Abstract
The mammalian target of rapamycin (mTOR) complex exerts a pivotal role in protein anabolism and cell growth. Despite its importance, few studies adequately address the complexity of phosphorylation of the mTOR protein itself to enable conclusions to be drawn on the extent of kinase activation following this event. In particular, a large number of studies in the skeletal muscle biology field have measured Serine 2448 (Ser2448) phosphorylation as a proxy of mTOR kinase activity. However, the evidence to be described is that Ser2448 is not a measure of mTOR kinase activity nor is a target of AKT activity and instead has inhibitory effects on the kinase that is targeted by the downstream effector p70S6K in a negative feedback loop mechanism, which is evident when revisiting muscle research studies. It is proposed that this residue modification acts as a fine-tuning mechanism that has been gained during vertebrate evolution. In conclusion, it is recommended that Ser2448 is an inadequate measure and that preferential analysis of mTORC1 activation should focus on the downstream and effector proteins, including p70S6K and 4E-BP1, along mTOR protein partners that bind to mTOR protein to form the active complexes 1 and 2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The mammalian Target of Rapamycin (mTOR) pathway is a central converging point of a diverse range of cellular growth signals, including hormones/growth factors, nutrients, energy status, and mechanical strain. The mTOR protein associates with other proteins to form mTOR complexes 1 and 2 (mTORC1 and mTORC2). mTORC1 is characterized by its association with Raptor (regulatory associated protein of mTOR), whilst mTORC2 is associated with Rictor (rapamycin insensitive companion of mTOR) [44]. Raptor and Rictor are both binding and scaffold proteins that direct mTOR activity towards specific substrates [49, 60, 80], therefore, providing specificity in determining cellular functionality. In the case of mTORC1, mTOR specificity is directed to p70 S6 Kinase (p70S6K) and 4E-BP1 [16, 37], which makes mTORC1 a central node regulating protein synthesis, cell size, and proliferation [36, 37, 98] that is highly responsive to nutrients and energy stress [7, 60]. On the other hand, mTORC2 directly phosphorylates AKT [81] regulating aspects of cell survival and the organization of actin cytoskeleton [80, 95].
4E-BP1 and p70S6K possess the conserved TOS (TOR signaling) motif that is essential for activation by mTOR [82, 83]; although mTOR exhibits differing kinase activity towards these targets [16, 20, 21]. Whilst mTOR-dependent phosphorylation of downstream effectors is a well-characterized signaling event [16, 82], there is frequent uncertainty regarding the functional significance of phosphorylation of the mTOR protein itself. The mTOR protein has multiple residues that can undergo phosphorylation. Phospho-specific antibodies have been developed and are frequently used in experimental and clinical studies. Antibodies that recognize the Ser2481 and Ser2448 phosphorylation residues of mTOR are widely reported in the skeletal muscle biology literature, from basic cell culture experiments to applied exercise/nutrition physiology studies in humans. Indeed, it has been fifteen years, since the first studies reported measurements of Ser2448 phosphorylation in muscle tissue [10, 76]. Since then, the biomedical and muscle research field has experienced a substantial growth of scientific papers reporting measurement of Ser2448 phosphorylation (Fig. 1) purportedly to assess mTOR kinase activation or as a proxy measure of induction of the mTOR pathway. However, the relevance of this phosphosite to the kinase activity of mTOR is frequently not considered. Thus, the role of the Ser2448 phosphorylation site in control of mTOR activity will be discussed herein and further how new insights in muscle physiology can be gained from revaluation of past studies utilizing this phosphorylation site.
Number of publications reporting Ser2448 per year. A timeline of key publications to understand the role of the Ser2448 residue of the mTOR protein is shown. Note that the first appearance of Ser2448 is in 1998; however, Bunn et al. studied the region, where mTOR Ser2448 is located, which is a key study for the understanding of the role of phosphorylation at this site
Where Ser2448 is located within mTOR?
The (m)TOR protein is a highly evolutionarily conserved serine/threonine kinase protein from yeast to humans [26, 35, 46]. Multiple phosphorylation sites are present on the mTOR protein structure including Serine 1261, Serine 1415, Serine 2159, Threonine 2164, Threonine 2446, Serine 2448, and Serine 2481 (Fig. 2) [32, 99]. The Ser1261 residue lies within mTOR HEAT-repeat motif, is phosphorylatable on both mTORC1/mTORC2, and its phosphorylation appears to increase mTORC1 kinase activity [1]. The Ser1415 is a recent discovered phosphorylatable site by IKKα directly following activation by AKT that increases mTORC1 activity [25]. The Ser2159 and Thr2164 residues lie within the kinase domain and phosphorylation of these residues promotes activation of mTORC1 [32]. The others remaining phosphorylatable mTOR residues (Thr2446, Ser2448, and Ser2481) are found in the FIT domain (Found In TOR) spanning TOR residues 2427 to 2516 [88, 99], between the kinase domain and the FATC domain [51, 99]. Initially, it was thought that Ser2481 and Ser2448 were phosphorylation residues specific for mTORC2 and mTORC1 respectively [23]. However, Ser2481 has already been found to be phosphorylated in mTORC1 [87] and Ser2448 has also been found to be phosphorylated in mTORC2, despite a lack of an apparent correlation with its kinase activity [78]. Phosphorylation of Ser2481 residue has been shown to be an autophosphorylation residue as a proxy of mTOR’s own activity [87].
Although TOR proteins contain several domains that are conserved throughout eukaryotic evolution, the small FIT domain (where Ser2448 dwells) is a relatively recent acquisition in vertebrate evolution [88, 99] showing some divergences in taxonomic groups. The mTOR homologous proteins in yeast (TOR) and drosophila (dTOR) lack the residue Ser2448 [23, 98]. Within the FIT domain lies the Negative Regulatory Domain (NRD) which is defined as TOR residues 2430 to 2450 [50, 85]. Due to its recent acquisition during vertebrate evolution, the Ser2448 residue and its contiguous area are potentially a fine-tuning mechanism for control of mTOR’s own activity.
Mechanisms of mTOR Ser2448 phosphorylation
It was originally proposed that Ser2448 was an AKT consensus phosphorylation target [51, 71, 85]. However, in vitro phosphorylation analysis failed to demonstrate direct phosphorylation by AKT, thus an undescribed protein kinase theoretically lying between AKT and mTOR was hypothesized to exist, which would be called TOR Kinase, or TORK [84], which has never been shown to exist. Instead, AKT is capable of activating mTORC1 by inhibition of its negative regulators. AKT phosphorylates and inactivates TSC1/2 that regulates Rheb activity (by stimulating GTP hydrolysis), which in turn binds to mTORC1 [57, 64]. In addition, AKT can phosphorylate PRAS40 [79, 91, 95]. Recently, it was also shown that AKT could additionally modulate mTOR activity, indirectly, via IKKα [25].
Evidence to further support the lack of direct signaling between AKT and Ser2448 mTOR is found in the skeletal muscle biology literature. The anabolic pathways leading to mTOR activation in response to mechanical stimulation were originally proposed to use the upstream pathway as growth factors (PI3K-AKT-mTOR) [9]. However, this has now convincingly been shown to be an AKT independent event [40, 47, 68, 75]. Therefore, if mechanical stimulation increases the phosphorylation of mTOR at Ser2448 [76] and p70S6K at Thr389 by a PI3K/AKT independent mechanism [33, 40, 54, 55, 65, 72, 73], then it is unlikely that phosphorylation of the mTOR Ser2448 site is a direct target of AKT. Indeed, Atherthon et al. [5], using stretching as a model for mechanical stimulation of myotubes, were able to demonstrate that phosphorylation of the mTOR downstream targets, p70S6K1 and 4EBP1, actually precedes phosphorylation of mTOR at Ser2448.
Direct phosphorylation of mTOR by AKT was proven not to be the case by two elegant studies, published in 2005, which demonstrated that Ser2448 is actually phosphorylated by the mTOR downstream effector p70S6K [19, 53], as part of a feedback loop mechanism. Subsequently, the synthesis of the first specific p70S6K inhibitor (PF-4708671) enabled further confirmation that Ser2448 is indeed part of mTOR-p70S6k feedback loop as this inhibitor simultaneously reduces the phosphorylation of the p70S6K downstream effector rpS6 and mTOR Ser2448 [74]. More recently, von Walden et al. [92] has shown that this p70S6K inhibitor increases the phosphorylation of Thr389 on p70S6K, which demonstrate that inhibition of p70S6K activity increased mTOR activity in muscle cells, presumably by removing this negative feedback loop. The discovery that mTOR is phosphorylated at Ser2448 by p70S6K explains how mTOR has been previously observed to be phosphorylated at Ser2448 following stimulation with anabolic stimuli, including insulin, amino acids, and mechanical strain. The following question then is: does phosphorylation of Ser2448 by p70S6K increases or repress mTOR activity?
Role of NRD and Ser2448 phosphorylation in the regulation of mTOR kinase activation
Phosphorylation of mTOR Ser2448 is a common marker used in the cancer [43] and skeletal muscle literature [77] as an index of mTOR kinase activity and/or pathway signaling. However, analysis of mTOR pathway activation solely by analysis of Ser2448 phosphorylation status in cancer studies has already been withdrawn by some researchers due to its unknown role [67]. Whilst heightened Ser2448 phosphorylation may be associated with mTORC1 activation under certain conditions, it does not imply a cause and effect relationship [14, 19, 53, 78]. In fact, there is no evidence that phosphorylation of Ser2448 increases mTOR activity. Rather, there is evidence from cell culture experiments utilizing mTOR mutant protein versions to suggest that the NRD domain has a negative effect on mTOR activity. Deletion of this region (amino acids 2433–2450) increases mTOR activity [31, 66, 85], although recently, others have found that deletion of a slightly extended region (2443–2486) has the opposite effect to lower mTOR activity [97], and substitution of Ser2448 for the non-phosphorylatable alanine or the phosphomimetic glutamate had no clear effect on mTOR kinase activity [19, 85].
The residue Thr2446, such as Ser2448, is another phosphorylatable site within the NRD. Initially, it was hypothesized that Thr2446 was a target of AKT along with Ser2448, that would promote mTOR activity [85]. However, Thr2446 was subsequently been shown to be a major target of AMPK, with its phosphorylation suppressing mTOR activity [18]. These two close residues Thr2446 and Ser2448 (separated by only one amino acid) are unlikely to have divergent effects on mTOR function. At the time that Thr2446 was shown to be a target of AMPK (in 2004), the main concept was that Ser2448 was a downstream target of AKT. Therefore, it was hypothesized that these sites would have opposing effects on mTOR activity, since insulin treatment increased mTOR Ser2448 but not Thr2446, and starvation increased Thr2446 but not Ser2448. However, the discovery that Ser2448 is a p70S6K-dependent site explains how both residues function as negative regulators of mTOR activity. The deletion of the NRD increases mTOR activity not because it exerts an inhibitory effect on mTOR kinase activity as previously hypothesized, but rather, because mTOR becomes refractory to energy and nutritional cues. This explains why the mutant TOR protein-lacking NRD region [31] is protected from growth factor deprivation and why prevention of phosphorylation at Thr2446 and Ser2448 via treatment with an antibody raised against the region 2433–2450 increases mTOR kinase activity [13, 66]. Not surprisingly then, the loss of Ser2448 phosphorylation has been linked to cancer progression when currently accepted knowledge in the role of Ser2448 would expect otherwise [70]. This indicates that failure of p70S6K to feedback to mTOR to fine-tune its activity may lead to a constitutively active mTOR resistant to p70S6K feedback signals.
Since mTOR Ser2448 is a residue target of p70S6K, to the best of our knowledge, no studies so far have shown that p70S6K phosphorylation at Thr389 is further increased after the initial phosphorylation activity of mTORC1 towards p70S6K, which would indicate a positive feedback. Recently, however, it has been shown that a short isoform of p70S6K can also interact with mTOR protein and increase mTOR kinase activity [6]. Although this short isoform of p70S6K can bind to mTOR, it cannot phosphorylate mTOR Ser2448 as it lacks a kinase activity. The mechanism proposed for the enhanced activation of mTOR by the short isoform of p70S6K was actually that this short isoform competes with p70S6K for the mTOR Ser2448 residue, thereby preventing the phosphorylation of Ser2448 and mitigating the negative feedback loop. The evidence presented points to a role of a negative feedback loop initiated by the activation of p70S6K. p70S6K has been previously shown to be the effector of an additional negative feedback loop by phosphorylating and inhibiting IRS-1 [90, 98]. Hence, p70S6K regulates mTOR pathway at multiple levels.
Alternative markers of mTOR activation
Contrary to the domain, where Ser2448 dwells, (m)TOR-associated proteins known to regulate mTOR kinase activity are conserved in yeast TOR and dTOR, including TSC/Rheb, Raptor, and GβL amongst others [3, 26, 35]. PRAS40, besides not being conserved in some yeast, has a corresponding homology in drosophila [79]. Furthermore, the TOR complexes 1 and 2 seem also to be highly conserved in eukaryotic evolution throughout yeast to man [26, 46]. In addition, two independent research groups have identified the leucine sensor within cell, the enzyme Leucyl tRNA synthethase (LRS) [12, 48]. This enzyme activates mTOR through the Rag GTPase. This is another conserved mechanism both found in TOR [12] and mTOR pathways [48]. These data suggest that mTOR regulation and function are largely dependent on the proteins that mTOR docks or associate with, rather than the phosphorylation of mTOR protein itself. Nevertheless, new phosphorylatable residues on mTOR have also been discovered and their roles in regulation of mTOR kinase activity investigated [1, 25, 32]. The residue Ser1261 phosphorylation promotes mTORC1 activity. It is not found in yeast and plants, but contrary to Ser2448, is found in drosophila (d)TOR as a phosphorylatable Threonine, suggesting that this might be a more conserved mechanism regarding mTOR activity than Ser2448 [1, 28]. The residue Ser1415 also appears to be highly conserved [25]. Two other residues within the kinase domain, Ser2159 and Thr2164, have also been discovered recently with positive roles in the mTOR kinase activity. Ser2159 is only found in vertebrates, but Thr2164 is conserved in flies, plants, and yeasts [32]. Thus, these residues could be better markers than Ser2448 in the future, although the kinase responsible for Ser1261, Ser2159, and Thr2164 phosphorylation are not yet known [32, 99].
Therefore, to better understand mTOR protein activity, researchers should aim for mechanisms that are more conserved throughout species that potentially represent a main mechanism of its regulation. In practical terms, it is recommended that researchers willing to determine a meaningful snap-shot of mTORC1 activation using western blotting should focus on classical mTORC1 downstream targets: p70S6K and 4E-BP1. In addition, partner proteins may also add valuable data as PRAS40 phosphorylation on AKT-dependent site and Rag proteins. Moreover, whenever possible, immunoprecipitation of mTORC1 and mTORC2 protein partners, Raptor, and Rictor, respectively, may help determine on which mTOR, the phosphorylation of Ser2448, is complexed with.
Reinterpretation of exercise and muscle studies in light of a mTOR-p70S6K-mTOR negative feedback loop
Data supporting the role of mTOR in skeletal muscle growth appears overwhelming. To date, studies have focused on the mTORC1 downstream effectors, p70S6K and 4E-BP1, by measuring the mTOR-dependent phosphorylation sites at Thr389 on p70S6K, and Thr37/46, Ser65, and Thr70 on 4E-BP1 [82, 83]. Data on Thr389 p70S6K in particular seem quite convincing that mTORC1 activity is increased following exercise and that nutrition may play a role enhancing that response. However, the same is not true for mTOR’s own phosphorylation state. Many studies fail to demonstrate a consistent and reproducible Ser2448 phosphorylation response to a plethora of stimuli, whilst the mTORC1-dependent site Thr389 on p70S6K does. Titration of insulin [42] and IGF-1 [61] results in a clear dose-dependent phosphorylation curve for p70S6K at Thr389, whereas mTOR at Ser2448 phosphorylation response is disconnected in both cases. In rat skeletal muscle, phosphorylation of p70S6K at Thr389 was found to increase during feeding and decrease during fasting, whilst a corresponding effect for mTOR Ser2448 was not observed [86]. Furthermore, numerous past works measuring both mTOR Ser2448 and p70S6K Thr389 phosphorylation status have demonstrated that muscle load and/or nutritional interventions have a far greater and more obvious stimulatory effect on p70S6K Thr389 than on mTOR Ser2448 [2, 4, 8, 11, 17, 24, 27, 29, 30, 33, 34, 38, 39, 41, 45, 55, 59, 65, 69, 89, 94]. Since the discovery of the identity of the Ser2448 kinase by Holz and Blenis, and Chiang and Abraham [19, 53] in 2005, it has been established that p70S6K, and not AKT, regulates phosphorylation of the mTOR Ser2448 site as part of a feedback loop mechanism. Although some lines of evidence had already pointed to this regulation having a suppressive effect, it was only recently that the work from Ben-Hur [6] made it clear that a negative feedback loop exists between p70S6K and mTOR Ser2448.
Studies in the muscle literature in particular can have different interpretations in light of the real significance of Ser2448 phosphorylation. This is particularly true for studies, where mTOR Ser2448 is analysed in isolation or when the phosphorylation state of Ser2448 appears in opposition to other downstream targets of the mTOR pathway. For example, Leger et al. [63] showed that 4E-BP1 and p70S6K phosphorylation were unaltered by 8 weeks of resistance exercise training. At the same time, phosphorylation of mTOR Ser2448 was found to be significantly increased. After 8 weeks of detraining, accompanied by a reduction in muscle mass, the phosphorylation at Ser2448 remained elevated, whilst p70S6K and 4E-BP1 phosphorylation were once again unchanged. The authors concluded that activation of the mTOR pathway was increased throughout training based solely on mTOR Ser2448. This was despite the fact that heightened phosphorylation persisting throughout detraining and despite the lack of an effect of training/detraining on p70S6K and 4E-BP1. In addition, other researchers, expecting to observe phosphorylation of mTOR Ser2448 preceding Thr389 on p70S6K and having found the opposite, have suggested that phosphorylation at Thr389 could be a mTOR-independent event following resistance exercise [58]. With the knowledge that p70S6K and not AKT is the kinase of the mTOR Ser2448 site, such results may need to be reinterpreted.
Understanding the role of Ser2448 being a negative regulator of mTOR activity, subject to p70S6K activity in a negative feedback manner, aids in providing insight into the dynamic signaling events in muscle following either loading and/or protein ingestion. Phosphorylation of p70S6K Thr389 has been shown to peak at 15 [65] and 30 min post exercise in humans [52] and in a resistance training model in rodents [11]. Others have also found robust phosphorylation of p70S6K Thr389 at the same early time points with no apparent response on mTOR Ser2448 [56]. Phosphorylation at Thr389 appears to peak no later than the first hour post exercise and decreases in the following hours [15, 62]. Phosphorylation of mTOR Ser2448 should increase afterwards [5] and may thus contribute to the decrease in Thr389 of p70S6K. Thus, in studies that have taken biopsy timepoints exceeding 1 h or upwards, it is tempting to speculate that the feedback loop mechanism (p70S6K-mTOR-p70S6K) may have already taken place. Consistent with this notion, Churchward-Venne et al. [22] have shown recently that Thr389 was not increased compared to baseline at 1.5 h post resistance exercise when at the same timepoint, the other mTORC1 downstream 4E-BP1 was highly phosphorylated, as well as the p70S6K downstream effector S6rp at Ser240/244 also, demonstrating that mTOR/p70S6K was activated before this timepoint. Convincingly, mTOR Ser2448 was also highly phosphorylated; indicating that the decrease or return to baseline levels of Thr389 could have been due to the negative feedback mechanism. These results all are consistent with Ser2448 being a negative feedback mechanism dependent upon of p70S6K activation.
Concluding remarks
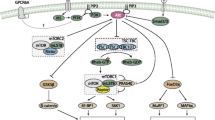
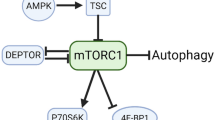
The Ser2448 residue of mTOR is part of a negative feedback loop mechanism in which activated p70S6K phosphorylates back onto mTOR to fine-tune its activity (Fig. 3). Whilst an increase in phosphorylation of this residue could potentially reflect an increase in the overall pathway, it does not apply for every cell and condition, as in fact it is rather a read-out of p70S6K activity and thus a late event in the pathway. The use of Ser2448 phosphorylation as the only marker of mTOR kinase activity or pathway activation should be discouraged. New phosphorylation residues have been discovered, which may have greater relevance towards mTOR activity. But as of yet their regulation is incompletely understood and their function should be better known to be widely used. mTOR seems to be mainly regulated as a complex, i.e., in conjunction with its partners (raptor, rictor, PRAS40, DEPTOR, Ragulator, Rheb, etc), rather than by its own phosphorylation. Moreover, because Ser2448 may also be phosphorylated on mTORC2, the significance of measuring global changes in Ser2448 phosphorylation is complicated further. On this basis, efforts to understand mTOR activity and/or activation in skeletal muscle should also focus on the mTOR partners and downstream targets.
p70S6K-mTORC1-p70S6k feedback loop mechanism. Cues that can modulate mTORC1 in the context of muscle cells are depicted. Red cross emphasizes that Ser2448 is not phosphorylated by Akt as previously thought, but by mTORC1’s downstream effector p70S6K. AKT although modulates mTOR activity, it does indirectly via TSC 1/2, and not directly via Ser2448 as previously though. Muscle contractions/exercise promote phosphorylation of mTOR targets (p70S6K and 4E-BP1), but also mTOR Ser2448. Since contractions can promote mTRO activity independently of AKT, Ser2448 cannot be a target of AKT, but as demonstrated by two independent studies (Chiang and Abraham, and Holz and Blenis) by p70S6K
In conclusion, it is necessary to state that mTOR Ser2448 phosphorylation: (1) is a target of p70S6K and not AKT; (2) is not a cause of p70S6K activity but rather its consequence; (3) has a negative regulatory effect on mTOR kinase activity, and (4) it does not necessarily mirror an increase in mTOR kinase activity nor should be taken as a measure of mTOR activation. This last item is the most important for researches using Ser2448 phosphorylation, since an erroneous assumption could generate misleading interpretations if other more proper markers are not used in parallel. Based on the present analysis, the interpretation from many studies in the muscle research literature assessing phosphorylation levels of mTOR at Ser2448 may need to be revaluated.
Abbreviations
- HEAT:
-
Acronym for four proteins (but not restricted to) that possess this specific domain:Huntingtin, Elongation factor 3, protein phosphatase 2A
- IRS-1:
-
Insulin receptor substrate-1
References
Acosta-Jaquez HA, Keller JA, Foster KG, Ekim B, Soliman GA, Feener EP, Ballif BA, Fingar DC (2009) Site-specific mTOR phosphorylation promotes mTORC1-mediated signaling and cell growth. Mol Cell Biol 29:4308–4324
Apro W, Wang L, Ponten M, Blomstrand E, Sahlin K (2013) Resistance exercise induced mTORC1 signaling is not impaired by subsequent endurance exercise in human skeletal muscle. Am J Physiol Endocrinol Metab 305:E22–E32
Aspuria PJ, Tamanoi F (2004) The Rheb family of GTP-binding proteins. Cell Signal 16:1105–1112
Atherton PJ, Etheridge T, Watt PW, Wilkinson D, Selby A, Rankin D, Smith K, Rennie MJ (2010) Muscle full effect after oral protein: time-dependent concordance and discordance between human muscle protein synthesis and mTORC1 signaling. Am J Clin Nutr 92:1080–1088
Atherton PJ, Szewczyk NJ, Selby A, Rankin D, Hillier K, Smith K, Rennie MJ, Loughna PT (2009) Cyclic stretch reduces myofibrillar protein synthesis despite increases in FAK and anabolic signalling in L6 cells. J Physiol 587:3719–3727
Ben-Hur V, Denichenko P, Siegfried Z, Maimon A, Krainer A, Davidson B and Karni R (2013) S6K1 alternative splicing modulates its oncogenic activity and regulates mTORC1. Cell Rep 3:103–115
Bhaskar PT, Hay N (2007) The two TORCs and Akt. Dev Cell 12:487–502
Blomstrand E, Eliasson J, Karlsson HK and Kohnke R (2006) Branched-chain amino acids activate key enzymes in protein synthesis after physical exercise. J Nutr 136:269S–73S
Bodine SC, Stitt TN, Gonzalez M, Kline WO, Stover GL, Bauerlein R, Zlotchenko E, Scrimgeour A, Lawrence JC, Glass DJ, Yancopoulos GD (2001) Akt/mTOR pathway is a crucial regulator of skeletal muscle hypertrophy and can prevent muscle atrophy in vivo. Nat Cell Biol 3:1014–1019
Bolster DR, Crozier SJ, Kimball SR, Jefferson LS (2002) AMP-activated protein kinase suppresses protein synthesis in rat skeletal muscle through down-regulated mammalian target of rapamycin (mTOR) signaling. J Biol Chem 277:23977–23980
Bolster DR, Kubica N, Crozier SJ, Williamson DL, Farrell PA, Kimball SR, Jefferson LS (2003) Immediate response of mammalian target of rapamycin (mTOR)-mediated signalling following acute resistance exercise in rat skeletal muscle. J Physiol 553:213–220
Bonfils G, Jaquenoud M, Bontron S, Ostrowicz C, Ungermann C, De Virgilio C (2012) Leucyl-tRNA synthetase controls TORC1 via the EGO complex. Mol Cell 46:105–110
Brunn GJ, Fadden P, Haystead TA, Lawrence JC, Jr (1997) The mammalian target of rapamycin phosphorylates sites having a (Ser/Thr)-Pro motif and is activated by antibodies to a region near its COOH terminus. J Biol Chem 272:32547–32550
Burgstaller S, Rosner M, Lindengrun C, Hanneder M, Siegel N, Valli A, Fuchs C, Hengstschlager M (2009) Tuberin, p27 and mTOR in different cells. Amino Acids 36:297–302
Burke LM, Hawley JA, Ross ML, Moore DR, Phillips SM, Slater GR, Stellingwerff T, Tipton KD, Garnham AP, Coffey VG (2012) Preexercise aminoacidemia and muscle protein synthesis after resistance exercise. Med Sci Sports Exerc 44:1968–1977
Burnett PE, Barrow RK, Cohen NA, Snyder SH, Sabatini DM (1998) RAFT1 phosphorylation of the translational regulators p70 S6 kinase and 4E-BP1. Proc Natl Acad Sci USA 95:1432–1437
Camera DM, West DW, Burd NA, Phillips SM, Garnham AP, Hawley JA and Coffey VG (2012) Low muscle glycogen concentration does not suppress the anabolic response to resistance exercise. J Appl Physiol (1985) 113:206–214
Cheng SW, Fryer LG, Carling D, Shepherd PR (2004) Thr2446 is a novel mammalian target of rapamycin (mTOR) phosphorylation site regulated by nutrient status. J Biol Chem 279:15719–15722
Chiang GG, Abraham RT (2005) Phosphorylation of mammalian target of rapamycin (mTOR) at Ser-2448 is mediated by p70S6 kinase. J Biol Chem 280:25485–25490
Choo AY, Blenis J (2009) Not all substrates are treated equally: implications for mTOR, rapamycin-resistance and cancer therapy. Cell Cycle 8:567–572
Choo AY, Yoon SO, Kim SG, Roux PP, Blenis J (2008) Rapamycin differentially inhibits S6Ks and 4E-BP1 to mediate cell-type-specific repression of mRNA translation. Proc Natl Acad Sci USA 105:17414–17419
Churchward-Venne TA, Breen L, Di Donato DM, Hector AJ, Mitchell CJ, Moore DR, Stellingwerff T, Breuille D, Offord EA, Baker SK, Phillips SM (2014) Leucine supplementation of a low-protein mixed macronutrient beverage enhances myofibrillar protein synthesis in young men: a double-blind, randomized trial. Am J Clin Nutr 99:276–286
Copp J, Manning G, Hunter T (2009) TORC-specific phosphorylation of mammalian target of rapamycin (mTOR): phospho-Ser2481 is a marker for intact mTOR signaling complex 2. Cancer Res 69:1821–1827
Cuthbertson D, Smith K, Babraj J, Leese G, Waddell T, Atherton P, Wackerhage H, Taylor PM, Rennie MJ (2005) Anabolic signaling deficits underlie amino acid resistance of wasting, aging muscle. FASEB J 19:422–424
Dan HC, Ebbs A, Pasparakis M, Van Dyke T, Basseres DS, Baldwin AS (2014) Akt-dependent activation of mTORC1 complex involves phosphorylation of mTOR (mammalian target of rapamycin) by IkappaB kinase alpha (IKKalpha). J Biol Chem 289:25227–25240
De Virgilio C, Loewith R (2006) The TOR signalling network from yeast to man. Int J Biochem Cell Biol 38:1476–1481
Deldicque L, Sanchez Canedo C, Horman S, De Potter I, Bertrand L, Hue L, Francaux M (2008) Antagonistic effects of leucine and glutamine on the mTOR pathway in myogenic C2C12 cells. Amino Acids 35:147–155
Dennis MD, Baum JI, Kimball SR, Jefferson LS (2011) Mechanisms involved in the coordinate regulation of mTORC1 by insulin and amino acids. J Biol Chem 286:8287–8296
Dreyer HC, Drummond MJ, Pennings B, Fujita S, Glynn EL, Chinkes DL, Dhanani S, Volpi E, Rasmussen BB (2008) Leucine-enriched essential amino acid and carbohydrate ingestion following resistance exercise enhances mTOR signaling and protein synthesis in human muscle. Am J Physiol Endocrinol Metab 294:E392–E400
Dreyer HC, Fujita S, Glynn EL, Drummond MJ, Volpi E, Rasmussen BB (2010) Resistance exercise increases leg muscle protein synthesis and mTOR signalling independent of sex. Acta Physiol (Oxf) 199:71–81
Edinger AL, Thompson CB (2004) An activated mTOR mutant supports growth factor-independent, nutrient-dependent cell survival. Oncogene 23:5654–5663
Ekim B, Magnuson B, Acosta-Jaquez HA, Keller JA, Feener EP, Fingar DC (2011) mTOR kinase domain phosphorylation promotes mTORC1 signaling, cell growth, and cell cycle progression. Mol Cell Biol 31:2787–2801
Eliasson J, Elfegoun T, Nilsson J, Kohnke R, Ekblom B, Blomstrand E (2006) Maximal lengthening contractions increase p70 S6 kinase phosphorylation in human skeletal muscle in the absence of nutritional supply. Am J Physiol Endocrinol Metab 291:E1197–E1205
Etheridge T, Atherton PJ, Wilkinson D, Selby A, Rankin D, Webborn N, Smith K, Watt PW (2011) Effects of hypoxia on muscle protein synthesis and anabolic signaling at rest and in response to acute resistance exercise. Am J Physiol Endocrinol Metab 301:E697–E702
Fingar DC, Blenis J (2004) Target of rapamycin (TOR): an integrator of nutrient and growth factor signals and coordinator of cell growth and cell cycle progression. Oncogene 23:3151–3171
Fingar DC, Richardson CJ, Tee AR, Cheatham L, Tsou C, Blenis J (2004) mTOR controls cell cycle progression through its cell growth effectors S6K1 and 4E-BP1/eukaryotic translation initiation factor 4E. Mol Cell Biol 24:200–216
Fingar DC, Salama S, Tsou C, Harlow E, Blenis J (2002) Mammalian cell size is controlled by mTOR and its downstream targets S6K1 and 4EBP1/eIF4E. Genes Dev 16:1472–1487
Fujita S, Abe T, Drummond MJ, Cadenas JG, Dreyer HC, Sato Y, Volpi E and Rasmussen BB (2007) Blood flow restriction during low-intensity resistance exercise increases S6K1 phosphorylation and muscle protein synthesis. J Appl Physiol (1985) 103:903–910
Glover EI, Oates BR, Tang JE, Moore DR, Tarnopolsky MA, Phillips SM (2008) Resistance exercise decreases eIF2Bepsilon phosphorylation and potentiates the feeding-induced stimulation of p70S6K1 and rpS6 in young men. Am J Physiol Regul Integr Comp Physiol 295:R604–R610
Goodman CA, Miu MH, Frey JW, Mabrey DM, Lincoln HC, Ge Y, Chen J, Hornberger TA (2010) A phosphatidylinositol 3-kinase/protein kinase B-independent activation of mammalian target of rapamycin signaling is sufficient to induce skeletal muscle hypertrophy. Mol Biol Cell 21:3258–3268
Gran P and Cameron-Smith D (2011) The actions of exogenous leucine on mTOR signalling and amino acid transporters in human myotubes. BMC Physiol 11:10-6793-11-10
Greenhaff PL, Karagounis LG, Peirce N, Simpson EJ, Hazell M, Layfield R, Wackerhage H, Smith K, Atherton P, Selby A, Rennie MJ (2008) Disassociation between the effects of amino acids and insulin on signaling, ubiquitin ligases, and protein turnover in human muscle. Am J Physiol Endocrinol Metab 295:E595–E604
Guertin DA, Sabatini DM (2007) Defining the role of mTOR in cancer. Cancer Cell 12:9–22
Guertin DA, Stevens DM, Thoreen CC, Burds AA, Kalaany NY, Moffat J, Brown M, Fitzgerald KJ, Sabatini DM (2006) Ablation in mice of the mTORC components raptor, rictor, or mLST8 reveals that mTORC2 is required for signaling to Akt-FOXO and PKCalpha, but not S6K1. Dev Cell 11:859–871
Haegens A, Schols AM, van Essen AL, van Loon LJ, Langen RC (2012) Leucine induces myofibrillar protein accretion in cultured skeletal muscle through mTOR dependent and -independent control of myosin heavy chain mRNA levels. Mol Nutr Food Res 56:741–752
Hall MN (2008) mTOR-what does it do? Transplant Proc 40:S5–S8
Hamilton DL, Philp A, MacKenzie MG, Baar K (2010) A limited role for PI(3,4,5)P3 regulation in controlling skeletal muscle mass in response to resistance exercise. PLoS One 5:e11624
Han JM, Jeong SJ, Park MC, Kim G, Kwon NH, Kim HK, Ha SH, Ryu SH, Kim S (2012) Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell 149:410–424
Hara K, Maruki Y, Long X, Yoshino K, Oshiro N, Hidayat S, Tokunaga C, Avruch J, Yonezawa K (2002) Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110:177–189
Hardt M, Chantaravisoot N, Tamanoi F (2011) Activating mutations of TOR (target of rapamycin). Genes Cells 16:141–151
Hay N, Sonenberg N (2004) Upstream and downstream of mTOR. Genes Dev 18:1926–1945
Holm L, van Hall G, Rose AJ, Miller BF, Doessing S, Richter EA, Kjaer M (2010) Contraction intensity and feeding affect collagen and myofibrillar protein synthesis rates differently in human skeletal muscle. Am J Physiol Endocrinol Metab 298:E257–E269
Holz MK, Blenis J (2005) Identification of S6 kinase 1 as a novel mammalian target of rapamycin (mTOR)-phosphorylating kinase. J Biol Chem 280:26089–26093
Hornberger TA, Stuppard R, Conley KE, Fedele MJ, Fiorotto ML, Chin ER, Esser KA (2004) Mechanical stimuli regulate rapamycin-sensitive signalling by a phosphoinositide 3-kinase-, protein kinase B- and growth factor-independent mechanism. Biochem J 380:795–804
Hulmi JJ, Tannerstedt J, Selanne H, Kainulainen H, Kovanen V and Mero AA (2009) Resistance exercise with whey protein ingestion affects mTOR signaling pathway and myostatin in men. J Appl Physiol (1985) 106:1720–1729
Hulmi JJ, Walker S, Ahtiainen JP, Nyman K, Kraemer WJ, Hakkinen K (2012) Molecular signaling in muscle is affected by the specificity of resistance exercise protocol. Scand J Med Sci Sports 22:240–248
Inoki K, Li Y, Xu T, Guan KL (2003) Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev 17:1829–1834
Kakigi R, Naito H, Ogura Y, Kobayashi H, Saga N, Ichinoseki-Sekine N, Yoshihara T, Katamoto S (2011) Heat stress enhances mTOR signaling after resistance exercise in human skeletal muscle. J Physiol Sci 61:131–140
Karagounis LG, Yaspelkis BB, 3rd, Reeder DW, Lancaster GI, Hawley JA, Coffey VG (2010) Contraction-induced changes in TNFalpha and Akt-mediated signalling are associated with increased myofibrillar protein in rat skeletal muscle. Eur J Appl Physiol 109:839–848
Kim DH, Sarbassov DD, Ali SM, King JE, Latek RR, Erdjument-Bromage H, Tempst P, Sabatini DM (2002) mTOR interacts with raptor to form a nutrient-sensitive complex that signals to the cell growth machinery. Cell 110:163–175
Kuemmerle JF (2003) IGF-I elicits growth of human intestinal smooth muscle cells by activation of PI3K, PDK-1, and p70S6 kinase. Am J Physiol Gastrointest Liver Physiol 284:G411–G422
Kumar V, Selby A, Rankin D, Patel R, Atherton P, Hildebrandt W, Williams J, Smith K, Seynnes O, Hiscock N, Rennie MJ (2009) Age-related differences in the dose-response relationship of muscle protein synthesis to resistance exercise in young and old men. J Physiol 587:211–217
Leger B, Cartoni R, Praz M, Lamon S, Deriaz O, Crettenand A, Gobelet C, Rohmer P, Konzelmann M, Luthi F, Russell AP (2006) Akt signalling through GSK-3beta, mTOR and Foxo1 is involved in human skeletal muscle hypertrophy and atrophy. J Physiol 576:923–933
Manning BD, Cantley LC (2003) Rheb fills a GAP between TSC and TOR. Trends Biochem Sci 28:573–576
Mascher H, Tannerstedt J, Brink-Elfegoun T, Ekblom B, Gustafsson T, Blomstrand E (2008) Repeated resistance exercise training induces different changes in mRNA expression of MAFbx and MuRF-1 in human skeletal muscle. Am J Physiol Endocrinol Metab 294:E43–E51
McMahon LP, Choi KM, Lin TA, Abraham RT, Lawrence JC, Jr (2002) The rapamycin-binding domain governs substrate selectivity by the mammalian target of rapamycin. Mol Cell Biol 22:7428–7438
Menon S, Manning BD (2008) Common corruption of the mTOR signaling network in human tumors. Oncogene 27(Suppl 2):S43–S51
Miyazaki M, McCarthy JJ, Fedele MJ, Esser KA (2011) Early activation of mTORC1 signalling in response to mechanical overload is independent of phosphoinositide 3-kinase/Akt signalling. J Physiol 589:1831–1846
Moore DR, Atherton PJ, Rennie MJ, Tarnopolsky MA, Phillips SM (2011) Resistance exercise enhances mTOR and MAPK signalling in human muscle over that seen at rest after bolus protein ingestion. Acta Physiol (Oxf) 201:365–372
Muller J, Ehlers A, Burkhardt L, Sirma H, Steuber T, Graefen M, Sauter G, Minner S, Simon R, Schlomm T, Michl U (2013) Loss of pSer2448-mTOR expression is linked to adverse prognosis and tumor progression in ERG-fusion-positive cancers. Int J Cancer 132:1333–1340
Nave BT, Ouwens M, Withers DJ, Alessi DR, Shepherd PR (1999) Mammalian target of rapamycin is a direct target for protein kinase B: identification of a convergence point for opposing effects of insulin and amino-acid deficiency on protein translation. Biochem J 344(Pt 2):427–431
O’Neil TK, Duffy LR, Frey JW, Hornberger TA (2009) The role of phosphoinositide 3-kinase and phosphatidic acid in the regulation of mammalian target of rapamycin following eccentric contractions. J Physiol 587:3691–3701
Parkington JD, Siebert AP, LeBrasseur NK, Fielding RA (2003) Differential activation of mTOR signaling by contractile activity in skeletal muscle. Am J Physiol Regul Integr Comp Physiol 285:R1086–R1090
Pearce LR, Alton GR, Richter DT, Kath JC, Lingardo L, Chapman J, Hwang C, Alessi DR (2010) Characterization of PF-4708671, a novel and highly specific inhibitor of p70 ribosomal S6 kinase (S6K1). Biochem J 431:245–255
Rasmussen BB (2011) The missing Akt in the mechanical regulation of skeletal muscle mTORC1 signalling and growth. J Physiol 589:1507
Reynolds TH, 4th, Bodine SC, Lawrence JC, Jr (2002) Control of Ser2448 phosphorylation in the mammalian target of rapamycin by insulin and skeletal muscle load. J Biol Chem 277:17657–17662
Rivas DA, Lessard SJ, Coffey VG (2009) mTOR function in skeletal muscle: a focal point for overnutrition and exercise. Appl Physiol Nutr Metab 34:807–816
Rosner M, Siegel N, Valli A, Fuchs C, Hengstschlager M (2010) mTOR phosphorylated at S2448 binds to raptor and rictor. Amino Acids 38:223–228
Sancak Y, Thoreen CC, Peterson TR, Lindquist RA, Kang SA, Spooner E, Carr SA, Sabatini DM (2007) PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol Cell 25:903–915
Sarbassov DD, Ali SM, Kim DH, Guertin DA, Latek RR, Erdjument-Bromage H, Tempst P, Sabatini DM (2004) Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr Biol 14:1296–1302
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM (2005) Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307:1098–1101
Schalm SS, Blenis J (2002) Identification of a conserved motif required for mTOR signaling. Curr Biol 12:632–639
Schalm SS, Fingar DC, Sabatini DM, Blenis J (2003) TOS motif-mediated raptor binding regulates 4E-BP1 multisite phosphorylation and function. Curr Biol 13:797–806
Scott PH, Lawrence JC, Jr (1998) Attenuation of mammalian target of rapamycin activity by increased cAMP in 3T3-L1 adipocytes. J Biol Chem 273:34496–34501
Sekulic A, Hudson CC, Homme JL, Yin P, Otterness DM, Karnitz LM, Abraham RT (2000) A direct linkage between the phosphoinositide 3-kinase-AKT signaling pathway and the mammalian target of rapamycin in mitogen-stimulated and transformed cells. Cancer Res 60:3504–3513
Shavlakadze T, Chai J, Maley K, Cozens G, Grounds G, Winn N, Rosenthal N, Grounds MD (2010) A growth stimulus is needed for IGF-1 to induce skeletal muscle hypertrophy in vivo. J Cell Sci 123:960–971
Soliman GA, Acosta-Jaquez HA, Dunlop EA, Ekim B, Maj NE, Tee AR, Fingar DC (2010) mTOR Ser-2481 autophosphorylation monitors mTORC-specific catalytic activity and clarifies rapamycin mechanism of action. J Biol Chem 285:7866–7879
Sturgill TW, Hall MN (2009) Activating mutations in TOR are in similar structures as oncogenic mutations in PI3KCalpha. ACS Chem Biol 4:999–1015
Thomson DM, Gordon SE (2006) Impaired overload-induced muscle growth is associated with diminished translational signalling in aged rat fast-twitch skeletal muscle. J Physiol 574:291–305
Tremblay F, Marette A (2001) Amino acid and insulin signaling via the mTOR/p70 S6 kinase pathway. A negative feedback mechanism leading to insulin resistance in skeletal muscle cells. J Biol Chem 276:38052–38060
Vander Haar E, Lee SI, Bandhakavi S, Griffin TJ, Kim DH (2007) Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat Cell Biol 9:316–323
von Walden F, Liu C, Aurigemma N and Nader GA (2016) mTOR signaling regulates myotube hypertrophy by modulating protein synthesis, rDNA transcription, and chromatin remodeling. Am J Physiol Cell Physiol 311:C663-C672
Wang L, Mascher H, Psilander N, Blomstrand E and Sahlin K (2011) Resistance exercise enhances the molecular signaling of mitochondrial biogenesis induced by endurance exercise in human skeletal muscle. J Appl Physiol (1985) 111:1335–1344
Wiza C, Nascimento EB, Ouwens DM (2012) Role of PRAS40 in Akt and mTOR signaling in health and disease. Am J Physiol Endocrinol Metab 302:E1453–E1460
Wullschleger S, Loewith R, Hall MN (2006) TOR signaling in growth and metabolism. Cell 124:471–484
Yang H, Rudge DG, Koos JD, Vaidialingam B, Yang HJ, Pavletich NP (2013) mTOR kinase structure, mechanism and regulation. Nature 497:217–223
Zhang H, Stallock JP, Ng JC, Reinhard C, Neufeld TP (2000) Regulation of cellular growth by the Drosophila target of rapamycin dTOR. Genes Dev 14:2712–2724
Zhang J, Gao Z, Yin J, Quon MJ, Ye J (2008) S6K directly phosphorylates IRS-1 on Ser-270 to promote insulin resistance in response to TNF-(alpha) signaling through IKK2. J Biol Chem 283:35375–35382
Zinzalla V, Sturgill TW and, Hall MN (2010) TOR complexes: composition, structure and phosphorylation. In: Hall MN, Tamanoi F (eds) The enzymes: structure. Function and regulation of TOR complexes from yeast to mammals. Academic Press, San Diego, pp 1–20
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Figueiredo, V.C., Markworth, J.F. & Cameron-Smith, D. Considerations on mTOR regulation at serine 2448: implications for muscle metabolism studies. Cell. Mol. Life Sci. 74, 2537–2545 (2017). https://doi.org/10.1007/s00018-017-2481-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2481-5