Abstract
Protein ubiquitination and deubiquitination participate in a number of biological processes, including cell growth, differentiation, transcriptional regulation, and oncogenesis. Ubiquitin C-terminal hydrolases (UCHs), a subfamily of deubiquitinating enzymes (DUBs), includes four members: UCH-L1/PGP9.5 (protein gene product 9.5), UCH-L3, UCHL5/UCH37, and BRCA1-associated protein-1 (BAP1). Recently, more attention has been paid to the relationship between the UCH family and malignancies, which play different roles in the progression of different tumors. It remains controversial whether UCHL1 is a tumor promoter or suppressor. UCHL3 and UCH37 are considered to be tumor promoters, while BAP1 is considered to be a tumor suppressor. Studies have showed that UCH enzymes influence several signaling pathways that play crucial roles in oncogenesis, tumor invasion, and migration. In addition, UCH families are associated with tumor cell sensitivity to therapeutic modalities. Here, we reviewed the roles of UCH enzymes in the development of tumors, highlighting the potential consideration of UCH enzymes as new interesting targets for the development of anticancer drugs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
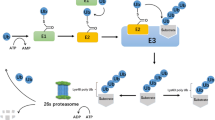
Ubiquitin (Ub) is a 76-aminoacid polypeptide that can covalently conjugate with protein substrates through the subsequent actions of three enzymes: ubiquitin-activating (E1), ubiquitin-conjugating (E2), and ubiquitin ligase (E3) enzymes. Ubiquitination process of a specific substrate protein requires selective E1, E2, and E3 [1,2,3]. However, only a subset of substrates that are ubiquitinated (such as Lys-48-linked polyubiquitination) are subsequently targeted to the 26S proteasome and degraded into small peptides. The ubiquitin-dependent proteasome degradation pathway (UPP) plays a crucial role in post-translational modification and degradation of proteins. In addition to the lysosomal degradation pathway, UPP is one of the main protein degradation pathways. Importantly, the discovery of Ub-mediated proteolysis was awarded the 2004 Nobel Prize in Chemistry [4,5,6,7,8].
In-depth research has revealed that protein ubiquitination is a highly reversible process. In some cases, deubiquitinating enzymes (DUBs), which are capable of removing Ub from protein substrates, protect proteins from degradation and release free Ub for recycling. However, in other cases, DUBs also enhance substrate degradation [9,10,11]. Protein ubiquitination and deubiquitination participate in a number of biological processes, including cell growth and differentiation, transcriptional regulation, and oncogenesis. Specifically, the dynamic relationship of these processes is important for the development of tumors [12,13,14].
Approximately 100 human DUBs have been identified, and these enzymes have been categorized into five subfamilies: ubiquitin-specific proteases/ubiquitin-specific processing proteases (USPs/UBPs), ubiquitin C-terminal hydrolases (UCHs), ovarian tumor proteases (OTUs), Josephin or Machado-Joseph disease protein domain proteases (MJDs), and Jab1/MPN domain-associated metalloisopeptidase (JAMM) domain proteins. Among these families, UBPs and UCHs have attracted a great deal of interest because of their various functions in cell behaviors [15,16,17,18]. To date, four UCH enzymes have been identified: UCH-L1/PGP9.5 (protein gene product 9.5), UCH-L3, UCHL5/UCH37, and BRCA1-associated protein-1 (BAP1) [19,20,21,22,23].
More attention has recently been paid to the relationship between UCH enzymes and malignancies. The effects of the four UCH members are quite complex, as these proteins play different roles in the progression of different tumors. For UCH-L1, researchers worldwide have failed to reach a consensus opinion on whether this enzyme is a promoter or suppressor in specific tumors. Herein, we reviewed the roles of UCH enzymes during the progression of tumors and illuminated the potential use of UCH enzymes as new interesting targets to develop anticancer drugs.
2 Biochemical characteristics of UCH enzymes
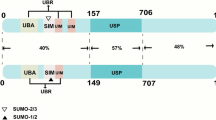
All UCH enzymes have a conserved catalytic domain (UCH-domain) comprising approximately 230 amino acids [24]. Studies on the crystal structures of UCH-L1, UCH-L3, and the UCH domain of UCH-L5 showed that all three UCHs contain an active-site crossover loop that is critical for the substrate specificity of the enzymes [20, 25,26,27,28,29,30,31,32,33] (Fig. 1).
Schematic representation of the UCH family. All UCH members have a conserved UCH-domain comprising approximately 230 amino acids. The C-terminal tail of UCH37 contains a KEKE motif, which plays the role of auto-inhibition via interacting with its UCH domain. The long C-terminal extension of BAP1 contains numerous binding sites for interaction proteins. UCH ubiquitin carboxyl hydrolase domain, KEKE KEKE motif, BARD1 BARD1 binding domain, HBM host cell factor 1 (HCF1) binding domain, FoxK1/2 forkhead transcription factors FoxK1/K2 binding domain BRCA1, BRCA1 binding domain, YY1 Ying Yang 1 binding domain, NLS nuclear localization signal
With only 223 amino acids, UCHL1 contains only one UCH domain and severs single amino acids or small peptides from the C-terminus of ubiquitin precursors. In dimeric form, UCH-L1 ligase activity produces Lys63-linked Ub chains to its substrates, which escape from the ubiquitin-dependent proteasome degradation pathway when polyubiquitinated via lysine 63 of Ub, leading to substrate stabilization. Reports have shown that the levels of α-synuclein are decreased, potentially reflecting one variant of UCHL1 (S18Y mutant) with reduced ubiquitin ligase activity, resulting in a reduced risk of Parkinson’s disease (PD). Another effect of UCHL1 is its capacity to bind to mono-ubiquitin (mono-Ub), leading to inhibition of mono-Ub degradation, which has been confirmed by experiments in neurons cells and gracile axonal dystrophy (gad) mice [34,35,36].
Although UCHL3 and UCHL1 have high homology, UCHL3 has differential biochemical features. UCHL3 displays hydrolyzing activity during the processing of both ubiquitin precursors and the poly-Ub chain from substrates [37, 38]. Moreover, UCHL3 cleaves Nedd8, an ubiquitin-like protein, from substrates, which is a unique feature of this enzyme. Unlike UCHL1, UCHL3 functions as a dimer-ubiquitin (di-Ub) stabilizer, which was confirmed by experiments in neurons cells and gad mice [39, 40].
UCH37 comprises an N-terminal UCH domain (residues 1–226) and C-terminal extension (residues 227–329) containing a KEKE motif (a group of amino acid residues with specific sequence). The C-terminal extension acts on the conserved catalytic region (UCH domain), which plays a role in autonomic inhibition [32]. hRpn13 is a subunit of the 26S proteasome. The N-terminal domain of hRpn13 binds to the polyubiquitin chain and hRpn2, while its C-terminal domain also contains a KEKE motif. The C-terminal tail of UCH37 binds to the C-terminal domain of hRpn13 through KEKE motifs, relieving UCH37 auto-inhibition. UCH37 is recruited to the proteasome through interactions with the KEKE motif, and this enzyme disassembles polyubiquitins from the distal subunit of the chain, through which ubiquitinated substrates avoid degradation [41,42,43,44].
BAP1, comprising 729 amino acids, is primarily localized in the nucleus [45]. In addition to the N-terminal UCH domain, which appears in all UCH enzymes, BAP1 has a long C-terminal extension containing numerous binding sites as interaction partners, including a host cell factor 1 (HCF1) binding domain (HBM), BRCA1, transcription factor Ying Yang 1 (YY1), forkhead transcription factors FoxK1/K2 binding domain, among others [46, 47]. The C-terminal tail of BAP1 also has nuclear localization signals (NLS), which are vital for its localization in the nucleus. BAP1 was named for its interactions with the RING finger domain of BRCA1, but it does not influence deubiquitination of BRCA1. Indeed, BAP1 can restrain the E3 ligase activity of the BRCA1–BARD1 heterodimeric complex by interacting with the RING domain of BARD1 (BRCA1-associated RING domain 1) [48, 49].
3 UCH members and cancers
3.1 UCHL1
UCHL1 is one of the most well studied UCH members. Previous studies focused on the role of UCHL1 in neurodegenerative disorders, particularly in Parkinson’s disease (PD) and Alzheimer’s disease (AD) [34, 36, 50, 51]. Decreased hydrolase activity could result in the accretion of some critical proteins, such as neurofibrillary and α-synuclein fibrils, which are associated with AD and PD, respectively [52,53,54,55]. Studies on the association between UCHL1 and neurodegeneration are ongoing, and several proteins interacting with UCHL1 have been identified in AD and other motor dysfunction diseases, including amyloid-β peptides (Aβ), synuclein, and amyloid beta (A4) precursor protein [56,57,58,59]. Subsequently, researchers have shown that some UCH-L1 variants are more susceptible to PD, as these variants cause α-synuclein accumulation in cells in vitro, which reflects the well-recognized hydrolase activity of UCH-L1. Researchers also showed that UCH-L1 possesses ligase activity. The dysfunction of UCHL1 ligase activity is also involved in neurodegeneration. Studies have illustrated that the S18Y mutant, a UCHL1 variant with decreased ubiquitin ligase activity, shows a positive effect associated with a lower PD risk through a reduction of α-synuclein levels [36, 60]. However, more recent studies have indicated a much lower possibility of the association between the S18Y mutant and PD risk in Asian population. Previously, researchers thought that UCH-L1’s ligase activity was resulted from certain dimeric form, but structure analysis showed no evidence of UCH-L1 existing as dimers. Furthermore, it is still unclear how UCH-L1 acts as a ligase from the structure perspective [61, 62].
The relationship between aberrant DNA methylation and malignant tumors has become a hot research topic over the last 20 years. Promoter CpG hypermethylation of UCHL1 is involved in several malignancies, including esophageal [63], gastric [64, 65], renal [66, 67], prostate [68, 69], head and neck squamous cell carcinoma [70], hepatocellular [71], ovarian [72, 73], nasopharyngeal [74], colorectal [75,76,77], and non-small cell lung cancers [78, 79]. In these malignant tumors, UCHL1 is decreased or silenced by promoter CpG hypermethylation, supporting the crucial effect of UCHL1 in tumor suppression. UCHL1 exerts tumor suppressor activities, primarily through activation of the p14ARF-p53 signaling pathway in breast cancer cells and nasopharyngeal carcinoma cells, thus restraining cell proliferation and inducing cancer cell apoptosis. Previous studies have demonstrated that UCHL1 can increase p53 and p14ARF through its hydrolase activity and downregulate MDM2 through its E3 ligase activity. UCHL1 subsequently upregulates the expression of p21 in hepatocellular carcinoma cells and p27 in lung cancer cell lines, which are inhibitors of cyclin-dependent kinases (CKD) [71, 74, 77, 80]. In addition, other potential roles for UCHL1 have been reported in hepatocellular carcinoma cells. Over-expression of UCHL1 can interdict cell evolution into mitosis from the G2 phase and hence suppress proliferation. In the caspase-dependent pathway, UCHL1 re-expression activates caspase-9 and subsequently initiates a caspase cascade and PARP cleavage, leading to DNA repair losses, cell disassembly, and apoptosis [71].
UCHL1 also shows high expression in other malignancies, such as breast cancer [81,82,83], cutaneous squamous cell cancer [84, 85], parathyroid carcinoma [86,87,88], melanoma [89, 90], and osteosarcoma [91, 92]. In contrast with previous studies, high UCHL1 mRNA levels were also detected in esophageal [93], gastric [94,95,96], non-small cell lung [97], and colorectal cancers [98,99,100] and were associated with aggressive phenotype and poor prognosis. For the same tumor, some studies have shown high UCHL1 expression, while other studies have shown low expression. The direct mechanism to this phenomenon remains unclear. The divergence may reflect different types of UCHL1 (wild or mutant), racial differences, the different pathway UCHL1 participate or other as yet unknown reasons. In such cases, UCHL1 influences several other pathways. Some studies have shown that UCHL1 activates Akt-mediated signaling depending on its hydrolase activity and subsequently promotes cell proliferation, migration, and invasion [89, 92, 97, 101]. Goto Y et al. showed that over-expression of UCHL1 made tumors more prone to distant metastasis, in association with poor prognosis in breast and lung cancer patients. The potential mechanism was subsequently examined, showing that UCHL1 can deubiquitinate HIF-1α, the regulatory subunit of hypoxia-inducible factor 1 (HIF-1), resulting in upregulation of HIF-1 signaling, consequently promoting metastasis [102]. A positive feedback exists between UCHL1 and β-catenin. UCHL1 can stabilize and upregulate β-catenin/TCF/Lef transcriptional activity, and β-catenin/TCF/Lef can reciprocally upregulate UCHL1. The expression of several genes, such as c-myc, cyclin D, c-jun, survivin, and other oncogenes, is induced by β-catenin/TCF/Lef-dependent transcription, indicating that β-catenin/TCF/Lef signaling can upregulate oncogenic cellular pathways [100, 103]. By contrast, UCHL1 interacts with Jun activation domain-binding protein-1 (JAB1), and this interaction could enhance the cytoplasmic transportation of P27. Subsequently, P27 is degraded in the cytoplasm, promoting cell proliferation [104]. In some cases, UCHL1 participates in signaling pathways that promote tumorigenesis. In other cases, UCHL1 plays a role in signaling pathways that inhibit tumorigenesis. Although each study had sufficient experimental data to support its conclusion, why did these studies have completely different results? Are these findings completely opposite as a result of different tumor types, or do these findings reflect different stages of tumor? It is likely that these differences may have no association with tumors and that UCHL1 is simply regulated by other more important factors that have not yet been identified. These deep mechanisms remain elusive, and the current understanding of the role of UCHL1 in cancer and signaling networks is superficial, prompting additional studies.
Some studies have shown that UCH-L1 can stabilize IκB-α, an inhibitor of nuclear factor-κB (NF-κB), through deubiquitination, resulting in NF-κB inactivation. Although there is no report showing the role of UCHL1 in tumors in the NF-κB pathway to date, the above-mentioned studies indicate that UCHL1 is a negative regulator of inflammatory responses, which provides new insights into the effects of UCHL1 on tumors via the NF-κB pathway [105,106,107,108,109]. A recent study showed that UCHL1 increased cellular reactive oxygen species (ROS) levels, indicating that UCHL1 is involved in oxidative stress. Studies have shown that UCH-L1 promoted melanoma cell invasion through upregulation of hydrogen peroxide generated via the deubiquitination of NADPH oxidase 4 (NOX4) [90].
Hence, the role of UCHL1 in tumors is complicated, and it remains controversial whether UCHL 1 is a tumor promoter or suppressor (Fig. 2). It is important to clarify whether UCHL1 has high expression or low expression in certain malignancies by expanding the clinical studies to determine the relationship between UCHL1 expression and tumorigenesis and prognosis. Whether UCHL1 can be used as a marker for early diagnosis of malignancy or as an evaluation indicator for prognosis has value in clinical practice. The signaling pathway in which UCHL1 participates and its transcriptional regulation remains unknown. Independent studies involving multiple signaling pathways have been conducted. If we put these pathways together and integrate them into a network through bioinformatics, it might be feasible to determine the role of UCHL1 in this vast network.
The role of UCHL1 in tumors is unclear. In the left column, UCHL1 plays a role as a suppressor, activating the P53 pathway via increasing P53 and decreasing MDM2. UCHL1 can initiate the caspase cascade and cause cell cycle arrest. Through the above functions, UCHL1 can inhibit cell proliferation and induce the apoptosis of cancer cells. In the right column, UCHL1 shows its tumor-promoting function. UCHL1 can upregulate β-catenin/TCF/Lef transcriptional activity and a positive feedback exists between UCHL1 and β-catenin. UCHL1 can activate Akt-mediated signaling and HIF-1 signaling, resulting in promotion of tumor invasion and metastasis. UCHL1 can enhance the cytoplasmic transportation of P27 by interacting with JAB1, and subsequently P27 is degraded in the cytoplasm, leading to promoting cell proliferation. In the middle column, UCHL1 can inactivate the NF-κB pathway and increase cellular reactive oxygen species (ROS) levels. These effects on the tumor through the NF-κB pathway or oxidative stress remain unclear. G2 cell cycle G2, M mitosis, NF-κB nuclear factor-κB, NOX4 NADPH oxidase 4, ROS reactive oxygen species, HIF hypoxia-inducible factor 1, JAB1 Jun activation domain-binding protein-1, L ligase activity, D deubiquitinating activity
3.2 UCHL3
Several in-depth studies for UCHL3 have been conducted on mammalian oocyte maturation. Aberrant cortical granule (CG) migration and meiotic spindle defects were observed in oocytes matured using the UCHL3 inhibitor [110,111,112,113]. However, there are few studies concerning the role of UCHL3 in tumors, and only a limited number of UCHL3 targets have been identified. Miyoshi Yet al. initially reported that UCHL3 was up-regulated in breast cancer [114]. Recently, Luo K et al. [115] elucidated that UCHL3 may be a novel regulator of DNA repair, which may be the potential mechanism of UCHL3 in oncogenesis. This study showed that RAD51 was deubiquitinated by UCHL3 and subsequently recruited to DNA double-strand breaks (DSBs) through interactions with BRCA2, a process that is critical for proper homologous recombination (HR), which is a major DSB repair pathway. Nevertheless, other studies have reported that UCHL3 was down-regulated in metastatic prostate cancer cell lines and showed that the knockdown of UCHL3 could promote epithelial-to-mesenchymal transition (EMT), resulting in cell invasion and metastasis [116]. Hence, the precise biological function of UCH-L3 in malignancies remains unknown. Does UCHL3 participate in different pathways in different tumors, and do these different roles reflect why UCHL3 has different expressions in different tumors? We first need to expand the sample size to determine whether UCHL3 is highly expressed or poorly expressed in malignancy and subsequently discuss the relationship between UCHL3 expression and tumorigenesis and prognosis. There are few studies concerning UCHL3 in signaling pathways, thus research in this area will likely start from scratch.
UCHL3 is unique in the UCH family for its deneddylation activity [39]. As cullins are the only identified target proteins for neddylation and since these proteins participate in cell-cycle control [117], UCHL3 may potentially act as a cell-cycle regulator. How does UCHL3 regulate the cell cycle? What cell-cycle factors are substrates for UCHL3? Are these cell-cycle factors regulated by UCHL3 through ubiquitination or deneddylation? None of the above questions have been answered. Thus, the role of UCHL3 in cell-cycle regulation is worth studying.
3.3 UCH37
Cohen and colleagues first discovered UCH37 in 1997 as a 19S-associated isopeptidase with a molecular mass of 37 kDa. These authors showed that UCH37 can selectively disassemble Lys48-linked poly-ubiquitin from the distal subunit of the chain in cells [118]. Subsequent studies have focused on how UCH37 deubiquitinates these substrates. UCH37 shows isopeptidase activity in the 19S proteasome regulatory complex, which is unique to the UCH family. hRpn13, component of the 19S particle, was bound to UCH37 via KEKE motifs in the C-terminal regions of hRpn13 and UCH37. UCH37 is recruited by the interaction of hRpn13 and UCH37 and activated to show deubiquitination activity [41, 42, 44, 119, 120]. Moreover, some studies have shown UCH37 combined with the human Ino80 chromatin-remodeling complex (hINO80) in the nucleus. These proteins bind via the C-terminal tail of UCH37 and the N-terminal domain of the hINO80 subunit (nuclear factor related to κB (NFRKB)). Although UCH37 maintains an inactive state in hINO80, this enzyme can be activated via the transient association of 19S regulatory particle- or proteasome-bound hRpn13 with hINO80 [43, 121,122,123]. In view of hINO80 as an ATP-dependent chromatin-remodeling complex that changes nucleosome positioning on DNA during both transcription and DNA repairmen [124], researchers have examined the function of UCH37 as a tumor regulator.
Rolén U et al. first reported upregulated expression of UCH37 in tumor tissues compared to adjacent normal tissues for cervical carcinoma [125]. In previous studies, we examined the carcinogenic character of UCH37. We first observed higher UCH37 expression among hepatocellular carcinoma (HCC) tissues compared with adjacent para-cancerous tissues and showed that UCH37 could be a predictor of HCC recurrence in HCC patients after radical resection [126]. Subsequently, we also observed higher UCH37 expression among esophageal squamous cell carcinoma (ESCC) [127] and epithelial ovarian cancer (EOC) patients [128] and observed the association between high UCH37 expression and poor clinical outcome. The multivariate analysis results revealed the predicting character of UCH37 for overall survival (OS) and disease-free survival (DFS) and potentially also for tumor recurrence. Recently, other studies have reported that UCH37 is involved in lung cancer [129] and pancreatic carcinoma [130], which further confirmed its potential roles in oncogenesis.
Recently, several studies have focused on the potential mechanism of UCH37 in malignancy. Thus far, the role of UCH37 in tumor via upregulation of transforming growth factor-β (TGF-β) signaling is relatively clear [130,131,132]. UCH37 can bind to Smad7 by competing with Smad ubiquitination regulatory factor 2 (Smurf2), a ubiquitin ligase, which prevents the formation of the Smurf2-Smad7 complex. As a result, the binding of Smurf2 and type I TGF-β receptor is inhibited and the type I TGF-β receptor is rescued from proteasomal degradation, leading to the upregulation of TGF-β signaling. Further studies on the relationship between UCH37 and TGF-β signaling showed that UCH37 knockdown selectively reduces the levels of certain TGF-β-dependent target genes, such as MMP-2 and PAI-1, which are crucial proteins in promoting tumor migration and invasion.
Other studies have reported other potential roles for UCH37. We performed functional proteomic analyses and screened proteins interacting with UCH37 in HCC and identified several meaningful downstream proteins, including glucose-regulated protein 78 (GRP78), a protein essential for cellular viability [133]. Subsequently, these studies showed that UCH37 could deubiquitinate PRP19 (essential RNA splicing factor) and promote migration and invasion of HCC cell [126]. Chen Z et al. illustrated that UCH37 silencing could trigger cell apoptosis in A549 cells (non-small-cell lung cancer cells) through activation of Bax/Bcl-2, caspase-9, and caspase-3 [129]. Christina S et al. showed that UCH37 deubiquitinates E2 promoter binding factor 1 (E2F1) and activates E2F1 transcriptional activity, resulting in proliferative and pro-apoptotic E2F1 target genes activation. Furthermore, E2F1 and UCH37 generate a positive feedback mechanism [134]. Recently, Han W et al. reported that UCH37 interacts with Tcf7 (one of Tcf/Lef family) to activate Wnt/β-catenin signaling and functions as a positive regulator of the Wnt/β-catenin pathway, similar to UCHL1 to some extent [135]. However, the above-mentioned pathways have only been sporadically reported, and further studies are needed for confirmation.
The downstream proteins of UCH37 have been extensively studied, but little focus has been paid to upstream regulation of UCH37. DNA methylation and microRNAs are important in regulating gene expression. Until recently, only three potential methylation sites in the UCH37 promoter have been predicted in cells. It is highly feasible that a total of three conserved sites and three poorly conserved sites serve as the target regions of miRNAs [136].
Increasing evidence for the important role of UCH37 in malignancies is surfacing. However, we only have limited knowledge of the effects of this enzyme on signaling. Additional studies to identify UCH37 substrates discuss how UCH37 deubiquitinates substrates, explore up-stream UCH37 regulation, and so on are in progress, and we look forward to observing how the studies unveil the deeper mechanism underlying the effects of UCH37 on oncogenesis.
3.4 BAP1
In 2010, the BAP1 gene was first reported as being somatically mutated in uveal melanoma with poor prognosis. It was reported that 26 of 31 metastasizing tumors had BAP1 mutations, including 13 out-of-frame deletions and 2 nonsense mutations, which led to premature protein termination, 6 missense mutations, 4 in-frame deletions, and 1 mutation, which was predicted to produce an abnormally extended BAP1 polypeptide [137]. Thus far, BAP1 has been implicated in several malignancies, including mesothelioma [138,139,140,141,142,143], uveal melanoma [144,145,146,147], cutaneous melanoma [148], cholangiocarcinoma [149,150,151,152], non-small-cell lung cancer [153, 154], clear cell renal carcinoma [155, 156], breast carcinoma [157, 158], and colorectal cancer [159]. BAP1 has been recognized as a tumor suppressor. Recently, Luchini C et al. [160] performed meta-analysis and discussed the relationship between BAP1 and malignancy. This study delivered two important conclusions. First, BAP1 acts as a tumor suppressor gene whose loss or mutation is associated with poor prognosis, such as increased all-cause mortality, cancer-specific mortality, and recurrence of cancer. This connection was identified in all types of analyzed tumors, except mesothelioma, for which controversial opinions about the effects of BAP1 have been reported. Some studies have reported that BAP1-mutated mesothelioma is associated with a longer survival time. From 1991 to 2014, Farzin et al. collected tissues from 229 thoracic mesothelioma patients and performed immunohistochemistry. This study showed that BAP1 loss corresponded to an improved median survival of 16.11 months (95% CI 12.16–20.06) versus 6.34 months (95% CI 5.34–7.34), p < 0.01 [161]. Baumann et al. compared survival among malignant mesothelioma patients with germline BAP1 mutations (N = 23) with that of all malignant mesothelioma patients (N = 10,556) recorded in the US Surveillance, Epidemiology, and End Results (SEER) data from 1973 to 2010. These authors showed that malignant mesothelioma patients with germline BAP1 mutations had an overall 7-times higher long-term survival, independent of sex and age [162]. This meta-analysis indicated the protective characteristic of BAP1 mutations in mesothelioma for all-cause mortality with a pooled RR of 0.92 (95% CI: 0.87–0.97) and HR adjusted for potential confounders of 0.52 (95% CI: 0.36–0.75). However, only the above two studies were included in this meta-analysis to analyze the relationship between BAP1 and malignant mesothelioma; therefore, the results may be biased. However, another study using a mouse model showed that BAP1 knockout mice are susceptible to the tumorigenic effects of asbestos compared to wild-type littermates (73 vs. 32%, respectively) [163]. Hence, additional studies are needed to ascertain the prognostic role of BAP1 in mesothelioma. Nevertheless, it is clear that a histopathological diagnosis using BAP1 immunohistochemistry will distinguish between malignant and benign mesothelial proliferations. Notably, malignant mesothelioma should be distinguished from reactive mesothelial proliferations using a highly specific method, detecting BAP1 impairment through immunohistochemistry and detecting p16 impairment through fluorescent in situ hybridization [164, 165]. Second, BAP1 mutations have been associated with high-grade renal and colorectal carcinoma. Furthermore, germline BAP1 mutations have been identified in familial neoplastic disease. Members of the BAP1-mutant families are more likely to develop malignancies, and some of the members possessed two or more tumors [48]. A recent study identified germline BAP1 mutations in sporadic melanoma. In this work, a total of 30 B.P. variants were identified, of which 27 variants were rare and 3 variants were common or polymorphisms [166].
Since BAP1 binds to the wild-type BRCA1 RING finger domain, it was believed that BAP1 acts as a suppressor gene by interacting with and deubiquitinating BRCA1. However, subsequent studies showed that BRCA1 was not the substrate of BAP1 for its deubiquitination activity. In recent years, additional studies have attempted to explain the molecular mechanism of BAP1 in tumors from other points of view. Studies have shown that BRCA1 forms a heterodimer through the combination of its RING domain with BRCA1-associated RING domain 1 (BARD1). This BRCA1–BARD1 complex exhibits E3 ubiquitin ligase activity, which regulates the DNA damage response (DDR) [167]. BAP1 modulates E3 ligase activity [168] via binding and deubiquitylating BARD1. Studies have shown that the loss of BAP1 impairs DDR and causes HeLa cells to become hypersensitive to ionizing radiation. In the nucleus, BAP1 interacts with host cell factor 1 (HCF1) [47], a cell cycle regulator, through its HCF1 binding motif (HBM), which is in the middle part of BAP1. Several transcription factors also contain HBM, such as Ying Yang 1 (YY1) [48], forkhead transcription factors FoxK1/K2 [46], O-linked N-acetylglucosaminetransferase (OGT) [169], and the E2F family [170]. BAP1 forms a ternary complex with HCF1 and transcription factors, which recruits HCF1 to specific promoters to regulate transcription and control cell proliferation [170]. In addition, some experiments have shown that BAP1 can directly interact with the E2F family and OGT, indicating that BAP1 may regulate cellular processes by deubiquitinating the E2F family and OGT [48, 170] (Fig. 3).
The function of BAP1 in regulating transcription. a BRCA1–BARD1 complex has E3 ubiquitin ligase activity that regulates the DNA damage response. BAP1 binds and deubiquitylates BARD1. b BAP1 forms a ternary complex with HCF1 and transcription factors, which recruits HCF1 to specific promoters to regulate transcription and control cell proliferation. c Histones are ubiquitinated by PRCs and deubiquitinated by PR-DUB, which plays crucial role in transcriptional balance. BARD1 BRCA1 associated RING domain 1, HCF1 host cell factor 1, PRCs polycomb-repressive complexes, PR-DUB polycomb group repressive deubiquitinase complex, ASXL1/2 additional sex combs
The polycomb group proteins are critical transcriptional regulators. These proteins contain polycomb-repressive complexes (PRCs) that ubiquitinate histones, leading to gene silencing. BAP1 interacts with additional sex combs (ASXL1/2) to form the polycomb group repressive deubiquitinase complex (PR-DUB). Transcriptional balances are maintained through histone ubiquitination by PRCs and deubiquitination by PR-DUB. Loss of BAP1 increased the ubiquitination level of histone 2A, promoted the deregulation of cell cycle progression and thus impeded cellular senescence [170, 171].
Therefore, BAP1 plays a role as suppressor in tumors, which is different from other UCH members. This is not only interesting but also worth further studies to illuminate the mechanisms resulting in this difference.
4 UCH members and treatment
Research on the application of UCH family members in tumor treatment is still preliminary. Thus far, studies focused on the relationship between the UCH family and tumor sensitivity to therapeutic modalities. There is no report concerning the UCH family as therapeutic targets or drugs. Similar to controversial opinions regarding the effect of UCHL1 in tumors (promoter or suppressor), contrasting conclusions exist concerning its role for the treatment of tumors. Jin Y et al. [83] showed that breast cancer patients with higher UCH-L1 expression after chemotherapy presented shorter overall survival (OS) and disease-free survival (DFS) than those with downregulated or unchanged UCH-L1 expression. However, in renal clear cell carcinoma (RCC), decitabine may suppress the proliferation of RCC cells by inhibiting UCHL1, interferon inducible protein 27 (IFI27), and cell division cycle-associated 2 (CDCA2), suggesting that UCHL1 expression may increase RCC cell sensitivity to decitabine [172]. Moreover, Brinkmann K et al. reported that UCHL1 strengthens cancer chemosensitivity in melanoma and colorectal cancer by stabilizing NOXA [173]. Given these disagreements, it is necessary to make clear whether UCHL1 is the target of cancer treatment. As mentioned above, promotion of CpG hypermethylation of UCHL1 is associated with several malignancies. Regulation of the DNA methylation of UCHL1 may be a promising treatment for cancers in the future.
BAP1 was reported to lead cancer cells to respond differently to radiotherapy and chemotherapy. Recent studies have shown that BAP1 loss enhances cancer cell sensitivity to radiotherapy, likely reflecting the impaired ability to repair double-stranded DNA breaks induced by these cellular stressors [174]. As to chemotherapy, BAP1 differently affects tumor cell sensitivity to different drugs. In uveal melanoma, BAP1 can make cells more vulnerable to histone deacetylase (HDAC) inhibitors, which can induce morphologic differentiation and cell-cycle arrest. Currently, several HDAC inhibitors are under phase II trial in metastatic renal cell carcinoma (RCC) patients, such as LBH589 (panobinostat), FK228 (depsipeptide), vorinostat, MS-275, belinostat, and entinostat (in combination with IL-2) [156]. These inhibitors may become potential therapeutic drugs in the future. However, another study indicated that BAP1 loss sensitized RCC cells to the PARP inhibitor olaparib [174]. Researchers are evaluating the combination of olaparib and AZD5363 (an AKT inhibitor) in a phase I study with advanced solid tumors patients, including cases of RCC refractory to standard therapy [156].
Regarding the impact of UCHL3 in cancer treatment, KunL et al. [115] showed that breast cancer patients with UCHL3 over-expression had poor survival and were more resistant to radiation and chemotherapy, while decreased UCHL3 sensitized cells to PARPi and radiotherapy, which may reflect the BRCA2-RAD51 pathway. These findings reveal the potential to make UCHL3 a new therapeutic target, overcoming resistance to standard therapy. In combination with existing therapies, UCHL3 targeting therapeutic interventions may hopefully improve treatment efficacy.
Thus far, the role of UCH37 in the treatment of tumors has not been reported, reflecting limited knowledge concerning the molecular mechanism of this enzyme in oncogenesis. First, the role of UCH37 in malignancy, which is a pre-condition for subsequent studies of the effects of UCH37 on cancer treatment, should be clarified. Furthermore, studies on the relationship between the expression of UCH37 and the therapeutic effect, and the sensitivity of UCH37 to chemotherapy drugs in vitro are needed.
As described above, studies on the effects of UCH family members in tumor treatment are needed. Future studies to totally understand the roles of UCH family members as potential therapeutic targets or drugs are ongoing.
5 Prospective
In summary, each member of the UCH family has its own characteristics in tumors. Despite numerous research studies on the association of UCH proteins and tumors, many questions remain unanswered. What causes the different effects of UCH family members in tumors? Why is UCHL1/UCHL3/UCH37 considered to be a tumor promoter while BAP1 is considered to be a tumor suppressor? Since UCH family members share some common pathways, is there a synergistic action between them during the development of tumors? Is there any relation between each of the pathways? What are the precise roles of these enzymes in the development of tumors? What are the precise substrates and up-stream regulators of the UCH family? Can UCH family members become potential therapeutic targets or drugs? The answers to the above questions may not only provide new insights into the oncogenesis but also open the new space for humankind in tumor treatments.
Abbreviations
- Aβ:
-
Amyloid-β peptides
- AD:
-
Alzheimer’s disease
- BAP1:
-
BRCA1-associated protein-1
- BARD1:
-
BRCA1-associated RING domain 1
- CDCA2:
-
Cell division cycle-associated 2
- CG:
-
Cortical granule
- CKD:
-
Cyclin-dependent kinases
- DDR:
-
DNA damage response
- DFS:
-
Disease-free survival
- di-Ub:
-
Dimers-ubiquitin
- DSBs:
-
Double-strand breaks
- DUBs:
-
Deubiquitinating enzymes
- E2F1:
-
E2 promoter binding factor 1
- EMT:
-
Epithelial-to-mesenchymal transition
- EOC:
-
Epithelial ovarian cancer
- ESCC:
-
Esophageal squamous cell carcinoma
- gad:
-
Gracile axonal dystrophy
- GRP78:
-
Glucose-regulated protein 78
- HBM:
-
HCF1 binding domain
- HCC:
-
Hepatocellular carcinoma
- HCF1:
-
Host cell factor 1
- HDAC:
-
Histone deacetylase
- HIF-1:
-
Hypoxia-inducible factor 1
- hINO80:
-
Human Ino80 chromatin-remodeling complex
- HR:
-
Proper homologous recombination
- IFI27:
-
Interferon inducible protein 27
- JAB1:
-
Jun activation domain-binding protein-1
- JAMM:
-
Jab1/MPN domain-associated metalloisopeptidase
- MJDs:
-
Josephin or Machado-Joseph disease protein domain proteases
- mono-Ub:
-
Mono-ubiquitin
- NF-κB:
-
Nuclear factor-κB
- NFRKB:
-
Nuclear factor related to κB
- NLS:
-
Nuclear localization signals
- NOX4:
-
NADPH oxidase 4
- OGT:
-
O-linked N-acetylglucosamine transferase
- OS:
-
Overall survival
- OTUs:
-
Ovarian tumor proteases
- PD:
-
Parkinson’s disease
- PGP9.5:
-
Protein gene product 9.5
- PRCs:
-
Polycomb-repressive complexes
- PR-DUB:
-
Polycomb group repressive deubiquitinase complex
- RCC:
-
Renal clear cell carcinoma
- ROS:
-
Reactive oxygen species
- Smurf2:
-
Smad ubiquitination regulatory factor 2
- TGF-β:
-
Transforming growth factor-β
- Ub:
-
Ubiquitin
- UCHs:
-
Ubiquitin C-terminal hydrolases
- UPP:
-
Ubiquitin-dependent proteasome degradation pathway
- USPs/UBPs:
-
Ubiquitin-specific proteases/ubiquitin-specific processing proteases
- YY1:
-
Ying Yang 1
References
Hershko, A., & Ciechanover, A. (1998). The ubiquitin system. Annual Review of Biochemistry, 67, 425–479.
Pickart, C. M. (2001). Mechanisms underlying ubiquitination. Annual Review of Biochemistry, 70, 503–533.
Finley, D., Ciechanover, A., & Varshavsky, A. (2004). Ubiquitin as a central cellular regulator. Cell, 116(Suppl. 2), S29–S32 2p following S32.
Komander, D., & Rape, M. (2012). The ubiquitin code. Annual Review of Biochemistry, 81, 203–229.
Liu, J., & Nussinov, R. (2013). The role of allostery in the ubiquitin-proteasome system. Critical Reviews in Biochemistry and Molecular Biology, 48(2), 89–97.
Varshavsky, A. (2012). The ubiquitin system, an immense realm. Annual Review of Biochemistry, 81, 167–176.
Xu, G., & Jaffrey, S. R. (2011). The new landscape of protein ubiquitination. Nature Biotechnology, 29(12), 1098–1100.
Welchman, R. L., Gordon, C., & Mayer, R. J. (2005). Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nature Reviews Molecular Cell Biology, 6(8), 599–609.
Neutzner, M., & Neutzner, A. (2012). Enzymes of ubiquitination and deubiquitination. Essays in Biochemistry, 52, 37–50.
Singhal, S., Taylor, M. C., & Baker, R. T. (2008). Deubiquitylating enzymes and disease. BMC Biochemistry, 9(Suppl. 1), S3.
Katz, E. J., Isasa, M., & Crosas, B. (2010). A new map to understand deubiquitination. Biochemical Society Transactions, 38(Pt 1), 21–28.
Sridhar, V. V., Kapoor, A., Zhang, K., Zhu, J., Zhou, T., Hasegawa, P. M., et al. (2007). Control of DNA methylation and heterochromatic silencing by histone H2B deubiquitination. Nature, 477(7145), 735–738.
Liu, H., Buus, R., Clague, M. J., & Urbe, S. (2009). Regulation of ErbB2 receptor status by the proteasomal DUB POH1. PLoS One, 4(5), e5544.
Reyes-Turcu, F. E., Ventii, K. H., & Wilkinson, K. D. (2009). Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annual Review of Biochemistry, 78, 363–397.
Nijman, S. M., Luna-Vargas, M. P., Velds, A., Brummelkamp, T. R., Dirac, A. M., Sixma, T. K., et al. (2005). A genomic and functional inventory of deubiquitinating enzymes. Cell, 123(5), 773–786.
Sowa, M. E., Bennett, E. J., Gygi, S. P., & Harper, J. W. (2009). Defining the human deubiquitinating enzyme interaction landscape. Cell, 138(2), 389–403.
Amerik, A. Y., & Hochstrasser, M. (2004). Mechanism and function of deubiquitinating enzymes. Biochimica et Biophysica Acta, 1695(1–3), 189–207.
Soboleva, T. A., & Baker, R. T. (2004). Deubiquitinating enzymes: their functions and substrate specificity. Current Protein & Peptide Science, 5(3), 191–200.
Lam, Y. A., Xu, W., DeMartino, G. N., & Cohen, R. E. (1997). Editing of ubiquitin conjugates by an isopeptidase in the 26S proteasome. Nature, 385(6618), 737–740.
Larsen, C. N., Krantz, B. A., & Wilkinson, K. D. (1998). Substrate specificity of deubiquitinating enzymes: ubiquitin C-terminal hydrolases. Biochemistry, 37(10), 3358–3368.
Doran, J. F., Jackson, P., Kynoch, P. A., & Thompson, R. J. (1983). Isolation of PGP 9.5, a new human neurone-specific protein detected by high-resolution two-dimensional electrophoresis. Journal of Neurochemistry, 40(6), 1542–1547.
Wilkinson, K. D., Lee, K. M., Deshpande, S., Duerksen-Hughes, P., Boss, J. M., & Pohl, J. (1989). The neuron-specific protein PGP 9.5 is a ubiquitin carboxyl-terminal hydrolase. Science, 246(4930), 670–673.
Jensen, D. E., Proctor, M., Marquis, S. T., Gardner, H. P., Ha, S. I., Chodosh, L. A., et al. (1998). BAP1: a novel ubiquitin hydrolase which binds to the BRCA1 RING finger and enhances BRCA1-mediated cell growth suppression. Oncogene, 16(9), 1097–1112.
Todi, S. V., & Paulson, H. L. (2011). Balancing act: deubiquitinating enzymes in the nervous system. Trends in Neurosciences, 34(7), 370–382.
Day, I. N., & Thompson, R. J. (1987). Molecular cloning of cDNA coding for human PGP 9.5 protein. A novel cytoplasmic marker for neurones and neuroendocrine cells. FEBS Letters, 210(2), 157–160.
Das, C., Hoang, Q. Q., Kreinbring, C. A., Luchansky, S. J., Meray, R. K., Ray, S. S., et al. (2006). Structural basis for conformational plasticity of the Parkinson’s disease-associated ubiquitin hydrolase UCH-L1. Proceedings of the National Academy of Sciences USA, 103(12), 4675–4680.
Zhou, Z. R., Zhang, Y. H., Liu, S., Song, A. X., & Hu, H. Y. (2012). Length of the active-site crossover loop defines the substrate specificity of ubiquitin C-terminal hydrolases for ubiquitin chains. Biochemical Journal, 441(1), 143–149.
Johnston, S. C., Larsen, C. N., Cook, W. J., Wilkinson, K. D., & Hill, C. P. (1997). Crystal structure of a deubiquitinating enzyme (human UCH-L3) at 1.8A° resolution. EMBO Journal, 16(13), 3787–3796.
Hirayama, K., Aoki, S., Nishikawa, K., Matsumoto, T., & Wada, K. (2007). Identification of novel chemical inhibitors for ubiquitin C-terminal hydrolase-L3 by virtual screening. Bioorganic and Medicinal Chemistry, 15(21), 6810–6818.
Navarro, M. F., Carmody, L., Romo-Fewell, O., Lokensgard, M. E., & Love, J. J. (2014). Characterizing substrate selectivity of ubiquitin C-terminal hydrolase-L3 using engineered α-linked ubiquitin substrates. Biochemistry, 53(51), 8031–8042.
Nishio, K., Kim, S. W., Kawai, K., Mizushima, T., Yamane, T., et al. (2009). Crystal structure of the de-ubiquitinating enzyme UCH37 (human UCH-L5) catalytic domain. Biochemical and Biophysical Research Communications, 390(3), 855–860.
Yao, T., Song, L., Xu, W., DeMartino, G. N., Florens, L., Swanson, S. K., et al. (2006). Proteasome recruitment and activation of the Uch37 deubiquitinating enzyme by Adrm1. Nature Cell Biology, 8(9), 994–1002.
Burgie, S. E., Bingman, C. A., Soni, A. B., & Phillips Jr., G. N. (2012). Structural characterization of human Uch37. Proteins, 80(2), 649–654.
Leroy, E., Boyer, R., Auburger, G., Leube, B., Ulm, G., Mezey, E., et al. (1998). The ubiquitin pathway in Parkinson's disease. Nature, 395(6701), 451–452.
Osaka, H., Wang, Y. L., Takada, K., Takizawa, S., Setsuie, R., Li, H., et al. (2003). Ubiquitin carboxy-terminal hydrolase L1 binds to and stabilizes monoubiquitin in neuron. Human Molecular Genetics, 12(16), 1945–1958.
Liu, Y., Fallon, L., Lashuel, H. A., Liu, Z., & Lansbury Jr., P. T. (2002). The UCH-L1 gene encodes two opposing enzymatic activities that affect alpha-synuclein degradation and Parkinson’s disease susceptibility. Cell, 111(2), 209–218.
Hemelaar, J., Borodovsky, A., Kessler, B. M., Reverter, D., Cook, J., Kolli, N., et al. (2004). Specific and covalent targeting of conjugating and deconjugating enzymes of ubiquitin-like proteins. Molecular and Cellular Biology, 24(1), 84–95.
Grou, C. P., Pinto, M. P., Mendes, A. V., Domingues, P., & Azevedo, J. E. (2015). The de novo synthesis of ubiquitin: identification of deubiquitinases acting on ubiquitin precursors. Scientific Reports, 5, 12836.
Frickel, E. M., Quesada, V., Muething, L., Gubbels, M. J., Spooner, E., Ploegh, H., et al. (2007). Apicomplexan UCHL3 retains dual specificity for ubiquitin and Nedd8 throughout evolution. Cellular Microbiology, 9(6), 1601–1610.
Misaghi, S., Galardy, P. J., Meester, W. J., Ovaa, H., Ploegh, H. L., & Gaudet, R. (2005). Structure of the ubiquitin hydrolase UCH-L3 complexed with a suicide substrate. Journal of Biological Chemistry, 280(2), 1512–1520.
Husnjak, K., Elsasser, S., Zhang, N., Chen, X., Randles, L., Shi, Y., et al. (2008). Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature, 453(7194), 481–488.
Schreiner, P., Chen, X., Husnjak, K., Randles, L., Zhang, N., Elsasser, S., et al. (2008). Ubiquitin docking at the proteasome through a novel pleckstrin-homology domain interaction. Nature, 453(7194), 548–552.
VanderLinden, R. T., Hemmis, C. W., Schmitt, B., Ndoja, A., Whitby, F. G., Robinson, H., et al. (2015). Structural basis for the activation and inhibition of the UCH37 deubiquitylase. Molecular Cell, 57(5), 901–911.
Jiao, L., Ouyang, S., Shaw, N., Song, G., Feng, Y., Niu, F., et al. (2014). Mechanism of the Rpn13-induced activation of Uch37. Protein & Cell, 5(8), 616–630.
Ventii, K. H., Devi, N. S., Friedrich, K. L., Chernova, T. A., Tighiouart, M., Van Meir, E. G., et al. (2008). BRCA1-associated protein 1 is a tumor suppressor that requires deubiquitinating activity and nuclear localization. Cancer Research, 68(17), 6953–6962.
Okino, Y., Machida, Y., Frankland-Searby, S., & Machida, Y. J. (2015). BRCA1-associated protein 1 (BAP1) deubiquitinase antagonizes the ubiquitin-mediated activation of FoxK2 target genes. Journal of Biological Chemistry, 290(3), 1580–1591.
Misaghi, S., Ottosen, S., Izrael-Tomasevic, A., Arnott, D., Lamkanfi, M., Lee, J., et al. (2009). Association of C-terminal ubiquitin hydrolase BRCA1-associated protein 1 with cell cycle regulator host cell factor 1. Molecular and Cellular Biology, 29(8), 2181–2192.
Carbone, M., Yang, H., Pass, H. I., Krausz, T., Testa, J. R., & Gaudino, G. (2013). BAP1 and cancer. Nature Reviews Cancer, 13(3), 153–159.
Mallery, D. L., Vandenberg, C. J., & Hiom, K. (2002). Activation of the E3 ligase function of the BRCA1/BARD1 complex by polyubiquitin chains. EMBO Journal, 21(24), 6755–6762.
Choi, J., Levey, A. I., Weintraub, S. T., Rees, H. D., Gearing, M., Chin, L. S., et al. (2004). Oxidative modifications and down-regulation of ubiquitin carboxyl-terminal hydrolase L1 associated with idiopathic Parkinson's and Alzheimer's diseases. Journal Biological Chemistry, 279(13), 13256–13264.
Son, O. L., Kim, H. T., Ji, M. H., Yoo, K. W., Rhee, M., & Kim, C. H. (2003). Cloning and expression analysis of a Parkinson's disease gene, Uch-L1, and its promoter in zebrafish. Biochemical and Biophysical Research Communications, 312(3), 601–607.
Barrachina, M., Castano, E., Dalfo, E., Maes, T., Buesa, C., & Ferrer, I. (2006). Reduced ubiquitin C-terminal hydrolase-1 expression levels in dementia with Lewy bodies. Neurobiology of Disease, 22(2), 265–273.
Gong, B., Cao, Z., Zheng, P., Vitolo, O. V., Liu, S., Staniszewski, A., et al. (2006). Ubiquitin hydrolase Uch-L1 rescues β-amyloid-induced decreases in synaptic function and contextual memory. Cell, 126(4), 775–788.
Cartier, A. E., Djakovic, S. N., Salehi, A., Wilson, S. M., Masliah, E., & Patrick, G. N. (2009). Regulation of synaptic structure by ubiquitin C-terminal hydrolase L1. Journal of Neuroscience, 29(24), 7857–7868.
Lederer, C. W., Torrisi, A., Pantelidou, M., Santama, N., & Cavallaro, S. (2007). Pathways and genes differentially expressed in the motor cortex of patients with sporadic amyotrophic lateral sclerosis. BMC Genomics, 8, 26.
Lakshmana, M. K., Chung, J. Y., Wickramarachchi, S., Tak, E., Bianchi, E., Koo, E. H., et al. (2010). A fragment of the scaffolding protein RanBP9 is increased in Alzheimer’s disease brains and strongly potentiates amyloidbeta peptide generation. FASEB Journal, 24(1), 119–127.
Zhou, Y., Gu, G., Goodlett, D. R., Zhang, T., Pan, C., Montine, T. J., et al. (2004). Analysis of alpha-synuclein-associated proteins by quantitative proteomics. Journal of Biological Chemistry, 279(37), 39155–39164.
Zhang, M., Deng, Y., Luo, Y., Zhang, S., Zou, H., Cai, F., et al. (2012). Control of BACE1 degradation and APP processing by ubiquitin carboxyl-terminal hydrolase L1. Journal of Neurochemistry, 120(6), 1129–1138.
Guglielmotto, M., Monteleone, D., Boido, M., Piras, A., Giliberto, L., Borghi, R., et al. (2012). Aβ1-42-mediated down-regulation of Uch-L1 is dependent on NF-κB activation and impaired BACE1 lysosomal degradation. Aging Cell, 11(5), 834–844.
Carmine Belin, A., Westerlund, M., Bergman, O., Nissbrandt, H., Lind, C., Sydow, O., et al. (2007). S18Y in ubiquitin carboxy-terminal hydrolase L1 (UCH-L1) associated with decreased risk of Parkinson's disease in Sweden. Parkinsonism & Related Disorders, 13(5), 295–298.
Sun, S., Zhao, Y., Jin, G., & Kang, H. (2014). Lack of association between UCHL1 S18Y gene polymorphism and Parkinson's disease in the Asian population: a meta-analysis. Neurological Sciences, 35(12), 1867–1876.
Bishop, P., Rocca, D., & Henley, J. M. (2016). Ubiquitin C-terminal hydrolase L1 (UCH-L1): structure, distribution and roles in brain function and dysfunction. The Biochemical Journal, 473(16), 2453–2462.
Mandelker, D. L., Yamashita, K., Tokumaru, Y., Mimori, K., Howard, D. L., Tanaka, Y., et al. (2005). PGP9.5 promoter methylation is an independent prognostic factor for esophageal squamous cell carcinoma. Cancer Research, 65(11), 4963–4968.
Yamashita, K., Park, H. L., Kim, M. S., Osada, M., Tokumaru, Y., Inoue, H., et al. (2006). PGP9.5 methylation in diffusetype gastric cancer. Cancer Research, 66(7), 3921–3927.
Wang, G., Zhang, W., Zhou, B., Jin, C., Wang, Z., Yang, Y., et al. (2015). The diagnosis value of promoter methylation of UCHL1 in the serum for progression of gastric cancer. BioMed Research International, 2015, 741030.
Kagara, I., Enokida, H., Kawakami, K., Matsuda, R., Toki, K., Nishimura, H., et al. (2008). CpG hypermethylation of the UCHL1 gene promoter is associated with pathogenesis and poor prognosis in renal cell carcinoma. The Joural of Urology, 180(1), 343–351.
Seliger, B., Handke, D., Schabel, E., Bukur, J., Lichtenfels, R., & Dammann, R. (2009). Epigenetic control of the ubiquitin carboxyl terminal hydrolase 1 in renal cell carcinoma. Journal of Translational Medicine, 7, 90.
Wang, Y., Yu, Q., Cho, A. H., Rondeau, G., Welsh, J., Adamson, E., et al. (2005). Survey of differentially methylated promoters in prostate cancer cell lines. Neoplasia, 7(8), 748–760.
Mitsui, Y., Shiina, H., Hiraki, M., Arichi, N., Hiraoka, T., Sumura, M., et al. (2012). Tumor suppressor function of PGP9.5 is associated with epigenetic regulation in prostate cancer--novel predictor of biochemical recurrence after radical surgery. Cancer Epidemiol Biomarkers Prevention, 21(3), 487–496.
Tokumaru, Y., Yamashita, K., Kim, M. S., Park, H. L., Osada, M., Mori, M., et al. (2008). The role of PGP9.5 as a tumor suppressor gene in human cancer. Internation Journal of Cancer, 123(4), 753–759.
Yu, J., Tao, Q., Cheung, K. F., Jin, H., Poon, F. F., Wang, X., et al. (2008). Epigenetic identification of ubiquitin carboxyl-terminal hydrolase L1 as a functional tumor suppressor and biomarker for hepatocellular carcinoma and other digestive tumors. Hepatology, 48(2), 508–518.
Okochi-Takada, E., Nakazawa, K., Wakabayashi, M., Mori, A., Ichimura, S., Yasugi, T., et al. (2006). Silencing of the UCHL1 gene in human colorectal and ovarian cancers. Internation Journal of Cancer, 119(6), 1338–1344.
Brait, M., Maldonado, L., Noordhuis, M. G., Begum, S., Loyo, M., et al. (2013). Association of promoter methylation of VGF and PGP9.5 with ovarian cancer progression. PLoS One, 8(9), e70878.
Li, L., Tao, Q., Jin, H., van Hasselt, A., Poon, F. F., et al. (2010). The tumor suppressor UCHL1 forms a complex with p53/MDM2/ARF to promote p53 signaling and is frequently silenced in nasopharyngeal carcinoma. Clinical Cancer Research, 16(11), 2949–2958.
Fukutomi, S., Seki, N., Koda, K., & Miyazaki, M. (2007). Identification of methylation-silenced genes in colorectal cancer cell lines: Genomic screening using oligonucleotide arrays. Scandinavian Journal of Gastroenterology, 42(12), 1486–1494.
Mizukami, H., Shirahata, A., Goto, T., Sakata, M., Saito, M., et al. (2008). PGP9.5 methylation as a marker for metastatic colorectal cancer. Anticancer Research, 28(5A), 2697–2700.
Abdelmaksoud-Dammak, R., Saadallah-Kallel, A., Miladi-Abdennadher, I., Ayedi, L., Khabir, A., et al. (2016). CpG methylation of ubiquitin carboxyl-terminal hydrolase 1 (UCHL1) and P53 mutation pattern in sporadic colorectal cancer. Tumour Biology, 37(2), 1707–1714.
Hibi, K., Liu, Q., Beaudry, G. A., Madden, S. L., Westra, W. H., Wehage, S. L., et al. (1998). Serial analysis of gene expression in non-small cell lung cancer. Cancer Research, 58(24), 5690–5694.
Kusakabe, M., Kutomi, T., Watanabe, K., Emoto, N., Aki, N., et al. (2010). Identification of G0S2 as a gene frequently methylated in squamous lung cancer by combination of in silico and experimental approaches. Internation Journal of Cancer, 126(8), 1895–1902.
Trifa, F., Karray-Chouayekh, S., Jmaa, Z. B., Jmal, E., Khabir, A., et al. (2013). Frequent CpG methylation of ubiquitin carboxyl-terminal hydrolase 1 (UCHL1) in sporadic and hereditary Tunisian breast cancer patients: clinical significance. Medical Oncology, 30(1), 418.
Lien, H. C., Wang, C. C., Lin, C. H., Lu, Y. S., Huang, C. S., et al. (2013). Differential expression of ubiquitin carboxy-terminal hydrolase L1 in breast carcinoma and its biological significance. Human Pathology, 44(9), 1838–1848.
Lien, H. C., Wang, C. C., Huang, C. S., Yang, Y. W., Kuo, W. H., et al. (2013). Ubiquitin carboxy-terminal hydrolase L1 may be involved in the development of mammary phyllodes tumors. Virchows Archiv, 462(2), 155–161.
Jin, Y., Zhang, W., Xu, J., Wang, H., Zhang, Z., et al. (2015). UCH-L1 involved in regulating the degradation of EGFR and promoting malignant properties in drug-resistant breast cancer. International Journal of Clinicl and Experimental Pathology, 8(10), 12500–12508.
Mastoraki, A., Ioannidis, E., Patsouris, E., Safioleas, M., & Aroni, K. (2009). PGP 9.5 expression in cutaneous keratoacanthomas and squamous cell carcinomas. Archives of Dermatological Research, 301(9), 653–658.
Mastoraki, A., Ioannidis, E., Apostolaki, A., Patsouris, E., & Aroni, K. (2009). PGP 9.5 and cyclin D1 coexpression in cutaneous squamous cell carcinomas. International Journal of Surgical Pathology, 17(6), 413–420.
Takano, T., Miyauchi, A., Matsuzuka, F., Yoshida, H., Nakata, Y., Kuma, K., et al. (2004). PGP9.5 mRNA could contribute to the molecular-based diagnosis of medullary thyroid carcinoma. European Journal of Cancer, 40(4), 614–618.
Howell, V. M., Gill, A., Clarkson, A., Nelson, A. E., Dunne, R., Delbridge, L. W., et al. (2009). Accuracy of combined protein gene product 9.5 and parafibromin markers for immunohistochemical diagnosis of parathyroid carcinoma. Journal of Clinical Endocrinology and Metabolism, 94(2), 434–441.
Truran, P. P., Johnson, S. J., Bliss, R. D., Lennard, T. W., & Aspinall, S. R. (2014). Parafibromin, galectin-3, PGP9.5, Ki67, and cyclin D1: using an immunohistochemical panel to aid in the diagnosis of parathyroid cancer. World Journal of Surgery, 38(11), 2845–2854.
Wulfänger, J., Biehl, K., Tetzner, A., Wild, P., Ikenberg, K., Meyer, S., et al. (2013). Heterogeneous expression and functional relevance of the ubiquitin carboxyl-terminal hydrolase L1 in melanoma. International Journal of Cancer, 133(11), 2522–2532.
Kim, H. J., Magesh, V., Lee, J. J., Kim, S., Knaus, U. G., & Lee, K. J. (2015). Ubiquitin C-terminal hydrolase-L1 increases cancer cell invasion by modulating hydrogen peroxide generated via NADPH oxidase 4. Oncotarget, 6(18), 16287–16303.
Liu, X., Zeng, B., Ma, J., & Wan, C. (2009). Comparative proteomic analysis of osteosarcoma cell and human primary cultured osteoblastic cell. Cancer Investigation, 27(3), 345–352.
Zheng, S., Qiao, G., Min, D., Zhang, Z., Lin, F., Yang, Q., et al. (2015). Heterogeneous expression and biological function of ubiquitin carboxy-terminal hydrolase-L1 in osteosarcoma. Cancer Letters, 359(1), 36–46.
Hibi, K., Kodera, Y., Ito, K., Akiyama, S., Shirane, M., & Nakao, A. (2004). Plasminogen activator inhibitor-1 is a downstream mediator of the PGP9.5-related oncogenic pathway in esophageal squamous cell carcinoma. Anticancer Research, 24(6), 3731–3734.
Mizukami, H., Goto, T., Kitamura, Y., Sakata, M., Saito, M., Ishibashi, K., et al. (2009). PGP9.5 was less frequently methylated in advanced gastric carcinoma. Hepato-Gastroenterology, 56(94–95), 1576–1579.
Yang, H., Zhang, C., Fang, S., Ou, R., Li, W., & Xu, Y. (2015). UCH-LI acts as a novel prognostic biomarker in gastric cardiac adenocarcinoma. International Journal of Clinicl and Experimental Pathology, 8(11), 13957–13967.
Gu, Y. Y., Yang, M., Zhao, M., Luo, Q., Yang, L., Peng, H., et al. (2015). The de-ubiquitinase UCHL1 promotes gastric cancer metastasis via the Akt and Erk1/2 pathways. Tumour Biology, 36(11), 8379–8387.
Orr, K. S., Shi, Z., Brown, W. M., O'Hagan, K. A., Lappin, T. R., Maxwell, P., et al. (2011). Potential prognostic marker ubiquitin carboxyl-terminal hydrolase-L1 does not predict patient survival in non-small cell lung carcinoma. Journal of Experimental and Clinical Cancer Research, 30, 79.
Akishima-Fukasawa, Y., Ino, Y., Nakanishi, Y., Miura, A., Moriya, Y., Kondo, T., et al. (2010). Significance of PGP9.5 expression in cancer-associated fibroblasts for prognosis of colorectal carcinoma. American Journal of Clinical Pathology, 134(1), 71–79.
Ma, Y., Zhao, M., Zhong, J., Shi, L., Luo, Q., Liu, J., et al. (2010). Proteomic profiling of proteins associated with lymph node metastasis in colorectal cancer. Journal of Cellular Biochemistry, 110(6), 1512–1519.
Zhong, J., Zhao, M., Ma, Y., Luo, Q., Liu, J., Wang, J., et al. (2012). UCHL1 acts as a colorectal cancer oncogene via activation of the β-catenin/TCF pathway through its deubiquitinating activity. International Journal of Molecular Medicine, 30(2), 430–436.
Hussain, S., Foreman, O., Perkins, S. L., Witzig, T. E., Miles, R. R., van Deursen, J., et al. (2010). The de-ubiquitinase UCH-L1 is an oncogene that drives the development of lymphoma in vivo by deregulating PHLPP1 and Akt signaling. Leukemia, 24(9), 1641–1655.
Goto, Y., Zeng, L., Yeom, C. J., Zhu, Y., Morinibu, A., Shinomiya, K., et al. (2015). UCHL1 provides diagnostic and antimetastatic strategies due to its deubiquitinating effect on HIF-1α. Nature Communications, 6, 6153.
Bheda, A., Yue, W., Gullapalli, A., Whitehurst, C., Liu, R., Pagano, J. S., et al. (2009). Positive reciprocal regulation of ubiquitin C-terminal hydrolase L1 and β-catenin/TCF signaling. PLoS One, 4(6), e5955.
Caballero, O. L., Resto, V., Patturajan, M., Meerzaman, D., Guo, M. Z., Engles, J., et al. (2002). Interaction and colocalization of PGP9.5 with JAB1 and p27(Kip1). Oncogene, 21(19), 3003–3010.
Takami, Y., Nakagami, H., Morishita, R., Katsuya, T., Cui, T. X., Ichikawa, T., et al. (2007). Ubiquitin carboxyl-terminal hydrolase L1, a novel deubiquitinating enzyme in the vasculature, attenuates NF-kappaB activation. Arteriosclerosis Thrombosis and Vascular Biology, 27(10), 2184–2190.
Ichikawa, T., Li, J., Dong, X., Potts, J. D., Tang, D. Q., Li, D. S., et al. (2010). Ubiquitin carboxyl terminal hydrolase L1 negatively regulates TNFalpha-mediated vascular smooth muscle cell proliferation via suppressing ERK activation. Biochemical and Biophysical Research Communications, 391(1), 852–856.
Sosna, J., Voigt, S., Mathieu, S., Kabelitz, D., Trad, A., Janssen, O., et al. (2013). The proteases HtrA2/Omi and UCH-L1 regulate TNF-induced necroptosis. Cell Communication and Signaling, 11, 76.
Karim, R., Tummers, B., Meyers, C., Biryukov, J. L., Alam, S., Backendorf, C., et al. (2013). Human papillomavirus (HPV) upregulates the cellular deubiquitinase UCHL1 to suppress the keratinocyte's innate immune response. PLoS Pathogens, 9(5), e1003384.
Zhang, H., Mao, X., Sun, Y., Hu, R., Luo, W., Zhao, Z., et al. (2015). NF-κB upregulates ubiquitin C-terminal hydrolase 1 in diseased podocytes in glomerulonephritis. Molecular Medicine Reports, 12(2), 2893–2901.
Kwon, J. (2007). The new function of two ubiquitin C-terminal hydrolase isozymes as reciprocal modulators of germ cell apoptosis. Experimental Animals, 56(2), 71–77.
Mtango, N. R., Sutovsky, M., Vandevoort, C. A., Latham, K. E., & Sutovsky, P. (2012). Essential role of ubiquitin C-terminal hydrolases UCHL1 and UCHL3 in mammalian oocyte maturation. Journal of Cellular Physiology, 227(5), 2022–2029.
Yi, Y. J., Sutovsky, M., Song, W. H., & Sutovsky, P. (2015). Protein deubiquitination during oocyte maturation influences sperm function during fertilisation, antipolyspermy defense and embryo development. Reproduction Fertility and Development, 27(8), 1154–1167.
Hennings, J. M., Zimmer, R. L., Nabli, H., Davis, J. W., Sutovsky, P., Sutovsky, M., et al. (2016). Improved murine blastocyst quality and development in a single culture medium compared to sequential culture media. Reproductive Sciences, 23(3), 310–317.
Miyoshi, Y., Nakayama, S., Torikoshi, Y., Tanaka, S., Ishihara, H., Taguchi, T., et al. (2006). High expression of ubiquitin carboxy-terminal hydrolase-L1 and -L3 mRNA predicts early recurrence in patients with invasive breast cancer. Cancer Science, 97(6), 523–529.
Luo, K., Li, L., Li, Y., Wu, C., Yin, Y., Chen, Y., et al. (2016). A phosphorylation-deubiquitination cascade regulates the BRCA2-RAD51 axis in homologous recombination. Genes & Development, 30(23), 2581–2595.
Song, H. M., Lee, J. E., & Kim, J. H. (2014). Ubiquitin C-terminal hydrolase-L3 regulates EMT process and cancer metastasis in prostate cell lines. Biochemical and Biophysical Research Communications, 452(3), 722–727.
Chiba, T., & Tanaka, K. (2004). Cullin-based ubiquitin ligase and its control by NEDD8-conjugating system. Current Protein and Peptide Science, 5(3), 177–184.
Lam, Y. A., DeMartino, G. N., Pickart, C. M., & Cohen, R. E. (1997). Specificity of the ubiquitin isopeptidase in the PA700 regulatory complex of 26 S proteasomes. Journal of Biological Chemistry, 272(45), 28438–28446.
Fang, Y., Fu, D., & Shen, X. Z. (2010). The potential role of ubiquitin c-terminal hydrolases in oncogenesis. Biochimica et Biophysica Acta, 1806(1), 1–6.
Randles, L., Anchoori, R. K., Roden, R. B., & Walters, K. J. (2016). The proteasome ubiquitin receptor hRpn13 and its interacting deubiquitinating enzyme Uch37 are required for proper cell cycle progression. Journal of Biological Chemistry, 291(16), 8773–8783.
Yao, T., Song, L., Jin, J., Cai, Y., Takahashi, H., Swanson, M. P., et al. (2008). Distinct modes of regulation of the Uch37 deubiquitinating enzyme in the proteasome and in the Ino80 chromatinremodeling complex. Molecular Cell, 31(6), 909–917.
Zediak, V. P., & Berger, S. L. (2008). Hit and run: Transient deubiquitylase activity in a chromatin-remodeling complex. Molecular Cell, 31(6), 773–774.
Chen, X., & Walters, K. J. (2015). Structural plasticity allows UCH37 to be primed by RPN13 or locked down by INO80G. Molecular Cell, 57(5), 767–768.
Cai, Y., Jin, J., Yao, T., Gottschalk, A. J., Swanson, S. K., Wu, S., et al. (2007). YY1 functions with INO80 to activate transcription. Nature Structural & Molecular Biology, 14(8), 872–874.
Rolen, U., Kobzeva, V., Gasparjan, N., Ovaa, H., Winberg, G., Kisseljov, F., et al. (2006). Activity profiling of deubiquitinating enzymes in cervical carcinoma biopsies and cell lines. Molecular Carcinogenesis, 45(4), 260–269.
Fang, Y., Fu, D., Tang, W., Cai, Y., Ma, D., Wang, H., et al. (2013). Ubiquitin C-terminal hydrolase 37, a novel predictor for hepatocellular carcinoma recurrence, promotes cell migration and invasion via interacting and deubiquitinating PRP19. Biochimica et Biophysica Acta, 1833(3), 559–572.
Chen, Y., Fu, D., Xi, J., Ji, Z., Liu, T., Ma, Y., et al. (2012). Expression and clinical significance of UCH37 in human esophageal squamous cell carcinoma. Digestive Diseases and Science, 57(9), 2310–2317.
Wang, L., Chen, Y. J., Xu, K., Wang, Y. Y., Shen, X. Z., & Tu, R. Q. (2014). High expression of UCH37 is significantly associated with poor prognosis in human epithelial ovarian cancer. Tumour Biology, 35(11), 11427–11433.
Chen, Z., Niu, X., Li, Z., Yu, Y., Ye, X., Lu, S., et al. (2011). Effect of ubiquitin carboxy-terminal hydrolase 37 on apoptotic in A549 cells. Cell Biochemistry and Function, 29(2), 142–148.
Cutts, A. J., Soond, S. M., Powell, S., & Chantry, A. (2011). Early phase TGFβ receptor signalling dynamics stabilised by the deubiquitinase UCH37 promotes cell migratory responses. International Journal of Biochemistry & Cell Biology, 43(4), 604–612.
Wicks, S. J., Haros, K., Maillard, M., Song, L., Cohen, R. E., Dijke, P. T., et al. (2005). The deubiquitinating enzyme UCH37 interacts with Smads and regulates TGF-β signaling. Oncogene, 24(54), 8080–8084.
Wicks, S. J., Grocott, T., Haros, K., Maillard, M., ten Dijke, P., & Chantry, A. (2006). Reversible ubiquitination regulates the Smad/TGF-β signalling pathway. Biochemical Society Transactions, 34(Pt 5), 761–763.
Fang, Y., Mu, J., Ma, Y., Ma, D., Fu, D., & Shen, X. (2012). The interaction between ubiquitin C-terminal hydrolase 37 and glucose-regulated protein 78 in hepatocellular carcinoma. Molecular and Cellular Biochemistry, 359(1–2), 59–66.
Mahanic, C. S., Budhavarapu, V., Graves, J. D., Li, G., & Lin, W. C. (2015). Regulation of E2 promoter binding factor 1 (E2F1) transcriptional activity through a deubiquitinating enzyme, UCH37. Journal of Biological Chemistry, 290(44), 26508–26522.
Han, W., Lee, H., & Han, J. K. (2017). Ubiquitin C-terminal hydrolase37 regulates Tcf7 DNA binding for the activation of Wnt signalling. Scientific Reports, 7, 42590.
Chen, Y. J., Ma, Y. S., Fang, Y., Wang, Y., Fu, D., & Shen, X. Z. (2013). Power and promise of ubiquitin carboxyl-terminal hydrolase 37 as a target of cancer therapy. Asian Pacific Journal of Cancer Prevention, 14(4), 2173–2179.
Harbour, J. W., Onken, M. D., Roberson, E. D., Duan, S., Cao, L., Worley, L. A., et al. (2010). Frequent mutation of BAP1 in metastasizing uveal melanomas. Science, 330(6009), 1410–1413.
Testa, J. R., Cheung, M., Pei, J., Below, J. E., Tan, Y., Sementino, E., et al. (2011). Germline BAP1 mutations predispose to malignant mesothelioma. Nature Genetics, 43(10), 1022–1025.
Joseph, N. M., Chen, Y. Y., Nasr, A., Yeh, I., Talevich, E., Onodera, C., et al. (2017). Genomic profiling of malignant peritoneal mesothelioma reveals recurrent alterations in epigenetic regulatory genes BAP1, SETD2, and DDX3X. Modern Pathology, 30(2), 246–254.
Leblay, N., Leprêtre, F., Le Stang, N., Gautier-Stein, A., Villeneuve, L., Isaac, S., et al. (2017). BAP1 is altered by copy number loss, mutation, and/or loss of protein expression in more than 70% of malignant peritoneal mesotheliomas. Journal of Thoracic Oncology, 12(4), 724–733.
Wu, D., Hiroshima, K., Yusa, T., Ozaki, D., Koh, E., Sekine, Y., et al. (2017). Usefulness of p16/CDKN2A fluorescence in situ hybridization and BAP1 immunohistochemistry for the diagnosis of biphasic mesothelioma. Annals of Diagnostic Pathology, 26, 31–37.
McCroskey, Z., Staerkel, G., & Roy-Chowdhuri, S. (2017). Utility of BRCA1-associated protein 1 immunoperoxidase stain to differentiate benign versus malignant mesothelial proliferations in cytologic specimens. Diagnostic Cytopathology, 45(4), 312–319.
Hida, T., Hamasaki, M., Matsumoto, S., Sato, A., Tsujimura, T., Kawahara, K., et al. (2017). Immunohistochemical detection of MTAP and BAP1 protein loss for mesothelioma diagnosis: Comparison with 9p21 FISH and BAP1 immunohistochemistry. Lung Cancer, 104, 98–105.
Shah, A. A., Bourne, T. D., & Murali, R. (2013). BAP1 protein loss by immunohistochemistry: a potentially useful tool for prognostic prediction in patients with uveal melanoma. Pathology, 45(7), 651–656.
Kalirai, H., Dodson, A., Faqir, S., Damato, B. E., & Coupland, S. E. (2014). Lack of BAP1 protein expression in uveal melanoma is associated with increased metastatic risk and has utility in routine prognostic testing. British Journal of Cancer, 111(7), 1373–1380.
Koopmans, A. E., Verdijk, R. M., Brouwer, R. W., van den Bosch, T. P., van den Berg, M. M., Vaarwater, J., et al. (2014). Clinical significance of immunohistochemistry for detection of BAP1 mutations in uveal melanoma. Modern Pathology, 27(10), 1321–1330.
van Essen, T. H., van Pelt, S. I., Versluis, M., Bronkhorst, I. H., van Duinen, S. G., Marinkovic, M., et al. (2014). Prognostic parameters in uveal melanoma and their association with BAP1 expression. British Journal of Ophthalmology, 98(12), 1738–1743.
Kumar, R., Taylor, M., Miao, B., Ji, Z., Njauw, J. C., J€onsson, G., et al. (2015). BAP1 has a survival role in cutaneous melanoma. Journal of Investigative Dermatology, 135(4), 1089–1097.
Jiao, Y., Pawlik, T. M., Anders, R. A., Selaru, F. M., Streppel, M. M., Lucas, D. J., et al. (2013). Exome sequencing identifies frequent inactivating mutations in BAP1, ARID1A and PBRM1 in intrahepatic cholangiocarcinomas. Nature Genetics, 45(12), 1470–1473.
Simbolo, M., Fassan, M., Ruzzenente, A., Mafficini, A., Wood, L. D., Corbo, V., et al. (2014). Multigene mutational profiling of cholangiocarcinomas identifies actionable molecular subgroups. Oncotarget, 5(9), 2839–2852.
Churi, C. R., Shroff, R., Wang, Y., Rashid, A., Kang, H. C., Weatherly, J., et al. (2014). Mutation profiling in cholangiocarcinoma: prognostic and therapeutic implications. PLoS One, 9(12), e115383.
Simbolo, M., Fassan, M., Mafficini, A., Lawlor, R. T., Ruzzenente, A., & Scarpa, A. (2016). New genomic landscapes and therapeutic targets for biliary tract cancers. Frontiers in Bioscience, 21, 707–718.
Fan, L. H., Tang, L. N., Yue, L., Yang, Y., Gao, Z. L., & Shen, Z. (2012). BAP1 is a good prognostic factor in advanced non-small cell lung cancer. Clinical and Investigative Medicine, 35(4), E182–E189.
Andrici, J., Parkhill, T. R., Jung, J., Wardell, K. L., Verdonk, B., Singh, A., et al. (2016). Loss of expression of BAP1 is very rare in non-small cell lung carcinoma. Pathology, 48(4), 336–340.
Gossage, L., Murtaza, M., Slatter, A. F., Lichtenstein, C. P., Warren, A., Haynes, B., et al. (2014). Clinical and pathological impact of VHL, PBRM1, BAP1, SETD2, KDM6A, and JARID1c in clear cell renal cell carcinoma. Genes, Chromosomes & Cancer, 53(1), 38–51.
Piva, F., Santoni, M., Matrana, M. R., Satti, S., Giulietti, M., Occhipinti, G., et al. (2015). BAP1, PBRM1 and SETD2 in clear-cell renal cell carcinoma: molecular diagnostics and possible targets for personalized therapies. Expert Review of Molecular Diagnostics, 15(9), 1201–1210.
Jensen, D. E., & Rauscher, F. J. (1999). Defining biochemical functions for the BRCA1 tumor suppressor protein: analysis of the BRCA1 binding protein BAP1. Cancer Letters, 143(Suppl. 1), S13–S17.
Coupier, I., Cousin, P. Y., Hughes, D., Legoix-Ne, P., Trehin, A., Sinilnikova, O. M., et al. (2005). BAP1 and breast cancer risk. Familial Cancer, 4(4), 273–277.
Tang, J., Xi, S., Wang, G., Wang, B., Yan, S., Wu, Y., et al. (2013). Prognostic significance of BRCA1-associated protein 1 in colorectal cancer. Medical Oncology, 30(2), 541.
Luchini, C., Veronese, N., Yachida, S., Cheng, L., Nottegar, A., Stubbs, B., et al. (2016). Different prognostic roles of tumor suppressor gene BAP1 in cancer: a systematic review with meta-analysis. Genes Chromosomes & Cancer, 55(10), 741–749.
Farzin, M., Toon, C. W., Clarkson, A., Sioson, L., Watson, N., Andrici, J., et al. (2015). Loss of expression of BAP1 predicts longer survival in mesothelioma. Pathology, 47(4), 302–307.
Baumann, F., Flores, E., Napolitano, A., Kanodia, S., Taioli, E., Pass, H., et al. (2015). Mesothelioma patients with germline BAP1 mutations have 7-fold improved long-term survival. Carcinogenesis, 36(1), 76–81.
Xu, J., Kadariya, Y., Cheung, M., Pei, J., Talarchek, J., Sementino, E., et al. (2014). Germline mutation of Bap1 accelerates development of asbestos-induced malignant mesothelioma. Cancer Research, 74(16), 4388–4397.
Hwang, H. C., Pyott, S., Rodriguez, S., Cindric, A., Carr, A., Michelsen, C., et al. (2016). BAP1 immunohistochemistry and p16 FISH in the diagnosis of sarcomatous and desmoplastic mesotheliomas. American Journal of Surgical Pathology, 40(5), 714–718.
Cigognetti, M., Lonardi, S., Fisogni, S., Balzarini, P., Pellegrini, V., Tironi, A., et al. (2015). BAP1 (BRCA1-associated protein 1) is a highly specific marker for differentiating mesothelioma from reactive mesothelial proliferations. Modern Pathology, 28(8), 1043–1057.
O'Shea, S.J., Robles-Espinoza, C.D., McLellan, L., Harrigan, J., Jacq, X., Hewinson, J., et al. (2017). A population-based analysis of germline BAP1 mutations in melanoma. Human molecular genetics. Pii: ddw403. https://doi.org/10.1093/hmg/ddw403.
Greenberg, R. A., Sobhian, B., Pathania, S., Cantor, S. B., Nakatani, Y., & Livingston, D. M. (2006). Multifactorial contributions to an acute DNA damage response by BRCA1/BARD1-containing complexes. Genes & Development, 20(1), 34–46.
Nishikawa, H., Wu, W., Koike, A., Kojima, R., Gomi, H., Fkuda, M., et al. (2009). BRCA1-associated protein 1 interferes with BRCA1/BARD1 RING heterodimer activity. Cancer Research, 69(1), 111–119.
Forma, E., Jozwiak, P., Brys, M., & Krzeslak, A. (2014). The potential role of O-GlcNAc modification in cancer epigenetics. Cell & Molecular Biology Letters, 19(3), 438–460.
Wang, A., Papneja, A., Hyrcza, M., Al-Habeeb, A., & Ghazarian, D. (2016). Gene of the month: BAP1. Journal of Clinical Pathology, 69(9), 750–753.
Daou, S., Hammond-Martel, I., Mashtalir, N., Barbour, H., Gagnon, J., Jannantuono, N. V., et al. (2015). The BAP1/ASXL2 histone H2A deubiquitinase complex regulates cell proliferation and is disrupted in cancer. Journal of Biological Chemistry, 290(48), 28643–28663.
Shang, D., Han, T., Xu, X., & Liu, Y. (2015). Decitabine induces G2/M cell cycle arrest by suppressing p38/NF-κB signaling in human renal clear cell carcinoma. International Journal of Clinical Experimental Pathology, 8(9), 11140–11148.
Brinkmann, K., Zigrino, P., Witt, A., Schell, M., Ackermann, L., Broxtermann, P., et al. (2013). Ubiquitin C-terminal hydrolase-L1 potentiates cancer chemosensitivity by stabilizing NOXA. Cell Reports, 3(3), 881–891.
Pena-Llopis, S., Vega-Rubin-de-Celis, S., Liao, A., Leng, N., Pavía-Jiménez, A., Wang, S., et al. (2012). BAP1 loss defines a new class of renal cell carcinoma. Nature Genetics, 44(7), 751–759.
Acknowledgements
This study was financially supported by grants from the National Nature Science Foundation of China (Nos. 81301820, 81472673, 81672720, 81672334), the Fund of Shanghai Science and Technology Commission (16ZR1406100) and the National Clinical Key Special Subject of China.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no financial conflicts of interest.
Rights and permissions
About this article
Cite this article
Fang, Y., Shen, X. Ubiquitin carboxyl-terminal hydrolases: involvement in cancer progression and clinical implications. Cancer Metastasis Rev 36, 669–682 (2017). https://doi.org/10.1007/s10555-017-9702-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-017-9702-0