Abstract
Fatty Liver Disease (FLD) is caused by the deposition of fat in liver cells and leads to deadly diseases such as liver cancer. Several FLD detection and characterization systems using machine learning (ML) based on Support Vector Machines (SVM) have been applied. These ML systems utilize large number of ultrasonic grayscale features, pooling strategy for selecting the best features and several combinations of training/testing. As result, they are computationally intensive, slow and do not guarantee high performance due to mismatch between grayscale features and classifier type. This study proposes a reliable and fast Extreme Learning Machine (ELM)-based tissue characterization system (a class of Symtosis) for risk stratification of ultrasound liver images. ELM is used to train single layer feed forward neural network (SLFFNN). The input-to-hidden layer weights are randomly generated reducing computational cost. The only weights to be trained are hidden-to-output layer which is done in a single pass (without any iteration) making ELM faster than conventional ML methods. Adapting four types of K-fold cross-validation (K = 2, 3, 5 and 10) protocols on three kinds of data sizes: S0-original, S4-four splits, S8-sixty four splits (a total of 12 cases) and 46 types of grayscale features, we stratify the FLD US images using ELM and benchmark against SVM. Using the US liver database of 63 patients (27 normal/36 abnormal), our results demonstrate superior performance of ELM compared to SVM, for all cross-validation protocols (K2, K3, K5 and K10) and all types of US data sets (S0, S4, and S8) in terms of sensitivity, specificity, accuracy and area under the curve (AUC). Using the K10 cross-validation protocol on S8 data set, ELM showed an accuracy of 96.75% compared to 89.01% for SVM, and correspondingly, the AUC: 0.97 and 0.91, respectively. Further experiments also showed the mean reliability of 99% for ELM classifier, along with the mean speed improvement of 40% using ELM against SVM. We validated the symtosis system using two class biometric facial public data demonstrating an accuracy of 100%.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Over the last two decades, liver-related mortality has ranked among the top 12 causes of death and has been repeatedly listed as the fourth leading cause of death among adults aged 45–54 [1]. The presence of an excess amount of fat in liver cells leads to Fatty Liver Diseases (FLD). The process of deposition of fat in the liver cells is called steatosis and this can be caused by metabolic syndrome, consumption of alcohol, obesity due to insulin resistance, and a variety of other factors. [1, 2]. FLD is further categorized into two types: alcoholic and non-alcoholic. The majority of the population of western nations suffering from FLD is afflicted by non-alcoholic FLD (NAFLD) [3]. FLD may lead to serious diseases like inflammation (steatohepatitis), cirrhosis and liver cancer. This disease is curable in the early stages and early detection of FLD has shown great success in patient to live long lifespan. Also, the cost of FLD detection is less compared to treatment of advanced liver diseases. Currently, liver biopsy is the gold standard for the detection of FLD. The biopsy technique is uncomfortable, suffers from sampling error, and is invasive [4]. Various non-invasive imaging techniques such as Compute Tomography (CT) and Magnetic Resonance Imaging (MRI) are available for FLD detection. CT suffers from the challenge of radiation risks [5], while MRI can only detect very small amount of fat [6]. The MRI technique works well while detecting fatty infiltration [1, 7]. An alternative to these modalities is Ultrasound (US) images, which are commonly adapted for FLD imaging [8]. The application of machine learning (ML) for US liver images shows sensitivity and specificity above 80% [9]. Therefore, US has become one of the most popular scanning techniques for FLD detection [10].
Two sets of methods have been proposed in literature for characterization of liver disease: (a) based on ML and (b) based on signal processing. Under the class of Symtosis for FLD detection, Suri and his team designed tissue characterization system [11] utilizing features like: Discrete wavelet Transform (DWT) [12], High order spectra (HOS) [13] and texture features [14], which were computed using US liver images which were then fed to Decision Tree (DT)-based classifier leading to an accuracy of 93.3%. Under the same class, Acharya et al. in 2014, proposed a Fuzzy Classifier for detection of Hashimoto Thyroiditis from thyroid images [15,16,17] using wavelet transform [18]. The system achieved an accuracy of 84.6%. In 2014, Subramanya et al. [19] achieved accuracy of 84.9% on US liver images using SVM classifier. Using signal processing approach, Ma et al. in 2015, developed kurtosis-based [20] scanning method for detection and grading of FLD in US liver images, demonstrating an accuracy of 81.2%. In 2016, Suri and his team (Saba et al. [21]) used Back Propagation Neural Network (BPNN) consisting of 10 hidden layers, and used 128 features extracted from US liver images using six different types of feature extraction algorithms. BPNN showed an accuracy of 97.6%.
Support Vector Machine (SVM) [22] is a widely used ML technique for supervised learning. SVMs apply two main techniques for stratification. First, it applies kernel methods to transform the problem from original input space to a high dimensional one, called the feature space, where linear separation of training samples belonging to different classes is possible. Second, it tries to find the best separating hyper-plane between the two classes. These ML systems utilize large number of ultrasonic grayscale features, pooling strategy for selecting the best features and several combinations of training and testing. As a result, they are computationally intensive, slow and do not guarantee high performance due to mismatch between grayscale features and classifier type. Keeping the computational speed and performance with respect to data size in mind, we present here Extreme Learning Machine (ELM) [23, 24] paradigm. The ELM trains a single layer feed forward neural network (SLFFNN) where the input-to-hidden layer weights are randomly initialized. The ELM only trains the hidden-to-output layer weights using the least square loss model that employs a closed form solution given by the Moore–Penrose pseudo-inverse [25]. In a least square sense, the error is minimized and likely to prove more accurate or at least comparable to iterative neural network models [21]. Further, ELM allows only a single stop shop for training weights and therefore, we believe that ELM is likely to be faster technique compared to SVM, and will reach minimum least square error in a single pass irrespective of the training data size. Since ELM is single layered neural network architecture unlike other neural network architectures [21], this requires low resource management and likely to show better performance. We thus hypothesize that ELM will be a better system for FLD risk stratification compared to conventional ML systems.
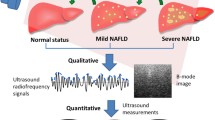
We provide a comprehensive analysis of the two methodologies in this paper using US liver dataset. We developed a computer aided system under the class of Symtosis for detection and stratification FLD-affected (diseased) and FLD-unaffected (controls or normal’s) liver images as shown in Fig. 1. The input US images are processed and partitioned before feeding them into the tissue characterization module. Four type of cross-validations (K = 2, 3, 5 and 10) are performed on the dataset before feeding. Since our data is limited, we additionally sub-sample the original images (S0) into four parts (S4) and sixty four parts (S8). Then, the tissue characterization module outputs a predicted risk based on ground truth (considered as biopsy reports) and cross-validation type. Finally, the predicted risk is evaluated against the ground truth that gives us the performance parameters. The Fig. 1 gives us an overall vision of the entire system. The system derives 46 features using Gabor, Gray-Level Co-occurrence Matrix (GLCM) and Gray Level Run Length matrix (GRLM). Since our scope of this paper is to purely understand and harness ELM and further to benchmark against SVM and back propagation neural network (BPNN), we thus have limited ourselves to handling only limited feature extraction without feature selection paradigms. Using K10 protocol on three kinds of data sets (S0, S4, S8), the system yields an accuracy of 92.4%, 94.8% and 96.7%, respectively, while the SVM-based system yields an accuracies as: 86%, 87.9%, and 89%, respectively. It is also observed that the accuracy values increase with an increase in K (cross-validation folds) and sub-sampling. We demonstrate a 40% improvement in ELM speed when compared against SVM. We further compared our architecture against BPNN [21], which is also designed using NN-based systems, showing comparable accuracy with efficient architecture and speed. The ELM-based tissue characterization system is also validated using biometric facial dataset where it achieves an accuracy of 100% across all cross-validation protocols showing greater degree of generalization compared to contemporary ML algorithms such as SVM.
In the following section 2, we discuss data demographics and US image acquisition protocol. Section 3 presents different feature extraction algorithms and establishes the mathematical foundation of the ELM paradigm. Experimental protocol is presented in section 4 and the results are presented in section 5. Benchmarking against conventional SVM-based classification is presented in section 6 while the discussion is presented in section 7. Conclusions are presented in section 8.
Data demographics, collection and preparation
demographics, ethics approval and gold standard
Sixty-three patients (36 abnormal and 27 normal) were collected after IRB approval by Instituto Superior Tecnico (IST), University of Lisbon, Portugal and written informed consent provided by all the patients. The images were retrospectively analyzed. The patient with normal body mass index was selected. The normal/abnormal US scanned images are as shown in Fig. 2. The gold standard or ground truth label for each patient (normal or abnormal) was determined by taking a liver biopsy and analyzing it in the tissue pathological laboratory [11].
Liver ultrasound scanning, data collection and preparation
The US scanning and analysis were done on the patients with the help of medical experts. A Philips CX 50 US machine was used for capturing US scanned images. The US scanner had frequency from 1 to 5 MHz and 160 piezoelectric elements of curved shape. The captured images were gray scale images with 1024 × 1024 pixels. Each gray scale image was stored as 8 bits/pixel resolution. The manufactures were provided a default computer interface for obtaining input. We used this interface for obtaining patient’s image data. We checked the setting and collaboration of US machine before obtaining input images. The standardization was done as per Qayyum et al. [26] approach. i.e., 20 patients with normal liver and normal body mass index (18.5–24.9) were called and US scanning was performed. The result of the image was then examined. Based on the results, standardization was done. US machine with image depth of 15 cm and frequency 3.5 MHz was used. The image had two focal zones with 7.5 cm in the central. The dynamic range for this experiment was set at 70 dB but the gain was changed based on the patient biotype. For all the examination of scanned images Time Gain Compensation (TGC) was fixed at the central point to remove this variable parameter. The fixed central position assists standardization of the protocol. Different transducer angles and orientations were used based on patient biotype to get the liver anatomical landmarks. Patients were kept in supine, comfortable position during scanning for avoiding major patient motion. The liver has a small left lobe (in the epigastric area) and a large right lobe (in the right hypochondrium) [27]. The effect of FLD disease can be viewed in both parts of the liver. Since right liver is the major liver part, we used scanned image of the right lobe liver. A region of interest (ROI) of 128 × 128 pixels along the medial axis was extracted from each image.
Sub-sampling of us data sets (S4 and S8)
Since the learning strategies of ELM-based Symtosis require faster generalization if the training samples be increased. We therefore subsample the original DICOM images using spatial transformation into two sets of data sets: S4 and S8. Examples of S4 and S8 images are as shown in Figs. 3 and 4.
Methodology
The working of classifiers in the Symtosis system shown in Fig. 1 has been discussed here. The main challenge in application of SVM is computational cost involved with finding support vectors in the training dataset. The application of kernel functions to find linear solution for non-linearly separable data in high dimensional space adds to the mathematical stress [28]. ELM solves the problem of classification in single iteration, i.e., removing the idea of an iterative approach. The internal architecture ELM-based tissue classification in Symtosis system that allows training of the SLFFNN in a single pass is shown in Fig. 5. It is seen that ELM combines generalized matrix inverse of an activation function (sometimes called as pseudo inverse matrix activation function matrix-shown in Appendix A) with the known targets to find the optimized hidden-to-output weights in a single iteration and thereby reducing computational cost of the system. Note that, the activation function consists of the combination of input features and randomized input-to-hidden layer weights. We perform feature extraction on US liver images and propose an ELM-based CADx system for the detection and risk stratification for FLD diseases [23, 24, 29]. We use three texture feature extraction algorithms namely Gabor, GLCM and GRLM features [30,31,32,33,34]. The Gabor feature extraction is based on the scale and direction of the pixel distribution in image using Gabor filters. GLCM extracts statistical second order features and finally, GRLM matrix calculates the neighboring pixel of a reference pixel and texture feature are then computed (Appendix C). The details of ELM architecture and mathematical foundation are discussed in subsection 3.1, while subsection 3.2 presents the tissue characterization algorithm. The details of feature extraction algorithms are given in subsection 3.3.
Three layered ELM architecture for training weights
Extreme learning machine (ELM) is a SLFFNN which can be trained in a single pass, making it faster compared to contemporary ML algorithms. There are three layers of neurons (or nodes) in SLFFNN, where weights between input and the hidden nodes are randomly initiated and then fixed without any iteration (so called input-to-hidden weights). The only weights that are to be learned are the weights between the hidden layer and the output layer. Since ELM learns the weights in single pass, it tends to reach a global optimum immediately. The architecture for ELM is shown in Fig. 5. It is a three layered architecture. The first layer accepts the input and forwards it to the hidden layer. The outputs from hidden layer are forwarded to the output layer.
Let the number of training input images be represented by vector P trg and testing images be represented as P tst . The label vectors corresponding to training images and testing images be represented as L trg and L tst . Let the weight vector be W from input-to-hidden layer. Let Q be the output of application of activation function to the input data. Let δ be the hidden-to-output layer vector of training weights. Then, the least square solution is given by \( \boldsymbol{Q}\hat{\boldsymbol{\delta}}-{\boldsymbol{L}}_{trg}=\underset{\delta }{\min}\boldsymbol{Q}\boldsymbol{\delta } -{\boldsymbol{L}}_{trg}, \) where \( \hat{\boldsymbol{\delta}} \) is the least squares solution of the Qδ = L trg . For a larger training dataset the smallest norm least squares solution of the linear system is given by: \( \hat{\boldsymbol{\delta}}={\boldsymbol{Q}}^{\dagger }{\boldsymbol{L}}_{trg}, \) where ,Q † is the Moore–Penrose [35] generalized inverse of matrix Q. The complete set of mathematical symbols and their meaning are given in Table 9. The mathematical derivation of ELM is given in detail in Appendix A.
Tissue characterization and risk stratification using ELM and SVM frameworks
The system for tissue characterization and risk stratification is based on the conventional ML system design, where, the input data is split into training and testing data sets for cross-validation protocol design. This can be seen in Fig. 6. This consists of two components: training-phase and testing-phase correspondingly shown as the left and right half of the Fig. 6. The training-phase generates the training weights or coefficients, while the testing-phase predicts the label class. The testing-phase is primarily the mirror image of training-phase, except that the training-phase uses ground truth labels along with grayscale features computed from training US liver images to generate the training weights (or training coefficients). The testing-phase then predicts the label class on the test images which is computed by transformation of testing features using the training weights. Note that both systems (ELM and SVM) adapt the same feature computation protocol. The ELM-based tissue characterization system uses SLFFNN that is comprised of three sets of neurons connected by lines carrying weights. The weights between input and the hidden neurons (input-to-hidden weights) are randomly initiated while the weights that are to be learned are the weights between the hidden layer and the output (hidden-to-output) layer. The number of input neurons equals the number of features extracted from an image. Empirically, the number of hidden layer neurons is taken as two hundred. All input neurons are connected to the hidden neurons.
The most common types of activation functions used in ELM are: sigmoid, sine hard limit, triangular basis function and radial basis function. The activation function used adapted in our experiment is sigmoid function. The number of output neurons set is based on the type of classification problem. Each of the hidden neurons is connected to the output layer neuron. Using the notations (as explained in the Appendix A), the least square solution can be converted into the algorithmic steps as presented in the pseudo code block shown below. Note that, if SVM (as explained in Appendix B) is adapted in Fig. 6, then, the maximum margin hyper-plane between two classes is found out from the computed support vectors obtained from the SVM during training-phase.
Feature extraction
The idea behind the feature extraction to compute a limited number of features to understand the power of ELM while benchmarking against SVM. Here, we discuss the feature extraction algorithms applied in our experiment i.e., Gabor, GLCM, GRLM. The choice of these features are based on the directions and scales combined with texture repeatability [36].
Gabor-based directional features
Gabor filter is edge detection filter and is the combination of Gaussian and complex-plane wave. Through this combination, it tries to diminish the uncertainty in both spatial and frequency domains. Application of dilations and rotations of this function produce alike Gabor filters. It helps in the alignment and scale-tunable edge and line detection. It helps in expanding an image and become contained in spatial frequency depiction. Gabor transform has an impulse response that can be represented by a sinusoidal wave (a plane wave for distinct frequency and aligned 2-D Gabor). The function is given as:
where, (p, q) represents the spatial-domain rectilinear coordinates, (U, V) are points that are the specific 2-D frequency of the complex sinusoid and (σ p , σ q ) depict the spatial extent and bandwidth of f. The Fig. 7 shows the Gabor filters used for feature extraction. A scale of 2 and 10 orientations were selected to define 20 Gabor features.
Gray level co-occurrence matrix
Gray Level Co-occurrence Matrix (GLCM) is a widely known methodology for texture extraction [37,38,39,40,41]. GLCM shows the spatial relationship of neighboring pixels. It calculates the occurrence of a pixel with a specific gray level or intensity compared to its neighbors in a number of directions. Features are calculated based on the statistical distribution of pixel intensities. GLCM based feature extraction uses second order statistics. The texture feature obtained in Co-occurrence matrix never directly uses for analysis. Gray level co-occurrence matrix calculates the probability of two pixel with gray level i, j which located in inter distance d direction, θ. The probability is represented by p(i, j | d, θ). The spatial relationship is represented in terms of angle θ and distance d. From the calculated probability we calculate features. A brief description of GLCM is given in Appendix C.1.
Gray level run length matrix
Gray Level Run Length Matrix (GRLM) is based on a set of collinear pixels that have the same gray level called Run Length Matrix (RLM). The main function of GRLM is to extract texture features and images of grey intensity pixels in a specific orientation from which the reference pixels are computed. The number of neighboring pixels with the same grey intensity in a particular direction is called run length represented as S(i, j | d, θ), which is the number of j neighboring pixels with the intensity i, in the direction θ. GRLM is further discussed in Appendix C.2. Description for all symbols are given in Appendix D.
Experimental protocol
We carry out cross-validation experimental protocol to analyze the strength of generalization for each methodology. The subsection 4.1 discusses the effect of four cross-validation protocols on stratification accuracy using all three kinds of data sets. In the subsection 4.2, we study the effect of percentage of data size on sub-sampling data on the system’s accuracy using various cross-validation protocols. Since ELM is a single pass algorithm, we inspect the time required by ELM and SVM in this experiment. Subsection 4.3 presents the comparative time analysis for ELM and SVM algorithms.
Experiment 1: Effect of training data size on accuracy using four CV protocols
The objective of this experiment is to understand the effect of training data size on the performance of risk stratification. The cross-validation protocol allows us to change the number of patients in the training data sets. We adapted four kinds of cross-validation protocols: K2, K3, K5 and K10 labeled as: 2 fold, 3 fold, 5 fold and 10 fold, respectively. Each fold is a part of the data set. In K2 cross-validation, the dataset is equally partitioned into two, where one part is used for training and the other part is used for testing. This process is the same for K3, K5, and K10, with data in KN being divided into N parts where N-1 parts are used for training and the remaining one part is used for testing. Each of the cross-validated datasets is the input into the classifier (ELM or SVM) for training and testing. The protocols are repeated twenty times randomly and average accuracy, sensitivity, specificity and time are recorded.
Experiment 2: Effect of training set size using sub-sampling strategy
It is important to understand the effect of the training data size on the ELM architecture. Since no iterations are involved unlike conventional NN or BPNN, the size of the training data can play a larger role in the computing the performance of the ELM system. We therefore sub-sampled the original databases (S0) into two kinds of sub-samplings called as: S4 and S8 datasets. In S4, images in S0 dataset were divided into 4 equal parts with each image representing one-fourth dimension of original image. The S4 dataset consists of 252 images. The S8 was obtained from S4 data set. This means 16 parts for each of the S4 data sets. Thus, S8 was: ×16 parts of 252, which is (252 × 16) 4032. So, S0 = 63, S4 = 63 × 4 = 252; S8 = 252 × 16 = 63x4x16 = 4032. The images are shown in Figs. 3 and 4. It is therefore required to run all CV protocols (i.e., K2, K3, K5 and K10 cross-validations) for all three kinds of data sets: S0, S4 and S8.
Experiment 3: Time comparison between ELM & SVM
Since Extreme Learning Machine comes from the ability to learn extremely fast, it is necessary to compute the time complexity of the ELM system for both training-phase and testing-phases. Thus, it requires computing the times for all CV protocols (i.e., K2, K3, K5 and K10 cross-validations) and for all three kinds of data sets (i.e, S0, S4 and S8), thus, leading to 12 time comparisons.
Results
This section provides the results of the three experiments carried out on the US liver dataset in ELM framework. Sub section 5.1 shows the effect of training data size using four CV Protocols. The results on the effect of training set size using sub-sampling strategy are shown in sub section 5.2. The timing analysis results are presented in sub section 5.3.
Experiment 1: Effect of training data size on accuracy using four CV protocols
If η sys is the system accuracy, k represents the cross-validation method i.e., K2, K3, K5 and K10, t represents index of trial numbers, T represents total number of trials, i represents index of data size, N L represents total size of the liver dataset, then the average accuracy for each cross-validation protocol, k, of the system can be mathematically expressed as:
A total of T = 20 trials are conducted. The average accuracy, sensitivity, specificity and timing for all protocols are as shown in Table 1. Note that same formula is applicable for SVM-based and ELM-based Symtosis systems. It is clearly seen that, ELM outperforms SVM for all cross-validations. ELM gives 92.4% accuracy with K10 cross-validation compared to SVM that gives only 86.42%. The average specificity and sensitivity is higher for ELM when compared with SVM. Results for S4 and S8 datasets are given in Table 10 and Table 11 in Appendix E.
Experiment 2: Effect of percentage of training data size during CV protocols
To know the effect of the training data size on the ELM architecture, we perform the experiment with varying data sizes. We performed this experiment on S0, S4 and S8 datasets. As the size of training dataset increases, the accuracy also increased. The S4 outperforms S0, while S8 outperforms S4 and S0 for all training dataset sizes. The accuracy obtained for SVM and ELM with different dataset sizes for each cross-validation is shown separately in Fig. 8.
Experiment 3: Time comparison between ELM & SVM
ELM is a fast learning neural network. The ELM gives better performance in terms of training and testing time. The time comparison between ELM and SVM classifier for S0 is given in Table 2, S4 in Table 3, and S8 in Table 4. The training time is less for K2 and more for K10, but for testing it is reversed. It happens because the data size in case of training increases from K2 to K10 and for testing it decreases from K2 to K10. The SVM has greater training and testing time for all types of cross-validations in all three datasets. The ELM architecture uses 2.1 milliseconds (ms) for testing and 9.3 ms in training for K10 cross-validation for S0. The testing time is almost negligible. The average speed-up improvement of ELM over SVM is 31% for S0. The testing time for S4 is 3.0 ms and training time is 10.3 ms for K10 cross-validation with ELM. The maximum testing time is for K2 which is 4.6 ms for ELM classifier. The time increased from S0 to S4 for ELM and SVM, but is still negligible. For SVM training time is 16.0 ms in K10 cross-validation. The speed-up achieved for ELM over SVM is approximate 47% for S4. When we consider S8 dataset SVM needs maximum 19.5 ms for training whereas ELM needs only 15.3 ms for training. The increase in performance speed for ELM over SVM for S8 is 41%. Overall, the average speed-up of ELM over SVM is approximately 40%. We further validated our ELM and SVM classification using 2-class biometric facial data (Appendix F).
Performance valuation
The performance of the ELM system is computed by plotting the ROC and AUC’s for all sets of CV protocols. We further record the performance attributes such as: accuracy, sensitivity, specificity and is presented in subsection 6.1. The reliability and stability analysis is evaluated in subsection 6.2.
ROC curves
The performance of the ELM was computed using the ROC curves using all three kinds of datasets: S0, S4 and S8. For each data set, we adapted four kinds of cross-validation protocols: K2, K3, K5 and K10. Thus we demonstrate 12 ROC curves spanned in 3 figures: Fig. 9 (A), (B), and (C), respectively. Note that in each combination of K and S, we compute ROC curves using the two sets of machine learning systems: ELM and SVM. They are represented by alphabets (a), (b), (c) and (d) in each of the three set of figures. The AUC for S0, S4 and S8 data set is shown in Table 1, Table 10 and Table 11. For each data set (S0, S4 and S8), K10 does the best off all the four cross-validation protocols and ELM shows superior performance when compared against SVM (0.97 vs. 0.91).
Reliability and stability analysis
In this subsection, reliability and stability analysis of ELM is done. This assessment is crucial because it gives an indication how the system performs under repeated conditions and also different conditions. This affirms the results produced are consistent and repeatable. The reliability index has been derived by observing the deviation of the classification accuracy with respect to its mean as the data size increases [33]. The reliability index \( {\zeta}_{N_L}\left(\%\right) \) is formulated as:
Where, \( {\mu}_{N_L} \) is the mean accuracy and \( {\sigma}_{N_L} \) represents the standard deviation of all accuracies for N L US liver images.
The stability assessment analyses how the system changes across repeated conditions. We do this by using a similar approach to dynamics of the control theory [34]. Firstly, a threshold stability criterion of 5% variation is defined. When a system varies more than 5%, it is said that the system is not stable. Next we calculate the standard deviation (SD) for each computation of different data sizes. If the SD is less than 5% we can declare that the system is stable. The reliability indices for all four K-fold protocols in Table 5 are above 0.95 indicating a strong reliability of the ELM classification system. We further validated our ELM and SVM classification using 2-class biometric facial data (Appendix F).
Discussion
This study proposed a reliable and fast Extreme Learning Machine (ELM)-based tissue characterization system (a class of Symtosis system) for stratification of FLD disease in US liver images. ELM was used to train SLFFNN. The input-to-hidden layer weights were randomly generated reducing computational cost. The only weights to be trained were hidden-to-output layer which was done in a single pass (without any iteration) making ELM faster compared to SVM model. ELM-based characterization system was benchmarked against previously developed SVM-based system. Note that same set of feature were applied to ELM and SVM systems. The common three sets of grayscale features were: GRLM, GLCM and Gabor. The main spirit of the study was to compare ELM vs. SVM. Since ELM is a NN-based system, we compared ELM against BPNN. It was demonstrated that by reducing the number of features to one-third and also reducing the number of hidden layers by one-third (as demonstrated by BPNN [21]), the ELM still yielded comparable accuracy and the speed several times faster. While Suri’s group [21, 40] have developed features maximizing close to 1000 features combined with feature selection methods such as PCA, FDA embedded with classifiers such as Bayesian, SVM, K-mean, etc., we have confined this study only to benchmark the ELM against SVM-based paradigm for tissue liver classification. We performed the scientific validation using biometric facial datasets shown in the Appendix F.
Benchmarking
There is not much literature covering CADx-based system for liver diagnosis and risk stratification. Suri and his team performed classification of US liver dataset [11] using Decision tree (DT) and detection of Hashimoto Thyroiditis using Fuzzy classifiers [25] (shown in Table 6). Three sets of features were computed which was then applied to the DT-based classifier. This constituted Higher Order Spectra (HOS), Texture and Discrete Wavelet Transform (DWT) with the assumption that the pixels are distributed non-linear in nature. Texture captured the various granular structures in the US liver images, which was ideal. Feature reduction was performed followed by DT-based classification yielding an accuracy of 93.3%. Douali et al. [42] used Case Based Fuzzy Cognitive Map (CBFCM) in the year 2013, for classification of FLD and achieved an accuracy of 91.9% for 162 patients. In the year 2014, Vanderbeck et al. [43] achieved an accuracy of 89.3% using SVM on 47 patients using 582 features. In the year 2014, Acharya et al. [29] proposed a Fuzzy Classifier (FC) for detection of Hashimoto Thyroiditis from US thyroid images. The features were extracted using wavelet transform. A total of 526 US images were used and the system achieved an accuracy of 84.6%. In 2014, Subramanya et al. [19] used 53 US liver images which were distributed among four different classes consisting of: 12 normal, 14 mild, 14 moderate and 13 severe. Six types of features were computed such as: First Order Statics (FOS), Gradient-based (Gr), Mutual Information-based (MI), GRLM, GLCM and Laws Texture. SVM was applied to achieve an average accuracy of 84.9%. Very recently, Suri and his team [21] achieved an accuracy of 97.6% using BPNN on US liver images. A short comparison of BPNN and ELM is discussed in the next subsection. More recently, Liu et al. [44] used a combination of liver capsule detection technique and trained Convolution Neural Network [45] model for feature extraction, and used SVM as classifier to achieve accuracy of 89.2%.
Our study used ELM for the classification process on three kinds of data sets: S0, S4, S8 data sets demonstrating the accuracies of: 92.4%, 94.8% and 96.7%, respectively. The K10 cross-validation outperforms other three cross-validations. It is observed from Table 1 that ELM accuracy is higher than SVM for all cross-validation protocols i.e., K2, K3, K5 and K1 (81.70, 82.70, 89.00 and 92.40 against 76.14, 75.40, 83.50, 86.42, percentage respectively). It is also observed from Tables 2, 3 and 4 that average speed-up of ELM over SVM is approximately 40% asserting the hypothesis that ELM is faster than SVM. The stability analysis from Table 5 shows that ELM is highly reliable and stable system. It is further noted, that ELM accuracy increases as the data size increases.
FC: Fuzzy Classifier; HOS: High Order Spectra; DWT: Wavelet Packet Decomposition; FOS: First Order Statistics; Gr: Gradient based features; MI: Moment invariant; Laws: Laws texture features; BG: Basic geometric, CBFCM: Case Based Fuzzy Cognitive Map
A short comparison on ELM Vs. BPNN
Since ELM and BPNN are both NN-based strategies, we therefore ensured that we compared them very closely. BPNN adapted by Suri produced an accuracy of 97.6% while ELM gave the best accuracy of 96.7%. From these observations (also shown in Table 6), it can be argued that BPNN achieved better accuracy compared to ELM, so BPNN would be a better classifier. However, the merits of ELM far outweigh BPNN. ELM is a single layer feed forward neural network, thus network complexity is much lower compared to BPNN, since BPNN can have multiple number of hidden layers and neurons (up to 10 layers in [21]). Second, BPNN convergence is far slower compared to ELM, because each weight in BPNN architecture is updated iteratively, and such iterations can be a larger number (say 1000), which can take more time unlike one iteration in ELM, thereby increasing computational complexity of the system. On the contrary, ELM achieves a comparable accuracy in a single pass, due to its simple matrix multiplications and single hidden layer. Moreover, ELM achieves accuracy difference less than 1 % (0.9%) with only 46 ordinary features, unlike BPNN, which takes three times more number of features (128). Overall, these merits rationalize the selection of ELM compared to BPNN and SVM which are categorized as conventional ML techniques.
A special note on ELM Vs. SVM
The SVM training is in two stages. i.e., in stage one, the input data is mapped to a higher dimensional feature space through a nonlinear feature mapping function or kernel functions and in the stage two, the optimization method is used to find maximum separating margin of two different classes in this feature space while minimizing the training errors. The optimization problem is quadratic and convex, and so it can be solved efficiently.
The ELM trains a SLFFNN in two main stages: (1) feature mapping and (2) linear parameters solving. In the first stage, the hidden layer weights are randomly initialized to map the input data into a feature space by some non-linear mapping functions i.e., sigmoid. In the second stage of ELM learning, the weights in hidden-to-output layer, denoted by δ are solved by minimizing the approximation error in the squared error sense. The ELM is basically a SLFFNN whose weights and biases in the first layer are randomly initialized and kept constant. The weights and optionally biases of the second layer are selected by minimizing the squared loss of predicted errors by using Moore–Penrose pseudo-inverse [25, 34]. The weights of hidden-to-output neurons are learned in a single step. The ELM requires a time proportional to the number of hidden neurons for datasets smaller than the size of hidden neurons [26].
However, ELM is different from SVM, as the input-to-hidden layer weights do need not be tuned. ELM provides a universal solution for regression, binary and multi-class classification where the least square solution is dependent only on input data and number of training samples [46]. The ELM computational complexity is much simpler than SVM since training of ELM only involves finding hidden-to-output layer weights which is obtained in single pass by multiplying of Moore-Penrose inverse of activation function output and the target. Therefore, computational complexity of the ELM is dependent on the number of hidden nodes for smaller dataset and requires a time proportional to the number of hidden neurons, which is much larger than the size of small datasets and whose evaluations require a single feed-forward pass. The SVM computational complexity increases with non-linearly separable data as it has to be solved in high-dimensional space using kernel functions. Also, the kernel functions of SVM vary from application-to-application while ELM provides a more generalized solution to the classification problem. It becomes more complex in SVM with an increase in size of training dataset since it involves finding support vectors from the entire training dataset involving huge number mathematical computations. Although, ELM and SVM employ the same cost function, the optimization constraint in case of ELM is milder compared to SVM, wherein, in the former case it employs the least square model for optimization while the latter uses highest separating margin approach. The ELM is faster compared to other classifiers because the input-hidden weights are constant; the model learns only the hidden-output weights, which is equivalent to learning a linear model [46, 47]. It was verified from experiments that ELM with random hidden nodes can run even up to ten times faster compared to SVM. From this assessment, it is correctly concluded and justified that ELM gives faster accuracy results compared to SVM.
The ELM employs constrained least square model for error minimization. It applies gradient descent derivative of error with respect to δ, in a single pass over the whole feature space [48,49,50], allowing smallest possible training error. In SVM, the application of all features allows presence of noisy data which does not allow it to converge it to a single optimized separating hyper-plane. Therefore, it is necessary for SVM to identify feature types and employ feature selection algorithms to remove noisy features for achieving higher accuracy. So thus can say with confidence that accuracy of ELM is better than SVM in absence of feature selection algorithms.
Strengths, weaknesses and future work
It is clearly seen that ELM is faster and more accurate compared to SVM, however, there is a need for testing ELM for Big Data applications [51] to know its actual strength. We also need to test ELM on abdomen [52] and other bio-inspired imaging applications [53]. Also a comparison is needed to be made with contemporary Deep Learning [54] techniques. The experimental scope of work is limited to basic feature extraction algorithms only. In future, we intent to propose for application of better feature selection algorithms leveraging on principle component analysis or discriminate analysis or mutual information-based. This ensures that other feature extraction methods can be adapted along with feature selection methods, however this study is focused on benchmarking of SVM against ELM for liver tissue characterization, keeping the paradigms in comparison framework. To improve classifier performance, it is proposed that larger training dataset be provided for efficient classification. The high classification accuracy of basic ELM model in this study entails us to study other versions of ELM for stratification of US liver images in future.
Conclusions
The study presented a superior strategy for FDL stratification using Extreme Learning Machine (ELM) and benchmarked against SVM. The ELM was based on single layer feed forward neural network where input-to-hidden layer weights are randomly generated reducing computational cost and hidden-to-output weights were only trained. Due to simpler architecture and single pass, ELM was faster compared to SVM. Further, since the least square’s paradigm was adapted, hence more accurate with lesser number of features. The Symtosis system adapted four types of K-fold cross-validation (K = 2, 3, 5 and 10) protocols on three kinds of data sizes: S0-original, S4-four splits, S8-sixty four splits (a total of 12 cases) using 46 types of grayscale features derived using Gabor, GLCM and GRLM feature sets. Using the K10 cross-validation protocol on S8 data set, ELM showed an accuracy of 96.75% compared to 89.01% for SVM, and correspondingly, the AUC: 0.97 and 0.91, respectively. Further experiments also showed the mean reliability of 99% for ELM classifier, along with the mean speed improvement of 40% using ELM against SVM. We validated the symtosis system using two class biometric facial public data demonstrating an accuracy of 100%.
Change history
07 December 2017
The original version of this article unfortunately contained a mistake. The family name of Rui Tato Marinho was incorrectly spelled as Marinhoe.
References
Saverymuttu, S.H., Joseph, A.E., and Maxwell, J.D., Ultrasound scanning in the detection of hepatic fibrosis and steatosis. Br Med J (Clin Res Ed). 292(6512):13–15, 1986.
Mohamed, W.S., Mostafa, A.M., Mohamed, K.M., and Serwah, A.H., The epidemiology of nonalcoholic fatty liver disease in adults by Clark, Jeanne M MD, MPH. J. Clin. Gastroenterol. 40:S5–S10, 2006.
Wieckowska, A., and Feldstein, A.E., Nonalcoholic fatty liver disease in the pediatric population: A review. Current opinion in pediatrics. 17(5):636–641, 2005.
Ratziu, V., Charlotte, F., Heurtier, A., Gombert, S., Giral, P., Bruckert, E., Grimaldi, A., and Capron, F., Thierry Poynard, and LIDO study group, sampling variability of liver biopsy in nonalcoholic fatty liver disease. Gastroenterology. 128(7):1898–1906, 2005.
Cheung, R.C., Complications of liver biopsy, gastrointestinal emergencies, gastrointestinal emergencies. In: Tham, T.C.K., Collins, J.S.A., and Soetikno, R. (Eds.). Blackwell, West Sussex, UK, pp. 72–79, 2009.
Saadeh, S., Younossi, Z.M., Remer, E.M., Gramlich, T., Ong, J.P., Hurley, M., Mullen, K.D., Cooper, J.N., and Sheridan, M.J., The utility of radiological imaging in nonalcoholic fatty liver disease. Gastroenterology. 123(3):745–750, 2009.
Wang, D., Fang, Y., Hu, B., Cao, H., B-scan image feature extraction of fatty liver. In Internet Computing for Science and Engineering (ICICSE), 2012 Sixth International Conference. (2012) 188–192.
Yajima, Y., Ohta, K., Narui, T., Abe, R., Suzuki, H., and Ohtsuki, M., Ultrasonographical diagnosis of fatty liver: Significance of the liver-kidney contrast. Tohoku. J. Exp. Med. 139(1):43–50, 1983.
Mathiesen, U.L., Franzen, L.E., Aselius, H., Resjö, M., Jacobsson, L., Foberg, U., Frydén, A., and Bodemar, G., Increased liver echogenicity at ultrasound examination reflects degree of steatosis but not of fibrosis in asymptomatic patients with mild/moderate abnormalities of liver transaminases. Dig. Liver Dis. 34(7):516–522, 2002.
Mendler, M.H., Bouillet, P., Le Sidaner, A., Lavoine, E., Labrousse, F., Sautereau, D., and Pillegand, B., Dual-energy CT in the diagnosis and quantification of fatty liver: Limited clinical value in comparison to ultrasound scan and single-energy CT, with special reference to iron overload. J. Hepatol. 28(5):785–794, 1998.
Acharya, U., and Rajendra, J., Suri, data mining framework for fatty liver disease classification in ultrasound: A hybrid feature extraction paradigm. Med. Phys. 39(7):4255–4264, 2012.
Shensa, M.J., The discrete wavelet transform: The discrete wavelet transform: Wedding the atrous and Mallat algorithms. IEEE. Trans. Signal Process. 40(10, 1992):2464–2482.
Bruce, L.M., Koger, C.H., and Li, J., Dimensionality reduction of hyperspectral data using discrete wavelet transform feature extraction. IEEE. Trans. Geosci. Remote. Sens. 40(10):2331–2338, 2002.
Manjunath, B.S., and Ma, W.Y., Texture features for browsing and retrieval of image data. IEEE. Trans. Pattern. Anal. Mach. Intellig. 18(8):837–842, 1996.
Ishibuchi, H., Nakashima, T., and Murata, T., Performance evaluation of fuzzy classifier systems for multidimensional pattern classification problems. IEEE. Trans. Syst. Man. Cybern. Part B (Cybern.). 29(5):601–618, 1999.
Kuncheva L., Fuzzy classifier design, Springer Science & Business Media; 2000 Apr 26.
Uebele, V., Abe, S., and Lan, M.-S., A neural-network-based fuzzy classifier. IEEE. Trans. Syst. Man. Cybern. 25(2):353–361, 1995.
Antonini, M., Barlaud, M., Mathieu, P., and Daubechies, I., Image coding using wavelet transform. IEEE. Trans. Image. Process. 1(2):205–220, 1992.
Subramanya, M.B., Kumar, V., Mukherjee, S., and Saini, M., A CAD system for B-mode fatty liver ultrasound images using texture features. J. Med. Eng. Technol. 39(2):123–130, 2015.
Ma, H.Y., Zhou, Z., Wu, S., Wan, Y.L., and Tsui, P.H., A computer-aided diagnosis scheme for detection of fatty liver in vivo based on ultrasound kurtosis imaging. J. Med. Syst. 40(1):33, 2016.
Saba, L., Dey, N., Ashour, A.S., Samanta, S., Nath, S.S., Chakraborty, S., Sanches, J., Kumar, D., Marinho, R., and Suri, J.S., Automated stratification of liver disease in ultrasound: An online accurate feature classification paradigm. Comput. Methods. Programs. Biomed. 130:118–134, 2016.
Vapnik VN, An overview of statistical learning theory, IEEE transactions on neural networks (10) (1999) 988–999.
Huang, G.-B., and Zhu, Q.-Y., Chee-Kheong Sie extreme learning machine: Theory and applications. Neurocomputing. 70(1):489–501, 2006.
Huang, G.-B., Zhu, Q.-Y., and Siew, C.-K., Extreme learning machine: A new learning scheme of feedforward neural networks, in neural networks, 2004. Proceedings. IEEE. Int. Joint. Conference. 2(2004):985–990, 2004.
Rao, C.R., and Mitra, S.K., Generalized inverse of matrices and its applications. Wiley, New York, 1971.
Qayyum, A., MR spectroscopy of the liver: Principles and clinical applications. Radiographics. 29(6):1653–1664, 2009.
Kadah, Y.M., Farag, A.A., Zurada, J.M., Badawi, A.M., and Youssef, A.M., Classification algorithms for quantitative tissue characterization of diffuse liver disease from ultrasound images. IEEE Trans. Med. Imaging. 15(4):466–478, 1996.
Chorowski, J., Wang, J., and Zurada, J.M., Review and performance comparison of SVM-and ELM-based classifiers. Neurocomputing. 128:507–516, 2014.
Acharya, U.R., Sree, S.V., Krishnan, M.M., Molinari, F., ZieleŸnik, W., Bardales, R.H., Witkowska, A., and Suri, J.S., Computer-aided diagnostic system for detection of Hashimoto thyroiditis on ultrasound images from a polish population. J. Ultrasound. Med. 33:245–253, 2014.
Mohanty, A.K., Beberta, S., and Lenka, S.K., Classifying benign and malignant mass using GLCM and GLRLM based texture features from mammogram. Int. J. Eng. Res. Appl. 1(3):687–693, 2011.
Mohanaiah, P., and Sathyanarayana, L., GuruKumar, image texture feature extraction using GLCM approach. Int J. Sci. Res. Publ. 3(5):1, 2013.
Herman, P., Comparative analysis of spectral approaches to feature extraction for EEG-based motor imagery classification. IEEE Trans. Neural Syst. Rehabil. Eng. 16(4):317–326, 2008.
Anuradha, K., Statistical feature extraction to classify oral cancers. J.Glob. Res. Comput. Sci. 4(2):8–12, 2013.
Barbu, T., Gabor filter-based face recognition technique. Proc. Rom. Acad. 11(3):277–283, 2010.
MacAusland, Ross, The Moore-Penrose Inverse and Least Squares, Math 420. Advanced Topics in Linear Algebra. (2014).
Mirmehdi M, Handbook of texture analysis, Imperial College Press. 2008.
Acharya, U.R., Mookiah, M.R., Sree, S.V., Afonso, D., Sanches, J., Shafique, S., Nicolaides, A., Pedro, L.M., e Fernandes, J.F., and Suri, J.S., Atherosclerotic plaque tissue characterization in 2D ultrasound longitudinal carotid scans for automated classification: A paradigm for stroke risk assessment. Med. Biol. Eng. Comput. 51(5):513–523, 2013.
Acharya, R.U., Faust, O., Alvin, A.P., Sree, S.V., Molinari, F., Saba, L., Nicolaides, A., and Suri, J.S., Symptomatic vs. asymptomatic plaque classification in carotid ultrasound. J. Med. Syst. 36(3):1861–1871, 2012.
Acharya, U.R., Faust, O., Sree, S.V., Molinari, F., Saba, L., Nicolaides, A., and Suri, J.S., An accurate and generalized approach to plaque characterization in 346 carotid ultrasound scans. IEEE. Trans. Instrum. Meas. 61(4):1045–1053, 2012.
Shrivastava, V.K., Computer-aided diagnosis of psoriasis skin images with HOS, texture and color features: A first comparative study of its kind. Comput Meth. Programs Biomed. 126:98–109, 2016.
Shrivastava, V.K., Londhe, N.D., Sonawane, R.S., and Suri, J.S., Reliable and accurate psoriasis disease classification in dermatology images using comprehensive feature space in machine learning paradigm. Expert Syst. Appl. 42:6184–6195, 2015.
Douali, N., Abdennour, M., Sasso, M., Miette, V., Tordjman, J., Bedossa, P., Veyrie, N., Poitou, C., Aron-Wisnewsky, J., Clément, K. and Jaulent, M.C. Noninvasive diagnosis of nonalcoholic steatohepatitis disease based on clinical decision support system. MedInfo. 192:1178, 2013.
Vanderbeck, S., Bockhorst, J., Komorowski, R., Kleiner, D.E., and Gawrieh, S., Automatic classification of white regions in liver biopsies by supervised machine learning. Hum. Pathol. 45(4):785–792, 2014.
Liu, X., Song, J.L., Wang, S.H., Zhao, J.W., and Chen, Y.Q., Learning to diagnose cirrhosis with liver capsule guided ultrasound image classification. Sensors. 17(1):149, 2017.
LeCun, Y., Bengio, Y., and Hinton, G. Deep learning. Nature. 521(7553):436–444, 2015.
Huang, G.-B., Extreme learning machine for regression and multiclass classification. IEEE. Trans. Syst. Man. Cybern. Part B (Cybern). 42(2):513–529, 2012.
Zhu, Q.-Y., Evolutionary extreme learning machine. Pattern. Recogn. 38(10):1759–1763, 2005.
Li Mao, Lidong Zhang, Xingyang Liu, Chaofeng Li, Hong Yang, Improved Extreme Learning Machine and Its Application in Image Quality Assessment. Mathematical Problems in Engineering. (2014).
Tang, J., Deng, C., and Huang, G.-B., Extreme learning machine for multilayer perceptron. IEEE. Trans. Neural Netw. Learn. Syst. 27(4):809–821, 2016.
Demuth, H.B., Beale, M.H., De Jess, O. and Hagan, M.T., Neural network design, Martin Hagan, 2014.
El-Baz A, Suri JS, Big Data in Medical Imaging, CRC Press, 2018 (to appear).
El-Baz AS, Saba L, Suri JS. Abdomen and thoracic imaging. Springer, 2014.
El-Baz A, Gimel’farb G, Suri JS, Stochastic modeling for medical image analysis, CRC Press, 2015.
Esses SJ, Lu X, Zhao T, Shanbhogue K, Dane B, Bruno M, Chandarana H. Automated image quality evaluation of T2-weighted liver MRI utilizing deep learning architecture, Journal of Magnetic Resonance Imaging, (2017).
Acknowledgements
The authors of National Institute of Technology Goa, India would like to acknowledge Ministry of Human Resource department, Government of India and MediaLab Asia, Ministry of Electronics and Information Technology, Government of India for their kind support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
None of the authors have any conflict of interest.
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Data was collected after IRB approval by Instituto Superior Tecnico (IST), University of Lisbon, Portugal and written informed consent provided by all the patients.
Additional information
This article is part of the Topical Collection on Image & Signal Processing
A correction to this article is available online at https://doi.org/10.1007/s10916-017-0862-9.
Appendices
Appendix A: ELM mathematical framework and solution
Let there be H hidden layer neurons and N output neurons. Let each training sample of P images be denoted as (a j , l j ), where each input image denoted as a j = [a j1, a j2, …, a jm ]T ∈ R m and each output label is denoted as l j = [l j1, l j2, …, l jN ]T ∈ R N. The output vector is denoted as L = [l 1, l 2, …, l j , …l P ]T. Further, the P images and their ground truth labels are divided into two parts for training and testing which can be represented by \( {\boldsymbol{P}}_{trg}=\left({\boldsymbol{a}}_j^{trg},{\boldsymbol{l}}_j^{trg}\right) \) for training and \( {\boldsymbol{P}}_{tst}=\left({\boldsymbol{a}}_k^{tst},{\boldsymbol{l}}_k^{tst}\right) \). Similarly, output vector L is also divided into two sets L trg and L tst . Each input layer neuron is connected to all hidden layer neurons. Let each hidden layer weight be denoted as a vector w i = [w 1i , w 2i , …, w mi ]T. Each of the connections or weights from hidden-to-output layer are denoted as δ i = [δ i1, δ i2, …, δ iN ]T connecting i th hidden node to the output nodes. A standard SLFFNN can be modeled as given by:
where, g is the activation function and b is the bias. This equation is written in more compact form which is given by:
where,
\( \boldsymbol{\delta} ={\left[\begin{array}{c}\hfill {\delta}_1^T\hfill \\ {}\hfill \begin{array}{c}\hfill \vdots \hfill \\ {}\hfill {\delta}_i^T\hfill \\ {}\hfill \vdots \hfill \end{array}\hfill \\ {}\hfill {\delta}_H^T\hfill \end{array}\right]}_{H\times N}\mathrm{and}\ {\boldsymbol{L}}_{trg}={\left[\begin{array}{c}\hfill {\boldsymbol{l}}_1^{trg}\hfill \\ {}\hfill \begin{array}{c}\hfill \vdots \hfill \\ {}\hfill {\boldsymbol{l}}_{\boldsymbol{j}}^{trg}\hfill \\ {}\hfill \vdots \hfill \end{array}\hfill \\ {}\hfill {\boldsymbol{l}}_{\#{\boldsymbol{P}}_{trg}}^{trg}\hfill \end{array}\right]}_{\#{\boldsymbol{P}}_{\boldsymbol{trg}}\times N}.\mathrm{The}\ \mathrm{cost}\ \mathrm{function}\ \mathrm{of}\ \mathrm{ELM}\ \mathrm{is}\ \mathrm{given}\ \mathrm{as}: \)
The objective is to find the minimum δ which minimizes the cost function E. By using Eq. 5, the Eq. 6 can also be written as:
where \( \hat{\boldsymbol{\delta}} \) is the least squares solution of the Qδ = L trg . If the number of hidden nodes is equal to the number of training samples (#P trg = H), the matrix Q is square and invertible. Therefore, with random weights w i and bias b i the training samples can be approximated with zero error. However, in maximum cases, number of training samples is larger than number of hidden nodes. So, the smallest norm least squares solution of the linear system is given by:
Where, Q † is the Moore–Penrose [35] generalized inverse of matrix Q. Thus the smallest training error can be reached by:
The trained hidden-to-output layer weights are then used in Testing-phase as shown in Fig. 5, to test the performance of the Symtosis model using test dataset P tst .
Appendix B: Support vector machine
Support Vector Machine (SVM) is a kernel based classification technique based on the maximum margin classifier. It transforms the original input data to high-dimensional feature space and tries to find the hyper-plane which maximizes the distance between data points of distinct classes. We consider a binary classification task with the training dataset designated as {(a i , L i ), i = 1, 2, …, l} where a i є R q is the input data for i th training sample and L i є [−1, +1] are the equivalent target values, l designates total number of samples and q is the input space dimension. The SVM model can be represented in feature space by following equation:

where, ℵ(x) represents kernel function, b represents bias and  is a weight vector which is normal to the hyper-plane. The decision rule is mathematically represented in the Eq. (11):
is a weight vector which is normal to the hyper-plane. The decision rule is mathematically represented in the Eq. (11):

The non-linear kernel function finds the maximum margin hyper-plane,  between the classes in a feature space. To find optimal hyper-plane, the Eq. (12) is minimized subject to Eqs. (10) and (11).
between the classes in a feature space. To find optimal hyper-plane, the Eq. (12) is minimized subject to Eqs. (10) and (11).

where, ϑ represents the trade-off between error and margin and ξ is a slack variable. By using Lagrangian multipliers (α) in dual form, the Eq. (12) can be transformed into following optimization problem:
The final decision function is given by following equation:
The parameters ωand γ define the separating hyperplane. The most general kernel functions are:
Where, deg. is the degree of kernel in Eq. 18 and (.) denotes the dot product.
Appendix C: Feature extraction
Haralick texture (GLCM)
GLCM calculates the following features shown Table 7 from the co-occurrence matrix calculated from the image.
Run length texture
GRLM feature extraction algorithm calculate features from the run length matrix as shown in Table 8.
Appendix D: Symbols table
Appendix E: Results of ELM/SVM classifier for S4 and S8 dataset
Appendix F: Scientific validation
Scientific validation is always an integrated component of the system design. For validation, one needs to run another set of liver data sets whose results are known a priori. Since such a clinical data is hard to obtain, we use facial biometric data set to test the classification accuracy. We do acknowledge Dr. Libor Spacek of Department of Computer Science, University of Essex for providing data on biometric facial dataset, namely Face94 [42]. This dataset consisting of male and female faces was experimented for validation using ELM/SVM.
Face94 data set
We have conducted experiments to validate our results using Face94 data set. The Face94 data set consists 153 individual images with various expressions and poses seated at a fixed distance from camera. There are 2 classes, male and female and total number of images are 2660, of which 2260 are male images and 400 are female images. A subset of images is given in Fig. 10. Cross-validation protocol (K2, K3, K5 and K10) is also performed to check generalization. The validation results are shown in Table 12. It is seen ELM gives 100% accuracy across all cross-validation protocols.
Rights and permissions
About this article
Cite this article
Kuppili, V., Biswas, M., Sreekumar, A. et al. Extreme Learning Machine Framework for Risk Stratification of Fatty Liver Disease Using Ultrasound Tissue Characterization. J Med Syst 41, 152 (2017). https://doi.org/10.1007/s10916-017-0797-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10916-017-0797-1