rhizoctonia solani and Binucleate Rhizoctonia from potato in South Africa. Plant Disease, 99, 1790–1802.
Naito, S. (2004). Rhizoctonia diseases: taxonomy and population biology (pp. 18–31). Japan-Argentina Joint Study, Buenos Aires, Argentina: Proceeding of the International Seminar on Biological Control of Soilborne Plant Diseases.
Nadarajah, K., Omar, N. S., Rosli, M. M., & Tze, O. S. (2014). Molecular Characterization and Screening for Sheath Blight Resistance Using Malaysian Isolates of Rhizoctonia solani. BioMed Research International. doi:10.1155/2014/434257
Ogoshi, A. (1975). Grouping of Rhizoctonia solani Kühn and their perfect stages. Review of Plant Protection Research, 8, 93–103.
Ogoshi, A. (1987). Ecology and pathogenicity of anastomosis and intraspecific groups of Rhizoctonia solani Kuhn. In R. J. Cook (Ed.), Annual Review of Phytopathology (p. 25). Palo Alto: Annual Reviews Inc..
Ogoshi, A. (1996). Introduction – the genus Rhizoctonia. In B. Sneh, S. Jabaji-Hare, S. Neate, & G. Dijst (Eds.), Rhizoctonia species: taxonomy, molecular biology, ecology, pathology, and disease control (pp. 1–9). The Netherlands: Kluwer Academic Publisher.
Ogoshi, A., Oniki, M., Araki, T., & Ui, T. (1983). Studies on the anastomosis groups of binucleate Rhizoctonia and their perfect states. Journal of the Faculty of Agriculture , 61, 244–260.Hokkaido University
Ohkura, M., Abawi, G. S., Smart, C. D., & Hodge, K. T. (2009). Diversity and aggressiveness of Rhizoctonia solani and Rhizoctonia-like fungi on vegetables in New York. Plant Disease, 93, 615–624.
Polizzi, G., Aiello, D., Castello, I., & Vitale, A. (2009a). First report of crown and root rot caused by Rhizoctonia solani AG-4 on Coprosma repens and C. lucida in Italy. Plant Disease, 93, 972.
Polizzi, G., Aiello, D., Vitale, A., Kato, M., & Hyakumachi, M. (2009b). First report of crown and root rot caused by binucleate Rhizoctonia AG-A on Dodonaea viscosa in Italy. Plant Disease, 93, 1347.
Polizzi, G., Aiello, D., Vitale, A., Lahoz, E., Nicoletti, R., & Hyakumachi, M. (2009c). First report of crown rot, stem rot, and root rot caused by binucleate Rhizoctonia AG-G on Viburnum tinus in Italy. Plant Disease, 93, 433.
Polizzi, G., Aiello, D., Castello, I., Guarnaccia, V., & Vitale, A. (2010a). First report of damping-off caused by Rhizoctonia solani AG-4 on Mediterranean fan palm in Italy. Plant Disease, 94, 125.
Polizzi, G., Aiello, D., Castello, I., Guarnaccia, V., & Vitale, A. (2010b). First report of crown rot and stem rot caused by Rhizoctonia solani AG-4 on marmalade bush in Italy. Plant Disease, 94, 486.
Polizzi, G., Aiello, D., Castello, I., Vitale, A., Kato, M., & Hyakumachi, M. (2010c). First report of crown and root rot caused by binucleate Rhizoctonia AG-A on Thryptomene saxicola in Italy. Plant Disease, 94, 275.
Polizzi, G., Aiello, D., Guarnaccia, V., Panebianco, A., & Formica, P. T. (2011a). First report of crown and root rot caused by Rhizoctonia solani AG-4 on banana passionflower (Passiflora mollissima) in Italy. Plant Disease, 95, 1194.
Polizzi, G., Aiello, D., Vitale, A., Guarnaccia, V., Panebianco, A., & Cirvilleri, G. (2011b). First report of damping-off caused by Rhizoctonia solani AG-4 on pink ipê (Tabebuia impetiginosa) in Italy. Plant Disease, 95, 78.
Pope, E. J., & Carter, D. A. (2001). Phylogenetic placement and host specificity of mycorrhizal isolates belonging to AG-6 and AG-12 in the Rhizoctonia solani species complex. Mycologia, 93,712–719.
Priyatmojo, A., Escopalao, V. E., Tangonan, N. G., Pascual, C. B., Suga, H., Kageyama, K., et al. (2001). Characterisation of a new subgroup of Rhizoctonia solani anastomosis group 1 (AG-1 ID), causal agent of a necrotic leaf spot on coffee. Phytopathology, 91, 1054–1061.
Rinehart, T. A., Copes, W. E., Toda, T., & Cubeta, M. A. (2007). Genetic characterisation of binucleate Rhizoctonia species causing web blight on azalea in Mississippi and Alabama. Plant Disease, 91, 616–623.
Saitou, N., & Nei, M. (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4, 406–425.
Schroeder, K. L., & Paulitz, T. C. (2008). Effect of inoculum density and soil tillage on the development and severity of Rhizoctonia root rot. Phytopathology, 98, 304–314.
Sharon, M., Kuninaga, S., Hyakumachi, M., & Sneh, B. (2006). The advancing identification and classification of Rhizoctonia spp. using molecular and biotechnological methods compared with the classical anastomosis grouping. Mycoscience, 47, 299–316.
Sharon, M., Kuninaga, S., Hyakumachi, M., Naito, S., & Sneh, B. (2008). Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience, 49, 93–114.
Sneh, B. (1998). Use of non-pathogenic or hypovirulent fungal strains to protect plants against closely related fungal pathogens. Biotechnology Advances, 16, 1–32.
Sneh, B., Burpee, L., & Ogoshi, A. (1991). Identification of Rhizoctonia species. The APS: St. Paul, Press.
Sneh, B., Burpee, L., & Ogoshi, A. (1998). Identification of Rhizoctonia species. St. Paul: The APS.
Stevens Johnk, J., & Jones, R. K. (2001). Differentiation of three homogeneous groups of Rhizoctonia solani anastomosis group 4 by analysis of fatty acids. Phytopathology, 91, 821–830.
Strausbaugh, C. A., Eujayl, I. A., Panella, L. W., & Hanson L. E. (2011). Virulence, distribution, and diversity of Rhizoctonia solani from sugar beet in Idaho and Oregon. Canadian Journal of Plant Pathology, 33, 210–226.
Sumner, D. R. (1985). Virulence of anastomosis groups of Rhizoctonia solani and Rhizoctonia-like fungi on selected germ plasm of snap bean, lima bean, and cowpea. Plant Disease, 69, 25–27.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., & Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30, 2725–2729.
Toda, T., Mushika, T., Hayakawa, T., Tanaka, A., Tani, T., & Hyakumachi, M. (2005). Brown-ring-patch disease: a new disease on bentgrass caused by Waitea circinata Var. circinata. Plant Disease, 89, 536–542.
Ueyama, I., Araki, Y., Kurogochi, S., Yoneyama, K., & Yamaguchi, I. (1990). Mode of action of the phenyl urea fungicide pencycuron in Rhizoctonia solani. Pesticide Science, 30, 363–365.
Virgen-Calleros, G., Olalde-Portugal, V., & Carling, D. E. (2000). Anastomosis groups of Rhizoctonia solani on potato in Central Mexico and potential for biological and chemical control. American Journal of Potato Research, 77, 219–224.
Yang, G. (2013). Identification of AG-V, a new anastomosis group of binucleate Rhizoctonia spp. from taro and ginger in yunnan province. 5th International Symposium on Rhizoctonia, Zhengzhou, China, August 21–24.
Yang, Y. G., & Wu, X. H. (2013). First report of potato stem canker caused by binucleate Rhizoctonia AG-A in Jilin Province. China Plant Disease, 97, 1246.
Yang, G. H., Naito, S., Ogoshi, A., & Dong, W. H. (2006). Identification, isolation frequency and pathogenicity of Rhizoctonia spp. causing the wire stem of red birch in China. Journal of Phytopathology, 154, 80–83.
Yang, Y. G., Zhao, C., Guo, Z. J., & Wu, X. H. (2015). Characterisation of a new anastomosis group (AG-W) of binucleate Rhizoctonia, causal agent for potato stem canker. Plant Disease, 99(12), 1757–1763.
Acknowledgements
This work was supported by MIUR project number PON01_01611 (SO.PRO.ME: Sustainable production of potted plants in Mediterranean environment). The authors thank Prof. Masao Arakawa (Faculty of Agriculture, Meijo University, Japan) for his contribution in the molecular characterisation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aiello, D., Guarnaccia, V., Formica, P.T. et al. Occurrence and characterisation of Rhizoctonia species causing diseases of ornamental plants in Italy. Eur J Plant Pathol 148, 967–982 (2017). https://doi.org/10.1007/s10658-017-1150-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-017-1150-8