Abstract
From 2007 to 2013, a disease of Welsh onion, causing leaf sheath rot and concomitant death of outer leaves was found in 20 fields in Hokkaido, Japan. We obtained 20 Rhizoctonia isolates from diseased tissues and identified them based on the number of nuclei, hyphal fusion reactions, and molecular techniques using specific PCR primers and sequence of the rDNA-ITS region. The 20 isolates consisted of 16 multinucleate and four binucleate isolates. Of the multinucleate isolates, five were found to be so far unknown and designated here as Rhizoctonia solani AG-4 hybrid subgroup between HG-I and HG-II. Others were identified as AG-1 IB (three isolates), AG-2-2 IIIB (two isolates), AG-4 HG-I (two isolates), AG-1 IC (one isolate), AG-2-1 (one isolate), AG-4 HG-II (one isolate) and AG-5 (one isolate). All four binucleate isolates were binucleate Rhizoctonia AG-U. Original symptoms were reproduced on all plants inoculated with these isolates. Thus, we revealed that as many as nine taxa of Rhizoctonia spp. were associated with the disease. This is the first report of leaf sheath rot of Welsh onion caused by Rhizoctonia spp.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Welsh onion (Allium fistulosum L.) is native to Southwest China and commonly cultivated in East Asian countries such as Japan, China and Korea (Ford-Lloyd and Armstrong 1993; Kumazawa and Katsumata 1965). In the 2014 growing season, approximately 22,900 ha were planted with Welsh onion in Japan. In Hokkaido, the northernmost island of Japan, 2-month-old seedlings of Welsh onion are raised in a greenhouse, transplanted into open fields from late April to mid June, then harvested 4 months later (from August to October). Leaf sheaths of the plants are covered three times with soil by hilling with an interval of 1 month to promote etiolated growth. Some farmers also cultivate the plants in greenhouses from autumn to the following spring. Welsh onion is usually cultivated continuously without rotation with other crops in Hokkaido. Leaf sheath rot and outer leaf death of Welsh onion were first found to occur in fields and a greenhouse in Hokkaido from 2007 to 2013, and Rhizoctonia spp. were consistently isolated from diseased tissues.
Rhizoctonia solani is a species complex separated by hyphal anastomosis interactions. Currently, the R. solani complex is divided into 13 anastomosis groups (AGs): AGs 1 through 13 (Carling et al. 2002). Several AGs are further divided into subgroups on the basis of pathogenicity, biochemical and genetic makers. Binucleate Rhizoctonia is also classified based on hyphal anastomosis interactions. To date, 18 AGs A through W, except for AG-J, M, T, N and V, are recognized (Sharon et al. 2008; Yang et al. 2015).
Leaf sheath rot of Welsh onion caused by Rhizoctonia spp. has not been reported so far, therefore, we aimed to confirm their pathogenicity, to identify and to characterize the isolates.
Materials and methods
Disease occurrence and fungal isolation
The disease was surveyed in 20 Welsh onion fields, including one greenhouse in major production areas in Hokkaido from 2007 to 2013 (Table 1). Rotted basal parts of leaves (Fig. 1b) were sampled from each field. A total of 40–60 leaf pieces (ca. 5 × 5 mm) were cut from ca. 10 leaves collected randomly from each field. Leaf pieces were washed in running water, blotted on sterilized filter paper, and then placed on 9-cm-diameter petri dishes containing potato dextrose agar (PDA) amended with 20 mg/L streptomycin sulfate. After incubation at 25 °C for 2 days, 24–30 isolates typical of Rhizoctonia spp. were selected for each field, transferred to water agar (WA) plates, and grown for 2 days at 25 °C for hyphal tip isolation. Hyphal tip isolates were placed on PDA plates (6 isolates per plate) and incubated at 25 °C. Ten days after incubation, one isolate was selected from isolates with similar cultural morphology to maintain as barley grain culture (Naito et al. 1993) at − 20 °C until use.
Leaf sheath rot and outer leaf death of Welsh onion caused by Rhizoctonia spp. a Outer leaf death on Welsh onion plants infected with Rhizoctonia solani AG-4 HG-I in Hokuto, Hokkaido in 2007. b Rot symptoms of leaf sheath and basal part of outer leaf near the soil line on diseased plant. c Outer leaf death of Welsh onion plant 7 days after inoculation with R. solani AG-4 hybrid subgroup between HG-I and HG-II isolate WY31. d–g Cultural morphology of four isolates after 22 days on PDA at 25 °C. d Isolate WH31 (AG-4 hybrid subgroup between HG-I and HG-II). e Isolate WH41 (AG-4 hybrid subgroup between HG-I and HG-II). f Isolate WLS11 (AG-4 HG-I). g Isolate WNn11 (AG-4 HG-II)
Identification of isolates
The isolates were examined for the number of nuclei per cell and cultural morphology as described previously (Misawa and Kuninaga 2010, 2013). Hyphal anastomosis tests were conducted according to Rinehart et al. (2007) with a modification. All the isolates were paired with tester isolates belonging to R. solani AGs 1 − 5 (Misawa et al. 2015) or with the binucleate Rhizoctonia AGs A through U, excluding AGs J, M, T and N (Misawa and Toda 2013). Isolates were grown on PDA for 3 days at 25 °C. Mycelial plugs (5 mm diameter) were taken from colony margins of each Welsh onion isolates and AG tester isolates and paired ca. 2 cm apart on glass slides (76 × 26 mm) coated with WA, placed on a paper towel moistened with distilled water in a plastic box (24 × 17 × 8.5 cm) and incubated at 25 °C. From 2 to 3 days after incubation, the overlapping portion of mycelia was observed microscopically, and fusion frequency was determined as follows: low, <30%; moderate, 30–50%; high, >50% (Sneh et al. 1991), and the interactions were scored as C0–C3 reactions based on the scale developed by Carling (1996).
PCR amplifications using AG- or subgroup-specific primers were used to detect R. solani AG-1 IA, IB and IC (Kuninaga 2003), AG-2-1 (Carling et al. 2002), AG-2-2 IIIB, IV and LP (Carling et al. 2002), AG-4 HG-I, HG-II and HG-III (Kuninaga 2003) and AG-5 (Arakawa and Inagaki 2014) (Table 2). These primers were designed for the internal transcribed spacer region on ribosomal DNA (rDNA-ITS). Their sequences, annealing temperatures, and product size are shown in Table 2. Fungal DNA was extracted using a DNeasy Plant Mini Kit (Qiagen, Chatsworth, CA, USA) from 10-day-old mycelia grown at 25 °C on PDA plates. PCR amplification was performed in 25 μL mixture containing 0.2 μL template DNA, 200 μM each of primer pairs, and 12.5 μL HotStarTaq Master Mix Kit (Qiagen) in a GeneAmp PCR system 9700 (Applied Biosystems, Foster City, CA, USA). Cycling conditions consisted of 95 °C for 15 min; followed by 30 cycles of denaturation at 94 °C for 40 s, annealing at the temperature specific for each primer pair for 1 min, and extension at 72 °C for 1 min; then a final extension for 5 min at 72 °C. PCR products were separated on 2% agarose gels, stained with ethidium bromide, and viewed using a UV transilluminator.
The rDNA-ITS regions of the binucleate isolates were amplified using primers ITS4 and ITS5 (White et al. 1990) since specific primers were only available for AGs A–F and K (Arakawa and Inagaki 2014). DNA was extracted as described above. PCR reactions consisted of 95 °C for 15 min; followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 1 min, and extension at 72 °C for 1 min; then final extension for 10 min at 72 °C. PCR products were purified using the Wizard SV Gel and PCR Clean-Up System (Promega, Madison, WI, USA) and rDNA-ITS region of the isolates were directly sequenced using the same primers for amplification. DNA was sequenced by Hokkaido System Science Co. Ltd (Sapporo, Japan).
A part of the rDNA-ITS region of R. solani AG-4 hybrid subgroup between HG-I and HG-II (AG-4 HG-I + II h.s.) isolate WLY21 was amplified using the subgroup-specific primer pairs for AG-4 HG-I or HG-II (described above) to compare its sequence with the sequences of R. solani AG-4 isolates deposited in the DDBJ/EMBL/GenBank databases. The PCR products were directly sequenced using the same primers for amplification.
Mycelial growth temperature relations and cultural morphology
Radial mycelial growth rate of the isolates on PDA was determined at different temperatures, i.e., 5, 10, 15, 20, 25, 30, 35 and 40 °C. The radius of each colony was measured at 24 h intervals until 72 h after incubation. Each treatment was made with three replicates. For description of culture morphology, a 6-mm-diameter agar disk from the edge of a PDA colony was placed in the center of a PDA plate and incubated at 25 °C for 22 days in the dark. The morphology of each culture was then evaluated in terms of color, zonation and formation of sclerotia.
Genetic similarity among field isolates of binucleate Rhizoctonia
To clarify genetic relationship among the field isolates of binucleate Rhizoctonia belonging to the same AG, we tested hyphal fusion between each isolate and the other isolates as described above.
Pathogenicity
Six-month-old Welsh onion plants (cv. Motokura) grown in 1/5000-a (15.9 cm diameter × 19.3 cm depth) plastic pots filled with commercial potting soil (Pottace, Katakura Chikkarin Co., Tokyo, Japan), which had been autoclaved at 121 °C for 60 min, were used for pathogenicity tests. Pots each with single plants were girdled around the mouth with a plastic film (30 × 60 cm), secured with thick plastic strings, and received soil-inoculum mixture up to 10 cm thick on the soil surface (Fig. 1c). The soil-inoculum mixture was prepared as follows: 6-mm-diameter mycelia disks were placed on sterilized wheat bran and incubated at 25 °C for 2 weeks; and 5 g of wheat bran culture were mixed with 1.5 L of sterilized soil and used as the soil-inoculum mixture. Inoculated plants were grown in a greenhouse with average temperatures of 19.4–25.0 °C. Sterilized wheat bran served as the control. Virulence on Welsh onion was determined 7 days after inoculation. Each inoculation test consisted of two replicate pots.
Results
Disease occurrence
Leaf sheath rot and outer leaf death of Welsh onion (Fig. 1a) were found on 10 cultivars in 20 fields varying in acreage from 40 to 36,000 m2 including one greenhouse in five cities i.e., Hokuto, Yakumo, Naganuma, Nanporo and Date, from early August to mid September, except Field 3 (Table 1). Plants cultivated in Field 3 (greenhouse) showed symptoms in December (Table 1). The disease occurred from 2 to 3 months after transplanting. Leaf sheaths of diseased plants turned brown and became soft 0–10 cm below the soil line. Outer leaves submerged in the soil were also diseased (Fig. 1b). Outer leaves ultimately died when they were girdled by lesions. Inner leaves, which were not in contact with soil, remained healthy (Fig. 1a). The incidence of plants with symptoms ranged between 5 and 100% (Table 1). All plants were diseased to various extents in six fields, i.e., Fields 6, 9, 10, 15, 19 and 20. Nine other fields, i.e., Fields 1, 2, 5, 11, 12, 13, 14, 17 and 18, were severely damaged with 50–90% diseased plants. One to five leaves were diseased per plant (Table 1).
Identification of isolates
Rhizoctonia-like fungi were frequently recovered from diseased tissues in all 20 fields (Table 3). All isolates collected from the same fields were similar to each other in culture morphology; therefore, one isolate each representing 20 fields was examined in detail. Hyphae lacked clamp connections, and young hyphae branched near the distal septum and were constricted near the base of branching. These features indicated that the isolates belonged to the genus Rhizoctonia (Ogoshi 1987). Sixteen isolates were multinucleate (3–18 nuclei), and hyphal diameter ranged from 5.2 to 11.3 μm (mean 7.3–8.6 μm) (Table 3). They were identified as R. solani, according to the descriptions of Ogoshi (1987). Four other isolates (isolates WLS21, WH71, WY11 and WNg11) had two nuclei per cell, and hyphal diameter ranged from 3.7 to 6.8 μm (mean 4.7–5.5 μm) (Table 3). These four isolates were identified as binucleate Rhizoctonia according to the descriptions of Ogoshi (1987).
Thirteen isolates of R. solani, i.e., WLS1, WLS11, WLY21, WH11, WH31, WH41, WH51, WY31, WNn11, WNn21, WNn31, WD11 and WD21, anastomosed with the tester isolate of AG-1 or AG-4 or AG-5 each with a high fusion frequency of more than 50% with their respective testers (Table 3). One isolate (isolate WLS91) and two isolates (isolates WLS81 and WH21) showed the C2 reaction, e.g., cell wall fused, with the AG-2-1 and AG-2-2 tester isolates, respectively, with high fusion frequencies of more than 50%, and the C1 reaction with tester isolates AG-2-2 and AG-2-1, respectively, with frequencies less than 30%, respectively (Table 3). All isolates failed to anastomose with other tester isolates.
Based on the results of the hyphal fusion tests (Table 3) and PCR with the specific primers (Fig. 2), three isolates (isolates WH51, WNn21 and WD11), one isolate (isolate WNn31), one isolate (isolate WLS91), two isolates (isolates WLS81 and WH21), two isolates (isolates WLS1 and WLS11), one isolate (isolate WNn11) and one isolate (isolate WD21) were identified as R. solani AG-1 IB, AG-1 IC, AG-2-1, AG-2-2 IIIB, AG-4 HG-I, AG-4 HG-II and AG-5, respectively (Table 3). DNA of five isolates (isolates WLY21, WH11, WH31, WH41 and WY31) that anastomosed with the AG-4 tester isolate, reacted with the primer sets both for AG-4 HG-I and HG-II, producing PCR products of the expected size (Fig. 2). Here, we refer to them as R. solani AG-4 HG-I + II h.s. Sequence of isolate WLY21 analyzed by AG-4 HG-I specific primers (deposited in GenBank as LC090072) was closely related to sequences from AG-4 HG-I isolates with identities of 99.7–100% (AB000018, KF907732–907733, KC405624–405628, FJ480862–480865, DQ102446, AY154307). Isolate WLY21 was then analyzed using AG-4 HG-II specific primers (deposited in GenBank as LC090073) to obtain sequence. The sequence was identical with those of AG-4 HG-II isolates (AB000008, JX843818, FM867593–867594, AF354373–354074, HQ629858–629861). These results indicate that isolate WLY21 has the rDNA-ITS regions of both AG-4 HG-I and AG-4 HG-II.
Agarose gel electrophoresis of PCR products amplified from DNA of Rhizoctonia solani isolates from Welsh onion. Amplifications were done using primer pairs designed for specific detection of R. solani AG-4 HG-I (lanes 1, 4 and 7), HG-II (lanes 2, 5 and 8) and HG-III (lanes 3, 6 and 9). Lane M 100-bp ladder. Genomic DNA from isolate WLS1 (lanes 1–3), isolate WNn11 (lanes 4–6) and isolate WLY21 (lanes 7–9)
All four binucleate Rhizoctonia isolates (isolates WLS21, WH71, WY11 and WNg11) had the C2 reaction with the AG-U tester, with a high fusion frequency and also with the AG-P tester, but with a low fusion frequency. They failed to react to other tester isolates. Sequences of the rDNA-ITS regions of the four binucleate isolates (deposited in GenBank as LC090068–090071) were closely related to sequences from AG-U isolates with identities of 99.9–100% (HQ269825) or 99.3–99.8% (HQ269809–269811, HQ269816, HQ269820, AB196664–196666, AB731590). These results agreed with the results from hyphal fusion tests, and these four isolates were identified as binucleate Rhizoctonia AG-U (Table 3). Twenty isolates obtained in this study were deposited in Genebank, National Agriculture and Food Research Organization (NARO) with the accession numbers as shown in Table 3.
Mycelial growth temperature relations and cultural morphology
Most isolates, including binucleate Rhizoctonia did not grow at 5 or 40 °C on PDA (Table 4). Growth was optimal at 25–30 °C in most isolates, and isolates of R. solani AGs 1 and 4 tended to grow faster (Table 4). The results agreed with Watanabe and Matsuda (1966), Ogoshi (1976), and Hyakumachi and Sumino (1984) for R. solani and with Priyatmojo et al. (2001) and Hyakumachi et al. (2005) for binucleate Rhizoctonia.
Morphology of 22-day-old PDA cultures of AG-4 HG-I + II h.s. isolates were compared with the related subgroups; all eight AG-4 isolates had slight aerial mycelia, and all five AG-4 HG-I + II h.s. isolates showed radial mycelia growth as was the case with AG-4 HG-I isolate WLS11 (Fig. 1d–f). While cultures of all five AG-4 HG-I + II h.s. isolates and AG-4 HG-I isolate WLS1 were creamy to pale brown and mealy (Fig. 1d–e), those of isolates WLS11 (AG-4 HG-I) and WNn11 (AG-4 HG-II) were creamy and mealy (Fig. 1f–g). Isolate WNn11 was characterized by a concentric mycelia growth pattern (Fig. 1g). A few medium-brown sclerotia were produced, which ranged from 0.2 to 0.5 mm across for isolates WH11 and WNn11 (Fig. 1g), 0.5–2 mm across for isolate WH41 (Fig. 1e) and 1–1.5 mm across (isolate WY31). Thus, mycelial growth temperature relations and cultural morphology of AG-4 HG-I + II h.s. isolates were similar to those of AG-4 isolates reported previously (Ogoshi 1976; Stevens Johnk and Jones 2001). Cultural morphology of isolates belonging to other AGs agreed with that in previous papers (Hyakumachi and Sumino 1984; Hyakumachi et al. 2005; Ogoshi 1976; Priyatmojo et al. 2001; Watanabe and Matsuda 1966) (data not shown).
Genetic similarity among field isolates of binucleate Rhizoctonia
The four binucleate Rhizoctonia AG-U isolates from Welsh onion exhibited the C2 reaction with each other, indicating that they were not clonal.
Pathogenicity
All 20 isolates consistently incited leaf sheath rot and outer leaf death on Welsh onion plants (Fig. 1c). The symptoms were similar to those of naturally diseased plants, and leaves attached to the infested soil died. The isolates were reisolated from inoculated, diseased plants. There was no significant difference in virulence among the 20 isolates. Control plants remained healthy, and the pathogen was not isolated.
Discussion
This study demonstrated that as many as nine groups of Rhizoctonia spp. (R. solani AG-1 IB, AG-1 IC, AG-2-1, AG-2-2 IIIB, AG-4 HG-I, AG-4 HG-II, AG-4 HG-I + II h.s., and AG-5, in addition to binucleate Rhizoctonia AG-U) were associated with the leaf sheath disease of mature Welsh onion plants. Young plants are more susceptible to the pathogen (Baker 1970), and several AGs and subgroups of Rhizoctonia spp. have been reported in Japan as damping-off pathogens of single crops, e.g., seven groups from sugarbeet seedlings, i.e., R. solani AG-1, AG-2-1, AG-2-2, AG-3, AG-4, AG-5 and unidentified binucleate Rhizoctonia (Naito et al. 1975) and four groups from broccoli seedlings, i.e., R. solani AG-1 IC, AG-2-1, AG-2-2 IIIB and AG-4 HG-I (Kubota and Abiko 1997; Kubota et al. 2009; Misawa et al. 2013; Yamauchi et al. 2009). In the case of mature plants, only root rot of Japanese radish was reported as the disease being caused by R. solani (five groups: AG-1, AG-2-1, AG-2-2, AG-4 and AG-5; Homma and Ishii 1984). The diversity of the Welsh onion leaf sheath rot pathogens suggests that mature Welsh onion plants are, at least in some plant parts, as susceptible to Rhizoctonia spp. as seedlings of other crops.
Since the rDNA-ITS region of R. solani AG-4 HG-I + II h.s. isolates could not be analyzed by direct sequencing due to their genetic heterogeneity, alternative methods were cloning and PCR amplification using the subgroup-specific primers. Using the latter method, we successfully distinguished AG-4 HG-I + II h.s. from the known three subgroups (HG-I, HG-II and HG-III). To our knowledge, there is no report on genetic heterogeneity among R. solani AG-4 subgroups. AG-4 HG-I + II h.s. is likely to incite diseases on other crops in the world. Since this fungus was almost indistinguishable from isolates of AG-4 HG-I and AG-4 HG-II on the basis of cultural morphology, the possibility of their coexistence in single fields cannot be ruled out. Further research on AG-4 HG-I + II h.s. is needed to clarify the geographic distribution and host range of this group. PCR using AG-4 HG-I + II specific primers would facilitate identification of this group.
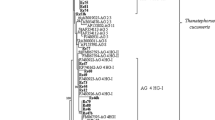
Field isolates are usually heterokaryons in AG-4 (Anderson et al. 1972), and the new subgroup AG-4 HG-I + II h.s. is one of the examples. Heterokaryosis and genetic heterogeneity normally occur through sexual recombination of genotypes; basidiospores of Thanatephorus cucumeris, the teleomorph of R. solani, germinate to develop primary mycelia, and hyphal fusion between primary mycelia results in heterogenic, secondary mycelia. Cubeta et al. (1993) paired many homokaryotic single-basidiospore isolates from different field AG-4 isolates to confirm heterokaryosis at the colony junction. We assume that inter-subgroup mating of primary mycelia between AG-4 HGs-I and -II to establish HG-I + II h.s.
Alternatively, Kuninaga et al. (2004) examined heterokaryosis in isolates of the R. solani AG-1 hybrid subgroup between IA and IE (AG-1 IA + IE h.s.), inciting common bean web blight. They obtained homokaryotic isolates of the pathogen using a protoplast method developed by Phillips (1993) to prepare homokaryons from heterokaryotic field isolates without basidiospore formation and identified genotypes; protoplast progenies were identified as AG-1 IA or AG-1 IE. These subgroups were then paired to generate AG-1 IA + IE h.s. Their results represent the basis of our assumption, although we have neither observed basidiospores of T. cucumeris nor found hybrid subgroups in AG-1 in Welsh onion fields.
In this study, we obtained four binucleate Rhizoctonia AG-U isolates from three main Welsh onion production cities (Hokuto, Yakumo and Naganuma) in 2012–2013. Hyphal fusion tests using these isolates demonstrated that they were not clonal, implying that AG-U was originally present in the soil of various regions of Hokkaido. AG-U was first isolated from roses with root and stem rot in Gifu, Japan, in 1999–2001 (Hyakumachi et al. 2005; Priyatmojo et al. 2001), and later recognized on other crops (Misawa and Toda 2013; Rinehart et al. 2007). Therefore, AG-U should be monitored on other hosts, and critical assessment of disease damage is necessary.
This study, along with our subsequent routine survey from 2006 to 2014 revealed the epidemiology of the disease except greenhouse cultivation (Field 3) as follows: (1) the disease occurs from early July (in 2006; data not shown) to mid September in Hokkaido with average daily temperatures, ranging from 19 to 23 °C; (2) plants become diseased 5 to 10 days after hilling, but disease does not develop further; (3) outer leaves in contact with soil are killed entirely, whereas inner leaves with no soil contact, remain healthy. (4) Because new inner leaves develop every week, healthy leaves gradually increase in number until next hilling; and (5) if the next hilling is done when the daily average temperature is more than 19 °C, the disease occurs again.
The maximum temperature for pathogenesis could not be clarified from field surveys. The maximum average daily temperature of 23 °C occurs in Hokkaido in mid August. Since optimal growth temperatures of the pathogens other than AG-2-1 were at 25–30 °C, it may be inferred that the disease occurs between 25 and 30 °C.
Although the damage varied among fields in terms of diseased plant ratio (Table 1), inoculation experiments failed to reveal any significant difference in virulence among the isolates. Since hilling is an important factor to promote disease occurrence, differences in field disease severity seems to correlate with cultural practices such as the height and period of hilling.
The only Rhizoctonia disease so far known to occur on Welsh onion in Japan has been damping off of seedlings up to 2 months old caused by Rhizoctonia spp. (Yamamoto and Uehara 1972). The pathogens, however, were not identified to the AG. The leaf sheath rot we report here occurs later than damping off, between 2 and 3 months after transplanting (4 to 5 months after sowing), and the symptoms are different. These features distinguish leaf sheath rot from damping off and represent a basis for recognizing leaf sheath rot as a new disease. We refer to this new disease as “leaf sheath rot of Welsh onion (Rhizoctonia youshou-fuhai-byo in Japanese)”.
References
Anderson JB, Stretton HM, Groth JV, Flentje NT (1972) Genetics of heterokaryosis in Thanatephorus cucumeris. Phytopathology 62:1057–1065
Arakawa M, Inagaki K (2014) Molecular markers for genotyping anastomosis groups and understanding the population biology of Rhizoctonia species. J Gen Plant Pathol 80:401–407
Baker KF (1970) Types of Rhizoctonia diseases and their occurrence. In: Parmeter JR (ed) Rhizoctonia solani: biology and pathology. University of California Press, Berkeley, pp 125–148
Carling DE (1996) Grouping in Rhizoctonia solani by hyphal anastomosis reaction. In: Sneh B, Jabaji-Hare S, Neate S, Dijst G (eds) Rhizoctonia species taxonomy, molecular biology, ecology, pathology and disease control. Kluwer, Dordrecht, pp 37–47
Carling DE, Kuninaga S, Brainard KA (2002) Hyphal anastomosis reactions, rDNA-internal transcribed spacer sequences, and virulence levels among subsets of Rhizoctonia solani anastomosis group-2 (AG-2) and AG-BI. Phytopathology 92:43–50
Cubeta MA, Briones-Ortega R, Vilgalys R (1993) Reassessment of heterokaryon formation in Rhizoctonia solani anastomosis group 4. Mycologia 85:777–787
Ford-Lloyd BV, Armstrong SJ (1993) Welsh onion Allium fistulosum L. In: Kalloo G, Bergh BO (eds) Genetic improvement of vegetable crops. Pergamon Press, Oxford, pp 51–58
Homma Y, Ishii M (1984) Anastomosis groups of Rhizoctonia solani Kühn responsible for various symptoms of root rot of Japanese radish (in Japanese with English summary). Bull Shikoku Agric Exp Stn 42:1–9
Hyakumachi M, Sumino A (1984) New morphological type (IC) in Rhizoctonia solani AG-1 isolated from the sugarbeet-manufactory-waste-soils and some of its characteristics (in Japanese with English summary). Jpn J Phytopathol 50:507–514
Hyakumachi M, Priyatmojo A, Kubota M, Fukui H (2005) New anastomosis groups, AG-T and AG-U, of binucleate Rhizoctonia spp. causing root and stem rot of cut-flower and miniature roses. Phytopathology 95:784–792
Kubota M, Abiko K (1997) Rhizoctonia solani Kühn isolated from damping-off broccoli (in Japanese). Proc Kansai Pl Prot 39:33–34
Kubota M, Tomioka K, Sato T (2009) Damping-off of broccoli caused by Rhizoctonia solani AG-1 IC (in Japanese). Annu Rep Kansai Pl Prot 51:27–28
Kumazawa S, Katsumata K (1965) Welsh onion. In: Kumazawa S (ed) The theory of vegetable and horticulture (in Japanese). Yokendo, Tokyo, pp 280–289
Kuninaga S (2003) Current situation of the taxonomy of Rhizoctonia solani (in Japanese). Plant Prot 57:219–222
Kuninaga S, Godoy-Lutz G, Yokosawa R (2004) ITS heterogeneity within an individual isolate of Rhizoctonia solani AG-1 (Abstract in Japanese). Jpn J Phytopathol 70:219
Misawa T, Kuninaga S (2010) The first report of tomato foot rot caused by Rhizoctonia solani AG-3 PT and AG-2-Nt and its host range and molecular characterization. J Gen Plant Pathol 76:310–319
Misawa T, Kuninaga S (2013) First report of white leaf rot on Chinese chives caused by Rhizoctonia solani AG-2-1. J Gen Plant Pathol 79:280–283
Misawa T, Toda T (2013) First report of black scurf on carrot caused by binucleate Rhizoctonia AG-U. J Gen Plant Pathol 79:86–88
Misawa T, Yamazaki K, Takada K (2013) Damping-off of broccoli caused by Rhizoctonia solani AG-2-1 (in Japanese with English summary). Annu Rep Plant Prot North Jpn 64:60–64
Misawa T, Kubota M, Sasaki J, Kuninaga S (2015) First report of broccoli foot rot caused by Rhizoctonia solani AG-2-2 IV and pathogenicity comparison of the pathogen with related pathogens. J Gen Plant Pathol 81:15–23
Naito S, Sugimoto T, Yamaguchi T, Fujisawa I (1975) Anastomosis groups of Rhizoctonia solani Kühn isolated from diseased sugar beet seedlings (in Japanese with English summary). Res Bull Hokkaido Natl Agric Exp Stn 111:25–35
Naito S, Yamaguchi T, Sugimoto T, Homma Y (1993) A simple method for the long-time culture storage of Rhizoctonia spp. on barley grains (in Japanese with English summary). Ann Rept Plant Prot North Jpn 44:20–23
Ogoshi A (1976) Studies on the grouping of Rhizoctonia solani Kühn with hyphal anastomosis and on the perfect stages of groups (in Japanese with English summary). Bull Nat Inst Agric Sci Ser C 30:1–63
Ogoshi A (1987) Ecology and pathogenicity of anastomosis and intraspecific groups of Rhizoctonia solani Kühn. Ann Rev Phytopathol 25:125–143
Phillips AJL (1993) The use of protoplasts for the preparation of homokaryons from heterokaryotic isolates of Rhizoctonia solani. Mycol Res 97:456–460
Priyatmojo A, Yotani Y, Hattori K, Kageyama K, Hyakumachi M (2001) Characterization of Rhizoctonia spp. causing root and stem rot of miniature rose. Plant Dis 85:1200–1205
Rinehart TA, Copes WE, Toda T, Cubeta MA (2007) Genetic characterization of binucleate Rhizoctonia species causing web blight on azalea in Mississippi and Alabama. Plant Dis 91:616–623
Sharon M, Kuninaga S, Hyakumachi M, Naito S, Sneh B (2008) Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience 49:93–114
Sneh B, Burpee L, Ogoshi A (1991) Identification of Rhizoctonia species. APS Press, St. Paul, pp 17–18
Stevens Johnk J, Jones RK (2001) Differentiation of three homogeneous groups of Rhizoctonia solani anastomosis group 4 by analysis of fatty acids. Phytopathology 91:821–830
Watanabe B, Matsuda A (1966) Studies on grouping Rhizoctonia solani Kühn pathogenic to upland crops (in Japanese with English summary). Bull Appointed Exp 7:1–131
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yamamoto T, Uehara H (1972) Damping-off of Welsh onion and onion and their control (in Japanese). Plant Prot 4:153–156
Yamauchi N, Miura Y, Shirakawa T (2009) Damping-off of broccoli caused by Rhizoctonia solani AG-4 HG-I (in Japanese with English summary). Annu Rep Plant Prot North Jpn 60:105–107
Yang YG, Zhao C, Guo ZJ, Wu XH (2015) Characterization of a new anastomosis group (AG-W) of binucleate Rhizoctonia, causal agent for potato stem canker. Plant Dis 99:1757–1763
Acknowledgements
This study was supported in part by Grants-in-Aid for Regional R&D Proposal-Based Program from Northern Advancement Center for Science & Technology of Hokkaido Japan. For help with this work, we thank staff members of Hokkaido Prefectural Oshima, Sorachi and Iburi Agricultural Extension Centers, Japan. We thank A. Sumino (Hokkaido Research Organization) for valuable comments and Dr. N. Matsumoto (Hokkaido University, Japan) for critically reading the manuscript and correcting the English.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence data reported are available in the DDBJ/EMBL/GenBank databases as accessions LC090068–090073.
Rights and permissions
About this article
Cite this article
Misawa, T., Kurose, D. & Kuninaga, S. First report of leaf sheath rot of Welsh onion caused by nine taxa of Rhizoctonia spp. and characteristics of the pathogens. J Gen Plant Pathol 83, 121–130 (2017). https://doi.org/10.1007/s10327-017-0706-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10327-017-0706-y