Abstract
Objectives
To explore systemic effects of mutations in the UDP-N-acetylmuramoylalanyl-d-glutamate 2,6-diaminopimelate ligase (MurE) of Corynebacterium glutamicum, that leads to extracellular l-lysine accumulation by this bacterium.
Results
The analysis of a mutant cohort of C. glutamicum strains carrying all possible 20 amino acids at position 81 of MurE revealed unexpected effects on cellular properties. With increasing l-lysine accumulation the growth rate of the producing strain is reduced. A dynamic flux balance analysis including the flux over MurE fully supports this finding and suggests that further reductions at this flux control point would enhance l-lysine accumulation even further. The strain carrying the best MurE variant MurE-G81K produces 37 mM l-lysine with a yield of 0.17 g/g (l-lysine·HCl/glucose·H2O), bearing no other genetic modification. Interestingly, among the strains with high l-lysine titers, strain variants occur which, despite possessing the desired amino acid substitutions in MurE, have regained close to normal growth and correspondingly lower l-lysine accumulation. Genome analyses of such variants revealed the transposition of mobile genetic elements which apparently annulled the favorable consequences of the MurE mutations on l-lysine formation.
Conclusion
MurE is an attractive target to achieve high l-lysine accumulation, and product formation is inversely related to the specific growth rate. Moreover, single point mutations leading to elevated l-lysine titers may cause systemic effects on different levels comprising also major genome modifications. The latter caused by the activity of mobile genetic elements, most likely due to the stress conditions being characteristic for microbial metabolite producers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
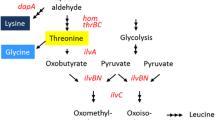
The UDP-N-acetylmuramoylalanyl-d-glutamate 2,6-diaminopimelate ligase activity, encoded by murE, is essential in bacteria (Mengin-Lecreulx et al. 1989). MurE links meso-diaminopimelate in an ATP-dependent manner to the dipeptide UDP-N-acetyl-α-d-muramoyl-l-alanyl-d-glutamate (UDP-MurNAc-l-Ala-d-Glu). After two additional elongation reactions of the tripeptide, the resulting pentapeptide becomes a constituent of peptidoglycan providing strength and rigidity of the cell wall for protection against the environment and to counteract internal osmotic pressure. However, in addition to being a substrate for MurE, meso-diaminopimelate is also a substrate for LysA, the diaminopimelate decarboxylase catalyzing the last step of l-lysine synthesis. Hence, meso-diaminopimelate is at a branch point of cell wall and l-lysine synthesis. This is, of course, of relevance, when increased l-lysine synthesis is required for production purposes. Currently, l-lysine is produced with mutants of Corynebacterium glutamicum or Escherichia coli on a scale of 2,200,000 t/a (Eggeling and Bott 2015). We recently identified in a biosensor-based single cell screening approach the C. glutamicum mutations MurE-G81E and MurE-L121F both resulting in increased l-lysine formation (Binder et al. 2012). We also introduced recombinant engineering for C. glutamicum which, in combination with the screening approach, was called RecFACS (Binder et al. 2013). Using a mixture of mutagenic oligonucleotides, RecFACS enabled the isolation in one single experiment of l-lysine-excreting mutants substituting G81 in MurE by 12 different amino acids. Inspection of available MurE structure models of Escherichia coli and Mycobacterium tuberculosis revealed that C. glutamicum-MurE-G81 is close to the nucleoside binding site of UDP-MurNAc-l-Ala-d-Glu (Basavannacharya et al. 2010). This might be the reason for the observed tolerated variability at this site. Driven by these findings we now wanted to know whether an even more increased tolerance to other amino acids at the specific MurE-G81 position is possible, and to take measure of systemic consequences of the cell including l-lysine production.
Materials and methods
For strain constructions we generated 20 pK19mobsacB derivatives carrying the same murE-fragment 2289521-2290250 (BA000036), but with different codons at nucleotide position 2290844-46 corresponding to all possible amino acids at position 81 of MurE, except native glycine. (Details are given in the supporting information—Supplementary Table 1). The plasmids were used to positively select for double crossover events with the genome of C. glutamicum ATCC13032 to perform allelic exchanges by homologous recombination (Schäfer et al. 1994). For each mutation, after the first positive selection for integration of the pK19mobsacB derivative, at least two independent clones were used for the second positive selection and loss of plasmid. Colony PCR and sequencing was used to assay whether the mutation was introduced in the final strain or the wild type situation retained.
Strain characterizations were performed in 48-well flowerplates using the BioLector system (m2p-labs GmbH, 52499 Baesweiler, Germany). To do this cells were pregrown in CGXII medium containing 4 % (w/v) glucose as substrate (Reyes and Eggeling 2005). This overnight culture was used as inoculum for the main culture which was grown on the same medium. Cultivations were done at 30 °C, 990 rpm and a throw of 3 mm. Growth was monitored online, and samples for l-lysine determination taken after 42 h when cells were in stationary phase.
Genomic DNA was sequenced by GATC Biotech AG (Konstanz, Germany). Raw sequencing reads were quality trimmed and mapped to the C. glutamicum ATCC13032 reference genome (BA000036) using CLC Genomics Workbench (Version 7.5.1, Qiagen Aarhus A/S). Single nucleotide variants and larger structural variants (SVs) were detected and manually analyzed. To verify assumed insertions of mobile elements, the reference sequence was adjusted and used for re-mapping of reads.
Dynamic flux balance analysis (DFBA) was performed using the previously published genome-scale model of C. glutamicum (Zelle et al. 2015). In short, DFBA extends flux balance analysis and enables the combined simulation of both intracellular and extracellular environments of microbial cultivation processes (for more details see for example Gomez et al. 2014). The DFBA model contains two coupled parts: (i) a dynamic part comprising mass balances for the extracellular substrate d-glucose, the C. glutamicum biomass and the product l-lysine, and (ii) a static optimization part for the simulation of intracellular fluxes with growth as cellular objective function. The uptake rate for d-glucose was modelled via simple Monod kinetics with vupt,max = 4.8 mmol/gCDW:h and K S = 5 mM. The network model was extended by one additional intermediate reaction step (v murE ) to allow for studying the effect of reduced UDP-N-acetylmuramoylalanyl-d-glutamate 2,6-diaminopimelate ligase activity on biomass growth and l-lysine formation. All simulations were performed using MATLAB (Mathworks, R2015a) in combination with the ODE45 solver for the solution of the dynamic model part and the COBRA toolbox with LP solver GLPK for the solution of the static optimization part.
Results and discussion
Using the pK19mobsacB system for allelic replacement of genomic regions (Schäfer et al. 1994), we generated strains derived from the type strain ATCC13032 carrying almost all possible amino acid exchanges in position 81 of MurE. Despite several attempts the substitution from glycine to tyrosine could not be obtained. This was surprising as clones carrying this specific mutation could be generated and isolated with RecFACS (Binder et al. 2013). However, the fact that viable mutants could be obtained in which G81 is substituted by amino acids spanning the entire range of charged, over polar to hydrophobic residues, suggests an exceptional flexibility of MurE at this particular site both from the structural and cellular view.
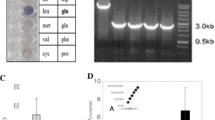
For each amino acid substitution, at least two independently generated clones were grown to determine l-lysine accumulation (Supplementary Table 2). Substitution of G by S, H, T, or A had hardly any influence on l-lysine formation, whereas exchange to Q, F, L, W, or R resulted in a strong increase of l-lysine accumulation up to 19 mM (Fig. 1). In these cases, both clones prepared independently for each amino acid substitution accumulated similar amounts of l-lysine. Interestingly, among the MurE variants accumulating more than 19 mM l-lysine in part large clonal variabilities could be observed. For instance, the two clones carrying the G81N substitution accumulated 23 and 4 mM l-lysine, respectively. A similar variance regarding l-lysine formation could be detected for the three clones with G81C or G81V substitutions. In contrast, both clones bearing the G81E or G81D substitutions in MurE exhibited comparable l-lysine accumulations at a very high level. Although two separately generated clones do not allow for deriving statistically significant conclusions from the obtained product titers, the performance of the latter two mutants could be related with the positively charged residues introduced in the UDP-N-acetylmuramyl-tripeptide synthetase. According to the homology model of Cg-MurE calculated with HHPred (Söding et al. 2005) based on the structure of Mt-MurE with dipeptide and ADP (Basavannacharya et al. 2010) position G81 is located close to the nucleoside binding site of UDP-MurNAc-l-Ala-d-Glu. It is not directly lining the substrate channel, but in the second shell within a short helix. Here in particular, large branched side chains and charged residues can disturb the orientation of this helix and thereby affecting the neighbouring residues of the UDP binding loop from T56 to F63 involved in substrate binding (Supplementary Fig. 1).
Effect of individual amino acid substitutions at position G81 in MurE on l-lysine production with C. glutamicum. The Asterisk denotes l-lysine formation obtained with two clones of the wild type with native glycine in position 81 of MurE. The strains carrying the specific amino acid replacements in MurE-G81 are given as single letters corresponding to the IUPAC code. In each case two independently derived clones were analyzed, except for substitution G81C and G81V, where experiments were performed for three clones. Data represent average values and standard deviations from three independent cultivations
The obtained results inevitably raise two questions: Are there cellular consequences of the drain of meso-diaminopimelate towards l-lysine, and what is the reason for the clonal variability observed? It is obvious to conclude from cell physiology and the MurE structure, that most of the substitutions of native G in position 81 of MurE reduce the enzyme activity. Light microscopic inspection revealed no phenotype of three selected mutants G81E, G81K, G81R, and none of these mutants exhibited altered sensitivity to vancomycin, benzylpenicillin, or ethambutol which are well known inhibitors of cell wall synthesis in C. glutamicum (Möker et al. 2004). However, we observed reduced growth rates of the three selected strains and therefore checked growth of all MurE-variants for a correlation with l-lysine accumulation (Supplementary Table 2). Interestingly, with increasing l-lysine titer the maximal specific growth rate is reduced. The dots in Fig. 2b show this correlation with the l-lysine titer given on the y-axis and the growth rate on the upper x-axis. A dynamic flux balance analysis was performed, coupling dynamic mass balances with an optimization part with growth as cellular objective function (Zelle et al. 2015). When this was done for a constant glucose uptake rate, and assuming a reduction of the MurE flux from 100 % (set as 1) for the C. glutamicum wild-type strain down to 70 % for the strain bearing the MurE-G81K mutation, an almost perfect fit of the simulated data and the experimental data could be obtained (Fig. 2b, solid line). Consequently, the model allows us to predict that a maximum l-lysine titer of 137 mM at a remaining MurE activity of 20 % is possible, suggesting that there is still room for a further improvement of l-lysine production by targeting MurE and the ensuing assembly line towards mureine. Interestingly, also a set of C. glutamicum strains with an engineered pentose phosphate pathway or an engineered glycolysis exhibited reduced growth at increased l-lysine accumulation (Becker et al. 2011). Whereas in the latter study the limiting growth could be due to the availability of building blocks or the uptake of sugar, in our study the favorite candidate limiting growth is the availability of the MurE product UDP-N-acetylmuramoylalanyl-d-glutamate 2,6-diaminopimelate.
Systemic effect of MurE activity reduction on growth and l-lysine production in C. glutamicum. a The reduction in MurE activity limits the availability of the precursor meso-diaminopimelate for peptidoglycan biosynthesis. b Correlation between extracellular l-lysine titer and maximum specific growth rates for the constructed MurE variants. The dots give the experimentally determined values, with the data for each individual clone of the cohorte also given as supplementary information (Supplementary Table 2). The solid line represents the results of the dynamic flux balance analysis. To do this, a constant glucose uptake rate of 4.8 mmol/gCDW/h is assumed and a reduction of the MurE flux from 100 % for wild-type MurE (set as 1) to 70 % for MurE-G81K, with the flux given on the lower x-axis. The model used enables the prediction of growth and l-lysine titers at even lower MurE fluxes with a maximum l-lysine titer of 137 mM at a remaining MurE flux of only 20 %. At still lower MurE fluxes, biomass limitation prevents an additional increase of the maximum obtainable l-lysine titer
A possible explanation for the unexpected clonal variability among clones accumulating higher l-lysine concentrations might be the occurrence of suppressor mutations resulting in either reduced decarboxylation of meso-diaminopimelate to l-lysine or reduced l-lysine export (Fig. 2a). However, no mutations in lysA, lysE, lysG, or their promoters could be found. We therefore performed paired-end genome sequencing of the clones C. glutamicum MurE-G81K2, C. glutamicum MurE-G81V3, both with low l-lysine titer, and C. glutamicum wild type ATCC13032 on an Illumina HiSeq 2000. The three strains shared 21 SNP´s, among them 12 transitions, seven transversions, and two deletions (compared to BA000036), apparently arisen during strain storage over 37 years in our laboratory. Strain V3 carried an additional silent mutation. Moreover, the strains exhibited structural variations due to mobile DNA elements. C. glutamicum has at least 24 insertion elements (IS) in its genome (Kalinowski et al. 2003), and variants of C. glutamicum ATCC13032 differing in prophage content are documented (Kalinowski 2005). Although we were unable to comprehensively fix all structural variations in the three strains we analyzed, some were unequivocally verified by adjusting the reference sequence according to assumed transposon insertion sites and subsequent mapping of sequencing reads (Fig. 3; Supplementary Table 3).
Insertion elements and their observed transposition in different C. glutamicum strains. In the C. glutamicum wild type (WT) used in the study, IS elements present in Cgl1066 and Cgl1580 have duplicated and inserted into Cgl1992 and Cgl2168, respectively. In C. glutamicum MurE-G81K2 an IS element in Cgl2479 has additionally duplicated and inserted between Cgl0530 and Cgl0531. In clone C. glutamicum MurE-G81V3 a new IS element is present in Cgl2265 either derived from that inserted in Cgl1580 or Cgl2549. Locations are approximately drawn to scale which is in megabase pairs (Mb)
In the C. glutamicum wild type we found IS sequences present within Cgl1066 duplicated that had transposed to Cgl1992. Similarly, Cgl1580 was duplicated and present in the possible promoter region of Cgl2168. Strain K2 exhibited a partial duplication of an IS element within Cgl2479 shifted to an intergenic region between Cgl0530 and Cgl0531. In strain V3 an additional IS sequence is present in Cgl2265, which could be due to either Cgl1580 or Cgl2549, the latter two exhibiting sequence identity. There is no apparent linkage of the allocated gene regions to l-lysine or cell wall synthesis. However, the data show that unexpected genome alterations in C. glutamicum during strain construction are frequent.
In summary, MurE is an attractive genomic target for increasing l-lysine synthesis. The l-lysine titers obtained with the MurE-variants presented in this study are in the range of C. glutamicum strains harboring mutations in the lysC-gene encoding the aspartate kinase, which is the key enzyme of the l-lysine synthesis pathway (Eggeling and Bott 2015). The conducted genome analyses revealed that the strain stability should be put more into focus during the construction of high-level producer strains. Undesirable IS-mediated mutagenesis occurs frequently during metabolic engineering of E. coli, which may influence strain physiology (Posfai et al. 2006). In fact, E. coli MSD42, genetically stabilized by removing all mobile DNA elements from E. coli MG1655, accumulated 303 mg l-threonine/l after introducing five specific mutations to increase synthesis of this amino acid, whereas introduction of the same mutations in E. coli MG1655 in parallel led to only 165 mg l-threonine/l (Lee et al. 2009). Therefore, presence and transposition of mobile elements might confuse data interpretation during metabolic engineering approaches, in particular when only single strains are analysed, making it a tempting option to have an IS-free variant of C. glutamicum available.
References
Basavannacharya C, Moody PR, Munshi T, Cronin N, Keep NH, Bhakta S (2010) Essential residues for the enzyme activity of ATP-dependent mure ligase from Mycobacterium tuberculosis. Protein Cell 1:1011
Becker J, Zelder O, Häfner S, Schröder H, Wittmann C (2011) From zero to hero—design-based systems metabolic engineering of Corynebacterium glutamicum for l-lysine production. Metab Eng 13:159–168
Binder S, Schendzielorz G, Stäbler N, Krumbach K, Hoffmann K, Bott M, Eggeling L (2012) A high-throughput approach to identify genomic variants of bacterial metabolite producers at the single-cell level. Genome Biol 13:R40
Binder S, Siedler S, Marienhagen J, Bott M, Eggeling L (2013) Recombineering in Corynebacterium glutamicum combined with optical nanosensors: a general strategy for fast producer strain generation. Nucleic Acids Res 41:6360–6369
Eggeling L, Bott M (2015) A giant market and a powerful metabolism: l-lysine provided by Corynebacterium glutamicum. Appl Microbiol Biotechnol 99:3387–3394
Gomez J, Höffner K, Barton P (2014) DFBAlab: a fast and reliable matlab code for dynamic flux balance analysis. BMC Bioinform 15:1–10
Kalinowski J (2005) The genomes of amino acid-producing Corynebacteria. In: Eggeling, Bott (eds) Handbook of Corynebacterium glutamicum. Taylor & Francis, Boca Raton, pp 37–56
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Krämer R, Linke B, McHardy AC, Meyer F, Möckel B, Pfefferle W, Pühler A, Rey DA, Rückert C, Rupp O, Sahm H, Wendisch VF, Wiegräbe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of l-aspartate-derived amino acids and vitamins. J Biotechnol 104:5–25
Lee JH, Sung BH, Kim MS, Blattner FR, Yoon BH, Kim JH, Kim SC (2009) Metabolic engineering of a reduced-genome strain of Escherichia coli for l-threonine production. Microb Cell Fact 8:2
Mengin-Lecreulx D, Parquet C, Desviat LR, Plá J, Flouret B, Ayala JA, van Heijenoort J (1989) Organization of the murE–murG region of Escherichia coli: identification of the murD gene encoding the d-glutamic-acid-adding enzyme. J Bacteriol 171:6126–6134
Möker N, Brocker M, Schaffer S, Krämer R, Morbach S, Bott M (2004) Deletion of the genes encoding the MtrA–MtrB two-component system of Corynebacterium glutamicum has a strong influence on cell morphology, antibiotics susceptibility and expression of genes involved in osmoprotection. Mol Microbiol 54:420–438
Pósfai G, Plunkett G 3rd, Fehér T, Frisch D, Keil GM, Umenhoffer K, Kolisnychenko V, Stahl B, Sharma SS, de Arruda M, Burland V, Harcum SW, Blattner FR (2006) Emergent properties of reduced-genome Escherichia coli. Science 312:1044–1046
Reyes O, Eggeling L (2005) Experiments. In: Eggeling, Bott (eds) Handbook of Corynebacterium glutamicum. Taylor & Francis, Boca Raton, pp 535–566
Schäfer A, Tauch A, Jäger W, Kalinowski J, Thierbach G, Pühler A (1994) Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145:69–73
Söding J, Biegert A, Lupas AN (2005) The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res 33:W244–W248
Zelle E, Nöh K, Wiechert W (2015) Growth and production capabilities of Corynebacterium glutamicum: interrogating a genome-scale metabolic network model. In: Burkowski A (ed) Corynebacterium glutamicum: from systems biology to biotechnological applications. Caister Academic Press, Erlangen, pp 39–54
Acknowledgments
JH was supported by a fellowship from the CLIB Graduate Cluster Industrial Biotechnology (http://www.graduatecluster.net/).
Supporting information
Supplementary procedures—allelic exchange of murE mutations and dynamic flux balance analysis.
Supplementary Table 1—List of plasmids and primers.
Supplementary Table 2—Lysine accumulations and growth rates of clones.
Supplementary Table 3—Structural variants detected by genome sequencing.
Supplementary Fig. 1—Structure of MurE of C. glutamicum.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hochheim, J., Kranz, A., Krumbach, K. et al. Mutations in MurE, the essential UDP-N-acetylmuramoylalanyl-d-glutamate 2,6-diaminopimelate ligase of Corynebacterium glutamicum: effect on l-lysine formation and analysis of systemic consequences. Biotechnol Lett 39, 283–288 (2017). https://doi.org/10.1007/s10529-016-2243-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-016-2243-8