Abstract
Purpose
Breast cancer (BC) is a heterogeneous disease characterised by variant biology, metabolic activity, and patient outcome. Glutamine availability for growth and progression of BC is important in several BC subtypes. This study aimed to evaluate the biological and prognostic role of the combined expression of key glutamine transporters, SLC1A5, SLC7A5, and SLC3A2 in BC with emphasis on the intrinsic molecular subtypes.
Methods
SLC1A5, SLC7A5, and SLC3A2 were assessed at the protein level, using immunohistochemistry on tissue microarrays constructed from a large well-characterised BC cohort (n = 2248). Patients were stratified into accredited clusters based on protein expression and correlated with clinicopathological parameters, molecular subtypes, and patient outcome.
Results
Clustering analysis of SLC1A5, SLC7A5, and SLC3A2 identified three clusters low SLCs (SLC1A5−/SLC7A5−/SLC3A2−), high SLC1A5 (SLC1A5+/SLC7A5−/SLC3A2−), and high SLCs (SLC1A5+/SLC7A5+/SLC3A2+) which had distinct correlations to known prognostic factors and patient outcome (p < 0.001). The key regulator of tumour cell metabolism, c-MYC, was significantly expressed in tumours in the high SLC cluster (p < 0.001). When different BC subtypes were considered, the association with the poor outcome was observed in the ER+ high proliferation/luminal B class only (p = 0.003). In multivariate analysis, SLC clusters were independent risk factor for shorter BC-specific survival (p = 0.001).
Conclusion
The co-operative expression of SLC1A5, SLC7A5, and SLC3A2 appears to play a role in the aggressive subclass of ER+ high proliferation/luminal BC, driven by c-MYC, and therefore have the potential to act as therapeutic targets, particularly in synergism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Altered metabolic profiles, one of the cancer hallmarks [1], has been substantiated for many years, and the significance of onco-metabolism and its exploitation for targeting cancers has recently attracted great attention. To maintain the unremitting proliferation, cancer cells reprogram their metabolism to provide nutrients required to support bioenergetics and biosynthetic pathways. Many cancer cells are highly reliant on amino acids for their growth, not only because they are precursors for nucleotide and protein synthesis, but also because they activate mammalian target of rapamycin complex1 (mTORC1) through nutrient signalling pathways which in turn regulates protein translation and cell growth [2].
Glutamine is the second primary metabolite to fuel cancer cell proliferation after glucose, as it is characterised by its ability to replenish the carbon pool of the mitochondrial tricarboxylic acid (TCA) cycle via α-ketoglutarate (α-KG) synthesis, in addition to its role for maintaining the redox balance [3]. Indeed, certain tumours may have glutamine-dependent cell growth or ‘glutamine addiction’ [4]. In breast cancer (BC), glutamine-dependent mechanisms can vary substantially between the molecular subtypes, as previous studies have revealed that the expression levels of glutamine metabolic enzymes, such as glutaminase (GLS), are mainly observed in triple-negative (TN), HER2+, and luminal B subtypes compared with luminal A tumours [5,6,7]. Specific genetic alterations may drive cancer cells to resist harsh metabolic stress conditions and nutrient deprivation including the upregulation of the glutamine transporters, solute carrier family 1 member 5 (SLC1A5), and solute carrier family 7 member 5 (SLC7A5) by the oncogene c-MYC [8, 9].
SLC7A5 is a sodium-independent amino acid transporter which imports large neutral amino acids such as leucine, isoleucine, tyrosine, and tryptophan, in exchange with intracellular glutamine [10]. It therefore functions in supplying amino acids to cancer cells as well as maintaining intracellular leucine which is considered a master regulator of mTORC1 signalling pathway [11, 12]. SLC7A5 requires a covalent association with the heavy chain of the membrane protein, solute carrier family 3 member 2 (SLC3A2), for its functional expression in plasma membrane [13]. We have previously described the potential utility of these two solute carriers as prognostic factors for the highly proliferative BC subtypes [14, 15]. Additionally, other studies have revealed the prognostic value of the co-operative expression of SLC7A5 and SLC3A2 in transitional cell carcinoma [16], squamous cell carcinoma of the lung [17], and pulmonary adenocarcinoma [18].
SLC1A5 is a sodium-dependent transporter which regulates the transport of neutral amino acids: alanine, serine, cysteine, threonine, and glutamine. It has been suggested that glutamine influx by SLC1A5 triggers essential amino acid entry via SLC7A5 exchanger. These exchange mechanisms balance the cytoplasmic pool of the amino acids without expending energy and therefore give a strong advantage for cancer cell proliferation [19]. SLC1A5 is over-expressed in several human neoplasms [7, 20,21,22]. In both esophageal squamous cell carcinoma [23] and laryngeal squamous cell carcinoma [24], co-expression of SLC7A5 and SLC1A5 is associated with a worse prognosis compared to expression of a single transporter, suggesting a potential functional coupling, at transportomic level, which supports tumour aggressiveness and progression.
While approximately 14 SLC members accept glutamine a substrate, SLC1A5 transporter and SLC7A5/SLC3A2 heterodimeric antiporter have a higher affinity for their substrates that confers them the potential to support the growth and survival of the aggressive tumour types.
In this study, we sought to understand the prognostic impact of the co-operative expression of the transporters, SLC1A5, SLC7A5, and SLC3A2, in a large BC cohort. We stratified patients in to clusters based on assessment of SLC1A5, SLC7A5, and SLC3A2 protein expression, to determine their clinicopathological and prognostic value with emphasis on the different molecular classes.
Materials and methods
Patient cohort
This study evaluated a well-characterised cohort of early-stage, primary operable, invasive BC patients aged ≤ 70 years. Patients (n = 2248) presented at Nottingham City Hospital between 1989 and 2006. Patient management was uniform and based on tumour characteristics by Nottingham Prognostic Index (NPI) and hormone receptor status. Patients within the NPI excellent prognostic group (score ≤ 3.4) received no adjuvant therapy, but those patients with NPI > 3.4 received tamoxifen if ER+ [± Goserelin (Zoladex) in case the patients were premenopausal]. Conversely, classical cyclophosphamide, methotrexate, and 5-fluorouracil (CMF) were used if the patients were ER− and fit enough to receive chemotherapy. None of the patients in this study received neoadjuvant therapy. Clinical history, tumour characteristics, information on therapy, and outcomes are prospectively maintained. Outcome data included development and time to distant metastasis (DM) and BC-specific survival (BCSS). The clinicopathological parameters for the BC series are summarised in (Supplementary Table 1).
Western blotting
Primary antibody specificity of SLC1A5 (1:250; HPA035240, Sigma-Aldrich, UK), SLC7A5 (1:200; EPR17573, Abcam, UK), and SLC3A2 (1:2000; HPA017980, Sigma-Aldrich, UK) were validated using western blotting in BC cell line lysates (American Type Culture Collection, Rockville, MD, USA) as previously described [14, 15]. A single band for all biomarkers was visualised at the correct predicted size (75 kDa, 40 kDa, and 80 kDa), respectively.
Tissue arrays and Immunohistochemistry
Tumour samples of 0.6 mm cores were arrayed [14, 25] and immunohistochemical (IHC) staining was performed on 4 µm TMA sections using the Novolink polymer detection system (Leica Biosystems, RE7150-K), as previously described [5].
Stained TMA sections were scanned using high-resolution digital images (NanoZoomer; Hamamatsu Photonics, Welwyn Garden City, UK), at ×20 magnification. Evaluation of staining for SLC1A5, SLC7A5, and SLC3A2 was based on a semi-quantitative assessment of cores’ digital images using a modified histochemical score (H-score) which includes an assessment of both the intensity and the percentage of stained cells [26]. Staining intensity was subjectively assessed as follows: 0, negative; 1, weak; 2, medium; 3, strong, and the percentage of the positively stained tumour cells was determined. The final H-score was calculated multiplying the percentage of positive cells (0–100) by the intensity (0–3), producing a total range of 0–300.
IHC staining and dichotomisation of the other biomarkers included in this study were as per previous publication [27]. ER and PgR positivity was defined as ≥ 1% staining. Immunoreactivity of HER2 was scored using standard HercepTest guidelines (Dako). Chromogenic in situ hybridisation (CISH) was used to quantify HER2 gene amplification in borderline cases using the HER2 FISH pharmDx™ plus HER2 CISH pharmDx™ kit (Dako) and was assessed according to the American Society of Clinical Oncology guidelines [28]. BC molecular subtypes were defined, based on tumour IHC profile and the Elston–Ellis [29] mitotic score as ER+/HER2− low proliferation (mitotic score 1), ER+/HER2− high proliferation (mitotic scores 2 and 3), HER2-positive class: HER2+ regardless of ER status, TN: ER−, PgR−, and HER2− [30].
Cluster analysis
Six cluster validity indices were calculated to determine the optimal number of clusters as previously described [32]. Two algorithms, partitioning around medoids (PAM) and K-means, were used to cluster tumours into the optimal number of clusters based on the H-score of SLC1A5, SLC7A5, and SLC3A2 as previously described [31].
K-means clustering is based on the initial setting of the cluster assignments, and for this study, we used a fixed initialisation obtained with hierarchical clustering. PAM is dependent on finding the k-representative objects (the so-called medoids) among the observations of the data set. K-clusters are constructed, after finding a set of k-medoids, through assigning each observation to its nearest medoid. Principle component analysis was applied to transform the original data space. Biplots were generated by plotting the points at their projected position on axes of the first and second principal components [32].
Statistical analysis
Statistical analysis was performed using SPSS 24.0 statistical software (SPSS, Inc., Chicago, IL, USA). The χ2 test was performed for inter-relationships between categorical variables. Survival curves were analysed by Kaplan–Meier with log rank test. This was performed with the BC-specific death, and those who died of other causes, alive, and lost to follow-up were censored. Cox’s proportional hazard method was performed for multivariate analysis to identify the independent prognostic factors. p values were adjusted using Bonferroni correction for multiple testing. A p value < 0.05 was considered significant. The study endpoints were 5-year BCSS or DM-free survival (DMFS). This study complied with reporting recommendations for tumour marker prognostic studies (REMARK) criteria [33].
This study was approved by the Nottingham Research Ethics Committee 2 under the title ‘Development of a molecular genetic classification of breast cancer’.
Results
Solute carrier protein expression in breast cancer
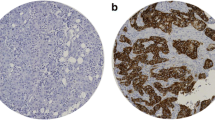
Expression of the three solute carriers was predominantly in the membrane of the invasive BC cells, with intensity levels varying from absent to high (Fig. 1a–c).
SLC protein expression in invasive breast cancer cores. a Low SLC cluster; from left to right, negative immunohistochemical (IHC) expression in SLC1A5, SLC7A5, and SLC3A2. b High SLC1A5 cluster; form left to right, positive IHC expression in SLC1A5 and negative IHC expression in SLC7A5 and SLC3A2. c High SLC cluster; from left to right, positive IHC expression in SLC1A5, SLC7A5, and SLC3A2
Clustering of SLC expression in breast cancer
To explore the prognostic value of the co-expression of SLC1A5, SLC7A5, and SLC3A2 in invasive BC, unsupervised clustering analysis was applied. Based on several indices, acceptable classification was applied by clustering the tumours into three groups. An appropriate separation between clusters was attained as indicated from the biplots (Supplementary Fig. 1a, b). The final three clusters were obtained from the overlap of the two clustering algorithms. The three clusters were characterised as follows: low SLCs (SLC1A5−/SLC7A5−/SLC3A2−), high SLC1A5 (SLC1A5+/SLC7A5−/SLC3A2−), and high SLCs (SLC1A5+/SLC7A5+/SLC3A2+) (Fig. 2).
SLC clusters correlate with the clinicopathological features and breast cancer subtypes
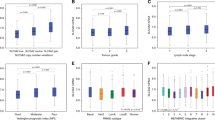
BCs in the high SLC and high SLC1A5 clusters were significantly associated with poor prognostic parameters including larger tumour size, higher histologic grade, poor NPI, and lymph node stage 3 (p < 0.001), whereas tumours in the Low SLC cluster were associated with good prognostic factors (Table 1).
When association with hormone receptor status was examined, both low SLC and high SLC1A5 clusters showed a significant association with hormone receptor (ER and PgR)-positive tumours, while the high SLC cluster included mainly tumours with hormone receptor-negative status (p < 0.001, Table 2). Tumours with HER2 positivity were more frequent in high SLC and high SLC1A5 clusters compared with low SLCs (p < 0.001, Table 2).
The prevalence of the three clusters in different BC subtypes was varied, where the low SLC cluster was primarily observed in ER+ low proliferation/luminal A tumours and the high SLC1A5 cluster was mainly found in the ER+ high proliferation/luminal B and HER2+ subtypes. In contrast, the poor prognostic TN class was strongly represented in patients having tumours with high SLCs (p < 0.001, Table 1).
High SLCs is associated with poor patient outcome in breast cancer
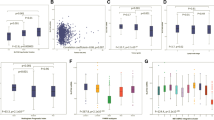
Clusters of the three solute carriers were significantly associated with patient outcome, where patients with tumours showing high SLCs had the worst survival compared to tumours with high SLC1A5, which had moderate outcome and low SLCs which showed the best survival (p < 0.001, Fig. 3a). Within the different molecular subtypes, the high SLC cluster was predictive of shorter BCSS but only in ER+ high proliferation/luminal B tumours (p = 0.003, Fig. 3b). There was no association between the SLC clusters and outcome in ER+ low proliferation/luminal A, HER2+, or TN tumours (Fig. 3c–e).
There was a similar observation regarding the association of SLC clusters with DMFS, where patients in the high SLC cluster had significantly shorter DMFS compared with the other clusters (p < 0.001, Fig. 4a). Similarly, the result is only observed in the ER+ high proliferation/luminal B class (p = 0.01, Fig. 4b). There was no association with the SLC clusters in predicting DMFS in ER+ low proliferation/luminal A, HER2+, or TN subtypes (Fig. 4c–e). In multivariate Cox regression analysis, the SLC clusters remained a predictor of shorter BCSS independent of tumour size, histologic grade, and lymph node stage in unselected tumours (p = 0.001, Table 3). The result remained significant in the ER+ high proliferation/luminal B tumours only (p = 0.01, Supplementary Table 2).
SLC clusters and other molecular biomarkers
The expression of c-MYC protein was primarily observed in the high SLC cluster (Table 2, p < 0.001). This observation remained significant only in ER+ high proliferation/luminal B tumours (p = 0.01, data not shown). Furthermore, patients in high SLC cluster with c-MYC-positive tumours showed shorter BCSS compared to those tumours that did not express c-MYC (Supplementary Fig. 2a, b). Low PIK3CA expression was predominant in low SLC cluster, while high SLC1A5 and high SLC clusters were mainly associated with positive PIK3CA expression (Table 2, p < 0.001). The association with poor outcome was only observed in patients with PIK3CA-positive expression and not with the cases who did not express this protein (Supplementary Fig. 3a, b). The proliferation marker, Ki67, was also significantly expressed in high SLC1A5 and high SLC clusters (Table 2, p < 0.001) and patients belonging to these clusters showed shorter BCSS when accompanied with the positive expression of Ki67 (Supplementary Fig. 4a, b). Conversely, mTORC1 was mainly observed in low SLC cluster (Table 2, p < 0.001) and the association with poor outcome showed no difference between mTORC+ and mTORC− tumours (Supplementary Fig. 5a, b).
Discussion
BC is a heterogeneous disease with various molecular subtypes differing in terms of tumour biology and patient outcome [34]. In addition, different subtypes exhibit disparity in their metabolic pathways, where ER− tumours can be distinguished from the ER+ based on their metabolic signatures. ER+/luminal tumours which comprise the majority of BC cases (approximately 75%) remain heterogeneous in terms of mortality rates and disease prognosis [35, 36]. There is thus a clear need to understand the biology of BC which then can help in exploring methods of prognostic and predictive stratification as well as identifying potential novel therapeutic targets.
Currently, the prognostic stratification of invasive BC is mainly based on the known clinicopathological parameters, such as tumour grade, stage, and hormone receptor status. Although a large number of biomarkers have been suggested to be prognostic markers in this tumour, other studies have demonstrated that combination of prognostic indicators is superior to recruiting single markers [5, 37, 38]. Therefore, applying cluster analysis is not only competent for identifying biologically relevant groups of biomarkers, but it can also allow more detailed prognostication and potentially provide optimal treatment strategies with the objective of improving patient outcome and quality of life.
Cancer cells, including BC, require a constant supply of nutrients to support their growth and proliferation. To do so, they need to elevate the expression levels of nutrient transporters in the plasma membrane, a process which has a strong relationship with tumorigenesis and drug resistance [39]. We have recently demonstrated that the solute carriers SLC7A5 and SLC3A2 are associated with poor prognosis particularly in the ER+ high proliferation subtype of BC [14, 15] and the co-expression of SLC7A5 and SLC3A2 in TNBC has been previously established [40]. However, studies that address the prognostic significance of the co-operative expression of those two proteins alongside the glutamine transporter SLC1A5 in BC, and their influence in the other molecular subtypes, remain limited.
This study has revealed differential expression levels of the three solute carriers across a large BC cohort which can be used to stratify the patients into three clusters, where the low SLC cluster did not show expression of any transporter proteins, high SLC1A5 cluster was derived by the high expression of SLC1A5 only, and the high SLC cluster included tumours with high expression of all three transporters. The prognostic value of the three clusters was varied, where patients with tumours in the high SLC1A5 and high SLC clusters were associated with poor clinicopathological parameters, including larger tumour size, higher grade, and poor NPI. In contrast, tumours with good prognostic characteristics were predominantly in the low SLC cluster. It was further established that patients with tumours in the high SLC cluster had the poorest outcome compared to those cases in the low SLC or high SLC1A5 clusters. This is anticipated, as the high SLC cluster embraces the overexpression of the three solute carriers together, where SLC1A5 mediates uptake of glutamine, while the SLC7A5-SLC3A2 heterodimeric complex uses the intracellular glutamine concentrations to adjust the essential amino acid cytoplasmic pool for metabolic demands and the subsequent activation of mTORC1 signalling pathway, suggesting a system of functional coupling between these transporters to confer the more aggressive character to BC cells. It is also known that the SLC7A5-SLC3A2 complex is a thyroid hormone transporter, and recently, Søgaard et al. have identified an association between hyperthyroidism and increasing BC risk [41]. It is noteworthy that SLC7A5 and SLC3A2 can be expressed independently. Furthermore, the single expression of SLC3A2, but not SLC7A5, can predict the poor patient outcome (data not shown). This is can be attributed to the extra role played by SLC3A2 in favouring cancer growth, as it can modulates integrin-induced signal transduction which derives malignant tumour cells’ behaviour including cell spreading and migration [42].
With respect to the ER+ BC subtypes, the three solute carriers were highly expressed in a subset of the highly proliferative ER+/luminal B tumours and were related with poor patient outcome and shorter DMFS. We noted that the single expression of SLC1A5, the driver of high SLC1A5 cluster, was not enough to predict the shorter BCSS in this subtype. To do so, it requires the concurrent expression of SLC7A5 and SLC3A2. This is most likely due to their heavier energy and nutrient requirements for cell survival and proliferation. Furthermore, we and others have previously demonstrated that SLC7A5 and SLC3A2 are significantly correlated with the proliferative marker Ki67 in thymic carcinomas [43], tongue cancer [44], neuroendocrine carcinoma of the lung [45], non-small cell lung cancer [46], and BC [14, 15, 40]. In this study, Ki67 was mainly expressed in tumours of high SLC1A5 and high SLC clusters, indicating that the studied solute carriers can support cancer cell proliferation, the main characteristic which distinguishes luminal B from luminal A subtypes.
The BC subtypes, TN and HER2+, were also commonly represented in cases demonstrating the high SLC cluster, in concordance with Kim et al. [6] and Furuya et al. [40], who studied the expression levels of SLC1A5 and SLC7A5/SLC3A2, respectively, within the BC molecular subtypes. However, neither the TN nor the HER2+ subtype showed a significant association between the SLC clusters and patient outcome.
The oncogene MYC has a robust influence in tumour cell metabolism through enhancing the uptake and utilisation of the necessary nutrients via upregulation of cellular transporters and enzymes supporting tumour growth and giving rise to aggressive cancer phenotypes [9, 47]. Previous studies have showed that MYC controls the expression of SLC1A5 and SLC7A5 via binding to specific promoters and enhancing protein transcription [8, 48]. Upon glutamine deprivation, activation factor 4 (ATF4), in co-ordination with MYC, activates the expression of both transporters directly to maintain intracellular levels of amino acids [7, 49, 50]. In addition, SLC7A5 can control MYC expression and constitute a positive feedback loop mechanism to encourage essential amino acid transport and tumorigenesis [51, 52]. The current study revealed that MYC not only regulates the concurrent expression of the three studied solute carriers, but it also conferred poorer outcome, in patients with tumours in the high SLC cluster, compared to lack of its expression, concluding that this cluster might play a role in endocrine therapy resistance as previous work by our group showing that MYC overexpression confers resistance to endocrine therapy [27].
Mutations in PIK3CA are known to have a relation to metabolic alterations in cancer cells. Hao et al. have demonstrated that PIK3CA mutations in colorectal cancer increase glutamine dependency by upregulating glutamate pyruvate transaminase 2 (GPT2), which enhances glutamine conversion to αkG to replenish TCA and assist ATP production through an ATF4-dependent mechanism [53]. Recent work carried out by our group has shown that PIK3CA is the driver for glutaminase 1 expression, which converts glutamine to glutamate, in the luminal B tumours (data unpublished). In this study, we showed that PIK3CA is primarily over-expressed in the high SLC1A5 and high SLC clusters and is therefore a potential driver of shorter BCSS in these clusters.
Previous research has detailed the role of amino acids in activating mTORC1, which in turn controls cell proliferation and prevent apoptosis in cancer cells [2]. This study, however, showed that mTORC1 was mainly expressed in low SLC cluster and its expression did not make a difference in patient outcome compared to its absence. This can be explained by a previous study, which reported that phosphorylation of mTORC1 at ser (2448), which included in this study, is stimulated by growth factors and it was mutually exclusive with mTORC1 phosphorylated at Thr (2446), which is regulated by amino acids [54]. In addition, Figueiredo et al. concluded that mTORC1 phosphorylated at Ser (2448) is an inadequate measure for mTORC1 kinase activity and the alternative should be the downstream and effector proteins, such as p70S6K [55].
With the increasing number of treatment strategies available for cancer patients, it is critical to find effective approaches which can support personalisation of care and allow tailored treatment plan for BC patients, taking into consideration its molecular subtypes. Inhibition of SLC1A5, via the inhibitor l-γ-glutamyl-p-nitroanilide (GPNA), decreases the growth of various cancer cell lines including TNBC [7, 56, 57]. Blocking SLC7A5, using 2-aminobicyclo-(2,2,1)-heptane-2-carboxylic acid (BCH), efficiently decreased colony formation of MDA-MB-231 TNBC cells [58]. In addition, the selective SLC7A5 inhibitor (JPH203/KYT-0353) is undergoing Phase I clinical trials (UMIN000016546) as a novel adjuvant treatment approach for solid tumours including BC [59, 60]. Although the consequences of blocking the action of these solute carriers in ER+ high proliferation tumours remain undetermined, our data suggest that the synergistic lethality targeting the three solute carriers may prove to be more effective in BC treatment.
Conclusion
This study reveals that the co-operative expression of SLC1A5, SLC7A5, and SLC3A2 was associated with the poor prognostic characteristics and poor patient outcome particularly in the ER+ high proliferation/luminal B subtype. Therefore, elaboration in understanding the biological diversity of BC is essential, with the linked development of classification strategies suitable for clinical use. Further functional assessment is essential to reveal the specific role played by solute carrier co-expression in the highly proliferative BC subclass of ER+ disease.
References
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674
Bar-Peled L, Sabatini DM (2014) Regulation of mTORC1 by amino acids. Trends Cell Biol 24(7):400–406. https://doi.org/10.1016/j.tcb.2014.03.003
DeBerardinis RJ, Cheng T (2010) Q’s next: the diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene 29(3):313–324. https://doi.org/10.1038/onc.2009.358
Wise DR, Thompson CB (2010) Glutamine addiction: a new therapeutic target in cancer. Trends Biochem Sci 35(8):427–433. https://doi.org/10.1016/j.tibs.2010.05.003
Craze ML, Cheung H, Jewa N, Coimbra ND, Soria D, El-Ansari R, Aleskandarany MA, Cheng KW, Diez-Rodriguez M, Nolan CC (2017) MYC regulation of Glutamine-Proline regulatory axis is key in Luminal B breast cancer. Br J Cancer. https://doi.org/10.1038/bjc.2017.387
Kim S, Kim DH, Jung WH, Koo JS (2013) Expression of glutamine metabolism-related proteins according to molecular subtype of breast cancer. Endocr Relat Cancer 20(3):339–348. https://doi.org/10.1530/erc-12-0398
van Geldermalsen M, Wang Q, Nagarajah R, Marshall AD, Thoeng A, Gao D, Ritchie W, Feng Y, Bailey CG, Deng N, Harvey K, Beith JM, Selinger CI, O’Toole SA, Rasko JE, Holst J (2016) ASCT2/SLC1A5 controls glutamine uptake and tumour growth in triple-negative basal-like breast cancer. Oncogene 35(24):3201–3208. https://doi.org/10.1038/onc.2015.381
Hayashi K, Jutabha P, Endou H, Anzai N (2012) c-Myc is crucial for the expression of LAT1 in MIA Paca-2 human pancreatic cancer cells. Oncol Rep 28(3):862–866. https://doi.org/10.3892/or.2012.1878
Wise DR, DeBerardinis RJ, Mancuso A, Sayed N, Zhang XY, Pfeiffer HK, Nissim I, Daikhin E, Yudkoff M, McMahon SB, Thompson CB (2008) Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc Natl Acad Sci USA 105(48):18782–18787. https://doi.org/10.1073/pnas.0810199105
Kanai Y, Segawa H, Miyamoto K, Uchino H, Takeda E, Endou H (1998) Expression cloning and characterization of a transporter for large neutral amino acids activated by the heavy chain of 4F2 antigen (CD98). J Biol Chem 273(37):23629–23632
Jewell JL, Kim YC, Russell RC, Yu FX, Park HW, Plouffe SW, Tagliabracci VS, Guan KL (2015) Metabolism. Differential regulation of mTORC1 by leucine and glutamine. Science 347(6218):194–198. https://doi.org/10.1126/science.1259472
Kimball SR (2001) Regulation of translation initiation by amino acids in eukaryotic cells. Prog Mol Subcell Biol 26:155–184
Yanagida O, Kanai Y, Chairoungdua A, Kim DK, Segawa H, Nii T, Cha SH, Matsuo H, Fukushima J, Fukasawa Y, Tani Y, Taketani Y, Uchino H, Kim JY, Inatomi J, Okayasu I, Miyamoto K, Takeda E, Goya T, Endou H (2001) Human L-type amino acid transporter 1 (LAT1): characterization of function and expression in tumor cell lines. Biochim Biophys Acta 1514(2):291–302
El Ansari R, Craze ML, Diez-Rodriguez M, Nolan CC, Ellis IO, Rakha EA, Green AR (2018) The multifunctional solute carrier 3A2 (SLC3A2) confers a poor prognosis in the highly proliferative breast cancer subtypes. Br J Cancer. https://doi.org/10.1038/s41416-018-0038-5
El Ansari R, Craze ML, Miligy I, Diez-Rodriguez M, Nolan CC, Ellis IO, Rakha EA, Green AR (2018) The amino acid transporter SLC7A5 confers a poor prognosis in the highly proliferative breast cancer subtypes and is a key therapeutic target in luminal B tumours. Breast Cancer Res 20(1):21. https://doi.org/10.1186/s13058-018-0946-6
Nakanishi K, Ogata S, Matsuo H, Kanai Y, Endou H, Hiroi S, Tominaga S, Aida S, Kasamatsu H, Kawai T (2007) Expression of LAT1 predicts risk of progression of transitional cell carcinoma of the upper urinary tract. Virchows Arch Int J Pathol 451(3):681–690. https://doi.org/10.1007/s00428-007-0457-9
Kaira K, Oriuchi N, Imai H, Shimizu K, Yanagitani N, Sunaga N, Hisada T, Ishizuka T, Kanai Y, Endou H, Nakajima T, Mori M (2009) Prognostic significance of L-type amino acid transporter 1 (LAT1) and 4F2 heavy chain (CD98) expression in early stage squamous cell carcinoma of the lung. Cancer Sci 100(2):248–254. https://doi.org/10.1111/j.1349-7006.2008.01029.x
Kaira K, Oriuchi N, Imai H, Shimizu K, Yanagitani N, Sunaga N, Hisada T, Ishizuka T, Kanai Y, Nakajima T, Mori M (2009) Prognostic significance of L-type amino acid transporter 1 (LAT1) and 4F2 heavy chain (CD98) expression in stage I pulmonary adenocarcinoma. Lung Cancer (Amst Neth) 66(1):120–126. https://doi.org/10.1016/j.lungcan.2008.12.015
Bhutia YD, Babu E, Ramachandran S, Ganapathy V (2015) Amino acid transporters in cancer and their relevance to “glutamine addiction”: novel targets for the design of a new class of anticancer drugs. Cancer Res 75(9):1782–1788. https://doi.org/10.1158/0008-5472.can-14-3745
Fuchs BC, Finger RE, Onan MC, Bode BP (2007) ASCT2 silencing regulates mammalian target-of-rapamycin growth and survival signaling in human hepatoma cells. Am J Physiol Cell Physiol 293(1):C55–C63. https://doi.org/10.1152/ajpcell.00330.2006
Li R, Younes M, Frolov A, Wheeler TM, Scardino P, Ohori M, Ayala G (2003) Expression of neutral amino acid transporter ASCT2 in human prostate. Anticancer Res 23(4):3413–3418
Witte D, Ali N, Carlson N, Younes M (2002) Overexpression of the neutral amino acid transporter ASCT2 in human colorectal adenocarcinoma. Anticancer Res 22(5):2555–2557
Honjo H, Kaira K, Miyazaki T, Yokobori T, Kanai Y, Nagamori S, Oyama T, Asao T, Kuwano H (2016) Clinicopathological significance of LAT1 and ASCT2 in patients with surgically resected esophageal squamous cell carcinoma. J Surg Oncol 113(4):381–389. https://doi.org/10.1002/jso.24160
Nikkuni O, Kaira K, Toyoda M, Shino M, Sakakura K, Takahashi K, Tominaga H, Oriuchi N, Suzuki M, Iijima M, Asao T, Nishiyama M, Nagamori S, Kanai Y, Oyama T, Chikamatsu K (2015) Expression of amino acid transporters (LAT1 and ASCT2) in patients with Stage III/IV laryngeal squamous cell carcinoma. Pathol Oncol Res 21(4):1175–1181. https://doi.org/10.1007/s12253-015-9954-3
Abd El-Rehim DM, Ball G, Pinder SE, Rakha E, Paish C, Robertson JF, Macmillan D, Blamey RW, Ellis IO (2005) High-throughput protein expression analysis using tissue microarray technology of a large well-characterised series identifies biologically distinct classes of breast cancer confirming recent cDNA expression analyses. Int J Cancer 116(3):340–350. https://doi.org/10.1002/ijc.21004
McCarty KS Jr, McCarty KS, Sr (1984) Histochemical approaches to steroid receptor analyses. Semin Diagn Pathol 1(4):297–308
Green AR, Aleskandarany MA, Agarwal D, Elsheikh S, Nolan CC, Diez-Rodriguez M, Macmillan RD, Ball GR, Caldas C, Madhusudan S, Ellis IO, Rakha EA (2016) MYC functions are specific in biological subtypes of breast cancer and confers resistance to endocrine therapy in luminal tumours. Br J Cancer 114(8):917–928. https://doi.org/10.1038/bjc.2016.46
Wolff AC, Hammond ME, Hicks DG, Dowsett M, McShane LM, Allison KH, Allred DC, Bartlett JM, Bilous M, Fitzgibbons P, Hanna W, Jenkins RB, Mangu PB, Paik S, Perez EA, Press MF, Spears PA, Vance GH, Viale G, Hayes DF (2013) Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol Off J Am Soc Clin Oncol 31(31):3997–4013. https://doi.org/10.1200/jco.2013.50.9984
Elston CW, Ellis IO (2002) Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. C. W. Elston & I. O. Ellis. Histopathology 1991; 19;403–410. Histopathology 41(3a):151–152, discussion 152–153
Senkus E, Kyriakides S, Ohno S, Penault-Llorca F, Poortmans P, Rutgers E, Zackrisson S, Cardoso F (2015) Primary breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol Off J Eur Soc Med Oncol 26(Suppl 5):v8–30. https://doi.org/10.1093/annonc/mdv298
Soria D, Garibaldi JM, Ambrogi F, Green AR, Powe D, Rakha E, Macmillan RD, Blamey RW, Ball G, Lisboa PJ, Etchells TA, Boracchi P, Biganzoli E, Ellis IO (2010) A methodology to identify consensus classes from clustering algorithms applied to immunohistochemical data from breast cancer patients. Comput Biol Med 40(3):318–330. https://doi.org/10.1016/j.compbiomed.2010.01.003
Edwards J (1991) A users guide to principal components. Wiley series in probability and mathematical statistics. Wiley, New York
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2005) REporting recommendations for tumour MARKer prognostic studies (REMARK). Br J Cancer 93(4):387–391. https://doi.org/10.1038/sj.bjc.6602678
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752. https://doi.org/10.1038/35021093
Dawson SJ, Rueda OM, Aparicio S, Caldas C (2013) A new genome-driven integrated classification of breast cancer and its implications. EMBO J 32(5):617–628. https://doi.org/10.1038/emboj.2013.19
Rakha EA, El-Sayed ME, Green AR, Paish EC, Powe DG, Gee J, Nicholson RI, Lee AH, Robertson JF, Ellis IO (2007) Biologic and clinical characteristics of breast cancer with single hormone receptor positive phenotype. J Clin Oncol Off J Am Soc Clin Oncol 25(30):4772–4778. https://doi.org/10.1200/jco.2007.12.2747
Makretsov NA, Huntsman DG, Nielsen TO, Yorida E, Peacock M, Cheang MC, Dunn SE, Hayes M, van de Rijn M, Bajdik C, Gilks CB (2004) Hierarchical clustering analysis of tissue microarray immunostaining data identifies prognostically significant groups of breast carcinoma. Clin Cancer Res Off J Am Assoc Cancer Res 10(18 Pt 1):6143–6151. https://doi.org/10.1158/1078-0432.ccr-04-0429
Tsutsui S, Ohno S, Murakami S, Kataoka A, Kinoshita J, Hachitanda Y (2003) Prognostic significance of the coexpression of p53 protein and c-erbB2 in breast cancer. Am J Surg 185(2):165–167
Januchowski R, Zawierucha P, Andrzejewska M, Rucinski M, Zabel M (2013) Microarray-based detection and expression analysis of ABC and SLC transporters in drug-resistant ovarian cancer cell lines. Biomed Pharmacother 67(3):240–245. https://doi.org/10.1016/j.biopha.2012.11.011
Furuya M, Horiguchi J, Nakajima H, Kanai Y, Oyama T (2012) Correlation of L-type amino acid transporter 1 and CD98 expression with triple negative breast cancer prognosis. Cancer Sci 103(2):382–389. https://doi.org/10.1111/j.1349-7006.2011.02151.x
Sogaard M, Farkas DK, Ehrenstein V, Jorgensen JO, Dekkers OM, Sorensen HT (2016) Hypothyroidism and hyperthyroidism and breast cancer risk: a nationwide cohort study. Eur J Endocrinol 174(4):409–414. https://doi.org/10.1530/eje-15-0989
Poettler M, Unseld M, Braemswig K, Haitel A, Zielinski CC, Prager GW (2013) CD98hc (SLC3A2) drives integrin-dependent renal cancer cell behavior. Mol Cancer 12:169. https://doi.org/10.1186/1476-4598-12-169
Kaira K, Oriuchi N, Imai H, Shimizu K, Yanagitani N, Sunaga N, Hisada T, Ishizuka T, Kanai Y, Endou H, Nakajima T, Mori M (2009) L-type amino acid transporter 1 (LAT1) is frequently expressed in thymic carcinomas but is absent in thymomas. J Surg Oncol 99(7):433–438. https://doi.org/10.1002/jso.21277
Toyoda M, Kaira K, Ohshima Y, Ishioka NS, Shino M, Sakakura K, Takayasu Y, Takahashi K, Tominaga H, Oriuchi N, Nagamori S, Kanai Y, Oyama T, Chikamatsu K (2014) Prognostic significance of amino-acid transporter expression (LAT1, ASCT2, and xCT) in surgically resected tongue cancer. Br J Cancer 110(10):2506–2513. https://doi.org/10.1038/bjc.2014.178
Kaira K, Oriuchi N, Imai H, Shimizu K, Yanagitani N, Sunaga N, Hisada T, Kawashima O, Iijima H, Ishizuka T, Kanai Y, Endou H, Nakajima T, Mori M (2008) Expression of L-type amino acid transporter 1 (LAT1) in neuroendocrine tumors of the lung. Pathol Res Pract 204(8):553–561. https://doi.org/10.1016/j.prp.2008.02.003
Kaira K, Oriuchi N, Imai H, Shimizu K, Yanagitani N, Sunaga N, Hisada T, Kawashima O, Kamide Y, Ishizuka T, Kanai Y, Nakajima T, Mori M (2010) Prognostic significance of L-type amino acid transporter 1 (LAT1) and 4F2 heavy chain (CD98) expression in surgically resectable stage III non-small cell lung cancer. Exp Ther Med 1(5):799–808. https://doi.org/10.3892/etm.2010.117
El Ansari R, McIntyre A, Craze ML, Ellis IO, Rakha EA, Green AR (2018) Altered glutamine metabolism in breast cancer; subtype dependencies and alternative adaptations. Histopathology 72(2):183–190. https://doi.org/10.1111/his.13334
Gao P, Tchernyshyov I, Chang TC, Lee YS, Kita K, Ochi T, Zeller KI, De Marzo AM, Van Eyk JE, Mendell JT, Dang CV (2009) c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458(7239):762–765. https://doi.org/10.1038/nature07823
Adams CM (2007) Role of the transcription factor ATF4 in the anabolic actions of insulin and the anti-anabolic actions of glucocorticoids. J Biol Chem 282(23):16744–16753. https://doi.org/10.1074/jbc.M610510200
Qing G, Li B, Vu A, Skuli N, Walton ZE, Liu X, Mayes PA, Wise DR, Thompson CB, Maris JM, Hogarty MD, Simon MC (2012) ATF4 regulates MYC-mediated neuroblastoma cell death upon glutamine deprivation. Cancer Cell 22(5):631–644. https://doi.org/10.1016/j.ccr.2012.09.021
Yue M, Jiang J, Gao P, Liu H, Qing G (2017) Oncogenic MYC activates a feedforward regulatory loop promoting essential amino acid metabolism and tumorigenesis. Cell Rep 21(13):3819–3832. https://doi.org/10.1016/j.celrep.2017.12.002
Sinclair LV, Rolf J, Emslie E, Shi YB, Taylor PM, Cantrell DA (2013) Control of amino-acid transport by antigen receptors coordinates the metabolic reprogramming essential for T cell differentiation. Nat Immunol 14(5):500–508. https://doi.org/10.1038/ni.2556
Hao Y, Samuels Y, Li Q, Krokowski D, Guan BJ, Wang C, Jin Z, Dong B, Cao B, Feng X, Xiang M, Xu C, Fink S, Meropol NJ, Xu Y, Conlon RA, Markowitz S, Kinzler KW, Velculescu VE, Brunengraber H, Willis JE, LaFramboise T, Hatzoglou M, Zhang GF, Vogelstein B, Wang Z (2016) Oncogenic PIK3CA mutations reprogram glutamine metabolism in colorectal cancer. Nat Commun 7:11971. https://doi.org/10.1038/ncomms11971
Cheng SW, Fryer LG, Carling D, Shepherd PR (2004) Thr2446 is a novel mammalian target of rapamycin (mTOR) phosphorylation site regulated by nutrient status. J Biol Chem 279(16):15719–15722. https://doi.org/10.1074/jbc.C300534200
Figueiredo VC, Markworth JF, Cameron-Smith D (2017) Considerations on mTOR regulation at serine 2448: implications for muscle metabolism studies. Cell Mol Life Sci 74(14):2537–2545. https://doi.org/10.1007/s00018-017-2481-5
Hassanein M, Qian J, Hoeksema MD, Wang J, Jacobovitz M, Ji X, Harris FT, Harris BK, Boyd KL, Chen H, Eisenberg R, Massion PP (2015) Targeting SLC1a5-mediated glutamine dependence in non-small cell lung cancer. Int J Cancer 137(7):1587–1597. https://doi.org/10.1002/ijc.29535
Wang Q, Hardie RA, Hoy AJ, van Geldermalsen M, Gao D, Fazli L, Sadowski MC, Balaban S, Schreuder M, Nagarajah R, Wong JJ, Metierre C, Pinello N, Otte NJ, Lehman ML, Gleave M, Nelson CC, Bailey CG, Ritchie W, Rasko JE, Holst J (2015) Targeting ASCT2-mediated glutamine uptake blocks prostate cancer growth and tumour development. J Pathol 236(3):278–289. https://doi.org/10.1002/path.4518
Liang Z, Cho HT, Williams L, Zhu A, Liang K, Huang K, Wu H, Jiang C, Hong S, Crowe R, Goodman MM, Shim H (2011) Potential biomarker of L-type amino acid transporter 1 in breast cancer progression. Nucl Med Mol Imaging 45(2):93–102. https://doi.org/10.1007/s13139-010-0068-2
Dickens D, Chiduza GN, Wright GS, Pirmohamed M, Antonyuk SV, Hasnain SS (2017) Modulation of LAT1 (SLC7A5) transporter activity and stability by membrane cholesterol. Sci Rep 7:43580. https://doi.org/10.1038/srep43580
Oda K, Hosoda N, Endo H, Saito K, Tsujihara K, Yamamura M, Sakata T, Anzai N, Wempe MF, Kanai Y, Endou H (2010) L-type amino acid transporter 1 inhibitors inhibit tumor cell growth. Cancer Sci 101(1):173–179. https://doi.org/10.1111/j.1349-7006.2009.01386.x
Acknowledgements
We thank the Nottingham Health Science Biobank and Breast Cancer Now Tissue Bank for the provision of tissue samples.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
El-Ansari, R., Craze, M.L., Alfarsi, L. et al. The combined expression of solute carriers is associated with a poor prognosis in highly proliferative ER+ breast cancer. Breast Cancer Res Treat 175, 27–38 (2019). https://doi.org/10.1007/s10549-018-05111-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-018-05111-w