Abstract
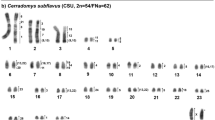
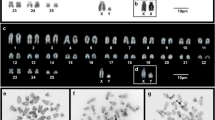
We established chromosome homology maps between Mus musculus (MMU) and five species of the Akodontini tribe, Akodon cursor (2n = 14, 15 and 16), A. montensis (2n = 24), A. paranaensis (2n = 44), A. serrensis (2n = 46) and Oligoryzomys flavescens (2n = 66) by Zoo-FISH (fluorescence in situ hybridization) using mouse chromosome-specific probes. The aims of this study were (1) to detect the chromosomal rearrangements responsible for the karyotype variation in this tribe and (2) to reconstruct the phylogenetic relationships among these species. We observed four common syntenic associations of homologous chromosome segments, of which the MMU 7/19 has been described previously in other rodents from Africa, Asia and Europe, and might represent a phylogenetic link between the Old World and Neotropical rodents. The remaining three associations (3/18, 6/12 and 8/13) have been observed exclusively in Neotropical rodents so far, which at present can be considered synapomorphic traits of this group. Five further mouse chromosomes (MMU 4, 9, 14, 18 and 19) were each found evolutionarily conserved as a separate syntenic unit. Our phylogenetic analysis using parsimony and heuristic search detected one consistent group, separating the Akodontini from other rodents.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bianchi NO, Reig OA, Molina OJ et al. (1971) Cytogenetics of the South American Akodont rodents (Cricetidae). I. A progress report of Argentinian and Venezuelan forms. Evolution 25: 724–736.
Cavagna P, Stone G, Stanyon R (2002) Black rat (Rattus rattus) genomic variability characterized by chromosome painting. Mamm Genome 13: 157–163.
Christoff AU, Fagundes V, Sbalqueiro IJ, Mattevi MS, Yonenaga-Yassuda Y (2000) Description of a new species of Akodon (Rodentia, Sigmodontinae) from Southern Brazil. J Mammal 81: 838–851.
de Oliveira EHC, Neusser M, Figueiredo WB et al. (2002) The phylogeny of howler monkeys (Alouatta, Platyrrhini): Reconstruction by multicolor cross-species chromosome painting. Chromosome Res 10: 669–683.
Engelbrecht A, Dobigny T, Robinson TJ (2006) Further insights into the ancestral murine karyotype: the construction of the Otomys-Mus comparison using chromosome painting. Cytogenet Genome Res 112: 126–130.
Espinosa MB, Reig OA (1991) Cytogenetics and karyosystematics of South American Oryzomyine rodents (Cricetidae, Sigmodontinae) III. Banding karyotypes of Argentinian Oligoryzomys. Z. Säugetierk 56: 306–317.
Fagundes V, Scalzi-Martin JM, Sims K et al. (1997) ZOO-FISH of a microdissection DNA library and G-banding patterns reveal the homeology between the Brazilian rodents Akodon cursor and A. montensis. Cytogenet Cell Genet 78: 224–228.
Ford CE, Hamerton JL (1956) A colchicine hypotonic citrate squash sequence for mammalian chromosomes. Stain Technol 31: 247–251.
Geise L, Canavez FC, Seuánez HN (1998) Comparative karyology in Akodon (Rodentia, Sigmodontinae) from Southeastern Brazil. Heredity 89: 158–163.
Geise L, Smith MF, Patton L (2001) Diversification in the genus Akodon (Rodentia, Sigmodontinae) in Southeastern South America: mitochondrial DNA sequence analysis. J Mammal 82: 92–101.
Guilly MN, Fouchet P, de Chamisso P, Schmitz A, Dutrillaux B (1999) Comparative karyotype of rat and mouse using bidirectional chromosome painting. Chromosome Res 7: 213–221.
Li T, O’Brien PCM, Biltueva L et al. (2004) Evolution of genome organizations of squirrels (Sciuridae) revealed by cross-specific chromosome painting. Chromosome Res 12: 317–335.
Liascovich R, Reig OA (1989) Low chromosomal number in Akodon cursor montensis Thomas, and karyologic confirmation of Akodon serrensis Thomas in Missiones, Argentina. J Mammal 70: 391–395.
Marshall LG (1979) A model of paleobiogeography of South American cricetine rodents. Paleobiology 5: 126–132.
Matsubara K, Nishida-Umehara C, Kuroiwa A, Tsuchiya K, Matsuda Y (2003) Identification of chromosome rearrangements between the laboratory mouse (Mus musculus) and the Indian spiny mouse (Mus platythrix) by comparative FISH analysis. Chromosome Res 11: 57–64.
Matsubara K, Nishida-Umehara C, Tsuchiya K, Nukaya D, Matsuda Y (2004) Karyotypic evolution of Apodemus (Muridae, Rodentia) inferred from comparative FISH analyses. Chromosome Res 12: 383–395.
Montes MA, Oliveira LFB, Langguth A, Mattevi MS (2008) Phylogeny and temporal diversification of the Neotropical rodent genus Akodon: Inferences from mitochondrial and nuclear DNA sequences data. Mol Phylogenet Evol (in press).
Musser GG, Carleton MD (1993 and 2005) Family Muridae. In: Wilson DE, Reeder DM, eds. Mammal Species of the World: a taxonomic and geographic reference, 2nd and 3rd edns. Washngton DC: Smithsonian Institution, pp. 501–753.
Nakamura T, Kuroiwa A, Nishida-Umehara C, Matsubara K, Yamada F, Matsuda Y (2007) Comparative chromosome painting map between two Ryukyu spiny rat species, Tokudaia osimensis and Tokudaia tokunoshimensis (Muridae, Rodentia). Chromosome Res 15: 799–806.
Rambau RV, Robinson TJ (2003) Chromosome painting in the African four-striped mouse Rhabdomys pumilio: Detection of possible murid specific contiguous segment combinations. Chromosome Res 11: 91–98.
Reig OA (1981) Teoria del origen y desarrollo de la fauna de mamíferos de America del Sur. Mus Muni. Cienc Nat ‘Lorenzo Scaglia’ 1: 1–161.
Reig AO (1984) Distribuição geográfica e história evolutiva dos roedores muroideos sul americanos (Cricetidae: Sigmodontinae). Braz J Genet 7: 333–65.
Romanenko SA, Perelman PL, Serdukova NA et al. (2006) Reciprocal chromosome painting between three laboratory rodent species. Mamm Genome 17: 1183–1192.
Romanenko SA, Volobouev VT, Perelman PL et al. (2007a) Karyotype evolution and phylogenetic relationships of hamsters (Cricetidae, Muroidea, Rodentia) inferred from chromosomal painting and banding comparison. Chromosome Res 15(3): 283–297.
Romanenko SA, Sitnikova NA, Serdukova NA et al. (2007b) Chromosomal evolution of Arvicolinae (Cricetidae, Rodentia). II. The genome homology of two mole voles (genus Ellobius), the field vole and golden hamster revealed by comparative chromosome painting. Chromosome Res doi:10.1007/s10577-007-1171-9.
Sbalqueiro IJ, Nascimento AP (1996) Occurrence of Akodon cursor (Rodentia, Cricetidae) with 14, 15 and 16 chromosome cytotypes in the same geographic area in Southern Brazil. Braz J Genet 19: 565–569.
Sbalqueiro IJ, Kiku M, Lacerda M, El Achkar D, Arnt LR (1986) Estudos cromossômicos em roedores da família Cricetidae coletados no Paraná. Cienc Cult 38: 926.
Seabright M (1971) A rapid banding technique for human chromosomes. Lancet 2: 971–972.
Silva MJ, Patton JL, Yonenaga-Yassuda Y (2006) Phylogenetic relationships and karyotype evolution in the sigmodontinae rodent Akodon (2n = 10 and 2n = 16) from Brazil. Genet Mol Biol 29: 469–474.
Sitnikova N, Romanenko SA, O’Brien PCM et al. (2007) Chromosomal evolution of Arvicolinae (Cricetidae, Rodentia). I. The genome homology of tundra vole, field vole, mouse and golden hamster revealed by comparative chromosome painting. Chromosome Res doi:10.1007/s10577-007-1137-y.
Smith MF, Patton JL (1991) Variation in mitochondrial cytochrome b sequence in natural populations of South American Akodontine Rodents (Muridae, Sigmidontinae). Mol Biol Evol 8: 85–103.
Smith MF, Patton JL (1993) The diversification of South-American murid rodents: evidence from mitochondrial-DNA sequence data for the Akodontine tribe. Biol J Linn Soc 50: 149–177.
Stanyon R, Yang F, Cavagna P et al. (1999) Reciprocal chromosome painting shows that genomic rearrangements between rat and mouse proceeds ten times faster then between humans and cats. Cytogenet Cell Genet 84: 150–155.
Stanyon R, Stone G, Garcia M, Froenicke L (2003) Reciprocal chromosome painting shows that squirrels, unlike murid rodents, have a highly conserved genome organization. Genomics 82: 245–249.
Stanyon R, Yang F, Morescalchi AM, Galleni L (2004) Chromosome painting in the long-tailed field mouse provides insights into the ancestral murid karyotype. Cytogenet Genome Res 105: 406–411.
Steppan SJ (1995) Revision of the Tribe Phyllotini (Rodentia: Sigmodontinae), with a phylogenetic hypothesis for the Sigmodontinae. Fieldiana Zool 80: 1–112.
Steppan SJ, Adkins RM, Anderson J (2004) Phylogeny and divergence-data estimates of rapid radiation in muroid rodents based on multiple nuclear genes. Syst Biol 53: 533–553.
Swofford DL (2001) PAUP*: Phylogenetic Analysis Using Parsimony (*and other methods), Version 4.0b10. Sunderland, MA: Sinauer Associates.
Thomas JW, Prasad AB, Summers TJ et al. (2002) Parallel construction of orthologous sequence-ready clone contig maps in multiple species. Genome Res 12: 1277–1285.
Yang F, O’Brien PCM, Ferguson-Smith MA (2000) Comparative chromosome map of the laboratory mouse and Chinese hamster defined by reciprocal chromosome painting. Chromosome Res 8: 219–227.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hass, I., Sbalqueiro, I.J. & Müller, S. Chromosomal phylogeny of four Akodontini species (Rodentia, Cricetidae) from Southern Brazil established by Zoo-FISH using Mus musculus (Muridae) painting probes. Chromosome Res 16, 75–88 (2008). https://doi.org/10.1007/s10577-007-1211-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-007-1211-5