Abstract
Polycomb group (PcG) proteins have recently been shown related to cancer development. The PcG protein EZH2 is involved in progression of prostate and breast cancers, and has been identified as a molecular marker in breast cancer. Nevertheless, the molecular mechanism by which PcG proteins regulate cancer progression and malignant metastasis is still unclear. PcG proteins methylate H3K27 in undifferentiated epithelial cells, resulting in the repression of differentiation genes such as HOX. FOXC1 is a member of the Forkhead box transcription factor family, which plays an important role in differentiation, and is involved in eye development. We discovered in this study that the expression of FOXC1 gene was negatively correlated to that of PcG genes, i.e., Bmi1, EZH2, and SUZ12, in MCF-7 and MDA-MB-231 cells. To investigate the regulatory effects of PcG proteins on FOXC1 gene, the two cell lines were transfected with either expression plasmids or siRNA plasmids of Bmi1, EZH2, and SUZ12, and we found that PcGs, especially EZH2, could repress the transcription of FOXC1 gene. Chromatin immunoprecipitation (ChIP) assay showed that histone methylation and acetylation modifications played critical roles in this regulatory process. When FOXC1 was stably transfected into MDA-MB-231 cells, the migration and invasion of the cells were repressed. Moreover, the tumorigenicity and the spontaneous metastatic capability regulated by FOXC1 were determined by using an orthotropic xenograft tumor model of athymic mice with the FOXC1-MDA-MB-231HM and the GFP-MDA-MB-231HM cells, and the results showed that FOXC1 in MDA-MB-231HM cells inhibited migration and invasion in vitro and reduced the pulmonary metastasis in vivo. Data presented in this report contribute to the understanding of the mechanisms by which EZH2 participates in tumor development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is a leading cause of cancer-related death in women, accounting for about 40,000 deaths in 180,000 women breast cancer patients per year in the US [1]. Advances have been made in the early detection and therapies of breast cancer, but the mortality of the patients with recurrences and metastases is still high [2]. Identification of new prognostic factors predicting tumor progression and metastasis at diagnosis remains a persistent research effort [3, 4].
Polycomb group (PcG) proteins were initially discovered as epigenetic silencers during embryogenesis, many of them have also been implicated in development and differentiation [5]. PcG proteins are assembled into multimeric complexes to exert their functions [6]. Biochemical and genetic studies indicate that PcG proteins exist in at least two separate protein complexes, i.e., the polycomb repressive complex 1 (PRC1) and the ESC-E(Z) complex, which function in a cooperative manner to maintain long-term gene silencing. The enhancer of zeste homologue 2 (EZH2) is a member of the PcG, and is involved in cell cycle regulation. PcG proteins have recently been suggested as candidates for targeted therapeutics of cancer [7]. Previous studies from tissue microarray and clinical tests indicate that the expression level of EZH2 was strongly correlated with breast cancer aggressiveness [8–11]. The PcG protein EZH2 is a histone methyltransferase involved in transcriptional repression, and EZH2 catalyzes the addition of methyl groups to histone H3 at lysine 27 (H3K27) [12]. PcG proteins methylate H3K27 in undifferentiated epithelial cells, resulting in the repression of differentiation genes such as HOX genes [13, 14]. Nevertheless, the molecular mechanisms by which PcG proteins regulate cancer progression and malignant metastasis are still unclear to date.
FOXC1 is a transcription factor playing an important role in regulation of ocular development [15, 16], and has been recognized as a potential regulator of differentiation [17]. In this study, we intended to investigate the regulatory effects of PcG proteins on FOXC1 gene, and how EZH2 promotes breast cancer cell proliferation and metastasis. We showed that PcG proteins EZH2, SUZ12, and Bmi1 participated in transcriptional inhibition of FOXC1. The histone acetylation and methylation modifications at FOXC1 promoter played an important role in FOXC1 expression. Overall, data from this study, as well as from the previous clinic tissue studies [10], suggested that the EZH2 expression was strongly associated with breast cancer aggressiveness. We also presented evidence that FOXC1 prevented breast cancer cell migration and invasion in vitro and in vivo. These results implicated that FOXC1, as a target of PcG, may potentially be a novel factor inhibiting the breast cancer aggression.
Materials and methods
Plasmid constructs
The expression vector of FOXC1 (pEGFP-N1-FOXC1) was generously provided by Dr. Fred B. Berry (Departments of Ophthalmology and Medical Genetics, University of Alberta, Canada). The expression vectors of EZH2, SUZ12, and Bmi1 were the gifts from Dr. Hong Chen (Department of Urology, University of Texas Southwestern Medical Center), Dr. Ru Cao (Department of Biochemistry and Biophysics Lineberger Comprehensive Cancer Center), and Dr. Dmitri (Oncology Research and Discovery Technologies, Novartis Institutes for BioMedical Research), respectively.
Cell culture and animals
MCF-7 and MDA-MB-231 cells were purchased from the Institute of Cell Biology, the Chinese Academy of Sciences. MCF-7 and MDA-MB-231 cells were maintained in Isocove’s modified Dulbecco’s medium and L15 medium (Gibco), respectively, supplemented with 10% fetal bovine serum, 100 μg/ml penicillin, and 100 μg/ml streptomycin. Transient transfection of MCF-7 and MDA-MB-231 cells was carried out using the LipofectAMINE 2000 (Invitrogen). Female athymic BALB/c-nu/nu mice, 4- to 6-week old, were obtained from the Shanghai Institute of Materia Medica of the Chinese Academy of Sciences, and housed in laminar-flow cabinets under specific pathogen-free conditions, with food and water ad libitum. All experiments on mice were conducted in accordance with the guidelines of National Institutes of Health (NIH, Bethesda, MD, USA) for the care and use of laboratory animals. The animal handling protocol was approved by Shanghai Medical Experimental Animal Care Committee (China).
RNA isolation and RT-PCR
Total RNA was isolated using the Trizol reagent (Invitrogen) following manufacturer’s instructions. One microgram RNA was used for cDNA synthesis using a reverse transcriptase reaction kit (Promega). The primers were 5′-CCCCCATGAGCGTGTACTC-3′ and 5′-ATGCCGTTCAGGGTGATCTTC-3′ for FOXC1, and 5′-TCGTGCGTGACATTAAGGAG-3′ and 5′-ATGCCAGGGTACATGGTGGT-3′ for β-actin.
Western blotting
Cells were lysed in ice-cold lysis buffer (75 mM NaCl, 2 mM EDTA, 1% Triton X-100, and protein inhibitor cocktail). Cytosolic extracts were resolved by SDS-PAGE in 12 or 10% gels dependent on the molecular weight of proteins, transferred to PVDF membranes, and blocked with PBS containing 0.1% Tween-20 (PBST) and 5% defatted milk for 1 h. The membranes were incubated with antibodies for 1 h at 37°C, washed with PBST and incubated with secondary antibodies for 30 min before chemiluminescent detection following the manufacturer’s instruction (ECL Detection System, Pierce). Polyclonal anti-SUZ12 (ab12073) and anti-FOXC1 antibodies were purchased from Abcam (Cambridge, USA). Monoclonal anti-EZH2 and anti-Bmi1 antibodies were purchased from Cell Signaling Technology (CST, Danvers, MA, USA). Monoclonal anti-β-actin antibody was purchased from Sigma (St. Louis, MO, USA).
RNA interference
The target sequences for EZH2, SUZ12, and Bmi1 genes were 5′-GACTCTGAATGCAGTTGCT-3′, 5′-GTCGCAACGGACCAGTTAA-3′, and 5′-TGATGCCACAACCATAATA-3′, respectively. Plasmids were constructed by inserting the target sequences into pSilencer4.1-CMVneo vector (Ambion Applied Biosystems, USA).
Chromatin immunoprecipitation assay
The protocol for chromatin immunoprecipitation (ChIP) was described elsewhere [18]. Briefly, the chromatin solution was precleared with 50 μl of protein A-agarose beads (Upstate Biotechnology). The soluble fraction was collected and 5 μg of anti-acetyl-histone H3 (Upstate Biotechnology), anti-acetyl-histone H4 (Upstate Biotechnology), anti-H3K27me3 (Upstate Biotechnology), anti-EZH2 (Cell Signal Technology), or anti-Bmi1 (Cell Signal Technology) antibodies was added. The precipitated chromatins were analyzed by PCR. Four pairs of primers (P1 to P4) that are specific to FOXC1 promoter regions were: P1, sense: 5′-CAGAACAGCATCCGCCACA-3′, antisense: 5′-CTCCTCCTTGTCCTTCACCG-3′; P2 sense: 5′-CCTGGGTGACGGATGCTCAA-3′, antisense: 5′-CAGACCGCCTTGCAGGAACT-3′; P3 sense: 5′-CACCCCTTGCCTTCATTTCG-3′, antisense: 5′-CCTCGTGCTGGCTCGCTCT-3′; and P4 sense: 5′-TATTCAGGTGACACGGAGATGC-3′, antisense: 5′-GGGTGGTTGGAATGGGACAG-3′.
Trans-well migration assay
The GFP-MDA-MB-231 cells and the FOXC1-MDA-MB-231 cells were starved overnight in assay media. Cells (1 × 105) were added to the top chambers of 24-well trans-well plates (BD Biosciences, 8 μm pore size), and assay media, with or without 10% FBS, were added to the bottom chambers. After overnight incubation, top (nonmigrated) cells were removed, and bottom (migrated) cells were fixed and stained with 4′,6-diamidino-2-phenylindole (5 μg/ml) to visualize nuclei. The number of migrating cells in 5 fields was counted under ×20 magnification, and the means for each chamber were determined.
Trans-well invasion chambers
The cell invasiveness was evaluated in vitro using Matrigel-coated semi-permeable modified Boyden inserts with a pore size of 8 μm (Becton–Dickinson/Biocoat, Bedford, MA). Cells were performed as described in “Trans-well migration assay”.
Tumorigenicity and metastasis assays in athymic mice
A novel breast cancer cell line that was derived from six cycles of pulmonary metastasis implantation into the mammary fat pad (MFP), designated MDA-MB-231HM (high metastasis), displayed significantly enhanced pulmonary metastatic potential compared with its parental counterparts, MDA-MB-231 cells [19]. The spontaneous metastatic capability of different cells was determined using an orthotropic xenograft tumor model in athymic mice, as described previously [19]. For mammary fat pad experiment, animals were divided into GFP-MDA-MB-231HM-implanted and FOXC1-MDA-MB-231HM-implanted groups, each containing seven mice. MDA-MB-231HM cells (2 × 106 cells) were injected orthotropically into the exposed axillary MFP of anesthetized athymic mice. For tail vein experiment, animals were grouped same as fat pad experiment, with five mice in each group. MDA-MB-231HM cells (2 × 105 cells) were injected orthotropically into the exposed axillary MFP of anesthetized athymic mice. Animals were monitored every 2 days for tumor growth and general health. Animals were killed and autopsied 6 weeks post-inoculation. The lungs used to evaluate the numbers of metastasis were fixed in Bouin’s solution for 24 h and stored in 100% ethanol. When the lungs restored to their inherent color, the white metastatic deposits were assessed macroscopically. To confirm the presence of lung metastases, sections were cut at 50 μm intervals and stained with H&E. In this study, the number of metastatic nodules on lung surface was counted. Two independent pathologists calculated the number of metastases.
Results
FOXC1 was negatively regulated by PcG
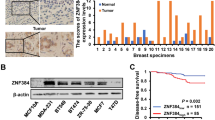
The polycomb group proteins, especially EZH2, were involved in progression of prostate cancer and breast cancer [10, 20, 21]. The expression levels of the PcG proteins EZH2, SUZ12, and Bmi1 were significantly higher in the more malignant MDA-MB-231 cells than in MCF-7 cells (Fig. 1a). To verify whether FOXC1 is regulated by PcGs, RT-PCR and western blotting were performed, and the results demonstrated that the level of FOXC1 expression was higher in MCF-7 cells than in MDA-MB-231 cells (Fig. 1b, c), implicating that the expression of FOXC1 was negatively correlated to PcGs in the two cell lines (Fig. 1). This was further supported by the results of the test of EZH2, Bmi1, and FOXC1 mRNA levels in ten different breast cell lines (Fig. S5). It can be seen that EZH2 and Bmi1 exhibited much lower expression in the breast cell lines listed at the left side of the figure (i.e., MCF10A, MCF7, T47D, and MDA-MB-468) than in the cell lines at the right side (i.e., MDA-MB435, MDA-MB-435H, MDA-MB-231, MD-MB-231H, BT-474, and SKBR3) (Fig S5a, b); meanwhile, FOXC1 showed much higher expression in the left side breast cell lines than in the right side cell lines (Fig S5c). Moreover, we showed that the mRNA (Fig. 2a) and protein (Fig. 2b) levels of FOXC1 were markedly decreased when EZH2, Bmi1, and SUZ12 were overexpressed in MCF-7 cells (Fig. 2c). To further investigate the roles of PcG in regulation of FOXC1, we transfected the siRNA plasmids targeting Bmi1, EZH2, and SUZ12 genes, respectively, into MDA-MB-231 cells. We found that both the FOXC1 mRNA (Fig. 2d) and protein (Fig. 2e) levels were upregulated when Bmi1, EZH2, and SUZ12 were knocked down (Fig. 2f).
Effects of overexpression and knockdown of PcG proteins on FOXC1 gene expression. MCF-7 cells were transfected with EZH2, SUZ12, and Bmi1 expression plasmids, and 48 h later the FOXC1 mRNA and protein levels were determined by PCR (a) and western blotting (b), respectively. The ectopic expression of EZH2, SUZ12, and Bmi1 proteins was confirmed by western blotting (c). MDA-MB-231 cells were transfected with EZH2, SUZ12, and Bmi1 siRNA, and 48 h later RT–PCR and western blotting were performed. The endogenous FOXC1 mRNA (d) and protein (e) levels were upregulated. f Western blotting verification of the interfering efficiency of EZH2, SUZ12, and Bmi1 siRNAs in MDA-MB-231 cells
PcG proteins repressed FOXC1 through histone acetylation and H3K27 tri-methylation at the promoter
Previous data implicated that Polycomb-Repressive Complex 1 (PRC1) and PRC2 repressed target gene expression through H3K27me3 (H3 lysine27 tri-methylation); and PcGs could also repress gene expression by histone acetylation modification [22]. We next examined the binding of PcG proteins, as well as histone acetylation and H3K27me3 modifications, at different regions of FOXC1 promoter (Fig. 3b) in MCF-7 and MDA-MB-231 cells. ChIP assays showed that the binding of endogenous EZH2 at the FOXC1 promoter was much weaker in MCF-7 cells than in MDA-MB-231 cells (Fig. 3b); meanwhile, the acetylation levels of H3 and H4 at these promoter regions were apparently increased, whereas the histone H3K27me3 level was decreased (Fig. 3b). Moreover, parallel ChIP assays demonstrated that the binding of EZH2 at the FOXC1 promoter was reduced in EZH2siRNA-MDA-MB-231 cells (Fig. 3c). The ChIP data also revealed that the H3K27me3 levels at these promoter regions were apparently reduced and the H3/H4 acetylation levels were increased when the endogenous EZH2 was interfered in MDA-MB-231 cells (Fig. 3c). The reduced binding of EZH2 to FOXC1 promoter mainly occurred at the far upstream regions (P3 and P4) of FOXC1 promoter, coincident with the reduction of H3K27me3 level at the same regions (Fig. 3c). However, the increased H3/H4 acetylation levels mainly occurred at the P1 and P2 regions near the transcriptional start site (Fig. 3c). These data indicated that the reduction of H3K27me3 and the increase of H3/H4 acetylation of the FOXC1 promoter coincided with the elevated expression of FOXC1 and the EZH2 binding to the promoter of FOXC1.
EZH2 regulated FOXC1 expression through H3K27 tri-methylation and H3/H4 acetylation. a Diagram of the 5′-flanking region of FOXC1 gene. P1–P3 denote the three promoter regions of FOXC1 gene analyzed in ChIP assays, and P4 is at the distal region. b ChIP assays in MDA-MB-231 (231) and MCF-7 (M7) cells. Chromatin was immunoprecipitated with anti-acetyl-H3, anti-acetyl-H4, anti-H3K27me3, anti-EZH2, and anti-Bmi1 antibodies, respectively. c ChIP assays in MDA-MB-231 (231-c) and MDA-MB-231 with EZH2 knockdown (siEZH2); chromatin was immunoprecipitated with anti-acetyl-H4, anti-H3K27me3, and anti-EZH2 antibodies, respectively. The amounts of precipitated endogenous FOXC1 promoter fragments were determined by PCR using the specific primer sets
FOXC1 prevented cell migration and invasion in MDA-MB-231 cells
Our results above showed that EZH2 bound to FOXC1 promoter and negatively regulated FOXC1 expression, implicating that FOXC1 might participate in breast cancer aggression. To validate this assumption, we analyzed the directional cell motility using wound closure and trans-well migration assays. As shown in Fig. 4a, FOXC1-MDA-MB-231 cells had only completed a half closure at 16 h compared to control GFP-MDA-MB-231 cells. Our trans-well migration assays also demonstrated that GFP-MDA-MB-231 cells migrated approximately three times faster than FOXC1-MDA-MB-231 cells (Fig. 4b). To examine the roles of FOXC1 in regulating cell invasion, we performed trans-well Matrigel invasion assay to assess the ability of the cells to invade through the Matrigel layer. As shown in Fig. 4c, FOXC1-MDA-MB-231 cells invaded much slower than GFP-MDA-MB-231 cells. These findings suggested that FOXC1 inhibited the invasion of MDA-MB-231 cells.
FOXC1 prevented MDA-MB-231 cell migration and invasion. a Confluent FOXC1-MDA-MB-231 and GFP-MDA-MB-231 cells were wounded and a representative micrograph for each well is shown (0, 4, 8, and 16 h after wounding). b FOXC1-MDA-MB-231 and GFP-MDA-MB-231 cells were plated in trans-well chambers in absence or presence of serum. After 24 h, the migrating cells were fixed, stained, and photographed at ×400 magnification under a microscope. The serum-free medium was used as a negative control. c FOXC1 prevented MDA-MB-231 cell invasion. Cell invasiveness was evaluated in vitro using Matrigel-coated semi-permeable modified Boyden inserts as described in the text
FOXC1 inhibited migration and invasion of MDA-MB-231HM cells in vitro, and inhibited the pulmonary metastasis in vivo
A highly metastatic variant of parental MDA-MB-231 cell line, designated MDA-MB-231HM, was a valuable model system for the study of the molecular events underlying breast cancer metastasis [23, 24]. We used this cell line to study the roles of FOXC1 in migration and invasion of breast cancer, and we found that both the FOXC1 mRNA (Fig. 5a) and protein (Fig. 5b) levels were lower in MDA-MB-231HM cells than in MDA-MB-231 cells. Our trans-well migration assays showed that the directional cell motility of FOXC1-MDA-MB-231HM cells was slower than GFP-MDA-MB-231HM cells, and this result was similar to that in MDA-MB-231 (Fig. 5d, e). In addition, we found that HER2 and vimentin, the two markers associated with metastasis, decreased remarkably when FOXC1 was overexpressed (Fig. 5c). These findings suggested that FOXC1 played an important role in the inhibition of invasion of MDA-MB-231HM cells.
FOXC1 prevented MDA-MB-231HM cell migration and invasion. FOXC1 expression was detected by RT-PCR (a) and western blotting (b) in MCF-7, MDA-MB-231 and MDA-MB-231HM cell, respectively. c FOXC1 prevented MDA-MB-231HM cell migration. FOXC1-MDA-MB-231 and GFP-MDA-MB-231 cells were plated in trans-well chambers as described above. d FOXC1 prevented MDA-MB-231HM cell invasion. Cell invasiveness was evaluated in vitro as described above. eThe whole cell lysates of FOXC1-MDA-MB-231HM and GFP-MDA-MB-231HM were prepared for western blotting detection of HER2 and Vimentin
Next, the effect of FOXC1 on metastasis was further assayed in athymic mice. The FOXC1-MDA-MB-231HM cells and the GFP-MDA-MB-231HM cells were implanted to the mammary fat pad and tail vein of the athymic mice. To study pulmonary metastasis, lungs were examined physically at autopsy, and overt surface metastases were inspected. For the mammary fat pad, surface metastases of GFP-MDA-MB-231HM cells were found in six out of seven mice in the GFP-MDA-MB-231HM implant groups, while only two were observed in seven mice in the FOXC1-MDA-MB-231HM group (Fig. 6b). For the tail vein injection group, lung surface metastases were found in four out of five mice in the GFP-MDA-MB-231HM implant group, and only one out of five mice was metastasized in the FOXC1-MDA-MB-231HM group (Fig. 6a). The majority of metastatic colonies were observed in GFP-MDA-MB-231HM implant mice, whereas only a few micrometastases were found in mice of FOXC1-MDA-MB-231HM groups (Fig. 6a). These results indicated that overexpression of FOXC1 was able to inhibit metastasis of MDA-MB-231HM cells in vitro and in vivo.
FOXC1 inhibited pulmonary metastasis of MDA-MB-231HM cells in vivo. a The tail vein experiment. Photomicrographs of micrometastases (arrows) in lung sections from mice injected with FOXC1-MDA-MB-231HM cells and GFP-MDA-MB-231HM cells for 4 weeks (H&E, ×200). b The mammary fat pad (MPF) experiment. Photomicrographs of micrometastases (arrows) of the tumor cell sections in MPF and lung sections in athymic mice bearing FOXC1-MDA-MB-231HM and GFP-MDA-MB-231HM cells for 6 weeks (H&E, ×200). The tumor cells in MPF exhibited little differences between FOXC1-MDA-MB-231HM and GFP-MDA-MB-231HM cell experiment groups, but the lung sections of GFP-MDA-MB-231HM group had more micrometastases (arrows) than that of FOXC1-MDA-MB-231HM group
Discussion
The target genes of PcG family proteins have been extensively investigated. By using a genome-wide location analysis in Drosophila melanogaster, more than 1,000 putative genes were identified that were regulated by PcG proteins [25]. Moreover, PcG proteins are highly conserved in other organisms, and they target many transcription factors such as HOX, SOX, and FOX families [26]. FOXC1 is a member of the Forkhead box transcription factor family, and it plays an important role in the regulation of ocular development [15, 16]. Results from this study validated that FOXC1 was indeed a target gene of the polycomb group protein family (Figs. 1, 2). We showed that all the PcG proteins tested, i.e., EZH2, SUZ12, and Bmi1, could repress the expression of FOXC1; among them EZH2 displayed the most significant inhibitory effect. Moreover, EZH2 played an important role in the progression of breast cancer [11]. We then chose EZH2 as a target for further study on the role of PcGs in regulation of FOXC1 gene. In Drosophila and mammals, PcG proteins commonly control gene expression by influencing the H3K27 methylation status [22, 27]. Besides, the histone deacetylase activity is also involved in gene silencing [28]. Our ChIP assays in this study demonstrated that EZH2 repressed FOXC1 expression through both H3K27 tri-methylation and histone acetylation (Fig. 3b). HDAC1 has been reported to be a component of the silencing complex, and it interacts cooperatively with polycomb group repressors to silence the target genes [28]. Our results implicated that HDACs may function as co-repressors of PcG proteins. However, it still remains an open question of which HDACs function and how to function in this process.
Interestingly, in our experiments, we detected the EZH2 binding and changes of H3K27 methylation even at the P4 distal region of FOXC1 promoter. We speculated that the EZH2 binding sites at FOXC1 promoter were mainly located at the relative distal regions (P3 and P4). Sen et al. [13] identified that H3K27me3 might be a mechanism of FOXC1 regulation by PcG proteins, and this coincided with our results. Moreover, reports from Muggerud et al. [29] showed that the methylation levels of FOXC1 gene displayed substantial differences between normal breast tissue and invasive tumors. Similarly, our ChIP assay also indicated that the methylation levels of FOXC1 was higher in MDA-MB-231 than in MCF-7 (Fig. 3b). PcG proteins lack a DNA binding domain, and they are usually recruited by specific transcription factors. An earlier study demonstrated that YY1 as a transcriptional factor could recruit the PcG proteins to genes’ promoters [30]. However, an analysis of FOXC1 gene detected no classic YY1 binding consensus sequences (data not shown). It is therefore possible that other transcription factors than YY1 may be involved in the recruitment of PcG proteins to FOXC1 gene. Alternatively, PcG proteins may also target the basic transcriptional machinery to directly inhibit the transcription initiation, as previously reported [31].
FOXC2 has been implicated to play a central role in promoting cancer cell invasion and metastasis [32]. Both FOXC1 and FOXC2 are members of the FOX family; however, they play distinct roles in development. FOXC1 was associated with Axenfeld-Rieger anomaly of the anterior eye chamber, while FOXC2 was associated with lymphedema-distichiasis [33]. Presumably, FOXC1 and FOXC2 may also exert different roles in breast cancer metastasis. We discovered in this study that ectopic expression of FOXC1 inhibited migration and invasion in both MDA-MB-231 and MDA-MB-231HM cells (Figs. 4, 5). By using a MDA-MB-231HM xenograft athymic mouse model, we showed that FOXC1 could inhibit the metastasis in both the mammary fat pad and tail vein implanted animal groups (Fig. 6). Results from both animal experiments suggested that FOXC1 could play an inhibitory role in the entire process of metastasis. Moreover, we found that FOXC1 could repress vimentin expression in MDA-MB-231HM cells (Fig. 5c). Vimentin is a signature of epithelial-mesenchymal transition (EMT), which is commonly associated with a high potential of tumor cell invasion [34]. Our results also revealed that FOXC1 could inhibit the expression of E-cadherin (Fig. S4), an epithelial marker that was downregulated in EMT progress. Based on these clues, we speculate that FOXC1 may inhibit metastasis by affecting the EMT progress, although this assumption requires further experiments to confirm. Muggerud et al. [29] reported that FOXC1 gene displayed a significantly increased methylation level in invasive breast tumors compared with normal breast tissues, with a concurrent decrease of FOXC1 expression. These data from the breast tissue were coincident with our results in breast cancer cells, in which MCF-7 cells expressed a higher level of FOXC1 than the malignant MDA-MB-231 cells (Fig. 1b, c). Concurrently, the H3K27 methylation of FOXC1 gene was lower in MCF-7 cells than in malignant MDA-MB-231 cells (Fig. 3b). Thus, both Muggerud’s and our results suggested that FOXC1 expression was negatively correlated with breast cancer progression. A previous study also showed that the expression of FOXC1 was lower in ovarian cancer than in normal primary endometrial [35]. Breast cancer and ovarian cancer behave similarly in the progression; therefore, it may be rational to comprehend that FOXC1 also played a similar role in inhibiting the breast cancer.
Altogether, experimental data presented in this report suggest that FOXC1 is a target of the polycomb group protein family, and PcG family proteins negatively regulate FOXC1 gene expression through H3K27 tri-methylation and H3/H4 acetylation of FOXC1 promoter. We identified downregulation of FOXC1 in highly invasive and potentially metastatic MDA-MB-231HM breast cancer cells compared with their parental MDA-MB-231 cells. When FOXC1 was stably transfected into MDA-MB-231 cells and MDA-MB-231HM cells, it prevented cell migration and invasion. Moreover, ectopic expression of FOXC1 in MDA-MB-231HM cells inhibited the pulmonary metastasis in both tail vein injection and in mammary fat pad implant athymic mice. Data presented in this report contribute to the understanding of the mechanisms by which EZH2 regulates tumor development, as well as the role of FOXC1 in the progression of breast cancer.
Abbreviations
- PcG:
-
Polycomb group
- EZH2:
-
The enhancer of zeste homologue 2
- H3K27:
-
Lysine 27 of histone H3
- ChIP:
-
Chromatin immunoprecipitation
- YY1:
-
Yin-Yang-1
- EMT:
-
Epithelial-mesenchymal transition
References
Jemal A, Murray T, Samuels A, Ghafoor A, Ward E, Thun MJ (2003) Cancer statistics. CA Cancer J Clin 53(1):5–26
Ellis M, Heyas D, Lippman M (2000) Treatment of metastatic disease. Dis Breast 2000:749–798
Hayes DF (2000) Do we need prognostic factors in nodal-negative breast cancer? Arbiter. Eur J Cancer 36(3):302–306
Yamashita J, Ogawa M, Shirakusa T (1995) Free-form neutrophil elastase is an independent marker predicting recurrence in primary breast cancer. J Leukoc Biol 57(3):375–378
Orlando V (2003) Polycomb, epigenomes, and control of cell identity. Cell 112(5):599–606
Shao Z, Raible F, Mollaaghababa R, Guyon JR, Wu CT, Bender W, Kingston RE (1999) Stabilization of chromatin structure by PRC1, a polycomb complex. Cell 98(1):37–46
Sparmann A, van Lohuizen M (2006) Polycomb silencers control cell fate, development and cancer. Nat Rev Cancer 6(11):846–856
Bracken AP, Pasini D, Capra M, Prosperini E, Colli E, Helin K (2003) EZH2 is downstream of the pRB-E2F pathway, essential for proliferation and amplified in cancer. EMBO J 22(20):5323–5335
Raaphorst FM, Meijer CJ, Fieret E, Blokzijl T, Mommers E, Buerger H, Packeisen J, Sewalt RA, Otte AP, van Diest PJ (2003) Poorly differentiated breast carcinoma is associated with increased expression of the human polycomb group EZH2 gene. Neoplasia 5(6):481–488
Kleer CG, Cao Q, Varambally S, Shen R, Ota I, Tomlins SA, Ghosh D, Sewalt RG, Otte AP, Hayes DF et al (2003) EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc Natl Acad Sci USA 100(20):11606–11611
Zeidler M, Kleer CG (2006) The polycomb group protein Enhancer of Zeste 2: its links to DNA repair and breast cancer. J Mol Histol 37(5–7):219–223
Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P, Jones RS, Zhang Y (2002) Role of histone H3 lysine 27 methylation in polycomb-group silencing. Science 298(5595):1039–1043
Sen GL, Webster DE, Barragan DI, Chang HY, Khavari PA (2008) Control of differentiation in a self-renewing mammalian tissue by the histone demethylase JMJD3. Genes Dev 22(14):1865–1870
Makarevich G, Leroy O, Akinci U, Schubert D, Clarenz O, Goodrich J, Grossniklaus U, Kohler C (2006) Different polycomb group complexes regulate common target genes in Arabidopsis. EMBO Rep 7(9):947–952
Mortemousque B, Amati-Bonneau P, Couture F, Graffan R, Dubois S, Colin J, Bonneau D, Morissette J, Lacombe D, Raymond V (2004) Axenfeld-Rieger anomaly: a novel mutation in the forkhead box C1 (FOXC1) gene in a 4-generation family. Arch Ophthalmol 122(10):1527–1533
Aldinger KA, Lehmann OJ, Hudgins L, Chizhikov VV, Bassuk AG, Ades LC, Krantz ID, Dobyns WB, Millen KJ (2009) FOXC1 is required for normal cerebellar development and is a major contributor to chromosome 6p25.3 Dandy-Walker malformation. Nat Genet 41(9):1037–1042
Lehmann OJ, Sowden JC, Carlsson P, Jordan T, Bhattacharya SS (2003) Fox’s in development and disease. Trends Genet 19(6):339–344
Wang X, Pan L, Feng Y, Wang Y, Han Q, Han L, Han S, Guo J, Huang B, Lu J (2008) P300 plays a role in p16(INK4a) expression and cell cycle arrest. Oncogene 27(13):1894–1904
Hou YF, Yuan ST, Li HC, Wu J, Lu JS, Liu G, Lu LJ, Shen ZZ, Ding J, Shao ZM (2004) ERbeta exerts multiple stimulative effects on human breast carcinoma cells. Oncogene 23(34):5799–5806
Bloushtain-Qimron N, Yao J, Snyder EL, Shipitsin M, Campbell LL, Mani SA, Hu M, Chen H, Ustyansky V, Antosiewicz JE et al (2008) Cell type-specific DNA methylation patterns in the human breast. Proc Natl Acad Sci USA 105(37):14076–14081
Kim JH, Yoon SY, Jeong SH, Kim SY, Moon SK, Joo JH, Lee Y, Choe IS, Kim JW (2004) Overexpression of Bmi-1 oncoprotein correlates with axillary lymph node metastases in invasive ductal breast cancer. Breast 13(5):383–388
Pirrotta V, Poux S, Melfi R, Pilyugin M (2003) Assembly of Polycomb complexes and silencing mechanisms. Genetica 117(2–3):191–197
Liu ZB, Hou YF, Di GH, Wu J, Shen ZZ, Shao ZM (2009) PA-MSHA inhibits proliferation and induces apoptosis through the up-regulation and activation of caspases in the human breast cancer cell lines. J Cell Biochem 108(1):195–206
Chang XZ, Li DQ, Hou YF, Wu J, Lu JS, Di GH, Jin W, Ou ZL, Shen ZZ, Shao ZM (2008) Identification of the functional role of AF1Q in the progression of breast cancer. Breast Cancer Res Treat 111(1):65–78
Schwartz YB, Kahn TG, Nix DA, Li XY, Bourgon R, Biggin M, Pirrotta V (2006) Genome-wide analysis of polycomb targets in Drosophila melanogaster. Nat Genet 38(6):700–705
Bracken AP, Dietrich N, Pasini D, Hansen KH, Helin K (2006) Genome-wide mapping of Polycomb target genes unravels their roles in cell fate transitions. Genes Dev 20(9):1123–1136
Grimaud C, Negre N, Cavalli G (2006) From genetics to epigenetics: the tale of polycomb group and trithorax group genes. Chromosome Res 14(4):363–375
Chang YL, Peng YH, Pan IC, Sun DS, King B, Huang DH (2001) Essential role of Drosophila Hdac1 in homeotic gene silencing. Proc Natl Acad Sci USA 98(17):9730–9735
Muggerud AA, Ronneberg JA, Warnberg F, Botling J, Busato F, Jovanovic J, Solvang H, Bukholm I, Borresen-Dale AL, Kristensen VN et al (2010) Frequent aberrant DNA methylation of ABCB1, FOXC1, PPP2R2B and PTEN in ductal carcinoma in situ and early invasive breast cancer. Breast Cancer Res 12(1):R3
Srinivasan L, Atchison ML (2004) YY1 DNA binding and PcG recruitment requires CtBP. Genes Dev 18(21):2596–2601
Dellino GI, Schwartz YB, Farkas G, McCabe D, Elgin SC, Pirrotta V (2004) Polycomb silencing blocks transcription initiation. Mol Cell 13(6):887–893
Mani SA, Yang J, Brooks M, Schwaninger G, Zhou A, Miura N, Kutok JL, Hartwell K, Richardson AL, Weinberg RA (2007) Mesenchyme Forkhead 1 (FOXC2) plays a key role in metastasis and is associated with aggressive basal-like breast cancers. Proc Natl Acad Sci USA 104(24):10069–10074
Hannenhalli S, Kaestner KH (2009) The evolution of Fox genes and their role in development and disease. Nat Rev Genet 10(4):233–240
Thiery JP (2002) Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2(6):442–454
Zhou Y, Kato H, Asanoma K, Kondo H, Arima T, Kato K, Matsuda T, Wake N (2002) Identification of FOXC1 as a TGF-beta1 responsive gene and its involvement in negative regulation of cell growth. Genomics 80(5):465–472
Acknowledgments
This study was supported by grants from The National Natural Science Foundation of China (30671184 and 30971613), The Program for Changjiang Scholars and Innovative Research Team (PCSIRT) in Universities (IRT0519), and The Fundamental Research Funds for the Central Universities.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Juan Du and Lin Li contributed equally to this study.
An erratum to this article is available at http://dx.doi.org/10.1007/s10549-016-3772-5.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10549_2011_1396_MOESM1_ESM.docx
Supplement Fig. 1. Generation of stable MDA-MB-231 cell lines with constitutive FOXC1 overexpression. MDA-MB-231 cells were transfected with FOXC1-pEGFP-N1 and pEGFP-N1 control plasmids using Lipofectamine-2000. After 48 h of incubation, cells were selected for 8 weeks using G418 (800 μg/ml). Western blots detecting the constitutive expression of FOXC1 and GFP in FOXC1-MDA-MB-231 and GFP-MDA-MB-231 cells with specific GFP antibody. (DOCX 121 kb)
10549_2011_1396_MOESM2_ESM.docx
Supplement Figs. 2, 3. Generation of stable MDA-MB-231HM cell line with FOXC1 overexpression. MDA-MB-231HM cells were transfected with FOXC1-pEGFP-N1 and pEGFP-N1 control plasmids using Lipofectamine-2000. After 48 h of incubation, cells were selected for 8 weeks using G418 (800 μg/ml). RT-PCR (S2) and western blotting (S3) detecting the constitutive expression of FOXC1 and GFP in FOXC1-MDA-MB-231HM and GFP-MDA-MB-231HM cells, were performed. (DOCX 99 kb)
10549_2011_1396_MOESM3_ESM.docx
Supplement Fig. 4. Knockdown of FOXC1 inhibited E-cadherin expression in MCF-7 cells. Cells were transfected with FOXC1siRNA1# and FOXC1siRNA2# vectors, and after 48 h, the whole cell lysates were extracted for western blotting. E-cadherin protein level was determined by western blotting (middle). The interfering efficiency of FOXC1siRNAs in MCF-7 cells was verified by western blotting (top). (DOCX 122 kb)
10549_2011_1396_MOESM4_ESM.docx
Supplement Fig. 5. Expression of EZH2, Bmi1, and FOXC1 in ten breast cells. Real-time RT-PCR assessments of EZH2 (A), Bmi1 (B) and FOXC1 (C) mRNAs in ten breast cancer cells are shown. The ten breast cell lines were as follows: MCF10A, MCF7, T47D, MDA-MB-468, MDA-MB435, MDA-MB-435H, MDA-MB-231, MD-MB-231H, BT-474, SKBR3. (DOCX 112 kb)
Rights and permissions
About this article
Cite this article
Du, J., Li, L., Ou, Z. et al. FOXC1, a target of polycomb, inhibits metastasis of breast cancer cells. Breast Cancer Res Treat 131, 65–73 (2012). https://doi.org/10.1007/s10549-011-1396-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-011-1396-3