Abstract
Visual or manual characterization and classification of atherosclerotic plaque lesions are tedious, error-prone, and time-consuming. The purpose of this study is to develop and design an automated carotid plaque characterization and classification system into binary classes, namely symptomatic and asymptomatic types via the deep learning (DL) framework implemented on a supercomputer. We hypothesize that on ultrasound images, symptomatic carotid plaques have (a) a low grayscale median because of a histologically large lipid core and relatively little collagen and calcium, and (b) a higher chaotic (heterogeneous) grayscale distribution due to the composition. The methodology consisted of building a DL model of Artificial Intelligence (called Atheromatic 2.0, AtheroPoint, CA, USA) that used a classic convolution neural network consisting of 13 layers and implemented on a supercomputer. The DL model used a cross-validation protocol for estimating the classification accuracy (ACC) and area-under-the-curve (AUC). A sample of 346 carotid ultrasound-based delineated plaques were used (196 symptomatic and 150 asymptomatic, mean age 69.9 ± 7.8 years, with 39% females). This was augmented using geometric transformation yielding 2312 plaques (1191 symptomatic and 1120 asymptomatic plaques). K10 (90% training and 10% testing) cross-validation DL protocol was implemented and showed an (i) accuracy and (ii) AUC without and with augmentation of 86.17%, 0.86 (p-value < 0.0001), and 89.7%, 0.91 (p-value < 0.0001), respectively. The DL characterization system consisted of validation of the two hypotheses: (a) mean feature strength (MFS) and (b) Mandelbrot's fractal dimension (FD) for measuring chaotic behavior. We demonstrated that both MFS and FD were higher in symptomatic plaques compared to asymptomatic plaques by 64.15 ± 0.73% (p-value < 0.0001) and 6 ± 0.13% (p-value < 0.0001), respectively. The benchmarking results show that DL with augmentation (ACC: 89.7%, AUC: 0.91 (p-value < 0.0001)) is superior to previously published machine learning (ACC: 83.7%) by 6.0%. The Atheromatic runs the test patient in < 2 s. Deep learning can be a useful tool for carotid ultrasound-based characterization and classification of symptomatic and asymptomatic plaques.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
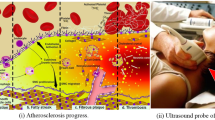
There are 17.9 million cardiovascular diseases (CVD) deaths globally and 647,000 in the USA per year [1, 2], which means a loss of one life every 37 s [3]. CVD's fundamental cause is atherosclerosis development with plaque formation in the vasculature, such as the coronary and carotid arteries [4]. Plaque rupture or plaque ulceration often results in the formation of a thrombus, which may embolize or occlude the lumen obstructing the blood flow causing myocardial infarction or stroke [5]. This study is focused on the characterization and classification of only carotid artery atherosclerotic plaques, a study classified under the topic of computer-aided diagnosis [6, 7]. This study will not dwell on the literature dealing with coronary artery plaques.
Several medical imaging modalities are used to visualize and screen the plaque, with the most common being magnetic resonance imaging (MRI) [8], computed tomography (CT) [9], and ultrasound (US) [10, 11]. Over the past decade, ultrasound has become an established norm as a first-line diagnostic modality in symptomatic patients and a powerful screening tool in asymptomatic individuals [12]. It is a safe, low-cost test [13], easy to use, has a small footprint, and is radiation-free [14]. Also, carotid ultrasound imaging at a resolution close to 0.2 mm provides the ability to study the texture of plaques and determine whether they are stable or unstable [15,16,17].
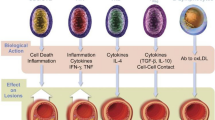
As seen on ultrasound, carotid plaque texture is variable and challenging to classify using the naked eye due to high inter-observer variability [18]. Symptomatic plaques tend to produce more significant stenosis, be more hypoechoic, have a large juxtaluminal black area close to the lumen without a visible echogenic cap, and discrete white areas hyperechoic areas compared with asymptomatic plaques [19, 20]. These findings have been identified in cross-sectional studies of symptomatic and asymptomatic patients, in a large prospective study of asymptomatic patients and subsequently verified by comparing histology with in vivo ultrasound imaging [21].
Because the pixel data is large and fuzzy, derived from the spatial ultrasound images, and machines have a better learning ability to handle linear and nonlinear variations in plaque distribution, the recent trend in artificial intelligence (AI) has been used to characterize and classify [22] plaques using machine learning (ML). This ML solution requires computing the grayscale features manually [23,24,25,26,27,28], which are then trained using a training classifier to generate offline signatures and patterns. These are then used to transform the test pattern to predict its class risk [29, 30]. Such ML-based solutions are ad-hoc, slow, and not generalized [30], besides lacking reliability and stability.
Deep learning (DL) technology has dominated all walks of life, particularly in radiological imaging [31,32,33]. This technology provides an alternative to the ML strategies, especially: (i) the ability to generate a down-sampled representation of the original pattern automatically (so-called feature maps), and (ii) dynamically adjust the variations in the grayscale contrast via the neural network layers of the DL architecture [34]. Lekadir et al. [35] developed a CNN model for the classification of the plaque components by extracting 90,000 patches from the 50 in-vivo ultrasound image and achieved a 0.90 correlation coefficient. The purpose of this study is to develop and design an automated carotid plaque characterization and classification system into binary classes, namely symptomatic and asymptomatic types via the deep learning (DL) framework implemented on a supercomputer.
We hypothesize that symptomatic plaque has tissue characteristics such as (a) hypoechoic regions, having a low grayscale median (GSM) as a result of a large lipid core, low calcium or intraplaque haemorrhage (IPH), and (b) more chaotic (heterogeneous) representation in the ultrasound scans [16, 17, 28] because of the frequent presence of neovascularization alternating with areas of collagen or lipid. This is contrary to asymptomatic plaques, which are often hyperechoic with higher GSM, because they have higher and diffuse collagen content, often with calcification and a small lipid core. We designed a novel carotid plaque tissue characterization and classification system using DL components of artificial intelligence (AI) based on such a hypothesis. The design overcomes the ML weaknesses such as (i) manual feature extraction and (ii) classification. The classification system's accuracy is determined using a K10 (90% training and 10% testing) cross-validation protocol. The characterization of the plaque is accomplished by (a) computing the mean feature strength (MFS) at different DL layers [36] and (b) fractal dimension exhibiting quantification of the randomness in these plaque images [37]. Subsequently, we benchmark the DL system against the previously developed ML system (on the same cohort), and finally, the system speed is optimized using the supercomputer framework. A work of similar.
This study's layout is as follows. Section 2 presents the patient demographics, data acquisition, and pre-processing. Section 3 depicts the AI architecture. Section 4 presents experimental protocol, AI parameters, and performance metrics. Section 5 shows the DL classification results. Section 6 presents the plaque characterization in a deep learning framework. Section 7 presents the discussion. The study concludes in Sect. 8.
Patient demographics, data acquisition, and pre-processing
The main components of this section primarily consist of (i) patient demographics and exclusion criteria, (ii) ultrasound data acquisition, and (iii) plaque delineation and data augmentation.
Patient demographics and exclusion criteria
We included 346 referred consecutive patients (mean age 69.9 ± 7.8 and 39% female); all these carotid duplex ultrasound scans were noted to have an internal carotid artery (ICA) stenosis of 50% to 99% (approval was obtained from the Institutional Ethics Committee, St. Mary's Hospital, Imperial College, London, UK) by experts. Out of these 346, 150 were asymptomatic with no neurological abnormalities. The remaining 196 had ipsilateral cerebral hemispheric symptoms (amaurosis fugax (AF), transient ischemic attacks, or stroke with good recovery) related to carotid artery atherosclerosis. Overall, there were 196 distributions, including 88 strokes, 70 TIA, and 38 AF. A neurologist noted the history of patients and their physical examination [20]. Note that this patient data were used in our previous machine learning work [23, 38].
Exclusion criteria
As per European Carotid Surgery Trialists (ECST) Collaborative Group 1991, it was recommended that surgery was harmful in patients with mild stenosis (0%–29%). Later in 1996, ECST-Collaborative Group 1996 recommended no benefit from surgery in patients with either 30%–49% stenosis or 50%–69% stenosis. In 1998, ECST recommended the use of Carotid Endarterectomy (CEA) for most patients with 80% stenosis. Thus, subjects excluded from the study were those with cardioembolic symptoms or distant symptoms (6 months). As per ECST, patients with 70%–99% stenosis showed a reduction in overall stroke risk. It was reported in [23, 38] that subjects having 70% to 99% stenosis and 50% to 69% stenosis highly benefited from CEA. The plaques that were less than 50% stenosis were eliminated from the study. This was because they were rarely associated with stroke, and therefore the inclusion of such plaque would have caused noise and bias. In this study, the process of characterization of the plaque was not blinded, mainly because the sonographers performed routine testing for grading of plaques and stenosis. However, the classification process was blinded to the person who processed the images since they did not know which plaques were symptomatic and asymptomatic.
Ultrasound data acquisition
The carotid scanning machine consisted of the following make and model: HDI 3000; Advanced Technology Laboratories, Seattle, WA, the USA having linear broadband width of 4–7 MHz (multi-frequency) transducer. Its resolution was 20 pixels per mm. The scanning was conducted at Irvine Laboratory for Cardiovascular Investigation and Research, Saint Mary’s Hospital, UK. With 40 years of experience in carotid ultrasound and a vascular surgery area, Professor Andrew Nicolaides carried out all the ultrasound image analysis and is discussed in our previous study [23]. It consisted of the following steps: (i) adjustment of the dynamic range of the US machine; (ii) the averaging of the frames; (iii) application of the time gain compensation (TGC), where the objective was to ensure that adventitia wall region of the anterior and posterior walls had similar brightness; (iv) the transfer function was kept linear while keeping the beam of the probe perpendicular to the lumen wall; (v) the depth (D) was adjusted to ensure that it had the largest plaque region in the image; (vi) Finally, the probe was calibrated in such a way that plaque close to adventitia was hyperechoic (bright region), which was ultimately used during the normalization.
Plaque delineation and data augmentation
Plaque delineation
The plaque delineation protocol’s objective was to manually trace the region-of-interest, so-called plaque, in the carotid artery's anterior and posterior walls. For tracing, we used “Plaque Texture Analysis software (PTAS)” (Iconsoft International Ltd, Greenford, London, UK) as adopted in previous studies [23, 38]. It offers two benefits: (i) one can normalize the images ensuring the median gray-level intensity of blood is in the range of 0–5 (dark intensities), and that of the adventitia layer was in the range of 180–190 (bright intensities) and (ii) the PTAS delineation system was user-friendly. This means post-normalization, the plaque could be outlined by the medical practitioner using a mouse, thereby saving it as a new file. Note that there were acoustic shadows for the calcified plaques, and they were not included in the delineation process. This way, the region of calcification and the plaque’s non-calcification components outside the acoustic shadow could be selected. Figure 1 shows exemplary cut sections of the manually delineated plaque.
Ultrasound plaque data augmentation
Because a deep learning system requires a relatively more extensive database (ranging from thousands to tens of thousands), we use a standardized protocol such as "augmentation". In this process, we use the random geometric transformation of the delineated plaque, such as (i) flipping, (ii) skewing, and (iii) rotating. The original size of the cohort is 346 contains 196 symptomatic and 150 asymptomatic plaques. Using the above augmentation procedure, the new cohort size of 2311 consists of 1191 symptomatic plaques and 1120 asymptomatic plaques. A total of 196 symptomatic plaques were converted to 1191 (804 were newly unique augmented to 1000, and then original 196 were added, yielding to 1,196. Five images were skewed and unacceptable; hence, they were removed from the cohort, leading to 1,191). A similar approach for the augmentation was adopted for the asymptomatic plaques. Thus, 150 plaques underwent augmentation to reach 1000, followed by an addition of 150 original plaques, leading to 1150. Since 30 skewed plaques were unacceptable, the finally tally consisted of removing these, accounting for 1120 asymptomatic plaques.
Artificial intelligence architectures
This section presents the two main pipeline architectures: Deep learning and Machine learning. Section 3.1 presents the DL architecture and Sect. 3.2 depicts ML architecture. The super computer specifications are presented in Sect. 3.3.
Characterization and classification stem out of the previously developed characterization system using machine learning (ML), applied to several applications based on signal and image processing such as arrhythmia [39], liver [40,41,42,43] breast tissue characterization [44] design of Thyroscan™ for thyroid tissue characterization [45,46,47,48,49], coronary plaque characterization [50], prostate tissue characterization using UroScan [51, 52], ovarian tissue characterization using GyneScan [53, 54], diabetes [55], skin cancer [56], left ventricle characterization [57], small vessel disease [58], and recently to carotid artery disease risk stratification using Atheromatic™ 1.0 [22, 27, 59, 60]. The above application was all ML-based, and the features in these methods were hand-crafted, and therefore required painful methods for feature extraction, feature selection, and optimization of the classification frameworks. Our proposed system's main novelty is designing the optimal DL architecture for tissue characterization and plaque classification. When combined, the complexity of the dataset and the hyperparameters play a vital role in selecting a number of convolution neural network (CNN) layers and the type of the layers [61] in a DL architecture.
Deep learning architecture
Our group has developed DL architectures before, which has taken up to 22 layers, which are typically meant to accept large sample sizes (considered in thousands) and to have bigger sized (W×H) images [40]. Since our cohort, size and image size are both moderate (i.e. 346 images and size varying from 55 × 43 pixel2 to 593 × 107 pixel2 without augmentation or cohort size of 2311, with augmentation). We chose a 13 layered CNN architecture having five convolution layers (CL), five average pooling layers (APL), two dense layers (DenL-1, DenL-2) and, one dropout layer as shown Fig. 2. We fine-tuned the hyper-parameters by changing the number of layers, type of layers, dropout rate, momentum rate, and learning rate. The last layer consists of the softmax layer that computes the categorical cross-entropy loss function (E) between the two classes symptomatic and asymptomatic and is mathematical as given in Eq. 1 as:
where p is the predicted probability of the plaque belonging to a particular class estimated using DL and y is the binary indicator for observed class, and "*" represents the product. The number of output features from the convolution process [36] given in Eq. 2, and the number of output features from the average pooling feature maps (APFM) given in Eq. 3.
where nin and nout are the numbers of input and output features, respectively, representing each CL. M is the convolution kernel size, P is the convolution padding size, S is the stride size (expressing the kernel movement), aout is the number of output features, w is the input feature map's width, and f is the kernel size. Table 1 shows the three columns depicting the name of the layers, the size of the feature maps, and the parameters used during training (per epoch). The DL system requires that the input size be the same. Therefore we converted the cut sections images to same-sized images by padding zeros but ensured that they do not get used during the DL process. Further, to consider the cut sections' grayscale pixels, we use a "mask image" of the same size as the cut section. This masked image is used for ensuring that the DL uses only the grayscale pixels of the cut sections.
Machine learning architecture
We benchmark the DL model against the popular ML models by comparing the attributes such as texture features, area-under-the-curve, and accuracy. Figure 3 shows the global architecture of the ML model. ML models' efficiency depends on the feature extracted and selected. We extracted histogram, Haralick, and Hu-moment features from the ultrasound scans, then we fed these features to linear discriminant analysis (LDA), k- nearest neighbors (k-NN) with K10 cross-validation protocol [62, 63].
Supercomputer specifications for DL architecture
We implemented our DL model using Tesla's NVIDIA-DGX v100-1. It contains 8 NVIDIA Tesla V100 graphical processing unit (GPU) accelerators connected through NVIDIA NVLink. All the GPUs are connected to form a cube-mesh network. This configuration is efficient on GPU load sharing. This is considered a state-of-the-art system in DL, and it provides unmatched performance for training. High-performance NVLink GPU interconnect improves the scalability of the DL-based training [64].
Statistical methods
ROC analysis
The final predicted labels are then compared against gold standards for performance evaluation using the receiving operating characteristics (ROC) and area-under-the-curve (AUC) using MEDCALC 17.0. Then p-values will be computed using T-test to highlight the significance of the predicted results.
Power analysis
We follow the standardized protocol for estimating the total samples needed for a certain threshold of the margin of error. The standardized protocol consisted of choosing the right parameters while applying the “power analysis.” Adapting the margin of error (MoE) to be 5%, the confidence interval (CI) to be 95%, the resultant sample size (n) was computed using the Eq. 4. Where z∗ represents the z-score value (1.64) from the table of probabilities of the standard normal distribution for the desired CI, and p̂ represents the data proportion (0.5). Plugging in the values, we obtain the number of samples 268 (as a baseline). Since the total number of samples in the input cohort consisted of 346 ultrasound scans, we were 29.1% higher than the baseline requirements.
Mean statistical performance
If ηDLK10(c) represents the accuracy of the DL method using K10 protocol for the combination c, η̅DLK10 represents the mean of the C combinations, and σK10DL represents the corresponding standard deviation, then these can be mathematically expressed as:
Following the similar notation for the ML-based strategy, we can compute the mean and standard deviation as follows:
The ML mean and standard deviation computation was conducted for two different kinds of validation data sets.
Experimental protocol, AI parameters, and performance metrics
Cross-validation protocol
Our experimental protocol consists of determining classification accuracy using a cross-validation (CV) paradigm that uses the K10 protocol (90% training and 10% testing). We run the CV protocol on both the datasets (default 346 plaques and augmented 2311 plaques).
Deep learning parameters
The following parameters are considered for training and testing the DL system: total epochs: 10,000, learning rate: 0.001, batch size: 32, regularization (L2): 0.001, and drop out: 0.5, optimizer: Adam.
Our DL lab experience [65] shows that during training, the total iterations for one combination is nearly 10,000–20,000. Since the image size was moderate, we empirically took a stable value of 10,000 epochs. Typically, the learning rate and regularization values are the same as adopted in the industry, which is 0.001 and 0.001, respectively. The batch size is 32 for training since the total data size was 311 (90% of 346) or 2080 (90% of 2311) for ten combinations of the K10 protocol.
Performance metrics
If ηDLK10(c) represents the accuracy of the DL method using K10 protocol for the combination c, η̅DLK10 represents the mean of the C combinations, and σK10DL represents the corresponding standard deviation, then these can be mathematically expressed as Eqs. 9, 10.
Following the similar notation for the ML-based strategy, we can compute the mean and standard deviation as follows Eqs. 11, 12.
The ML mean and standard deviation computation was conducted for two different kinds of validation data sets.
Results
There are two sets of experimental results. First is our DL system's performance evaluation (PE) using cross-validation protocol, and second, plaque characterization based on the two hypotheses. Section 5.1 presents the PE, while Sect. 5.2 discusses the characterization. Lastly, we validated our DL architecture by adapting the most widely used facial biometric data set.
Deep learning data analysis and benchmarking against machine learning
DL classification accuracy with and without augmentation
We implemented the DL architecture using 13-layered CNN consisting of the last layer as the softmax layer, as shown earlier in Fig. 2. The output layer gives us the probability that the predicted risk belongs to either symptomatic or asymptomatic classes, categorically (binary) estimated. The K10 protocol shows the best accuracy and AUC (computed using MEDCALC 17.0) of 86.17% and 0.86 (p-value < 0.0001) without augmentation, while 89.7% and 0.91 (p-value < 0.0001) with augmentation, respectively. To better understand the model performance, we compute every combination's real-time accuracy at the end of the 500th step value.
Benchmarking of deep learning against machine learning
Benchmarking protocol consists of a comparison of a DL system against the ML system on the same cohort. Further, we compare our novel DL system against the previous ML system published using the same plaque data [20, 35]. Using the K10 cross-validation protocol for both AI methods, the results can be seen in Fig. 4. While the accuracy of the ML system was 84.05%, the DL accuracy was.
a Bar charts showing the accuracy comparison between (i) machine learning (light gray), (ii) Deep learning w/o augmentation (DL w/o Aug) (dark gray color) and (iii) deep learning w/ augmentation (DL w/ Aug) (black color). b ROC curves showing AUC comparison: ML (0.83, p-value < 0.0001) vs. DL w/o Aug (0.86, p-value < 0.0001) systems showing an improvement of 3.61%. c ROC curves showing AUC comparison: ML (AUC: 0.83, p-value < 0.0001) vs. DL w/ Aug (AUC: 0.91, p-value < 0.0001) showing an improvement of 8%.
86.17% (w/o augmentation), 89.7% (w/ augmentation) (as shown in Fig. 4 (a)). The corresponding AUC for ML and DL systems was 0.83 (p-value < 0.0001) and 0.86 (p-value < 0.0001) (w/o augmentation), 0.91 (p-value < 0.0001) (w/ augmentation) respectively. Note the important point is that the methodology for design and implementation for ML requires exact steps, unlike DL systems. The offline training system was optimized, each time the features were manually computed. We will discuss the key differences in the discussion section further.
Validation of the DL and ML systems
Because the gold standard of the plaque images is based on the clinician's experience and lightning conditions under which the images are manually characterized, the DL systems are always vulnerable to slight variations inaccuracies. Therefore to further test the DL architecture, we used the most-widely "biometric facial data" [42] with a robust categorical gold standard. This database consisted of 1440 images with 72 classes, having 20 samples per class. Since the DL model consisted of two classes in the output layer, we changed the DL model output layer to 72 nodes for BFD experimental only. By applying the K10 protocol, the system's accuracy was 99.84% with AUC 0.99 (p-value < 0.0001). This was benchmarked using the ML system, yielding an accuracy of 97.9% and an AUC of 0.95 (p-value < 0.0001), almost comparable to the DL method. We validated the proposed model performance with a diagnostics odds ratio (DOR); it was observed that the DOR of DL was higher than ML.
Plaque characterization in a deep learning framework
Characterization of plaque requires to establish the (a) intensity distribution and (b) roughness (or chaotic behaviour) of the plaque area. We hypothesize that images of symptomatic and vulnerable plaques are (a) more hypoechoic (darker) and (b) have a more patchy (chaotic) representation of grayscale compared with asymptomatic plaques [66, 67]. Given this hypothesis, we must observe and justify these two features as part of the characterization process. Further, we must also determine if these can differentiate between symptomatic and asymptomatic plaques. Our computation shows GSM for symptomatic (25.67 ± 26.27) is higher than GSM for asymptomatic (3.53 ± 10.38) by ~ 86% (CI: 24.33 to 29.91, (p-value < 0.0001). It has a higher standard deviation and needs an automated method based on DL to characterize the plaque. This section is developed on establishing the two crucial components of the hypothesis.
Hypothesis 1: intensity distribution
The CNN has 13 layers, where the first eight layers are low-level features (CL-1, APL-1, CL-2, APL2 CL-3, APL-3, CL-4, and APL-4). Since asymptomatic plaque has higher collagen/high calcium (hyperechoic) content, the first eight layers must catch the textured plaque's bright surface. On the contrary, the low-level features (LLF) will not catch the high lipids/low calcium of symptomatic plaque. As we penetrate the deep layers of CNN (from the 9th layer to the 13th layer), CNN should have the ability to catch high lipid/low calcium (hypoechoic) features of the symptomatic plaque as part of high-level features (HLF).
The feature map's strength was used to characterize the intensity distribution, an accurate representation of the plaque type (symptomatic vs. asymptomatic). These strengths are nothing but the maps of the plaque features and must correspond as "feature maps." To represent this in a deep learning framework, we compute these feature maps for both data sets (symptomatic and asymptomatic). This is accomplished by running the classic CNN model for both symptomatic and asymptomatic deck and computing the strengths at all layers' output points.
Quantification of feature maps at different DL stages
We hypothesized the DL-2 layer would give the strength of the feature maps of the model. The strength of our model's feature maps in the K10 protocol is shown in Fig. 5a. The 2D FM is then converted to 1D FM for the rest of the two DL layers. Such vectors are computed for all the images for each class (symptomatic and asymptomatic). The strength of both classes is compared by quantifying the vector length corresponding to each of the classes. The separation of the classes can be seen, picked up by the DL system shown in Fig. 5b.
Justification of higher MFS of symptomatic plaques against MFS of asymptomatic plaques
Another way to justify as to why MFS (symptomatic plaques) > MFS (asymptomatic plaques), one can understand the intensity distribution of the lobes of the histograms of these plaques. Shown below in Fig. 6a, b are the symptomatic and asymptomatic histograms, depicting the lobes A1, A2 for symptomatic, and lobes B1 and B2 for asymptomatic. Note that side lobes of A1 and A2 are also considered while computing the main lobes' area. This means lobe A1 extends from the grayscale range 0–200, and lobe A2 extends from 200 to 255. Giving the same reasoning, lobes B1 and B2 are being depicted. The areas of these symptomatic lobes A1, A2, A1 + A2, and asymptomatic lobes B1, B2, and B1 + B2 are shown in table Table 2. As can be seen, the symptomatic lobe areas are more significant than the asymptomatic lobe areas by 32.86%, 31.05%, and 32.73%, respectively. This justifies the deep learning model's ability to show the MFS of the symptomatic class to be higher than the MFS of the asymptomatic class. This further validates the deep learning architecture design in terms of the number of layers of CNN. Note that the number of plaques in symptomatic and asymptomatic plaques is 196:150. On computing the lobe area per plaque (as seen in the row R5), symptomatic dominates compared to asymptomatic by 32.73%, which further attests to our hypothesis.
Visual representation of the visual feature maps using non-augmented data as example
A feature map from every layer is computed after training the model by loading the trained weight file. Collected all the filters mean output of 3 channels (RGB). Figures 7b and c represent an example feature map in layer 3 containing 64 filters. That is a grid of filter output with four rows and 16 columns (16 × 4 = 64). We use all the images from the training data for symptomatic (176 images) and asymptomatic (135 images) classes for mean value computation. We computed the mean feature map view of the symptomatic and asymptomatic class. In Fig. 7b and c, the purple block is a filtered image, and the features learned by the model at layer three shown in turquoise color, texture features of the grayscale images shown in the yellow band.
Hypothesis 2: chaotic distribution
The second hypothesis is well established in the literature. It states that the plaque assumes more randomness (chaotic or heterogeneous) in symptomatic patients than asymptomatic, as explained in the introduction. One elegant way of quantifying the plaque randomness is by representing the plaque as a chaotic behavior using Mandelbrot's fundamental equation of chaotic measurement so-called Fractal Dimension (FD) and symbolized as capital D. With this as an assumption, we, therefore, compute the FD for symptomatic and asymptomatic class pools. If the FD of symptomatic is higher than the FD of asymptomatic, our hypothesis holds on the plaque's characterization assumption leading to the validity of DL-based classification. Using Mandelbrot's equation, we, therefore, follow the standardized equation and algorithm to compute the D for both pools, and this can be seen in Fig. 8
D is the dimensionless quantity of self-similar objects, N is the number of boxes that cover the pattern, and r is the magnification. Note here that log is taken as a Napier log to the base "e." The D value for symptomatic and asymptomatic are 1.45 (derived from 176 images) and 1.36 (derived from 135 images), having [CI: 0.05 to 0.09], respectively, validating that D (Symptomatic) > D (Asymptomatic).
Atheromatic plaque segregation index using deep learning
This index aims to see how DL perceives the separation between the symptomatic and asymptomatic classes. We thus pass the training data sets through the DL system to compute the mean feature strength (MFS) as presented before. This MFS is computed for both symptomatic and asymptomatic data sets, and the percentage difference is considered the Atheromatic Plaque Segregation Index [23]. The separation further justifies the classification accuracy of ~ 90%. Atheromatic plaque separation index (APSI) is mathematically given in Eq. 14, computed using ML and DL strategies, as shown in Table 3.
The Table 3 shows that the APSI of the DL model is higher compared to the ML model. In the DL model, the feature learning is balanced between the two classes, whereas in the ML, it is not balanced.
Discussion
Our work is one of the first kinds in classifying symptomatic vs. asymptomatic carotid plaques using deep learning architecture. Our implementation consisted of 13-layered CNN architecture with and without augmentation. The CNN model yields the probability of the predicted risk belonging to categorical symptomatic or asymptomatic classes. Using the K10 protocol, the system showed accuracy and AUC of 86.17% and 0.86 (p-value < 0.0001) without augmentation, and 89.7% and 0.91 (p-value < 0.0001) with augmentation, respectively. We showed an improvement of 3.61%, when comparing ML vs. DL w/o augmentation (0.83, p-value < 0.0001 vs. augmentation 0.86, p-value < 0.0001), and an improvement of 8%, when comparing ML and DL w/o augmentation (0.83, p-value < 0.0001 vs. 0.91, p-value < 0.0001). The main cause of the AI model's success is the ability to select the low-level and high-level features representing the carotid plaque using the combination of convolution layers and average pooling layers followed by the dense layer combination based on neural network. Further, our system consisted of less number of weights, which gave an added advantage in terms of speed and stability [65].
Our DL model exhibited better accuracy and AUC compared to machine learning and demonstrated an accuracy increase of 6.0% (p-value < 0.0001) compared with the previously published work [23, 38]. We validated the DL system by running the algorithm on the facial biometric data [68] set, yielding 99% accuracy. We further validated the DL system using the widely accepted animal data set (ASSIRA) [69], yielding 97.56% accuracy.
A note on unbalanced dataset between symptomatic and asymptomatic plaques
Our dataset consisted of two classes of lesion types, symptomatic (196 Images) and asymptomatic (150 Images) plaques, and there was a slight imbalance in the class size. This was because they were obtained from consecutive patients referred for ultrasound scanning. We used two different strategies to avoid overfitting: (i) L2 regularisation in the dense layer of the DL system and (ii) probability of dropout layer in control. The probability of the dropout layer was 0.5 and this helped in preventing overfitting. This helped in maintaining the weights for balancing in each layer of the DL system.
Benchmarking against techniques available in the literature
We benchmarked our model with the existing study (representing from R1 to R5). Table 4 contains the eight attributes, namely reference/year (author with the year), plaque (represents input type), data size (represents the size of the cohort), feature extracted, type of classifier, ML vs. DL, ACC (accuracy in %), and AUC (area-under-the-curve) with the p-values.
Christodoulou et al. [70] (R1) extracted texture-based features using statistical methods in the carotid ultrasound scans (CUS). The authors fed these features to self-organizing map (SOM) and k-NN classifiers and achieved an accuracy of 73.1%, 68.8%, and AUC of 0.753 and 0.738, respectively. For this study, the authors' used plaque cut sections. Acharya et al. [23] extracted texture features from the delineated plaque using 346 CUS. SVM classifier was adapted yielding an accuracy of 83% (p-value = 0.0001). The same group in [71] showed texture and discrete wavelet transform (DWT) features derived from 346 delineated plaque cut sections using CUS. Using the SVM model with the RBF kernel, the author achieved an accuracy of 83.7% (p < 0.0001). In [60], the authors adopted two different cohorts, having sizes 346 and 146, taken from the UK and Portugal. Trace-transform and texture-based features were computed and fed to the fuzzy SVM classifier models, yielding an accuracy of 93.1% and 85.3%, respectively. Gastounioti et al. [72] developed several CAD schemes on 56 patients' ultrasound carotid plaque scans for classification using the SVM model and extracted features using Fisher Discrimination Ratio, yielding an accuracy of 88%. Recently, Skandha et al. [73]. characterized and classified the delineated plaques from the UK database (the same 346 patients as taken in [60]) using 3-D optimized deep convolution neural networks (DCNN) consisting of 11 layers, yielding an accuracy of 95.66%. Our proposed study adopted a different set ML and DL paradigm on the same UK database yielding an accuracy of 86.17% (w/o augmentation) and 89.7% (w/ augmentation), respectively. The corresponding AUC was 0.86 and 0.91 (p < 0.0001), respectively. The idea behind design of another DL and ML model was to compare and contrast with already existing DL models. Table 4 summarizes our DL model against previous works [23, 60, 70,71,72,73].
A special note on supercomputer hardware comparison to the local machine
We studied the impact of hardware resources on in-depth learning methodology. We adopted the supercomputer DGX V100, located in Bennett University (BU), Gr. Noida, India. The table below (Table 5) gives the specifications of the supercomputers. An essential point of difference between the local computer and the DGX model are (a) architecture (Dual 20 core vs. single core) and (b) clock speed (8.8 GHz vs. 2.6 GHz). Using this updated hardware, we compared the speed of training on a supercomputer with the local computer. Supercomputer (with 8 GPU) took 2 min per epoch during the DL training, while it took 28 min on the local computer.
Strengths, weakness, and extensions
We successfully demonstrated the classification of symptomatic and asymptomatic plaque using 13 layers of CNN architecture. This study is the first of its kind to adopt a supercomputer paradigm for the classification of symptomatic and asymptomatic plaque images. The system was validated using two established data sets. As our institution has multiple investigators using the supercomputer, we could use 6 out of the 8 GPUs available while running our protocols.
Despite the strengths of the system, there are certain limitations. A DL framework's training system is always challenging due to the many iterations (or epochs) required. Since the number of combinations was ten and three kinds of data sets, arranging these tasks on a shared supercomputer was challenging. The pilot study shows encouraging results.
Since the AI field is continuously changing, more modelling and computer-aided diagnosis systems can be exploited and tried in this framework [6, 74]. Because the dataset's size was not very large, one could further improve accuracy by augmentation. Additionally, AI methods using transfer learning could be adapted to prevent repetitive training. As part of the extension, effort can fuse US plaque with MRI for cross-modality validation using standardized registration methods [75] in the big data framework [76]. Lastly, the plaque characterization can be compared against higher-order spectra [39] and role of different US scanners [57].
Conclusion
This is the first study of its kind to characterize and classify carotid plaques into symptomatic and asymptomatic categories using a deep learning paradigm and implemented on a supercomputer. The deep learning system accuracy and AUC was 89.7% and 0.91 (p-value < 0.0001), respectively, showing an improvement of 6.0% compared with previous methods. Using the hypothesis that symptomatic plaque is heterogeneous and has a more chaotic representation by ultrasound, we demonstrated tissue characterization using (a) mean strengths of the feature map and (b) Mandelbrot's fractal dimension. We used the Atheromatic separation index to demonstrate the class separation. The system was developed on the supercomputer with 8 GPU configurations and took less than two seconds per image during the online test patient prediction. The system validated against two widely accepted datasets showing consistent results for the DL architecture.
References
Benjamin EJ et al (2019) Heart disease and stroke Statistics-2019 update a report from the American Heart Association. Circulation. https://doi.org/10.1161/CIR.0000000000000659
Fryar, C.D., T.-C. Chen, and X. Li, Prevalence of uncontrolled risk factors for cardiovascular disease: United States, 1999–2010. 2012: US Department of Health and Human Services, Centers for Disease Control and ….
Heron, M.P., Deaths: leading causes for 2015. 2017.
Suri JS, Kathuria C, Molinari F (2010) Atherosclerosis disease management. Springer Science & Business Media, USA
Sirimarco G et al (2013) Carotid atherosclerosis and risk of subsequent coronary event in outpatients with atherothrombosis. Stroke 44(2):373–379
El-Baz A, Suri JS (2011) Lung imaging and computer aided diagnosis. CRC Press, USA
Radeva, P. and J.S. Suri, Vascular and Intravascular Imaging Trends, Analysis, and Challenges, Volume 2; Plaque characterization, by Radeva, Petia; and Suri, Jasjit S.. ISBN: 978–0–7503–1999–7. IOP ebooks. Bristol, UK: IOP Publishing, 2019, 2019.
Liu Y et al (2019) Size of carotid artery intraplaque hemorrhage and acute ischemic stroke: a cardiovascular magnetic resonance Chinese atherosclerosis risk evaluation study. J Cardiovasc Magn Reson 21(1):36
Chien JD et al (2013) Demographics of carotid atherosclerotic plaque features imaged by computed tomography. Journal of Neuroradiology 40(1):1–10
Laine A, Sanches JM, Suri JS (2012) Ultrasound Imaging: Advances and Applications. Springer, USA
Liu, K. and J.S. Suri, Automatic vessel indentification for angiographic screening. 2005, Google Patents.
Aichner F et al (2009) High cardiovascular event rates in patients with asymptomatic carotid stenosis: the REACH Registry. Eur J Neurol 16(8):902–908
Viswanathan V et al (2020) Low-cost preventive screening using carotid ultrasound in patients with diabetes. Front Biosci (Landmark Ed) 25:1132–1171
Saba L et al (2014) Multi-modality atherosclerosis imaging and diagnosis. Springer, USA
Kotsis V et al (2018) Echolucency-based phenotype in carotid atherosclerosis disease for risk stratification of diabetes patients. Diabetes Res Clin Pract 143:322–331
Nicolaides AN et al (2005) Effect of image normalization on carotid plaque classification and the risk of ipsilateral hemispheric ischemic events: results from the asymptomatic carotid stenosis and risk of stroke study. Vascular 13(4):211–221
Nicolaides AN et al (2002) Ultrasound plaque characterisation, genetic markers and risks. Pathophysiol Haemost Thromb 32(5–6):371
Hussain MA et al (2018) Association between statin use and cardiovascular events after carotid artery revascularization. Journal of the American Heart Association 7(16):e009745
Nicolaides AN et al (2010) Asymptomatic internal carotid artery stenosis and cerebrovascular risk stratification. J Vasc Surg 52(6):1486-1496.e5
Kakkos SK et al (2013) The size of juxtaluminal hypoechoic area in ultrasound images of asymptomatic carotid plaques predicts the occurrence of stroke. J Vasc Surg 57(3):609–618
Paraskevas KI, Nicolaides AN, Kakkos SK (2020) Asymptomatic Carotid Stenosis and Risk of Stroke (ACSRS) study: what have we learned from it? Annals of Translational Medicine. https://doi.org/10.21037/atm.2020.02.156
Sharma AM et al (2015) A review on carotid ultrasound atherosclerotic tissue characterization and stroke risk stratification in machine learning framework. Current atherosclerosis reports 17(9):55
Acharya UR et al (2012) Atherosclerotic risk stratification strategy for carotid arteries using texture-based features. Ultrasound Med Biol 38(6):899–915
Acharya, U.R., et al. Carotid far wall characterization using LBP, Laws' Texture Energy and wall variability: A novel class of Atheromatic systems. in 2012 Annual International Conference of the IEEE Engineering in Medicine and Biology Society. 2012. IEEE.
Acharya, U.R., et al. Carotid ultrasound symptomatology using atherosclerotic plaque characterization: a class of Atheromatic systems. in 2012 Annual International Conference of the IEEE Engineering in Medicine and Biology Society. 2012. IEEE.
Acharya U et al (2013) Computed tomography carotid wall plaque characterization using a combination of discrete wavelet transform and texture features: A pilot study. Proc Inst Mech Eng [H] 227(6):643–654
Acharya UR et al (2013) Understanding symptomatology of atherosclerotic plaque by image-based tissue characterization. Comput Methods Programs Biomed 110(1):66–75
Araki T et al (2017) Stroke risk stratification and its validation using ultrasonic Echolucent Carotid Wall plaque morphology: a machine learning paradigm. Comput Biol Med 80:77–96
Araki T et al (2016) PCA-based polling strategy in machine learning framework for coronary artery disease risk assessment in intravascular ultrasound: A link between carotid and coronary grayscale plaque morphology. Comput Methods Programs Biomed 128:137–158
Saba L et al (2017) Plaque Tissue Morphology-Based Stroke Risk Stratification Using Carotid Ultrasound: A Polling-Based PCA Learning Paradigm. J Med Syst 41(6):98
Saba L et al (2019) The present and future of deep learning in radiology. Eur J Radiol 114:14–24
Khanna NN et al (2019) Rheumatoid arthritis: atherosclerosis imaging and cardiovascular risk assessment using machine and deep learning–based tissue characterization. Current atherosclerosis reports 21(2):7
Biswas M et al (2019) State-of-the-art review on deep learning in medical imaging. Frontiers in bioscience (Landmark edition) 24:392–426
Huang X et al (2017) Evaluation of carotid plaque echogenicity based on the integral of the cumulative probability distribution using gray-scale ultrasound images. PLoS ONE 12(10):e0185261
Lekadir K et al (2017) A Convolutional Neural Network for Automatic Characterization of Plaque Composition in Carotid Ultrasound. IEEE journal of biomedical and health informatics 21(1):48–55
Liu, B., et al. Feature generation by convolutional neural network for click-through rate prediction. in The World Wide Web Conference. 2019.
Yasar F, Akgunlu F (2005) Fractal dimension and lacunarity analysis of dental radiographs. Dentomaxillofacial radiology 34(5):261–267
Acharya UR et al (2012) An accurate and generalized approach to plaque characterization in 346 carotid ultrasound scans. IEEE Trans Instrum Meas 61(4):1045–1053
Martis RJ et al (2013) Application of higher order statistics for atrial arrhythmia classification. Biomed Signal Process Control 8(6):888–900
Biswas M et al (2017) Symtosis: A Liver Ultrasound Tissue Characterization and Risk Stratification in Optimized Deep Learning Paradigm. Comput Methods Programs Biomed 155:165–177
Kuppili V et al (2017) Extreme Learning Machine Framework for Risk Stratification of Fatty Liver Disease Using Ultrasound Tissue Characterization. J Med Syst 41(10):152
Saba L et al (2016) Automated stratification of liver disease in ultrasound: An online accurate feature classification paradigm. Comput Methods Programs Biomed 130:118–134
Acharya UR et al (2012) Data mining framework for fatty liver disease classification in ultrasound: a hybrid feature extraction paradigm. Med Phys 39:4255–4264
Singh BK et al (2017) Risk stratification of 2D ultrasound-based breast lesions using hybrid feature selection in machine learning paradigm. Measurement 105:146–157
Acharya U et al (2013) Diagnosis of Hashimoto’s thyroiditis in ultrasound using tissue characterization and pixel classification. Proc Inst Mech Eng [H] 227(7):788–798
Acharya UR et al (2012) Non-invasive automated 3D thyroid lesion classification in ultrasound: a class of ThyroScanTM systems. Ultrasonics 52(4):508–520
Acharya UR et al (2011) Cost-effective and non-invasive automated benign & malignant thyroid lesion classification in 3D contrast-enhanced ultrasound using combination of wavelets and textures: a class of ThyroScanTM algorithms. Technology in cancer research & treatment 10(4):371–380
Acharya UR et al (2014) A review on ultrasound-based thyroid cancer tissue characterization and automated classification. Technology in cancer research & treatment 13(4):289–301
Molinari F et al (2010) Characterization of single thyroid nodules by contrast-enhanced 3-D ultrasound. Ultrasound Med Biol 36(10):1616–1625
Banchhor SK et al (2017) Wall-based measurement features provides an improved IVUS coronary artery risk assessment when fused with plaque texture-based features during machine learning paradigm. Comput Biol Med 91:198–212
Pareek G et al (2013) Prostate tissue characterization/classification in 144 patient population using wavelet and higher order spectra features from transrectal ultrasound images. Technology in cancer research & treatment 12(6):545–557
McClure P et al (2014) In-vitro and in-vivo diagnostic techniques for prostate cancer: a review. J Biomed Nanotechnol 10(10):2747–2777
Acharya UR et al (2015) Ovarian tissue characterization in ultrasound: a review. Technology in cancer research & treatment 14(3):251–261
Acharya UR et al (2013) Ovarian tumor characterization and classification using ultrasound: A new online paradigm. Ovarian neoplasm imaging. Springer, New York p, pp 413–423
Maniruzzaman M et al (2017) Comparative approaches for classification of diabetes mellitus data: Machine learning paradigm. Comput Methods Programs Biomed 152:23–34
Shrivastava VK et al (2016) Computer-aided diagnosis of psoriasis skin images with HOS, texture and color features: a first comparative study of its kind. Comput Methods Programs Biomed 126:98–109
Acharya UR et al (2013) Automated classification of patients with coronary artery disease using grayscale features from left ventricle echocardiographic images. Comput Methods Programs Biomed 112(3):624–632
Cuadrado-Godia E et al (2018) Cerebral small vessel disease: A review focusing on pathophysiology, biomarkers, and machine learning strategies. Journal of stroke 20(3):302
Suri, J.S., Imaging based symptomatic classification and cardiovascular stroke risk score estimation. 2011, Google Patents.
Acharya UR et al (2013) Atherosclerotic plaque tissue characterization in 2D ultrasound longitudinal carotid scans for automated classification: a paradigm for stroke risk assessment. Med Biol Eng Compu 51(5):513–523
Monkam P et al (2018) CNN models discriminating between pulmonary micro-nodules and non-nodules from CT images. Biomedical engineering online 17(1):96
Acharya, U.R., et al. Atheromatic™: Symptomatic vs. asymptomatic classification of carotid ultrasound plaque using a combination of HOS, DWT & texture. in 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society. 2011. IEEE.
Than JC et al (2017) Lung disease stratification using amalgamation of Riesz and Gabor transforms in machine learning framework. Comput Biol Med 89:197–211
Li A et al (2019) Evaluating modern GPU interconnect: Pcie, nvlink, nv-sli, nvswitch and gpudirect. IEEE Trans Parallel Distrib Syst 31(1):94–110
Sanagala, S.S., et al. A Fast and Light Weight Deep Convolution Neural Network Model for Cancer Disease Identification in Human Lung (s). in 2019 18th IEEE International Conference On Machine Learning And Applications (ICMLA). 2019. IEEE.
Ho SSY (2016) Current status of carotid ultrasound in atherosclerosis. Quantitative imaging in medicine and surgery 6(3):285
Kurosaki Y et al (2017) Asymptomatic carotid T1-high-intense plaque as a risk factor for a subsequent cerebrovascular ischemic event. Cerebrovascular Diseases 43(5–6):250–256
Rujirakul K, So-In C (2014) and B. A parallel expectation-maximization PCA face recognition architecture. The Scientific World Journal, Arnonkijpanich, PEM-PCA, p 2014
Golle, P. Machine learning attacks against the Asirra CAPTCHA. in Proceedings of the 15th ACM conference on Computer and communications security. 2008.
Christodoulou CI et al (2003) Texture-based classification of atherosclerotic carotid plaques. IEEE Trans Med Imaging 22(7):902–912
Acharya UR et al (2011) An accurate and generalized approach to plaque characterization in 346 carotid ultrasound scans. IEEE Trans Instrum Meas 61(4):1045–1053
Gastounioti A et al (2014) A novel computerized tool to stratify risk in carotid atherosclerosis using kinematic features of the arterial wall. IEEE journal of biomedical and health informatics 19(3):1137–1145
Skandha S et al (2020) 3-D optimized classification and characterization artificial intelligence paradigm for cardiovascular/stroke risk stratification using carotid ultrasound-based delineated plaque: Atheromatic™ 2.0. Computers Biol Med 125:103958
Acharya R, Ng YE, Suri JS (eds) (2008) Image modeling of the human eye. Artech House, MA USA
Narayanan, R., et al. MRI-ultrasound registration for targeted prostate biopsy. in 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro. 2009. IEEE.
El-Baz A, Suri JS (2019) Big Data in Multimodal Medical Imaging. CRC Press, USA
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Dr. Deepak L. Bhatt discloses the following relationships - Advisory Board: Cardax, CellProthera, Cereno Scientific, Elsevier Practice Update Cardiology, Level Ex, Medscape Cardiology, MyoKardia, PhaseBio, PLx Pharma, Regado Biosciences; Board of Directors: Boston VA Research Institute, Society of Cardiovascular Patient Care, TobeSoft; Chair: American Heart Association Quality Oversight Committee; Data Monitoring Committees: Baim Institute for Clinical Research (formerly Harvard Clinical Research Institute, for the PORTICO trial, funded by St. Jude Medical, now Abbott), Cleveland Clinic (including for the ExCEED trial, funded by Edwards), Contego Medical (Chair, PERFORMANCE 2), Duke Clinical Research Institute, Mayo Clinic, Mount Sinai School of Medicine (for the ENVISAGE trial, funded by Daiichi Sankyo), Population Health Research Institute; Honoraria: American College of Cardiology (Senior Associate Editor, Clinical Trials and News, ACC.org; Vice-Chair, ACC Accreditation Committee), Baim Institute for Clinical Research (formerly Harvard Clinical Research Institute; RE-DUAL PCI clinical trial steering committee funded by Boehringer Ingelheim; AEGIS-II executive committee funded by CSL Behring), Belvoir Publications (Editor in Chief, Harvard Heart Letter), Canadian Medical and Surgical Knowledge Translation Research Group (clinical trial steering committees), Duke Clinical Research Institute (clinical trial steering committees, including for the PRONOUNCE trial, funded by Ferring Pharmaceuticals), HMP Global (Editor in Chief, Journal of Invasive Cardiology), Journal of the American College of Cardiology (Guest Editor; Associate Editor), K2P (Co-Chair, interdisciplinary curriculum), Level Ex, Medtelligence/ReachMD (CME steering committees), MJH Life Sciences, Population Health Research Institute (for the COMPASS operations committee, publications committee, steering committee, and USA national co-leader, funded by Bayer), Slack Publications (Chief Medical Editor, Cardiology Today’s Intervention), Society of Cardiovascular Patient Care (Secretary/Treasurer), WebMD (CME steering committees); Other: Clinical Cardiology (Deputy Editor), NCDR-ACTION Registry Steering Committee (Chair), VA CART Research and Publications Committee (Chair); Research Funding: Abbott, Afimmune, Amarin, Amgen, AstraZeneca, Bayer, Boehringer Ingelheim, Bristol-Myers Squibb, Cardax, Chiesi, CSL Behring, Eisai, Ethicon, Ferring Pharmaceuticals, Forest Laboratories, Fractyl, Idorsia, Ironwood, Ischemix, Lexicon, Lilly, Medtronic, MyoKardia, Pfizer, PhaseBio, PLx Pharma, Regeneron, Roche, Sanofi, Synaptic, The Medicines Company; Royalties: Elsevier (Editor, Cardiovascular Intervention: A Companion to Braunwald’s Heart Disease); Site Co-Investigator: Biotronik, Boston Scientific, CSI, St. Jude Medical (now Abbott), Svelte; Trustee: American College of Cardiology; Unfunded Research: FlowCo, Merck, Novo Nordisk, Takeda. Dr. Jasjit S. Suri is with AtheroPoint™, focused in the area of stroke/cardiovascular imaging.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Saba, L., Sanagala, S.S., Gupta, S.K. et al. Ultrasound-based internal carotid artery plaque characterization using deep learning paradigm on a supercomputer: a cardiovascular disease/stroke risk assessment system. Int J Cardiovasc Imaging 37, 1511–1528 (2021). https://doi.org/10.1007/s10554-020-02124-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10554-020-02124-9