Abstract
Condensin is an integral component of the mitotic chromosome condensation machinery, which ensures orderly segregation of chromosomes during cell division. In metazoans, condensin exists as two complexes, condensin I and II. It is not yet clear what roles these complexes may play outside mitosis, and so we have examined their behaviour both in normal interphase and in premature chromosome condensation (PCC). We find that a small fraction of condensin I is retained in interphase nuclei, and our data suggests that this interphase nuclear condensin I is active in both gene regulation and chromosome condensation. Furthermore, live cell imaging demonstrates condensin II dramatically increases on G1 nuclei following completion of mitosis. Our PCC studies show condensins I and II and topoisomerase II localise to the chromosome axis in G1-PCC and G2/M-PCC, while KIF4 binding is altered. Individually, condensins I and II are dispensable for PCC. However, when both are knocked out, G1-PCC chromatids are less well structured. Our results define new roles for the condensins during interphase and provide new information about the mechanism of PCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromosome condensation at mitosis is one of the most fascinating, visually dynamic and important processes in eukaryotic cells (Belmont 2006). Correct packaging of chromosomes is essential for ensuring orderly and even segregation of chromosomes during cell division and for many other essential cell functions including gene regulation. In multicellular eukaryotes, the 10-nm chromatin fibres that predominate during interphase eventually compact and thicken to form the 700-nm-wide chromatids of the metaphase chromosome (Belmont 2006, Maeshima et al. 2014).
Recent data show that mitotic chromosome condensation is not smooth and progressive as is commonly thought but rather a discontinuous process driven by alternating accumulation and release of mechanical stress (Liang et al. 2015). Perhaps surprisingly, when the volume occupied by chromosomes is measured, chromosomes are found to be only two- to threefold more condensed during mitosis than during interphase (Vagnarelli 2012).
Chromosome condensation is often viewed as a hierarchical series of events. Histone modifications are thought to play a leading role in condensation, but they work in tandem with non-histone proteins that help both to compact and organise higher order structures (Saitoh et al. 1994, Hudson et al. 2009, Moser and Swedlow 2011, Wilkins et al. 2014). These non-histone proteins comprise the chromosome scaffold, a loose network that is capable of tethering chromosome loops when histones are extracted (Paulson and Laemmli 1977, Paulson 1989). Core components of the chromosome scaffold include the condensins, topoisomerase II and chromokinesin (KIF4) (Samejima et al. 2012). These proteins display distinct and overlapping axial localisation in mitotic chromosomes. How they are able to cooperate with histones to organise mitotic chromosomes is still not fully clear.
Condensins I and II are pentameric complexes, each composed of the catalytic subunits SMC2 and SMC4, which are both ATPases, plus three auxiliary regulatory subunits (Hirano et al. 1997, Ono et al. 2003). The auxiliary polypeptides in condensin I are CAP-G, CAP-D2 and CAP-H; in the condensin II complex, they are CAP-G2, CAP-D3 and CAP-H2 (for reviews, see Ono et al. 2003, Hirota et al. 2004, Bazile et al. 2010, Hirano 2014, Frosi and Haering 2015). Condensin II is predominately nuclear during interphase and is bound to the chromosomes during mitosis. By contrast, condensin I is commonly viewed as being excluded from the nucleus until nuclear envelope breakdown (NEBD), which marks the transition from prophase to pro-metaphase early in mitosis (Maeshima and Laemmli 2003, Hirota et al. 2004, Gerlich et al. 2006). Mitotic chromosomes are wider and shorter, with a diffuse chromosome scaffold, when they are depleted of condensin I, while condensin II-depleted chromosomes retain more well-defined scaffolds but the chromosomes are more elongated and appear to lack axial rigidity (Green et al. 2012). Little information is available about what role the condensins play at the end of anaphase during chromosome decondensation or how condensins are redistributed to their cellular compartments.
In this study of chicken DT40 cells, we focus on two poorly understood aspects of chromosome condensation. First, we examine condensin behaviour during interphase using quantitative live cell imaging in conjunction with 3D reconstruction. This approach allows the roles of condensin I and II in G1 cells to be spatially and temporally dissected. Remarkably, we identify a pool of condensin I present in interphase nuclei which mediates chromosome condensation and gene regulation. Furthermore, we find that the cellular level of CAP-D3 (and thus condensin II) dramatically increases as cells pass from mitosis to G1-phase, implying an important role for condensin II at this stage of the cell cycle. These results give important new insights into the behaviour of the condensins during interphase and establish foundations for the second and major focus of this study: understanding the mechanism of premature chromosome condensation (PCC).
PCC occurs when chromosomes are induced to condense outside mitosis in G1-, S- or G2-phase. PCC has gained considerable attention in recent years with studies showing that it can drive chromosome rearrangements and can lead to certain diseases (Gotoh and Durante 2006, Ravi et al. 2012). Despite the increased interest, PCC remains poorly understood. There are conflicting hypotheses regarding the events that lead to PCC as well as those that follow it. In addition, the mechanism of PCC and the proteins involved are ill defined. For example, it is not yet clear whether the process recruits the key players involved in mitotic chromosome condensation.
PCC was first studied over 35 years ago when Johnson and Rao showed that an interphase cell fused with a mitotic cell will undergo NEBD and chromosome condensation (Johnson and Rao 1970). PCC chromosomes have distinct morphologies depending on the cell cycle phase. G1-PCC consist of single elongated chromatids, S-PCC appear as “pulverised chromosomes” and G2-PCC closely resemble mitotic metaphase chromosomes (Stevens et al. 2010, Ravi et al. 2012).
It is becoming increasingly clear that PCC does not just occur in artificial situations but also can happen naturally and lead to major genomic rearrangements and is observed in cancer (Williams et al. 1976, Hittelman et al. 1984, Zhang et al. 2013). One theory maintains that chromosome fragmentation (also called pulverisation or shattering) is a phenomenon restricted to cells that are already in mitosis or cells that are in S-phase and undergo PCC (Stevens et al. 2010, Crasta et al. 2012).
In this study, we define how the scaffold proteins behave in cells undergoing PCC. Previously, Ono et al. established a protocol for analysis of condensins in PCC cells and elegantly showed condensin II was necessary for late S-phase PCC in HeLa cells, with late S-phase PCC chromatids losing structure in CAP-H2 RNAi cells (Ono et al. 2013). We have extended this study by examining the roles of condensins I and II in G1 and G2/M-PCC chromosomes for the first time using our chicken DT40 conditional condensin knockouts. Surprisingly, chromosomes in PCC show mislocalisation of the chromosome scaffold protein KIF4. We find both condensins I and II show strong axial staining in G1-PCC chromatids, reminiscent of their mitotic chromosome localisation. Furthermore, using conditional knockouts of condensin I and II, we show that G1 prematurely condensed chromosomes appear to form normally when either condensin I or condensin II is depleted but not when both are depleted.
Material and methods
Cell culture, synchronisation and Calyculin A treatment
Chicken lymphoma B DT40 cell culture was performed as described previously (Buerstedde and Takeda 1991). CAP-H-GFP-streptavidin-binding peptide (SBP) H2B-RFP, CAP-D3-GFP-SBP H2B-RFP, CAP-H-GFP-SBP and CAP-D3-GFP-SBP rescue cell lines were grown in RPMI medium supplemented with 200 ng/ml doxycycline (dox). The SBP tag also is fused to a His-tag, S-peptide referred to collectively as the triple affinity purification (TrAP) tag (Hudson et al. 2008), but for simplicity and as this study only used SBP, we have described the fusion as GFP-SBP. Depletion of tetracycline-repressible CAP-H, CAP-D3 or SMC2 (Green et al. 2012, Kim et al. 2013) was induced by addition of 200 ng/ml dox for 36 h (CAP-H depletion) or 24 h (CAP-D3 and SMC2 depletion), respectively. For mitotic (M-phase) controls, cells were blocked with 500 ng/ml nocodazole (Noc) for 8 h. For a G1/S block, cells were treated with 4 mM thymidine (TdR) for 14 h as described previously (Kim et al. 2013). For G1 synchronisation by serum starvation, cells were maintained in serum-free medium for 48 h before being released into medium supplemented with 10 % FBS. For induction of PCC, synchronised or unsynchronised cells were treated for 1 h with 50 nM Calyculin A (Cal A) (LC Laboratories, Woburn, MA, USA).
Live cell imaging, microscopy and analysis
CAP-H-GFP-SBP H2B-RFP and CAP-D3-GFP-SBP H2B-RFP cells were studied by live cell imaging as described previously (Green et al. 2012, Bakhrebah et al. 2015). In brief, image stacks of whole cells were taken with 0.2-μm Z sections at 2-min intervals for 4–16 h using a DeltaVision widefield deconvolution microscope (Applied Precision) equipped with an environment control chamber. During live cell imaging, cells were maintained at 37 °C in a 5 % CO2 atmosphere. Images were deconvolved, aligned and projected in 2D using SoftWoRx 4.1. For CAP-H-GFP and CAP-D3-GFP intensity measurements, all images were automatically processed in the Fiji distribution of ImageJ using a custom written macro (Schindelin et al. 2012). Details of the analysis are as follows: Channels for the protein of interest and the DNA marker (H2B) were separated. A Gaussian blur (sigma = 1) was applied to the DNA channel and a specific cell of interest was selected from the end of the series using the wand tool. The selected cell was binarised and used as a mask to analyse the intensity of the protein of interest throughout the whole time series. Both mean and integrated intensity for each channel were recorded for each time point. Their ratios were calculated and standardised to a user-selected starting reference point. Immunofluorescent images were captured using either a Zeiss Axio Imager M1 microscope, a DeltaVision widefield deconvolution microscope or a Zeiss LSM 780 confocal microscope and processed by AxioVision 4.7 (Zeiss), SoftWoRx 4.1 or Zeiss ZEN 2012, respectively. For cell size analysis, whole cells and nuclei were measured using the polygon function of SoftWoRx 4.1. For 3D modelling, 200 × 0.1-μm Z section images were recorded and analysed using the volume modelling mode of SoftWoRx 4.1 and IMARIS 8.1.2. Line profiles across the cells were analysed by Fiji distribution of ImageJ. Box plots were generated using beeswarm R package (https://cran.r-project.org/web/packages/beeswarm/index.html). Statistical analyses were conducted using Student’s t test (unpaired). P values less than 0.001 were considered as indicating statistically significant differences. The hypergeometric test in Fig. 3c was performed by phyper function in R. The probability p values less than 0.05 were considered as indicating statistically significant differences.
Immunofluorescent staining and EdU labelling
Chromosome spreading and immunofluorescent staining were performed as previously described (Earnshaw et al. 1989). In brief, cells were treated hypotonically for 7 min before fixation with methanol/acetic acid (3:1). Fixed cells were dropped onto slides and stained with rabbit anti-CAP-H (1:400, Vagnarelli et al. 2006), rabbit anti-CAP-D2 (1:400), rabbit anti-KIF4 (1:400), rabbit anti-topo IIα (1:1000, Samejima et al. 2008), rabbit anti-centromere protein O (CENP-O) (1:400, Kwon et al. 2007), or mouse anti-SBP (1:400, Green et al. 2012). Secondary antibodies were donkey anti-rabbit Alexa Fluor 488 (1:1000, Invitrogen) and goat anti-mouse Alexa Fluor 594 (1:1000, Invitrogen). For mouse anti-Lamin B1 staining (1:400), cells were fixed with 4 % paraformaldehyde (PFA), permeabilised with 0.3 % Triton X-100 and blocked with 3 % BSA in PBS. Cells were mounted with VectaShield containing DAPI (Vector Laboratories). For chromosome morphology staining in Fig. 7a, cells were prefixed with 1 % PFA for 10 min and resuspended in 0.2 % Triton X-100 in PBS. For method 1, cells were cytospun onto slides, postfixed with 2 % PFA at room temperature for 15 min, permeabilised with 0.5 % Triton X-100 and mounted with VectaShield containing DAPI. For methods 2–6, cells were treated with a hypotonic KCl solution at 37 °C for 10 min, then prefixed with 1 % PFA, cytospun and postfixed with 2 % PFA (method 2), or cytospun and fixed with 1 % PFA in KCM (120 mM KCl, 20 mM NaCl, 10 mM Tris–HCl, 0.5 mM EDTA and 0.1 % Triton X-100) or 1 % PFA in PBS (methods 3 and 4) or fixed with methanol/acetic acid (method 5), before mounting with VectaShield containing DAPI. In order to detect newly synthesised DNA by 5-ethynyl-2'-deoxyuridine (EdU) labelling, cells were grown in medium containing 10 μM EdU for 0.5 h followed either by Cal A treatment or immediate fixation. EdU was detected with Click-iT EdU imaging kits (Invitrogen) according to the manufacturer’s procedure.
3D FISH protocol and statistical analyses
Three-dimensional fluorescent in situ hybridization (3D FISH) was performed as described by Cremer et al. (2008, 2005). In brief, slides with CAP-HOFF cells (+dox 36 h) or CAP-HON cells were incubated in 0.3 × PBS for 1 min; cells were then fixed in 4 % paraformaldehyde/0.3 × PBS for 10 min. After 20-min incubation in 0.5 % Triton X/PBS, the slides were incubated in 20 % glycerol/PBS at 4 °C overnight. The slides were frozen in liquid nitrogen for 30 s and thawed on a piece of tissue paper on the next day and was repeated four times for 3 × 5 min in 0.05 % Triton X/PBS, followed by an incubation in 0.1 N HCl. Next, the slides were washed 2 × 1 min in 2 × SCC and an overnight incubation in 50 % formamid/2 × SCC at 4 °C. The slides were incubated in RNAase/2 × SSC solution for 30 min at 37 °C before hybridisation. Zrep and rDNA probes are unique and specific to chromosome Z in the chicken DT40 cells. The Zrep probe is specific to a heterochromatic region and 1.6 kb repeat unit arranged in 24-kb blocks and spanning 15–20 Mb (Hori et al. 1996). The rDNA probe recognises the 18S gene of a euchromatic locus, and its length is 1.8 kb repeats covering 200 kb (Kim et al. 2013). These probes were labelled via a 30-μl nick translation reaction containing 2 μg DNA, 15 μl Biotin-dNTP mix (0.4 mM), 3 μl 10 × NEB buffer-2, 3 μl BSA (10×), 4 μl DNase (40 mU/μl), 1 μl DNA Pol I (20 U/μl) and H2O. Both probes were denatured at 95 °C for 5 min and co-denatured with the slides at 85 °C for 5 min, then incubated at 37 °C for 3 days in a humid chamber. After hybridisation, the slides were washed stringently and incubated with Avidin-conjugated Alexa 488 (1:500). DNA were stained with DAPI. The slides were analysed by a DeltaVision widefield deconvolution microscope. For volume measurement in Fig. 3a, 200 × 0.1-μm Z section images were recorded from 30 × 2 cells (Zrep experiment) and 15 × 2 cells (rDNA experiment) in CAP-HON and CAP-HOFF cells, respectively. These images were further analysed using the volume modelling mode of SoftWoRx 4.1, Zrep and rDNA and their corresponding nuclei were 3D reconstructed. The ratio of Zrep volume to nuclei volume and the ratio of rDNA volume to nuclei volume in CAP-HON and CAP-HOFF cells were subjected to statistical analysis and shown as box plots using beeswarm R package.
Flow cytometry
FACS analysis was performed as previously described (Kim et al. 2013) and analysed using FACSCalibur (Becton Dickinson) and Cell Quest (Becton Dickinson). For apoptosis analysis, Annexin V (Bio Vision, Inc) was used according to the manufacturer’s instruction.
Protein extraction, Western blotting and nuclear extraction
For analysis of whole cell proteins, cells were lysed with lysis buffer consisting of 50 mM Tris–HCl pH 7.4, 250 mM NaCl, 0.5 % NP-40, 30 μg/ml RNase A and protease inhibitors (Roche) before sonication with a Bioruptor (Diagenode). Protein (40 μg) from each sample was subjected to SDS-polyacrylamide gel electrophoresis in 4–15 % Mini-Protean TGX gradient gels (Bio-Rad) and immunoblotted as previously described (Kim et al. 2013). Primary antibodies for Western blotting were mouse anti-SBP (1:2000, Hudson et al. 2008), rabbit anti-CAP-H (1:2500, Hudson et al. 2008), rabbit anti-SMC2 (1:2500, Saitoh et al. 1994), rabbit anti-CAP-D3 (1:2500, Green et al. 2012), rabbit anti-CAP-D2 (1:2500, Hudson et al. 2008) and mouse anti-α tubulin (1:1000, Sigma). Secondary antibodies were goat anti-rabbit IgG-HRP and rabbit anti-mouse IgG-HRP (1:10,000). Nuclear extraction was conducted according to the protocol of Wang and Caldwell (2006). In brief, cells were pelleted by centrifugation and the pellet was resuspended in 5× packed cell volume (PCV) equivalent of isotonic lysis buffer consisting of 10 mM Tris-HCl, pH 7.5, 2 mM MgCl2, 3 mM CaCl2 and 0.32 M sucrose supplemented with a 1/100 protease inhibitor cocktail (Roche; Complete Mini, EDTA-free) and 1 mM DTT. After 15-min incubation on ice, 10 % IGEPAL CA-630 (Sigma-Aldrich) was added to a final concentration of 0.3 %. After centrifugation, the supernatant (cytoplasmic fraction) was collected. The pellet was resuspended in 2/3 × PCV equivalent of extraction buffer consisting of 20 mM HEPES, pH 7.7, 1.5 mM MgCl2, 0.42 M NaCl, 0.2 mM EDTA and 25 % (v/v) glycerol supplemented with a 1/100 protease inhibitor cocktail and 1 mM DTT. The pellet was agitated for 15 min at 700 rpm followed by 15 min at 1400 rpm and then centrifuged for 15 min at 9400 rpm. The supernatant (nuclear fraction) was carefully transferred to a freshly chilled tube. Both cytoplasmic and nuclear fractions were recovered for protein concentration measurement and Western blotting.
RNA-seq, data analysis and bioinformatics
Chicken DT40 CAP-H knockout (KO) cells were treated with or without dox (100 ng/ml) for 36 h. Total RNA samples were extracted using a mirVanaTM miRNA Isolation Kit (Thermo Fisher Scientific) and subjected to sequencing using an Illumina Hiseq2000 platform. Library preparation and Illumina sequencing were performed by Macrogen (South Korea), with ~33 million 2 × 100 bp (paired end) reads per sample, which allows ~6.5× coverage. The sequence tags were spliced-mapped onto the chicken genome galGal4 using Tophat and Bowtie2 (Langmead and Salzberg 2012) following quality test using FASTQC. Differential expression of genes was analysed using the Bioconductor v2.3 package edgeR v3.2.3 (Robinson et al. 2010). Tag enrichment in NCBI RefSeq genes was calculated (Pruitt et al. 2012) between dox-treated (CAP-HOFF) and untreated cells (CAP-HON) using edgeR exact test with tag-wise dispersion estimation. P values were calculated and corrected using the Benjamini-Hochberg method. After the list of differential expression (DE) genes was obtained, a GO enrichment analysis was performed and the biological processes, cellular components and molecular functions associated with those DE genes were examined through the DAVID Bioinformatics Resources 6.7 (Young et al. 2010). The enrichment of the GO terms was calculated by the Fisher’s exact test. Additionally, the enriched networks, molecular functions and canonical pathways were identified in the Ingenuity Pathways Analysis (IPA, Ingenuity Systems, and www.ingenuity.com) platform (Mayburd et al. 2006, Pospisil et al. 2006). In the IPA analysis, the p values were determined by Fisher’s exact test. To elucidate the interactions among the DE genes with CAP-H enrichment, the STRING 10 software was applied, which builds functional protein-association networks based on compiled available experimental evidence (Szklarczyk et al. 2011). Heatmaps were plotted using the gplots R package (http://cran.r-project.org/web/packages/gplots/index.html). The data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus (Edgar et al. 2002, Barrett et al. 2013) and are accessible through GEO Series accession number GSE74892 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE74892).
Results
Analysis of the localisation of condensin I and II in interphase chicken DT40 cells
Our primary interest was to observe condensin behaviour during PCC, but it was necessary to first establish how condensins I and II are localised in interphase cells under normal circumstances. Much effort has been devoted to studying condensin behaviour during mitosis (using both fixed and live cell imaging), but little is known about the kinetics in interphase. Condensin I and II localisation were therefore examined in real time in chicken DT40 cells containing histone H2B-red fluorescent protein (RFP) and also containing either CAP-H/condensin I or CAP-D3/condensin II fused with green fluorescent protein (GFP) and the affinity tag GFP-SBP. These cells have the wild-type CAP-H and CAP-D3 switched off and are rescued by CAP-H-GFP-SBP and CAP-D3 GFP-SBP transgenes, respectively, with no apparent defects (Kim et al. 2013, Bakhrebah et al. 2015). Figure 1 shows that the behaviour of condensin I and II in chicken cells is similar to that described for other metazoan systems during mitosis (Ono et al. 2003, Hirota et al. 2004, Ono et al. 2004, Gerlich et al. 2006). Condensin I (CAP-H) accumulates on chromosomes following nuclear envelope breakdown and leaves chromatin rapidly after telophase (Fig. 1a), while condensin II (CAP-D3) is present on the chromatin throughout interphase as well as during mitosis (Fig. 1b).
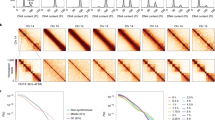
Spatial and temporal localisation of condensins during the cell cycle. Asynchronous CAP-H-GFP-SBP H2B-RFP and CAP-D3-GFP-SBP H2B-RFP cells were subjected to live cell imaging as described in the “Materials and methods”. For purposes of analysis, the time courses are split into two. Series 1 covers cells from interphase (G2) to metaphase, while series 2 covers from metaphase to anaphase and then to interphase (G1). Prometaphase is defined as the point of nuclear envelope breakdown. Anaphase is defined as the point of chromatid separation. In series 2, only one daughter was analysed. a CAP-H-GFP H2B-RFP live cell imaging. The ratio of mean nuclear CAP-H-GFP intensity to mean H2B-RFP intensity during entry into mitosis is plotted as a function of time (series 1, bottom left panel) relative to prophase, which occurred at 21:59 min in the time course. The ratio of mean nuclear CAP-H intensity to mean H2B-RFP intensity during exit from mitosis is plotted as a function of time (series 2, bottom right panel) relative to anaphase, which occurred at 98 min in the time course. b CAP-D3-GFP H2B-RFP live cell imaging is shown. The ratio of mean nuclear CAP-D3 intensity to mean H2B-RFP intensity during entry into mitosis is plotted as a function of time (series 1, bottom left panel) relative to prophase, which occurred at 16 min in the time course. The ratio of mean nuclear CAP-D3 intensity to mean H2B-RFP intensity during exit from mitosis is plotted as a function of time (series 2, bottom right panel) relative to anaphase, which occurred at 52 min in the time course. Error bars represent standard error of the mean (SEM), n = 15
Closer examination of the transition from mitosis to interphase for both condensins gave some surprising results. Quantitation of GFP fluorescence reveals that condensin II in fact increases in G1 nuclei relative to M, suggesting that it has an important functional role at this stage (Fig. 1b, series 2). Interestingly, we also find that condensin I remains in the G1-phase nuclei at a low level (Fig. 1a, series 2).
The presence of condensin I in interphase nuclei was also reported recently in a genome-wide analysis of condensin in human MCF-7 cells (Li et al. 2015). ChIP-seq analyses revealed that condensin I was bound to specific sites in G0/G1 cells synchronised to 80–95 %. Furthermore, binding to some sites such as ER-α was diminished during mitosis (Li et al. 2015). Since those results, and ours, challenge the commonly held view that condensin I is excluded from the interphase nuclei (Hirota et al. 2004, Gerlich et al. 2006), more exhaustive experiments were required.
To further test whether condensin I is present in interphase nuclei, we used fixed cell preparation epifluorescence deconvolution or confocal combined with 3D reconstruction and also examined the proteins in isolated nuclei by Western blotting. Interphase cells containing CAP-H-GFP-SBP and H2B-RFP were fixed with PFA and optically sectioned for 3D reconstruction. Orthogonal (XY, XZ and YZ) views clearly reveal a small pool of CAP-H signal in the nuclei, which was shown by quantitative analysis to comprise 20–30 % of the total CAP-H-GFP intensity in the cell (Fig. 2a). Line profile analysis across the cell in a single section also indicates a small pool of CAP-H signal in the nucleus. Interestingly, the pool of nuclear interphase condensin seems to be excluded from nucleoli (Fig. 2a–c).
A small pool of condensin I is retained in the interphase nuclei of chicken DT40 cells. a Asynchronous CAP-H-GFP H2B-RFP cells were fixed with 4 % PFA. Images of optical sections (0.2 μm) were recorded and analysed by Imaris 8.1.2. Orthogonal (XY, XZ and YZ) views are shown. The absolute GFP intensity in the nuclei (“nuclei”) was quantitated using the threshold signal in DAPI staining to define the nuclear periphery, and the total absolute GFP intensity of the whole cells (“total”) was also determined. Error bars represent 95 % confidence intervals (CI), n = 18. Line profiles across the cell were plotted and are shown at the bottom right. Scale bar represents 5 μm. b Asynchronous WT cells were fixed with 4 % PFA, stained with anti-CAP-H or anti-CAP-D2 antibodies and examined by confocal microscopy. Representative images are shown. Inserts are higher magnification of selected cells. Line profiles across the cell were plotted and are shown at the right. Scale bar represents 5 μm. c Asynchronous HCT116 (human) cells were fixed with 4 % PFA, stained with anti-hCAP-G antibody and examined by confocal microscopy. Representative images are shown. Insets show higher magnification views of quantitated nuclei. Scale bar represents 5 μm. d Protein components of condensin I were analysed by Western blotting. CAP-HOFF (CAP-H knock out cells), WT and CAP-H-GFP-SBP cells were synchronised with thymidine in interphase. Cytoplasmic and nuclear proteins were extracted, separated on SDS gels, blotted and probed with rabbit anti-CAP-D2, mouse anti-SBP, rabbit anti-CAP-H, mouse anti-α-tubulin and rabbit anti-histone H3 antibodies, respectively. Ponceau staining is shown in Online Resource 1d as a loading control. It is noted that the CAP-H antibody recognised a non-specific band at approximately the same size as the CAP-H-GFP-SBP. To simplify this figure, blots for mouse-anti-SBP which specifically detects CAP-H-GFP-SBP are shown
CAP-H-GFP-SBP cells were also examined using confocal microscopy, which uses a pinhole to reject out of focus light resulting in a thin optical section. These results clearly show some CAP-H-GFP signal in the nuclei (Online Resource 1a). Moreover, CAP-H-GFP-SBP cells were fixed, co-stained with anti-CAP-H and anti-SBP antibodies and viewed using confocal microscopy (Online Resource 1b). The resulting images show similar CAP-H/condensin I staining to those seen with CAP-H-GFP (Fig. 2a).
In further experiments, cells were fixed with PFA and stained with anti-Lamin B1 antibodies. Optical sectioning and orthogonal (XY, XZ and YZ) views reveal a pool of condensin I in nuclei, inside an intact ring of Lamin B1 (Online Resource 1c). In the line profile, a small pool of CAP-H-GFP signal can be detected inside the intact ring of Lamin B1, although its intensity is lower than that outside of the Lamin B1 ring (Online Resource 1c). Analysis of confocal data of CAP-H and CAP-D2 in wild-type chicken DT40 cells gave very similar results (Fig. 2b), suggesting that this observation is not due to the effect of the transgenic cell line. To alleviate any concern that this identification may be specific to chicken DT40 cells, human HCT116 cells were fixed and stained with anti-hCAP-G antibody. Here too, hCAP-G signal (indicating condensin I) can be seen inside the nucleus (Fig. 2c). We found that the nuclear condensin I signal is stronger relative to the cytoplasmic signal in WT chicken DT40 cells and human HCT116 cells than in CAP-H-GFP-SBP cells, suggesting CAP-H-GFP-SBP cells have more CAP-H in the cytoplasm. This is likely due to constitutive expression of CAP-H in the rescued cells producing excess protein that is unable to form the condensin I complex and enter the nucleus. Indeed, our previous pulldowns indicated CAP-H in excess of other condensin I subunits in the same CAP-H-GFP cell line (Kim et al. 2013). Nonetheless, these results clearly point to a pool of condensin I being retained in the interphase nucleus.
Finally, to test for the presence of condensin I in interphase nuclei using a different approach, nuclei were isolated from thymidine-blocked cells and immunoblotted with antibodies to condensin I subunits (CAP-H and CAP-D2). The immunoblots (Fig. 2d) confirm that condensin I is present in interphase nuclei. They also show that the proportion of condensin I present in the nuclei is about 20 % in WT DT40 cells, similar to that determined by microscopy (Fig. 2a). In the controls, α-tubulin is only detected in the cytoplasmic fraction, while histone H3 can only be detected in the nuclear fraction (Fig. 2d), showing that there is no cross-contamination between cytoplasmic and nuclear extracts. Consistent with immunofluorescence (IF) data in Fig. 2a–c, Online Resource 1a, b, the relative amount of condensin I in the nuclear fraction is greater in WT cells than in CAP-H-GFP-SBP cells.
In the course of live cell imaging studies of CAP-H-GFP and CAP-D3-GFP in H2B-RFP cells, we noted that the measured cell diameter and the behaviour of CAP-H could be used to establish the cell cycle phase of a specific cell. Live cell imaging was performed over 15 h using CAP-H-GFP/H2B-RFP and CAP-D3-GFP/H2B-RFP cells, and cell diameters were plotted as a function of time (Online Resource 2a). In cells shortly after cytokinesis, most of the cell diameters are less than 9 μm with CAP-H-GFP signal diffused throughout the whole cells, whereas cells in later phases of the cell cycle have cell diameters greater than 10 μm. This suggests that CAP-H-GFP localisation together with cell diameter could be a good indicator of the cell cycle stage, especially for G1 cells. These experiments establish important parameters for defining cell type (see below).
Significantly, normalised GFP intensity shows that the nuclear CAP-H-GFP signal gradually decreases after completion of cytokinesis until it reaches a level lower than at the beginning of this time course, presumably in late S-phase. This indicates possible removal of CAP-H from the nuclear pool to the cytoplasm pool as the cells move from a transcriptional to a replicative phase (Online Resource 2b). Together, these data describe new roles for condensin in interphase and also establish essential knowledge for understanding the behaviour of the condensins during PCC.
These results agree with the recent study using ChIP-seq analysis, mentioned above, which found that condensin I also exists in interphase nuclei from human cells (Li et al. 2015). They suggest that earlier reports confining chromosome-bound condensin I to mitotic cells overlooked a smaller pool bound to DNA during interphase. These results have important implications for the role of condensin I in gene regulation and are highly relevant to the PCC analyses described below.
Role of condensin I in interphase nuclei
Next, we were interested to find out what role condensin I plays in the structural organisation of genomic regions in the interphase nucleus. 3D FISH studies were conducted using Zrep and 18S rDNA probes to analyse cells with and without CAP-H (condensin I) (Lau et al. 2014). Zrep consists of a 10-kb repeating unit of heterochromatin on the single copy Z chromosome in the female chicken DT40 cell line. The entire array covers 24 kb, which spans 15–20 Mb. The euchromatic 18S rDNA is a 1.8-kb repeat array that maps morphologically to the Z chromosome making it an ideal complement to the Zrep. Another important feature of this analysis is condensin I is highly enriched at the 18S rDNA loci in chicken DT40 but undetectable in our analyses at the Zrep loci (Kim et al. 2013). Importantly for this study, our previous analysis of CAP-H KO cells showed that a time point (36 h + dox) can be chosen where no mitotic segregation or ploidy defects are observed, yet the CAP-H protein is entirely depleted (Green et al. 2012). After 36-h dox treatment, CAP-H was completely depleted (Online Resource 3a left panel). No ploidy defects were detected in the FACS profile in CAP-HOFF cells (Online Resource 3a right panel). Annexin V was further incorporated with PI for FACS analyses. Percentage of apoptotic cells (Annexin V positive and PI negative) in CAP-HON and CAP-HOFF samples are comparable and relatively low (7.7 vs 8.5 %) (Online Resource 3b). Further, damage-dependent phosphorylation of histone variant H2AX (γ-H2AX) foci was observed in adriamycin-treated CAP-HON cells but neither in untreated CAP-HON nor in CAP-HOFF cells (Online Resource 3c). These studies using asynchronous cells defined the time point (36 h + dox) could be further used for 3D FISH and RNA-seq experiments below.
Our analysis shows that in the absence of CAP-H, volumes increase significantly in size for the Zrep region and an even greater effect is observed at the rDNA region (Fig. 3a). This is perhaps not unexpected given that CAP-H/condensin I is known to bind strongly to the rDNA locus in chicken DT40 and in all other species tested (Wang et al. 2005, Kim et al. 2013, Huang et al. 2013). What is most surprising is that such a dramatic condensation defect is seen in interphase DNA in the absence of condensin I. These data further highlight that condensin I is present and also suggest it has a function in interphase nuclei.
Functional studies of CAP-H in interphase cells. a CAP-H KO cells were grown in the absence (CAP-HON) or presence (CAP-HOFF) of dox for 36 h in order to deplete CAP-H. Cells were collected for 3D FISH analysis using probes for Zrep and rDNA. Optical sections of images were taken and 3D reconstruction carried out. Representative 2D projected Zrep, DAPI and rDNA DAPI images and 3D reconstruction images are shown. The volumes of Zrep (green), rDNA (green) and whole nuclei (red) were measured and their ratios (Vol Zrep/Vol nuclei or Vol rDNA/Vol nuclei) were plotted. Scale bar represents 5 μm. Error bars represent 95 % confidence intervals (CI), n = 12. b CAP-H KO cells were grown in the absence or presence of dox for 36 h in order to deplete CAP-H. These cells were collected for RNA extraction and RNA-seq was conducted. An MA plot displays the log-fold changes with differentially expressed (DE) genes highlighted in red; blue lines indicate twofold changes at false discovery rate (FDR) <0.05, FC means fold change. CPM means counts per million. A summary of DE genes is shown at the right, with FDR cutoff <0.05. c A total of 116 DE genes were found to be overlapped with the set of genes differentially expressed when CAP-H is enriched. To obtain this set of genes, 712 differentially expressed genes with FDR <0.05 and fold change ≥ 2 in Fig. 3b were compared with 2360 CAP-H enrichment genes with CAP-H associated with their promoters from our previous condensin I Chip-seq studies (Kim et al. 2013). A Venn diagram is shown at the left. Hypergeometric test shows p = 0.02166379 (Berkopec 2007). These 116 DE gene products were analysed with STRING 10, and the confidence view of interaction for the ribosomal pathway is shown
Since condensin I occupies some territories inside nuclei during interphase, its removal might be expected to affect gene expression globally. Our previous genome-wide analysis found a total of 4369 CAP-H sites in the chicken DT40 genome, of which 2360 were associated with promoters of active genes from a total of 17,148 total promoters in DT40 (Kim et al. 2013). RNA-seq was therefore conducted using conditional CAP-H knockout cells (CAP-HON/OFF cells) to see if these CAP-H enriched promoters were specifically misregulated following CAP-H removal. After normalisation, a total of 3798 genes were found to be differentially expressed with 2495 downregulated and 1303 upregulated (Fig. 3b).
The human homologs were identified and analysed for 693 of the genes that were significantly misregulated in CAP-HOFF with at least a twofold change. Many of these human homologs were found to be involved in prostate cancer signalling or in the estrogen-dependent breast cancer signalling pathway (Online Resource 4a, b). In total, we found approximately 70 % (483/693) of these genes are involved in cancer and about 20 % (143/693) are involved in gene expression, consistent with our previous genome-wide condensin mapping studies (Kim et al. 2013). Tellingly, a recent analysis also found that condensins I and II play key roles in regulating the estrogen receptor during interphase in human cells, suggesting that this mechanism is conserved (Li et al. 2015).
A significant correlation was found between differentially expressed genes in CAP-HOFF cells in this analysis with the set of CAP-H enriched genes from our previous genome-wide condensin mapping studies of CAP-H/condensin I (Kim et al. 2013) (Fig. 3c). A total of 116 genes were found to be misregulated in the absence of CAP-H that normally bind to their promoter regions (Fig. 3c). A hypergeometric test was performed to determine whether more CAP-H bound genes are deregulated following CAP-H removal than would be expected by chance. The test shows the probability of obtaining 116 genes is 0.021, which indicated that this observation is statistically significant rather than by chance (Berkopec 2007). The corresponding heat map is displayed in Online Resource 5. GO enrichment studies showed that these 116 genes are involved in ribosome, ribonucleoprotein complex and RNA binding functions (Online Resource 4c). The fact that many ribosome pathway components were identified (Fig. 3c) is interesting because it is similar to previous studies in Drosophila (Smith et al. 2013). These experiments indicate that removal of CAP-H leads to significant misregulation of gene expression, suggesting a key role for condensin I in transcriptional control during interphase.
Categories of PCC in DT40 cells
In this study, Cal A was used to induce PCC. Cal A is a highly cytotoxic metabolite originally isolated from the marine sponge Discodermia calyx (Kato et al. 1988), and various studies have shown that it is a cell-permeable inhibitor of protein phosphatases 1 and 2A (Ishihara et al. 1989, Fagerholm et al. 2010). Cal A is a very useful drug for studying PCC because it directly mimics the phenotypes observed when PCC is induced by fusing mitotic and interphase cells (Gotoh 2009, Paulson and Mause 2013). It does so rapidly and homogeneously and at very low doses, so it is ideal for the large-scale biochemical experiments necessary to understand PCC. Indeed, recent studies highlight this drug as the most potent way to induce (and examine) PCC in a tractable cell biological system (Ravi et al. 2012, Ono et al. 2013).
In asynchronous DT40 cells treated with Cal A, G1-, S- and G2/M-PCC chromatids can be observed in chromosome spreads as shown in Fig. 4a. G1-PCC chromatids are single, thin and wiggly; S-PCC shows a “pulverised chromosome” appearance characterised by partially condensed but discontinuous dots; G2/M-PCC yields thick, rod-shape condensed chromosomes with paired chromatids. It is thought G2-PCC chromosomes are similar to chromosomes in early prophase (Stevens et al. 2010, Ono et al. 2013, Gallego-Paez et al. 2014), but in general, they may not be distinguishable from condensed mitotic chromosomes. For this reason, we have broadened the classification to G2/M in our study. Another study (Gallego-Paez et al. 2014) has classified the short, stubby chromosomes (representing 7 % of total PCC chromosomes in our study, Fig. 4a) as G2, but we feel we cannot exclude the possibility some are from M-phase; hence, these are labelled then as G2/M. These highly distinctive short “butterfly like” chromosomes are specific to PCC and not seen in normal mitotic cells.
PCC in chicken DT40 cells. a Asynchronous WT cells were treated with 50 nM Cal A for 1 h and then hypotonically swollen and fixed with methanol/acetic acid (3:1). Chromosomes were spread onto the slide and stained with DAPI. Representative G1-, S- and G2/M-PCC chromosomes are shown with their respective percentage of cell population. Control (no Cal A) mitotic is shown on the right. Scale bar represents 5 μm. b Asynchronous CAP-H-GFP-SBP cells were treated with or without Cal A for 1 h. Cells were collected for fixation with methanol/acetic acid and further processed for immunofluorescent staining of CENP-O (green) and DNA staining with DAPI (blue). Representative G1-PCC chromatids are shown in the upper right panels and mitotic chromosomes from cells without Cal treatment are shown as a control in the lower right panels. Insets show higher magnification views of single chromatids in G1 or paired, duplicated sister chromatids in M. Scale bar represents 5 μm. G1 synchronised PCC cells are shown in the left panel for comparison. c After 47.5-h serum starvation, WT cells were given a pulse of EdU for 0.5 h before being washed and released into fresh medium with 10 % serum and 50 nM Cal A as shown in the schematic diagram on the right top panel. After 1-h Cal A treatment, cells were collected for FACS (right middle panel) and IF stained for EdU incorporation, as shown in the bottom right panel. As a control, asynchronous WT cells were given a pulse of EdU 0.5 h before being fixed and further processed for IF staining. Representative G1, mid-S, late-S and M cells with EdU (red) and DAPI (blue) staining are shown in the left panel
In order to confirm the identification of G1-PCC, chromosome spreads were viewed by immunofluorescence microscopy following staining with an anti-centromere protein-CENP-O antibody. We found a unique single dot corresponding to each chromosome in PCC cells where wiggly, thin single chromatids were seen, confirming that these are unreplicated G1-PCC chromosomes (Fig. 4b). By contrast, double dots were observed in the replicated sister chromatids in metaphase cells. For subsequent characterisations, these results establish a clear demarcation between different types of PCC chromosomes based on morphology.
Condensin I and II localise to PCC chromosomes
The condensin I and II complexes are core scaffold proteins that play a pivotal role in chromosome condensation and shaping. We were interested to see how they behave during PCC. We first examined whether condensin subunits are present in G1-PCC chromatids. Due to the low percentage of G1 cells in asynchronous culture (Franklin and Sale 2006) (Fig. 4a), serum starvation was used to synchronise cells in G0/G1. Serum was then re-added and the cells were treated with Cal A to induce G1-PCC, as previously performed by Gallego-Paez et al. (2014).
Flow cytometry data (Fig. 4c) showed that the serum-starved cells had 2N DNA content, confirming that they were in G0/G1. The synchronised cells showed no EdU labelling, thus confirming tight synchronisation in G1-phase and the absence of any S-phase cells. In a control unsynchronised culture, strong EdU signals were seen in S-phase cells (Fig. 4c).
Strikingly, we found that both condensin I and condensin II were associated with chromatids undergoing Cal A-induced G1-PCC in the CAP-H-GFP-SBP cells using anti-SBP and CAP-D3 antibody (Fig. 5a). In a reciprocal experiment, we obtained similar results in the CAP-D3-GFP-SBP cells using anti-SBP and CAP-H antibodies (Online Resource 6a). Condensin I has previously been found in G1-PCC chromosomes (Ono et al. 2013), but our data shows that condensin II is also present. Both condensins I and II appear on the central axis of the single chromatid, though condensin I staining appears to be stronger. However, even in normal mitotic chromosomes from chicken DT40 cells, condensin I is significantly more enriched compared to condensin II (Ohta et al. 2010).
Behaviour of chromosome scaffold proteins in PCC cells. a Asynchronous CAP-H-GFP-SBP cells were treated with 50 nM Cal A for 1 h. Cells were collected, fixed with methanol/acetic acid (3:1) and co-stained with anti-SBP (red) to detect CAP-H-GFP-SBP and anti-CAP-D3 (green) antibodies. DNA was stained with DAPI (blue). Representative G1-PCC chromosomes are shown. Inserts are higher magnifications of single chromatids. Scale bar represents 5 μm. b CAP-H-GFP-SBP cells were treated as described in Fig. 5a. Cells were collected, fixed and co-stained with KIF4 (green) and SBP (red) antibodies. DNA was stained with DAPI (blue). Representative G1-PCC, G2/M-PCC and control mitotic chromosomes (from cells without Cal A treatment) are shown. Inserts are higher magnifications of single chromatids and duplicated sister chromatids. Scale bar represents 5 μm. c CAP-H-GFP-SBP cells were treated as described in Fig. 5a. Cells were collected, fixed and co-stained with topo IIα (green) and SBP (red) antibodies. DNA was stained with DAPI (blue). Representative G1-PCC, G2/M-PCC and control mitotic chromosomes (from cells without Cal A treatment) are shown. Inserts are higher magnifications of single chromatids and duplicated sister chromatids. Scale bar represents 5 μm
Topo II and condensin colocalise in PCC chromosomes, but KIF4 binding is altered
Although the behaviour of the condensins during PCC was the main focus of this study, we also examined two other key scaffold components, KIF4 and topo IIα, which cooperate with the condensins in shaping mitotic chromosomes (Samejima et al. 2012). Cells containing CAP-H-GFP-SBP were stained with anti-SBP and co-stained with anti-KIF4 or anti-topo IIα antibodies.
In Fig. 5b, the control mitotic chromosome spread shows characteristic axial KIF4 and CAP-H (condensin I) staining. However, more discontinuous KIF4 staining is evident in G1-PCC chromosome spreads, suggesting some mislocalisation of KIF4 in G1-PCC. More noticeably, in G2/M-PCC, KIF4 appears diffuse instead of axial along the condensed chromatids, significantly different from the axial CAP-H staining.
Topoisomerase IIα (topo IIα), another partner of condensin in shaping mitotic chromosome structure, maintained its characteristic axial staining in both G1-PCC and G2/M-PCC chromosome spreads (Fig. 5c). In contrast to KIF4, topo IIα and condensin I broadly overlap and appear continuous in G1-PCC. They both retain their distinct axial localisation, reminiscent of mitotic chromosomes.
We performed live cell imaging of DT40 cells that harboured a H2B-GFP construct as they were undergoing PCC to monitor condensation in real time and to determine the end stage of Cal A-treated cells as a control. Online Resource 7a (lower series) shows that DNA begins to condense at around 54 min after the start of the Cal A treatment, and DNA fragmentation with cell membrane disintegration is seen at around 210 min after Cal A addition. Towards the end of the time course, the H2B-GFP signal becomes much dimmer and appears in smaller pieces, indicating chromosome fragmentation (Online Resource 7a, upper series).
Further analysis of Cal A-treated cells using live cell imaging shows that all cells undergo discernible PCC, followed by chromosome shattering, regardless of what cell cycle stages they were in at the start of the Cal A treatment (Online Resource 7b). At later time points (Cal A post 4 h), the DIC (lower panel) shows the cell is dissolved with the matching H2B signal (top panel) showing chromosome fragmentation. This was not apparent in cells treated with Cal A for less than 3 h. We note the fragmentation was always accompanied by cell disintegration, implying chromosome fragmentation is likely to be a result of the cell disintegrating and being exposed to external nucleases.
Damage-dependent phosphorylation of histone variant H2AX (γ-H2AX) was also observed in Cal A-treated cells, suggesting the presence of DNA damage in these prematurely condensed chromosomes (Online Resource 7c). Double strand breaks cause accumulation of γ-H2AX foci, and we found these present in about 94 % of Cal A-treated cells, in all phases of the cell cycle. These results further demonstrate that PCC results in chromosome breakage (El Achkar et al. 2005). Interestingly, Rad51, a central player in homologous recombination (HR), was present in synchronised S-phase cells (obtained by release from a hydroxyurea block) and showed dimmer signals in asynchronous interphase cells but not in the PCC cells (Online Resource 7d).
Interphase pool of condensin I concentrates on PCC chromosomes before NEBD
We were next interested in understanding whether the pool of condensin I that binds to PCC chromosomes comes from the population of condensin I inside the nucleus or from the cytoplasmic pool. In other words, does condensin I bind to PCC chromosomes before or after NEBD?
To perform this analysis, it was first necessary to determine how much time elapses before NEBD occurs after G1 cells are treated with Cal A. We quantitated the NEBD time with Cal A treatment using live cell imaging of CAP-H-GFP-SBP/H2B-RFP cells. Cells with much more obvious cytoplasmic CAP-H-GFP signal and a cell diameter more than 9 μm had significantly shorter NEBD time (50 min) than cells with diffused CAP-H-GFP signal and a cell diameter less than 9 μm (75 min) (Fig. 6a). This suggests that G1 cells require at least 75 min of Cal A treatment before NEBD occurs. Moreover, the smaller the cells are at the beginning of the Cal A treatment, and thus the earlier in the cell cycle they are, the longer the time before NEBD occurs. Similar results were obtained with live cell imaging of CAP-D3-GFP-SBP/H2B-RFP cells (Online Resource 6b). These data suggested 60 min post-Cal A treatment as the optimal time to examine G1-PCC cells in order to find cells that are undergoing Cal A-induced chromosome condensation but also have an intact nuclear envelope.
G1-PCC cells with intact nuclear membranes. a CAP-H-GFP-SBP H2B-RFP cells were subjected to live cell imaging (2-min intervals) starting within 7 min after the addition of 50 nM Cal A. NEBD times (min) were measured from the time that Cal A was added to the time point when NEBD occurred. NEBD times are plotted according to where CAP-H-GFP was localised at t = 0 min (either cytoplasmic or diffused CAP-H) and what the cell diameter was at t = 0 min (<9 or >9 μm). The difference in NEBD time is significant in both cases, with p = 7.36 × 10−8 and p = 1.48 × 10−10, respectively. Error bars represent 95 % confidence intervals (CI), n = 45. Micrographs at the right show representative images for the first time point and the NEBD time point. b Asynchronous CAP-H-GFP-SBP cells were serum starved for 48 h to obtain G1-synchronised cells and then released into medium with or without Cal A treatment. Cell samples were collected at t = 0.5, 1, 1.5 and 2 h, fixed with PFA, immunostained for Lamin B1 (red) and mounted with DAPI to visualise DNA (blue). Endogenous CAP-H-GFP was preserved and images were taken. Representative images are shown. For cells treated with Cal A for 1 and 1.5 h, stacked images were taken, processed for 3D reconstruction and visualised as 3D models (right panel) (refer to “Material and methods” for details). Scale bars represent 5 μm. c Asynchronous CAP-D3-GFP-SBP cells were serum starved for 48 h and released into medium with or without Cal A treatment. Cell samples were collected at t = 0.5, 1 and 2, fixed with PFA, immunostained for Lamin B1 (red) and mounted with DAPI to visualise DNA (blue). Endogenous CAP-D3-GFP was preserved and images were taken. Representative images are shown. For cells treated with Cal A for 1 and 2 h, 3D models (right panel) were constructed as described in Fig. 6b
Figure 6b shows that the nuclear membrane starts to break down at 1.5 h and is completely disassembled after 2 h of Cal A treatment G1-synchronised CAP-H-GFP-SBP cells, with the nuclear membrane visualised by Lamin B1 staining. Once nuclear envelope breakdown has occurred, the majority of the cytoplasmic pool of CAP-H floods into the nuclei. By contrast, the nuclear membrane still remains intact with only 1 h Cal A treatment. Nevertheless, condensed G1-PCC chromatids are seen in these nuclei and are associated with a small amount of CAP-H. 3D reconstruction confirms the integrity of the nuclear membrane in G1-PCC cells with only 1 h of Cal A treatment (Fig. 6b). In similar experiments using CAP-D3-GFP-SBP cells, we found intact nuclear membranes in cells treated with Cal A for 1 h, but the CAP-D3 signal is entirely associated with the G1-PCC chromosomes inside the nuclei in this case (Fig. 6c).
These data indicate that the pool of condensin I in interphase nuclei (Figs. 1 and 2) is responsible for the axial staining in Cal A-induced PCC chromosomes and is distinct from the cytoplasmic pool that only gains access to the chromosomes after NEBD.
PCC chromosomes are structurally compromised and fragile
It is known that the intrinsic structure of the chromosomes is compromised in cells depleted of KIF4 (Samejima et al. 2012), and results presented above (Fig. 5) show that KIF4 is mislocalised in G1-PCC and G2/M-PCC cells. These observations lead to the prediction that PCC chromosomes should not be as structurally sound as mitotic chromosomes.
To test this prediction, both normal mitotic and PCC chromosomes were stressed under different conditions. We found that hypotonically treated PCC cells that were cytospun without prefixation treatment showed fragile G2/M-PCC chromosomes and poor chromosome resolution (bottom panel, methods 3 and 4, Fig. 7a) compared to controls (top panel). Consistent chromosome resolution in the PCC cells was found with methanol/acetic acid (3:1) after hypotonic treatment (methods 5 and 6, Fig. 7a). Only in methods 5 and 6 were G1-PCC chromosomes detected, suggesting they could not structurally withstand the mechanical force when cytospinning was employed (methods 1–4, Fig. 7a).
Fragility of PCC chromosomes and requirement for condensin I or II in PCC. a Asynchronous DT40 cells were treated with 50 nM Cal A for 1 h (bottom panel). Top panel shows mitotic chromosomes in the corresponding conditions. Cells were cytospun (methods 1, 2, 3 and 4) or dropped onto slides (methods 5 and 6) and stained with DAPI. Representative images for the various procedures are shown. Chromosome fragility, condensed chromosome morphology and detection of G1-PCC chromosomes are graded and summarised in the table. b Conditional knockout cells CAP-H KO, CAP-D3 KO and SMC2 KO were cultured in medium with 200 ng/ml dox for 36, 24 and 24 h, to deplete them of condensin I, condensin II and both condensins, respectively. These cells were then treated with Cal A for 1 h, and asynchronous WT cells were treated similarly as controls. Cells were collected, fixed with methanol/acetic acid and mounted with DAPI. Representative G1-PCC, early S-PCC and G2/M-PCC are shown in the centre panel. Representative mitotic chromosomes (from cells not treated with Cal A) are shown as controls in the left panel for WT, CAP-H KO (CAP-H OFF), CAP-D3 KO (CAP-D3 OFF) and SMC2 KO (SMC2 OFF). Representative images of PCC for cells synchronised in G1 are shown in the right panel. CAP-H KO, CAP-D3 KO and SMC2 KO cells were cultured in medium without serum but with 200 ng/ml dox in order to synchronise the cells in G1 and turn off CAP-H, CAP-D3 and SMC2, respectively. Cells were released by adding serum, treated with Cal A for 1 h, collected, fixed and further mounted with DAPI. Scale bar represents 5 μm. c Western blotting of cells used in Fig. 7b verifies that CAP-H, CAP-D3 and SMC2 have been completely turned off
Either condensin I or II is sufficient for PCC
We previously reported contrasting roles for condensin I and II complexes in organising mitotic chromosomes (Green et al. 2012). To further understand the roles of the two condensin complexes in PCC cells, we utilised DT40 conditional KO cells. Knocking out CAP-H removes condensin I (but not condensin II) from chromosomes, knocking out CAP-D3 removes condensin II only and knocking out SMC2 removes both condensins I and II (Hudson et al. 2003, Green et al. 2012, Bakhrebah et al. 2015).
PCC was induced by 1-h Cal A treatment in asynchronous WT, CAP-H KO, CAP-D3 KO and SMC2 KO cells (Fig. 7b). Western blotting confirmed complete depletion of CAP-H, CAP-D3 and SMC2, respectively (Fig. 7c). As expected in control cells without Cal A treatment (Green et al. 2012), depletion of CAP-H showed short and fat chromosomes, depletion of CAP-D3 showed long, thin and wiggly chromosomes, while depletion of SMC2 showed disorganised and loose chromosomes. Condensed chromatids in G1-PCC and hypercondensed chromosomes in G2/M-PCC were still found in CAP-H KO, CAP-D3 KO and SMC2 KO cells. There appears to be little difference in structure between G2/M-PCC and G1-PCC when CAP-H/condensin I or CAP-D3/condensin II is knocked down separately (Fig. 7b).
These data indicate that removal of a specific condensin (I or II) does not hinder formation of PCC chromosomes. However, some sharpness is lost in G2/M-PCC when both condensins I and II are removed (SMC2OFF). In this case, a more blurred or fuzzy chromosome appearance is observed that is similar to, but less extreme than, the mitotic defect in SMC2OFF cells (control). A more pronounced defect is observed for G1-PCC chromosomes in SMC2OFF cells, with the chromosomes dramatically losing resolution compared to G1-PCC in WT, CAP-HOFF or CAP-D3OFF cells.
These results suggest a redundancy of function between condensins I and II in PCC chromosome formation. Very similar results are seen for G1-PCC chromosomes using cells synchronised by serum starvation (Fig. 7b, right hand panel).
Modified nuclear condensin I binds G1-PCC chromosomes
Colocalisation experiments were performed in reciprocal cell lines to further confirm condensin I and II localisation in G1-PCC cells before NEBD (Fig. 8a). Four channels using GFP and endogenous antibodies to colocalise condensin I and II in each of the condensin KO cell lines (i.e. CAP-H-GFP and CAP-D3-GFP cell lines) were used. CAP-H-GFP- or CAP-D3-GFP-expressing cells induced for G1-PCC by Cal A were fixed with PFA to preserve the endogenous GFP and stained for immunofluorescence with Lamin B1 antibodies. CAP-H-GFP-SBP cells were co-stained using CAP-D3 (condensin II) antibodies, and the CAP-D3-GFP-SBP cells were co-stained using CAP-H (condensin I) antibodies, respectively. Similarly, in both transgenic cell lines, a small pool of CAP-H and a large portion of CAP-D3 signal can be detected in the nuclei while the nuclear membrane clearly remains intact (Fig. 8a), which is consistent with Fig. 6b, c. This result further suggests that both condensin II and the small fraction of nuclear pool condensin I are involved in G1-PCC. This raised a pertinent question of whether or not condensin I is modified in the G1-PCC cells.
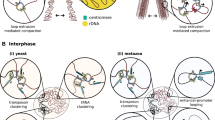
Residual CAP-H in interphase nuclei and modification of CAP-H. a For the upper panels, asynchronous CAP-H-GFP cells were treated with 50 nM Cal for 1 h. Cells were collected, fixed with PFA and further processed with anti-CAP-D3 (red) and Lamin B1 (cyan) antibody co-staining. Cells were stained with DAPI to stain DNA (blue). The endogenous CAP-H-GFP signal was preserved. Representative images are shown. Optical sections were taken and used for 3D reconstruction; these are visualised as 3D models in the right panel (see “Material and methods”). Similarly for the lower panels, asynchronous CAP-D3-GFP cells were treated with 50 nM Cal for 1 h. Cells were collected, fixed with PFA, further processed for anti-CAP-H (red) and Lamin B1 (cyan) antibody co-staining and mounted with DAPI to visualise DNA (blue). Endogenous CAP-D3-GFP signal was preserved. Representative images are shown. 3D models were reconstructed from stacked images and are shown in the right panel. Scale bar represents 5 μm. b Whole cell extracts (left panel) from asynchronous WT cells and from asynchronous, S-phase blocked (TdR) and G1-PCC CAP-H-GFP-SBP (G1-PCC) cells were separated by SDS-PAGE and detected with rabbit anti-CAP-H (upper panel) and mouse anti-α-tubulin antibodies (bottom panel), respectively (note: asterisk represents a non-specific band). Interphase CAP-H-GFP-SBP cells were obtained by blocking with 4 mM Thymidine (TdR) for 14 h, and G1-PCC cells were obtained as described in Fig. 4c. These cells were collected for cytoplasmic (Cyto) and nuclear (NE) extraction as described in the “Materials and methods” and were separated by SDS-PAGE and blotted with rabbit anti-CAP-H (upper right panel) and mouse anti-α-tubulin (lower right panel), respectively. Note a slower migrating single band is detected in nuclear extract in G1-PCC cells. c Model. In the normal cell cycle, shortly after cytokinesis, a small pool of condensin I is still present in the nuclei (G1), which decreases gradually when cells enter into S and G2/M-phases, suggesting a more prominent role in gene transcription. Following NEBD after G2 phase, the cytoplasmic pool of condensin I floods into the nuclei and assists condensin II in mitotic chromosome folding and organisation. Condensin II is retained in the nuclei and in contrast to condensin I gradually increases until S-phase. In G1-PCC, both the nuclear pool of phosphorylated condensin I and condensin II contribute to single condensed chromatid formation. After G1, chromosome shattering occurs in early S-phase (S-PCC) when the condensation machinery seemingly collides with the replication machinery and hyper-chromosome condensation becomes evident in G2/M cells (G2/M-PCC). A dotted line surrounds the G1-PCC nucleus as we cannot exclude the possibility based on our cytological studies that some holes might exist in the nuclear envelope
Next, whole cell extracts were prepared from asynchronous WT cells, asynchronous CAP-H-GFP-SBP cells, thymidine-blocked CAP-H-GFP-SBP cells and G1-PCC CAP-H-GFP-SBP cells. These samples were subjected to gel electrophoresis followed by Western blotting using anti-CAP-H antibodies. Slower migration of the CAP-H band is clearly detected in G1-PCC cells (Fig. 8b, left panel), suggesting that CAP-H may be phosphorylated during the PCC process (Ono et al. 2013).
In a further experiment, cytoplasmic and nuclear/chromosomal proteins were separately extracted from thymidine-blocked cells and from G1-PCC cells and analysed by gel electrophoresis and Western blotting with anti-CAP-H antibodies (Fig. 8b, right panel). As a control, α-tubulin is only detected in the cytoplasmic fraction, and histone H3 is only detected in the nuclear fraction, demonstrating that there is no cross-contamination between nuclear and cytoplasmic samples. It is evident the pool of condensin I in G1-PCC isolated nuclei is comparable to thymidine-arrested cells, suggesting (but not ruling out) the interphase condensin I pool is constant during PCC and that shuttling from the cytoplasm to the nucleus is not likely. Future experiments using fluorescence recovery after photobleaching (FRAP) are therefore required to definitively resolve this point.
Our experiment on isolated nuclei shows in both the thymidine-blocked cells and the G1-PCC cells, a faint CAP-H band is detected in nuclei while the majority of CAP-H is found in the cytoplasm (Fig. 8b, right panel). However, the CAP-H band in the nuclear fraction of the G1-PCC cells is shifted to slightly slower mobility compared to both the cytoplasmic (soluble) fraction in the G1-PCC cells and the nuclear fraction of CAP-H in the thymidine-blocked cells. This suggests that the small amount of CAP-H in the nuclear (chromosome bound) pool may be post-translationally modified in the G1-PCC cells, possibly by phosphorylation.
Importantly, these results show condensin I axial binding to G1-PCC chromosomes comes from the existing pool of condensin I in interphase nuclei before NEBD. The presence of only a single upper band CAP-H band in double thymidine-blocked nuclei further suggests that possible phosphorylation of condensin I in this pool activates condensin I and allows the complex to participate in G1-PCC chromosome formation.
Discussion
Our data sheds a new light on the roles of condensins I and II both in the normal cell cycle and in cells undergoing PCC. In the first part of this study, we have used live cell imaging to analyse condensin behaviour in interphase cells. The results reveal new and important characteristics of the condensin complexes during interphase and also the role of condensins in cells undergoing PCC.
Condensin plays a role in interphase nuclear organisation
The first important observation is that some condensin I is present in interphase nuclei. In metazoans, it has generally been thought that only condensin II is found in the nucleus, but this notion was recently challenged (Li et al. 2015). Although no microscopy analyses were performed, their studies of human cells using ChIP-sequencing provided evidence that condensin I is enriched on interphase chromosomes and in particular at promoters and enhancers (Li et al. 2015). Interestingly, one of the earliest analyses of vertebrate condensin I using microscopy described the complex as largely cytoplasmic in interphase, with some staining seemingly evident in the nuclei (Ono et al. 2004). However, the two most definitive and highly cited studies (Hirota et al. 2004, Gerlich et al. 2006), which used a combination of quantitative live cell imaging and fixed cell analysis, reported that condensin I is excluded from the nucleus during interphase in human cells.
Using microscopy and biochemical analysis, we find that a pool of condensin I does indeed exist in interphase nuclei. One could argue that this pool consists merely of condensin I that was trapped in the nuclei when the nuclear envelope re-formed at the end of the previous mitosis. This is not likely to be the case, however. Our data show that the condensin I molecules in the interphase nucleus are not just bystanders but have possible roles in the condensation of interphase chromosomes and also in gene regulation. The data shows that there is a reduction in the amount of condensin I in the nucleus as cells progress from G1 to S-phase, suggesting that condensin I’s primary role during interphase is in gene regulation. Furthermore, our live cell imaging also shows condensin II dramatically increases on interphase nuclei, suggesting it also plays a role in gene transcription.
Many genome-wide maps of condensin for a variety of species are now available and find condensin is most enriched at promoters of highly expressed Pol II genes, tRNA, and structural regions such as centromeres and directly relevant to this study, rDNA loci (Kranz et al. 2013, Dewari and Bhargava 2014, Nakazawa et al. 2015). While our 3D FISH analyses of interphase nuclei find a correspondingly high loss of condensation in the highly condensin I-enriched rDNA following removal of CAP-H (and thus condensin I), it also found significant decondensation in the chicken-specific heterochromatic Zrep region that is seemingly devoid of condensin I. At a simplistic level, the loss of condensation in the Zrep region after condensin I removal appears counterintuitive; however, condensin I in interphase can mediate long range interactions (Li et al. 2015), and chromosome folding itself involves multiple overlapping processes. Furthermore, the described ChIP-seq maps highlight the most enriched regions, and a more recent paper in Schizosaccharomyces pombe has shown that condensin I is indeed present at lower levels throughout the entire genome (Sutani et al. 2015). Tellingly, a number of recent papers have highlighted condensin II’s role in interphase chromosome condensation (Bauer et al. 2012, George et al. 2014, Wallace et al. 2015), but our data suggest condensin I is also an important player.
Although our experiments used CAP-H depleted cells at a time point where there are no segregation defects or polyploidy observed and are suggestive of a specific interphase defect, it cannot be ruled out that a percentage of interphase cells might have subtle chromosome defects from the previous mitosis. However to date, there is no evidence condensin I plays a role in chromosome decondensation during mitosis. Future experiments that incorporate ultra-rapid depletion strategies such as degron (Nishimura et al. 2009) will be able to answer this definitively.
The presence of condensin I in interphase nuclei appears not to be species-specific, since it occurs in both chicken DT40 cells and human HCT116 cells (this study) as well as in human MCF-7 breast cancer cells (Li et al. 2015). Curiously, there appears to be species differences in yeast, which contain only condensin I. In Schizosaccharomyces pombe, condensin is in the cytoplasm during interphase and jumps onto the DNA at mitosis (Sutani et al. 1999), while in Saccharomyces cerevisiae, condensin is in the nucleus throughout the cell cycle (Freeman et al. 2000). Although our studies indicate that the majority of condensin I exists outside the nucleus, the data clearly point to a nuclear pool of the complex during interphase. It remains to be determined whether this is a universal feature of all multicellular eukaryote systems, but at the very least the earlier observations and conclusions regarding the purely cytoplasmic localisation of condensin I might need to be revisited.
Condensin behaviour and function in PCC
During normal mitosis, chromosomes lacking one of the condensins lose their characteristic shape, with condensin I-depleted chromosomes becoming wider and shorter and condensin II-depleted chromosomes becoming longer and thinner. To determine if prematurely condensed chromosomes are similarly influenced by the absence of individual condensin complexes, we examined Cal A-induced PCC in cells depleted of condensin I (CAP-HOFF) or condensin II (CAP-D3OFF).
We found no evidence that either condensin I or II, individually, is necessary for chromosome condensation in either G1- or G2/M-PCC. Furthermore, there are no obvious differences in morphology between the PCC chromosomes of wild-type and condensin I- or condensin II-depleted cells (Fig. 7b). In fact, when cells are depleted of just one of the condensins, the PCC chromosomes induced with Cal A more closely resemble wild-type (control) metaphase chromosomes than the metaphase chromosomes of cells in which the same condensin complex has been knocked out. This is most strikingly illustrated in G2/M-PCC when condensin II is depleted (CAP-D3OFF). When the condensin II complex is depleted, metaphase chromosomes are longer and thinner than normal, but in G2/M-PCC short, longitudinally condensed chromosomes are evident (Fig. 7b). This suggests that the condensin II defect is totally overridden by the PCC conditions.
When both condensins are disrupted by depleting the cells of SMC2 (SMC2OFF), a clear difference is seen in the case of G1-PCC. The chromosomes appear to be less well defined and presumably less well condensed. Interestingly, histone H1 in mammalian cells is not phosphorylated in G1-PCC (Paulson and Mause 2013), and the same is likely to be true in DT40 cells. Thus, the incomplete chromosome condensation seen in G1-PCC with SMC2OFF may be due to the combined effect of condensin depletion and lack of histone H1 phosphorylation.
Condensins I and II display their distinct axial localisation in G2/M-PCC chromosomes, reminiscent of their mitotic localisation. Yet when either or both condensins are removed the very distinct short, squat morphology of G2/M-PCC chromosomes is retained, suggesting a heightened axial compaction activity in this phase. This activity appears to be independent or can override condensins in G2/M-PCC and likely relates to the mistiming of condensation in this phase. Further experiments are needed to determine what is the force driving the hypercondensation activity observed in G2/M-PCC.
Our study finds both condensins I and II localise axially to chicken DT40 G1-PCC chromosomes, whereas a previous study in human lymphoblast cells finds only condensin I is axial in G1-PCC (Ono et al. 2013). The lymphoblast analysis described the relative ratio of CAP-H2 in G2/G1 chromosomes and shows the G2 signal was twice that of G1, which implies but does not rule out G1-PCC staining for condensin II. Indeed, there is a faint axial signal in this study for condensin II on G1-PCC chromosomes, although there is no doubt our condensin II signal is obvious compared to at best very weak in the lymphoblast study. Why then the difference? In our study, we used the very specific anti-SBP antibody to localise condensin II and the anti-CAP-D3 antibody, both of which showed similar axial localisation. While our study used the standard hypotonic methanol/acetic acid fixation chromosome spread method for immunofluorescence, the previous study (Ono et al. 2013) established a modified fixation/spreading protocol, which used a PFA fix, 0.2 % Triton X-100 permeabilisation followed by cytospinning and refixing and then further permeabilisation with 0.5 % Triton X-100. We would argue that methanol/acetic acid fixation better preserves chromosome morphology and could easily account for observed differences in both studies. Our own attempts at cytospinning even prefixed cells resulted in the loss of the fragile G1 chromosome morphology (Fig. 7a). This fragility might be peculiar to chicken DT40 chromosomes but nonetheless meant we could not replicate these conditions for condensin I G1-PCC localisation in our system. The differences could relate to species used as we found that condensin II in chicken is approximately 1/10 of total condensin in mitotic chromosomes using SILAC quantitative proteomics (Ohta et al. 2010), whereas condensins I and II are in roughly equal proportions in nuclear extracts from human (Ono et al. 2003). However, we are inclined to believe the difference in the strong signal we observe for condensin II in G1-PCC chromatids reflects likely differences in antibody and fixation techniques rather than species differences.
Our results indicate that condensins are not the sole drivers of chromosome condensation in PCC or in normal mitotic chromosome condensation. However, this should by no means be taken to indicate that they are not involved. It is possible that there are multiple redundant mechanisms of chromosome condensation and that one or more of these mechanisms are dispensable as long as the others remain intact. Elimination of two or more, however, may begin to show an effect. For example, condensins I and II may work in tandem with histone phosphorylation so that, as mentioned above, eliminating both condensins and histone H1 phosphorylation in G1-PCC leads to defective chromosome condensation (Paulson and Mause 2013).
It is very likely that protein phosphorylation plays a role in chromosome condensation, since PCC is induced by Cal A, a potent inhibitor of protein phosphatases 1 and 2A (PP1 and PP2A). PP1 is responsible for the dephosphorylation of several major phosphoproteins at the end of mitosis, including histones H1 and H3, nuclear lamins and the retinoblastoma protein pRb (Paulson 2007, Paulson et al. 1996). We have previously proposed that PP1 is phosphorylated and downregulated at the onset of mitosis by Cdk1/cyclin B or other protein kinases and that inactivation of protein phosphatases including PP1 may be just as important a feature of the onset of mitosis as activation of protein kinases (Paulson 2007). This would explain why Cal A and other cell-permeable inhibitors of PP1 are able to induce PCC even in the G1-phase, when cyclin B has not yet been synthesised (Paulson and Mause 2013).
Conclusion
Our data explains a paradoxical finding from our genome-wide map of condensin I in mitotic cells in chicken DT40 cells (Kim et al. 2013). This study found that the majority of condensin I bound to the promoters of highly expressed genes during mitosis. Why would a complex bind a gene regulatory region at a time when transcription is all but inactive? Our finding that some condensin I is retained after mitosis and into the following cell cycle provides a logical explanation for this anomaly and suggests retained condensin I in G1 acts to fine tune transcription. Furthermore, our data show the interphase nuclear pool of condensin I is further reduced as the cell progresses from the G1 to S-phase, which is consistent with a role in gene regulation. 3D FISH analyses show removing condensin I affects euchromatic loci containing the highly expressed 18s rDNA genes but surprisingly finds the silent heterochromatic Zrep region is also affected. Our study in chicken cells and the recent findings using human cells (Li et al. 2015) point to a new player in gene regulation.
Previous studies found condensin II, but not condensin I removal, affected late S-phase chromatids (Ono et al. 2013), while our study shows removing condensins I and II together affects G1-PCC chromatid formation but not G2/M-PCC. These data suggest condensin activity in PCC varies or is altered between the different stages of the cell cycle. Why this is so remains unclear and requires further experiments. Our results suggest there is significant redundancy in the condensation machinery. The idea of multiple mechanisms in chromosome condensation is consistent with the notion of chromosome condensation consisting of two overlapping processes: (1) compaction and (2) assembly of higher order structure. Histone modifications could contribute more to the former while the condensins contribute more to the latter, with overlap in both directions. Certainly other factors involved. Pertinently, an elegant cell-free system was recently used to define the minimum set of purified proteins required to reconstitute a mitotic chromosome in vitro. Only six factors were necessary: the core histones, nucleoplasmin, Nap1, FACT, topoisomerase II and condensin I (Shintomi et al. 2015). Remarkably, aside from condensin I, all other proteins were unmodified suggesting the interplay between histone modification and non-histone proteins to build a mitotic chromosome is still far from being understood. Nonetheless, enormous progress has been made.
Abbreviations
- 3D FISH:
-
Three-dimensional fluorescent in situ hybridization
- Cal A:
-
Calyculin A
- CENP-O:
-
Centromere protein O
- Dox:
-
Doxycycline
- DE:
-
Differential expression
- EdU:
-
5-Ethynyl-2'-deoxyuridine
- IF:
-
Immunofluorescence
- NEBD:
-
Nuclear envelope breakdown
- PCC:
-
Premature chromosome condensation
- PFA:
-
Paraformaldehyde
- SBP:
-
Streptavidin-binding peptide
- Topo IIα:
-
Topoisomerase IIα
References
Bakhrebah M, Zhang T, Mann JR, Kalitsis P, Hudson DF (2015) Disruption of a conserved CAP-D3 threonine alters condensin loading on mitotic chromosomes leading to chromosome hypercondensation. J Biol Chem 290:6156–67
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A, Lee H, Zhang N, Robertson CL, Serova N, Davis S, Soboleva A (2013) NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res 41:D991–5
Bauer CR, Hartl TA, Bosco G (2012) Condensin II promotes the formation of chromosome territories by inducing axial compaction of polyploid interphase chromosomes. PLoS Genet 8, e1002873
Bazile F, St-Pierre J, D'Amours D (2010) Three-step model for condensin activation during mitotic chromosome condensation. Cell Cycle 9:3243–55
Belmont AS (2006) Mitotic chromosome structure and condensation. Curr Opin Cell Biol 18:632–8
Berkopec A (2007) HyperQuick algorithm for discrete hypergeometric distribution. J Discret Algorithm 5:341–347
Buerstedde JM, Takeda S (1991) Increased ratio of targeted to random integration after transfection of chicken B cell lines. Cell 67:179–88
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, Nezi L, Protopopov A, Chowdhury D, Pellman D (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482:53–8
Cremer M, Grasser F, Lanctot C, Muller S, Neusser M, Zinner R, Solovei I, Cremer T (2008) Multicolor 3D fluorescence in situ hybridization for imaging interphase chromosomes. Methods Mol Biol 463:205–39
Cremer M, Weierich C, Solovei I (2005) Multicolour 3D-FISH in vertebrate cells (PROT23) [Online]. http://www.epigenesys.eu/en/protocols/fluorescence-microscopy/181-multicolour-3d-fish-in-vertebrate-cells
Dewari PS, Bhargava P (2014) Genome-wide mapping of yeast histone chaperone anti-silencing function 1 reveals its role in condensin binding with chromatin. PLoS ONE 9, e108652
Earnshaw WC, Ratrie H 3rd, Stetten G (1989) Visualization of centromere proteins CENP-B and CENP-C on a stable dicentric chromosome in cytological spreads. Chromosoma 98:1–12
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–10
EL Achkar E, Gerbault-Seureau M, Muleris M, Dutrillaux B, Debatisse M (2005) Premature condensation induces breaks at the interface of early and late replicating chromosome bands bearing common fragile sites. Proc Natl Acad Sci U S A 102:18069–74
Fagerholm AE, Habrant D, Koskinen AM (2010) Calyculins and related marine natural products as serine-threonine protein phosphatase PP1 and PP2A inhibitors and total syntheses of calyculin A, B, and C. Mar Drugs 8:122–72
Franklin R, Sale JE (2006) 2D cell cycle analysis. Subcell Biochem 40:405–8
Freeman L, Aragon-Alcaide L, Strunnikov A (2000) The condensin complex governs chromosome condensation and mitotic transmission of rDNA. J Cell Biol 149:811–24
Frosi Y, Haering CH (2015) Control of chromosome interactions by condensin complexes. Curr Opin Cell Biol 34:94–100
Gallego-Paez LM, Tanaka H, Bando M, Takahashi M, Nozaki N, Nakato R, Shirahige K, Hirota T (2014) Smc5/6-mediated regulation of replication progression contributes to chromosome assembly during mitosis in human cells. Mol Biol Cell 25:302–17
George CM, Bozler J, Nguyen HQ, Bosco G (2014) Condensins are required for maintenance of nuclear architecture. Cells 3:865–82
Gerlich D, Hirota T, Koch B, Peters JM, Ellenberg J (2006) Condensin I stabilizes chromosomes mechanically through a dynamic interaction in live cells. Curr Biol 16:333–44
Gotoh E (2009) Drug-induced premature chromosome condensation (PCC) protocols: cytogenetic approaches in mitotic chromosome and interphase chromatin. Methods Mol Biol 523:83–92
Gotoh E, Durante M (2006) Chromosome condensation outside of mitosis: mechanisms and new tools. J Cell Physiol 209:297–304
Green LC, Kalitsis P, Chang TM, Cipetic M, Kim JH, Marshall O, Turnbull L, Whitchurch CB, Vagnarelli P, Samejima K, Earnshaw WC, Choo KH, Hudson DF (2012) Contrasting roles of condensin I and condensin II in mitotic chromosome formation. J Cell Sci 125:1591–604
Hirano T (2014) Condensins and the evolution of torsion-mediated genome organization. Trends Cell Biol 24:727–33
Hirano T, Kobayashi R, Hirano M (1997) Condensins, chromosome condensation protein complexes containing XCAP-C, XCAP-E and a Xenopus homolog of the Drosophila Barren protein. Cell 89:511–21
Hirota T, Gerlich D, Koch B, Ellenberg J, Peters JM (2004) Distinct functions of condensin I and II in mitotic chromosome assembly. J Cell Sci 117:6435–45
Hittelman WN, Menegaz SD, Mccredie KB, Keating MJ (1984) Premature chromosome condensation studies in human leukemia: 5. Prediction of early relapse. Blood 64:1067–73
Hori T, Suzuki Y, Solovei I, Saitoh Y, Hutchison N, Ikeda JE, Macgregor H, Mizuno S (1996) Characterization of DNA sequences constituting the terminal heterochromatin of the chicken Z chromosome. Chromosom Res 4:411–26
Huang K, Jia J, Wu C, Yao M, Li M, Jin J, Jiang C, Cai Y, Pei D, Pan G, Yao H (2013) Ribosomal RNA gene transcription mediated by the master genome regulator protein CCCTC-binding factor (CTCF) is negatively regulated by the condensin complex. J Biol Chem 288:26067–77
Hudson DF, Marshall KM, Earnshaw WC (2009) Condensin: architect of mitotic chromosomes. Chromosom Res 17:131–44
Hudson DF, Ohta S, Freisinger T, Macisaac F, Sennels L, Alves F, Lai F, Kerr A, Rappsilber J, Earnshaw WC, Spread PSSM (2008) Molecular and genetic analysis of condensin function in vertebrate cells. Mol Biol Cell 19:3070–9
Hudson DF, Vagnarelli P, Gassmann R, Earnshaw WC (2003) Condensin is required for nonhistone protein assembly and structural integrity of vertebrate mitotic chromosomes. Dev Cell 5:323–36
Ishihara H, Martin BL, Brautigan DL, Karaki H, Ozaki H, Kato Y, Fusetani N, Watabe S, Hashimoto K, Uemura D et al (1989) Calyculin A and okadaic acid: inhibitors of protein phosphatase activity. Biochem Biophys Res Commun 159:871–7
Johnson RT, Rao PN (1970) Mammalian cell fusion: induction of premature chromosome condensation in interphase nuclei. Nature 226:717–22
Kato Y, Fusetani N, Matsunaga S, Hashimoto K (1988) Calyculins, potent antitumour metabolites from the marine sponge Discodermia calyx: biological activities. Drugs Exp Clin Res 14:723–8
Kim JH, Zhang T, Wong NC, Davidson N, Maksimovic J, Oshlack A, Earnshaw WC, Kalitsis P, Hudson DF (2013) Condensin I associates with structural and gene regulatory regions in vertebrate chromosomes. Nat Commun 4:2537
Kranz AL, Jiao CY, Winterkorn LH, Albritton SE, Kramer M, Ercan S (2013) Genome-wide analysis of condensin binding in Caenorhabditis elegans. Genome Biol 14:R112
Kwon MS, Hori T, Okada M, Fukagawa T (2007) CENP-C is involved in chromosome segregation, mitotic checkpoint function, and kinetochore assembly. Mol Biol Cell 18:2155–68
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–9
Lau AC, Nabeshima K, Csankovszki G (2014) The C. elegans dosage compensation complex mediates interphase X chromosome compaction. Epigenetics Chromatin 7:31
Li W, Hu Y, Oh S, Ma Q, Merkurjev D, Song X, Zhou X, Liu Z, Tanasa B, He X, Chen AY, Ohgi K, Zhang J, Liu W, Rosenfeld MG (2015) Condensin I and II complexes license full estrogen receptor alpha-dependent enhancer activation. Mol Cell 59:188–202
Liang Z, Zickler D, Prentiss M, Chang FS, Witz G, Maeshima K, Kleckner N (2015) Chromosomes progress to metaphase in multiple discrete steps via global compaction/expansion cycles. Cell 161:1124–37
Maeshima K, Imai R, Tamura S, Nozaki T (2014) Chromatin as dynamic 10-nm fibers. Chromosoma 123:225–37
Maeshima K, Laemmli UK (2003) A two-step scaffolding model for mitotic chromosome assembly. Dev Cell 4:467–80
Mayburd AL, Martlinez A, Sackett D, Liu H, Shih J, Tauler J, Avis I, Mulshine JL (2006) Ingenuity network-assisted transcription profiling: identification of a new pharmacologic mechanism for MK886. Clin Cancer Res 12:1820–7
Moser SC, Swedlow JR (2011) How to be a mitotic chromosome. Chromosom Res 19:307–19
Nakazawa N, Sajiki K, Xu X, Villar-Briones A, Arakawa O, Yanagida M (2015) RNA pol II transcript abundance controls condensin accumulation at mitotically up-regulated and heat-shock-inducible genes in fission yeast. Genes Cells 20:481–99
Nishimura K, Fukagawa T, Takisawa H, Kakimoto T, Kanemaki M (2009) An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat Methods 6:917–22
Ohta S, Bukowski-Wills JC, Sanchez-Pulido L, Alves Fde L, Wood L, Chen ZA, Platani M, Fischer L, Hudson DF, Ponting CP, Fukagawa T, Earnshaw WC, Rappsilber J (2010) The protein composition of mitotic chromosomes determined using multiclassifier combinatorial proteomics. Cell 142:810–21
Ono T, Fang Y, Spector DL, Hirano T (2004) Spatial and temporal regulation of condensins I and II in mitotic chromosome assembly in human cells. Mol Biol Cell 15:3296–308
Ono T, Losada A, Hirano M, Myers MP, Neuwald AF, Hirano T (2003) Differential contributions of condensin I and condensin II to mitotic chromosome architecture in vertebrate cells. Cell 115:109–21
Ono T, Yamashita D, Hirano T (2013) Condensin II initiates sister chromatid resolution during S phase. J Cell Biol 200:429–41
Paulson JR (1989) Scaffold morphology in histone-depleted HeLa metaphase chromosomes. Chromosoma 97:289–95
Paulson JR (2007) Inactivation of Cdk1/Cyclin B in metaphase-arrested mouse FT210 cells induces exit from mitosis without chromosome segregation or cytokinesis and allows passage through another cell cycle. Chromosoma 116:215–25
Paulson JR, Laemmli UK (1977) The structure of histone-depleted metaphase chromosomes. Cell 12:817–28
Paulson JR, Mause E (2013) Calyculin A induces prematurely condensed chromosomes without histone H1 phosphorylation in mammalian G1-phase cells. Adv Biol Chem 3:8
Paulson JR, Patzlaff JS, Vallis AJ (1996) Evidence that the endogenous histone H1 phosphatase in HeLa mitotic chromosomes is protein phosphatase 1, not protein phosphatase 2A. J Cell Sci 109(Pt 6):1437–47
Pospisil P, Iyer LK, Adelstein SJ, Kassis AI (2006) A combined approach to data mining of textual and structured data to identify cancer-related targets. BMC Bioinforma 7:354
Pruitt KD, Tatusova T, Brown GR, Maglott DR (2012) NCBI Reference Sequences (RefSeq): current status, new features and genome annotation policy. Nucleic Acids Res 40:D130–5
Ravi M, Nivedita K, Pai GM (2012) Chromatin condensation dynamics and implications of induced premature chromosome condensation. Biochimie
Robinson MD, Mccarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–40
Saitoh N, Goldberg IG, Wood ER, Earnshaw WC (1994) ScII: an abundant chromosome scaffold protein is a member of a family of putative ATPases with an unusual predicted tertiary structure. J Cell Biol 127:303–18
Samejima K, Ogawa H, Cooke CA, Hudson DF, Macisaac F, Ribeiro SA, Vagnarelli P, Cardinale S, Kerr A, Lai F, Ruchaud S, Yue Z, Earnshaw WC (2008) A promoter-hijack strategy for conditional shutdown of multiply spliced essential cell cycle genes. Proc Natl Acad Sci U S A 105:2457–62
Samejima K, Samejima I, Vagnarelli P, Ogawa H, Vargiu G, Kelly DA, De Lima Alves F, Kerr A, Green LC, Hudson DF, Ohta S, Cooke CA, Farr CJ, Rappsilber J, Earnshaw WC (2012) Mitotic chromosomes are compacted laterally by KIF4 and condensin and axially by topoisomerase II alpha. J Cell Biol 199:755–70
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–82
Shintomi K, Takahashi TS, Hirano T (2015) Reconstitution of mitotic chromatids with a minimum set of purified factors. Nat Cell Biol 17:1014–23
Smith HF, Roberts MA, Nguyen HQ, Peterson M, Hartl TA, Wang XJ, Klebba JE, Rogers GC, Bosco G (2013) Maintenance of interphase chromosome compaction and homolog pairing in Drosophila is regulated by the condensin cap-h2 and its partner Mrg15. Genetics 195:127–46
Stevens JB, Abdallah BY, Regan SM, Liu G, Bremer SW, Ye CJ, Heng HH (2010) Comparison of mitotic cell death by chromosome fragmentation to premature chromosome condensation. Mol Cytogenet 3:20
Sutani T, Sakata T, Nakato R, Masuda K, Ishibashi M, Yamashita D, Suzuki Y, Hirano T, Bando M, Shirahige K (2015) Condensin targets and reduces unwound DNA structures associated with transcription in mitotic chromosome condensation. Nat Commun 6:7815
Sutani T, Yuasa T, Tomonaga T, Dohmae N, Takio K, Yanagida M (1999) Fission yeast condensin complex: essential roles of non-SMC subunits for condensation and Cdc2 phosphorylation of Cut3/SMC4. Genes Dev 13:2271–83
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork P, Jensen LJ, VON Mering C (2011) The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39:D561–8
Vagnarelli P (2012) Mitotic chromosome condensation in vertebrates. Exp Cell Res 318:1435–41
Vagnarelli P, Hudson DF, Ribeiro SA, Trinkle-Mulcahy L, Spence JM, Lai F, Farr CJ, Lamond AI, Earnshaw WC (2006) Condensin and Repo-Man-PP1 co-operate in the regulation of chromosome architecture during mitosis. Nat Cell Biol 8:1133–42
Wallace HA, Klebba JE, Kusch T, Rogers GC, Bosco G (2015) Condensin II regulates interphase chromatin organization through the Mrg-binding motif of Cap-H2. G3 (Bethesda) 5:803–17
Wang BD, Eyre D, Basrai M, Lichten M, Strunnikov A (2005) Condensin binding at distinct and specific chromosomal sites in the Saccharomyces cerevisiae genome. Mol Cell Biol 25:7216–25
Wang YD, Caldwell RB (2006) Isolation of nuclear and cytoplasmic proteins from DT40 cells. Subcell Biochem 40:439–40
Wilkins BJ, Rall NA, Ostwal Y, Kruitwagen T, Hiragami-Hamada K, Winkler M, Barral Y, Fischle W, Neumann H (2014) A cascade of histone modifications induces chromatin condensation in mitosis. Science 343:77–80
Williams DM, Scott CD, Beck TM (1976) Premature chromosome condensation in human leukemia. Blood 47:687–93
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14
Zhang CZ, Leibowitz ML, Pellman D (2013) Chromothripsis and beyond: rapid genome evolution from complex chromosomal rearrangements. Genes Dev 27:2513–30
Acknowledgments
The authors thank Dr. Matthew Burton for providing flow cytometry and microscopy technical support and Dr. Kathryn Marshall for careful reading of the manuscript.
This work was supported by NHMRC project grant GNT1030358 and GNT1047009 and by the Victorian Government’s Operational Infrastructure Support Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Additional information
Responsible Editor: Tatsuo Fukagawa, Ph.D
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
A small pool of condensin I is retained in the interphase nuclei of chicken DT40 cells. a The slide in Fig. 2a was examined by confocal microscopy (Zeiss LSM780). Representative images are shown, demonstrating that some CAP-H-GFP-SBP is present in the interphase nuclei. Line profiles across the cell were plotted and are shown at the right. Scale bar represents 2 μm. b Asynchronous CAP-H-GFP-SBP cells were fixed with 4 % PFA and co-stained with mouse anti-SBP and rabbit anti-CAP-H antibodies. DNA was stained with DAPI and the cells were examined by confocal microscopy (Zeiss LSM 780). Line profiles across the cell were plotted and are shown at the right. Scale bar represents 10 μm. Inserts are higher magnification of selected cell. c Asynchronous CAP-H-GFP cells were fixed with PFA and stained with Lamin B1 antibodies. Optical sections (0.2 μm) were imaged and analysed by Imaris 8.1.2. Orthogonal (XY, XZ and YZ) views of CAP-H-GFP and Lamin B1 show clearly that CAP-H-GFP is present inside the nuclear envelope. Line profiles across the cell were plotted and are shown at the top right. Scale bar represents 5 μm. d Ponceau staining of the blot in Fig. 2d is shown as a loading control. (PDF 432 kb)
Online Resource 2
Determination of cell cycle stages in live cell imaging. a CAP-H-GFP H2B-RFP and CAP-D3-GFP H2B-RFP cells (n = 7) were subjected to live cell imaging for 15 h with 2-min intervals. Cell size during the timecourse was determined by measuring the cell diameter (μm) using the polygon function in SoftWoRx 4.1 and plotted as a function of time. CAP-H-GFP and CAP-D3-GFP cellular localisations were monitored and this is reflected in the background colour of the graph. Cell cycle stages were determined accordingly. b, c CAP-H-GFP H2B-RFP cells were subjected to live cell imaging for 15 h with 2-min intervals. Normalised CAP-H-GFP intensity was plotted with time according to the description in Fig. 1a, T = 0 min was set as a completion of anaphase. Note the level of CAP-H in the nucleus drops over time. Representative images in different cell cycle stages are shown in b. H2B-RFP intensity during the timecourses were normalised to anaphase single cell H2B-RFP intensity at T = 0 min, respectively. Normalised H2B-RFP intensity was plotted with time in c. Cell cycle stages were determined based on the change of normalised H2B-RFP intensity (i.e. double H2B) and DIC morphology. Error bars represent standard error of the mean (SEM), n = 15. (PDF 589 kb)
Online Resource 3
Complete depletion of CAP-H. a CAP-H KO cells were grown in the absence (CAP-HON) or presence (CAP-HOFF) of dox for 36 h in order to deplete CAP-H. Cells were collected for Western blotting to detect CAP-H and α-tubulin, and FACS analysis. b CAP-HON and CAP-HOFF (+ dox 36 h) cells were collected for Annexin V staining and subjected for FACS analysis. c CAP-HON and CAP-HOFF (+ dox 36 h) cells were collected and fixed with 4 % PFA. Immunofluorescent staining was used to visualise γ-H2AX (red). DNA was stained with DAPI (blue). CAP-HON cells treated with Adriamycin 6 h were used as a positive control for γ-H2AX staining. Representative images are shown. Three different experiments were performed. γ-H2AX foci were counted and scored as 0, 1, 2, 3, 4 and ≥5 foci from 100 cells in each experiment. The average percentage of cells with 0, 1, 2, 3, 4 and ≥5 γ-H2AX foci is shown in the right. (PDF 682 kb)
Online Resource 4
Functional studies of CAP-H in interphase. a Human homologs of 693 DT40 genes that are significantly misregulated in CAP-H KO were analysed with Ingenuity Pathway Analysis (IPA). The top five canonical pathways are shown at the left. The top five diseases and biological functions are shown at the right. b Significant DE genes in this study that are involved in the prostate cancer signalling pathway are shown in red. c 116 significantly DE genes with CAP-H enrichment were analysed with DAVID Bioinformatics Resources 6.7. The top nine GO terms are shown. (PDF 380 kb)
Online Resource 5
Heat map of 116 misregulated genes directly due to CAP-H binding removal. Heat map representing 116 DE genes with CAP-H enrichment in comparison between CAP-HON and CAP-HOFF samples displays significant gene expression change. Each gene is represented by a row of coloured boxes, and each replicate is represented by a column. Red indicates gene expression up-regulation, blue indicates gene expression down-regulation. (PDF 178 kb)
Online Resource 6
Condensins I and II in the G1-PCC cells. a G1 synchronised CAP-D3-GFP-SBP cells were treated with 50 nM Cal A for 1 h. Cells were collected, fixed with methanol/acetic acid (3:1) and co-stained with mouse anti-SBP and rabbit anti-CAP-H antibodies to detect CAP-D3-GFP-SBP (green) and CAP-H (red). DNA was stained with DAPI (blue). Representative G1-PCC chromosomes are shown. Scale bar represents 5 μm. b CAP-D3-GFP H2B-RFP cells were subjected to live cell imaging. NEBD times (min) were plotted according to cell sizes (cell diameter <9 μm or >9 μm) at t = 0 min with p = 6.72E-13 as described in Fig. 6a. Error bars represent 95 % confidence intervals (CI), n = 30. (PDF 149 kb)
Online Resource 7
Chromosome shattering in the PCC cells. a WT H2B-GFP cells with and without Cal A treatment were subjected to live cell imaging for 7 h at 2-min intervals. Live cell imaging was started within 7 min of Cal A addition. At t = 0 min, G1 cells (cell diameters <9 μm, based on Online Resource 2a), G2/M cells, S cells (cell diameters >9 μm, based on Online Resource 2a) and anaphase cells were selected for monitoring. b Live cell imaging using H2B-GFP of G1-PCC, S-PCC and anaphase-PCC cells. Note the shattering in the final frame for each cell. c, d CAP-H-GFP-SBP cells were treated as described in Fig. 4c, collected and fixed with PFA. Immunofluorescent staining was used to visualise (c) γ-H2AX (red) or (d) RAD51 (red). DNA was stained with DAPI (blue). CAP-H-GFP-SBP cells blocked with hydroxyurea (HU) for 16 h and then released were used as a positive control for RAD51 staining. Representative images are shown. (PDF 1332 kb)
Rights and permissions
About this article
Cite this article
Zhang, T., Paulson, J.R., Bakhrebah, M. et al. Condensin I and II behaviour in interphase nuclei and cells undergoing premature chromosome condensation. Chromosome Res 24, 243–269 (2016). https://doi.org/10.1007/s10577-016-9519-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-016-9519-7