Abstract
Ustilago scitaminea is the causal agent of sugar-cane smut disease. There is, however, no genetic transformation method for it. Here we report the development of an efficient mutagenesis method based on Agrobacterium tumefaciens-mediated transformation. To improve transformation efficiency, a range of conditions, including the codon-usage preference of the selection marker gene, promoters and the culture conditions for transformation were optimized. A strong promoter to drive marker gene expression, optimized codon usage of selection marker gene, controlled water content and pH of co-culture medium were critical factors affecting transformation efficiency. Our findings provide a useful tool for genetic analysis of this important plant pathogen.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ustilago scitaminea Syd. (also known as Sporisorium scitamineum) is the causal agent for sugar-cane smut, which substantially decreases sucrose content, cane yield and sugar yield (Ferreira and Comstock 1989). First reported in South Africa in 1877, the disease has since spread to nearly all sugar-cane-growing regions in the world (Sundar et al. 2012). Like U. maydis, U. scitaminea is bisexual and dimorphic. It can be forced to mate with a compatible U. maydis haploid strain and the hybrid strain is no longer pathogenic to maize or sugar-cane (Bakkeren and Kronstad 1996). U. scitaminea produces a glycolipid biosurfactant with several potential biotechnological applications (Morita et al. 2011). As a crop with important bio-energy and food applications, sugar-cane has attracted considerable new research interests, including the pathobiology of U. scitaminea. However, the lack of a stable transformation method has become a major roadblock in the understanding of pathobiology, development of effective disease control strategies and industrial applications of this fungus.
Agrobacterium tumefaciens-mediated transformation (ATMT) (Bundock et al. 1995) is a major breakthrough in mutagenesis and transformation of fungal organisms (de Groot et al. 1998; Mullins et al. 2001; Michielse et al. 2005; Shi et al. 2012). Nevertheless, transformation of many fungal species remains a serious challenge. A range of factors affect transformation efficiency in a species-dependent manner. For example, the selection marker genes derived from bacteria are often expressed poorly in eukaryotes due to the differences in codon utilization patterns (Weng et al. 2006; Liu et al. 2013). Two recent reports imply that the protein expression level of selection cassettes is a major limiting factor (Abbott et al. 2013; Liu et al. 2013). In this study, our preliminary results showed that the method for transformation of U. maydis was not suitable for transformation of U. scitaminea. Thus, we examined various factors that could affect transformation efficiency and developed a successful ATMT for U. scitaminea by optimization of the condon usage of the selection marker gene and co-culture/selection conditions. In addition, we found that a strong promoter for driving marker gene expression is also key for successful transformation.
Materials and methods
Strains
Ustilago scitaminea strain S10, isolated from Sugarcane F134 in Guangzhou, China, was maintained on YPD medium. Agrobacterium tumefaciens strain AGL1 (Lazo et al. 1991) was used for its transformation. Escherichia coli XL1-Blue (Agilent Technologies, USA) was used to recombinant DNA work.
DNA constructs
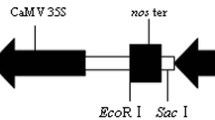
Information on oligonucleotides used is in Supplementary Table 1. Binary plasmid pEX1GPD-EGFP was described previously (Liu et al. 2013). Plasmid pEX0 was constructed by inserting the annealed products of oligonucleotides LoxP1 and LoxP2 into the EcoRI and XbaI sites of pPZP200. The original hygromycin phosphotransferase gene (hpt) of E. coli and a synthetic codon-optimized version of it (hpt-2) (GenBank Accession Number JF412803), both linked to the 35S transcriptional terminator of cauliflower mosaic virus (CMV), were amplified using oligonucleotide pair HPTU/T35SL and HPT2U/T35SL, respectively. The PCR products were blunt-ended with T4 DNA polymerase, digested with BspHI and phosphorylated at the 3′-end with T4 polynucleotide kinase. The gpdA promoter from a locally isolated strain of Aspergillus niger (SG1) was amplified by using oligonucleotides ANGPDU and ANGPDL while the A. niger tefA promoter was amplified with ANTEFU and ANTEFL. Both primer pairs were designed on the genome sequence of A. niger CBS 513.88 (GenBank Accession Number AM270359 and AM270408). The SalI–NcoI double-digested promoter fragment of gpdA, hpt:35S or hpt-2:35S fragment prepared above and the EcoRV–XhoI digested pEX0 were ligated to create pEX3 and pEX4, respectively. Similarly, ligation of tefA, hpt:35S or hpt-2:35S and EcoRV–XhoI digested pEX0 created pEX5 and pEX6, respectively. Plasmid pAN-GFP was created by inserting the gpdA promoter fragment derived from pAN7-1 and an egfp:trpC fragment into pUC18. Subsequently, pEX3GPDA-EGFP, pEX4GPDA-EGFP, pEX5GPDA-EGFP and pEX6GPDA-EGFP (Fig. 1) were constructed by inserting the SacI fragment containing the gpdA::egfp:trpC fragment from pAN-GFP into the SpeI site of pEX3, pEX4, pEX5 and pEX6, respectively. To construct pEX2GPDA-EGFP, the gpd::hpt-2:35S cassette was first created by replacing the hpt with hpt-2 in a pGH1 derivative pGHS3 (Ji et al. 2010), the product of which was digested with SpeI and XhoI and the gpd::hpt-2:35S cassette was inserted between the SacII–XhoI sites in pEX6GPDA-EGFP. All plasmids used in this study were shown in Supplementary Table 2.
T-DNA region of three improved binary vectors. All vectors were based on the pPZP200 backbone. hpt-2 codon-optimized hygromycin resistance gene; Pro U. maydis gpd promoter in pEX2GPDA-EGFP, or A. niger gpdA promoter in pEX4GPDA-EGFP, or A. niger tefA promoter in pEX6GPDA-EGFP; egfp coding sequence of enhanced green florescence protein gene; gpdA A. nidulans gpd promoter; 35S cauliflower mosaic virus 35S gene terminator; trpC A. nidulans trpC terminator (for egfp). LB and RB are the left and right border sequence of T-DNA respectively. Unique restriction sites are shown
Fungal transformation
ATMT was performed essentially as described (Ji et al. 2010; Mullins et al. 2001). U. scitaminea was cultured in YPD medium until the OD600 value was 0.5–0.8; then 150 μl was mixed in an Eppendorf tube with a pre-induced Agrobacterium culture (100 μl) before being spread evenly on to a 0.45 μm Hybond N membrane disc (GE Lifesciences, USA) that was placed on an induction medium (IM) plate (pH 5–5.9). Co-culture was performed at 24 °C in the dark for 48–96 h. Subsequently, membranes were transferred onto YPD plates containing 300 μg cefatoxime ml−1 and 50–200 μg hygromycin B ml−1 to select for transformants. Both IM and YPD plates contained at least 2.5 % (w/v) agar and were air-dried for 30–40 min before use.
Southern blotting analysis
Total DNA was extracted as described previously (Ji et al. 2010). Standard procedures were used for Southern blotting.
Fluorescence microscopy
Transformants were grown on YPD broth. Cells at the early growth stage were examined under a microscope (Nikon, Japan) equipped with GFP-L filter (GFP Band pass, Ex 480/40 DM 505 BA 510). Transformants on selection plates were observed with a stereomicroscope (Nikon, Japan) equipped with epifluorescence and GFP-L filter. Images were captured with a DS-5MC digital camera.
Results and discussion
Optimization of conditions for ATMT of Ustilago scitaminea
A sensitivity test of U. scitaminea strain S10 to antibiotics showed that its growth could be completely inhibited with 25 μg hygromycin B ml−1 in YPD agar medium. However, transformation was not successful in several attempts using the method and vectors that had been previously described for U. maydis (Ji et al. 2010). We noticed that the selection membranes developed a confluent watery layer of cells when S10 cells were co-cultured with Agrobacterium tumefaciens AGL1. We speculated that the development of confluent watery cell layer might account for the failure in transformation. Consequently, agar was increased to 2.5 % (w/v) from original 1.5 % (w/v) in both co-culture and selection medium and co-culture membranes on agar plates were air-dried for 30–40 min. This resulted in the appearance of a few antibiotic-resistant transformants when pEX1GPD-EGFP with a selection pressure of 50 μg hygromycin B ml−1 was used. However, 73 % (11 out of 15) of the resistant colonies were found to be false-positives when they subcultured in same concentration of hygromycin B (not shown).
When the selection pressure was increased to 100–200 μg hygromycin B ml−1, GFP-positive transformants were increased (up to 86 %) although this delayed the appearance of transformants to 6–8 days (Fig. 2a, b). Subsequently, additional parameters were assessed to optimize the transformation efficiency by ATMT using a GFP tracking vector, pEX1GPD-EGFP. In ATMT of U. scitaminea, no transformants were observed when acetosyringone (AS) was omitted. Consistent with many previous reports (de Groot et al. 1998; Michielse et al. 2005), AS is essential for fungal transformation because fungi are usually not able to synthesize the compound that is essential for inducing virulence genes in Agrobacterium. Co-culture pH had a drastic effect on transformation. No transformants were observed when co-cultured at pH 5.0 or pH 5.9 (Table 1); the optimal pH was approx. 5.5, at which point approx. 300 % more hygromycin B-resistant transformants were observed than at pH 5.3. However, the optimal pH for transformation of U. maydis is 5.7 (Ji et al. 2010). The sensitivity to co-culture medium pH values was also reported in the ATMT of a number of fungi (Soltani et al. 2008).
Expression of GFP in hygromycin-resistant cells of U. scitaminea. a Colonies derived from pEX1GPD-EGFP under fluorescent microscope 8 days after selection against 100 μg hygromycin B ml−1; b: as in a but with 200 μg hygromycin B ml−1. d, f Wild-type S10 cells and cells transformed with pEX2GPDA-EGFP and imaged under epifluorescent mode; c, e: cells of d and f and imaged under DIC mode. Scale bars represent 2 mm in a and b and 50 μm in c–f
Despite all the optimization efforts, the method remained unsatisfactory for routine transformation work due to the low and unstable efficiency as well as the long selection process. Therefore, the original hpt gene encoding a hygromycin B phosphotransferase was optimized. An analysis of codon usage patterns in three highly expressed proteins of U. scitaminea, i.e. actin (ACT1) (GenBank Accession Number FJ514819), elongation factor 1-alpha (EF1α) (GenBank Accession Number DQ352829) and glyceraldehyde-3-phosphate dehydrogenase (GPD1), revealed high frequency (73.4–78.7 %) of C or G at the third position of codons (Fig. 3). Based on the codon utilization pattern and GC content, a codon-optimized hpt gene (hpt-2) was synthesized, in which the GC content was increased to 62.4 % from the original 57.4 %, and the GC content at the third position of the codons was increased to 83.3 % (Fig. 3). Subsequently, three new GFP tracking vectors, pEX2GPDA-EGFP, pEX4GPDA-EGFP and pEX6GPDA-EGFP containing the codon-optimized hpt-2 under the regulation of strong fungal promoters were constructed. All showed drastically improved transformation efficiency. Among them, pEX4GPDA-EGFP showed the highest transformation efficiency, reaching up to 159 CFU/plate on average with 86 % transformants being GFP positive (Table 2). Importantly, hygromycin-resistant transformants were clearly visible after 3 days, 3–4 days earlier than those obtained with the original hpt gene. The combined effects of strong promoter, such as gpdA or tefA promoter from the Aspergillus species, and codon matching of selection marker genes are consistent with recent reports on the transformation of red yeasts (Abbott et al. 2013; Liu et al. 2013).
GC contents of the coding sequences and the third position of codons of abundantly expressed proteins in U. scitaminea and selection markers. EF1α, GPD1 and ACT1, hpt and hpt-2 indicate coding sequence for elongation factor 1-alpha, glyceraldehyde-3-phosphate dehydrogenase, Actin, original E. coli hpt and codon-optimized hpt-2, respectively
T-DNA integration patterns and stability of integrated T-DNA
We analyzed restriction patterns of 18 independent transformants of pEX1GPD-EGFP by Southern blotting. Strong hybridization bands appeared in all transformants with a single copy insertion and random site integration, no signal was present in the wild type strain (Fig. 4). After 10 successive subcultures on non-selective media, all GFP-positive transformants re-established on YPD agar plates were resistant to 200 μg hygromycin B ml−1 and showed strong GFP expression (Fig. 2f). A single insertion of T-DNA with random site integration in fungal genome by ATMT will facilitate subsequent isolation of tagged genes (Michielse et al. 2005; Soltani et al. 2008). Our results indicate that the ATMT system can be effectively used in molecular genetic studies of this fungus.
In conclusion, we have developed a simple and reliable method for transformation of U. scitaminea based on the codon-optimized hpt-2 gene coupling with a strong promoter, a higher concentration, of agar in the medium controlled membrane dryness and optimized medium pH. The egfp reporter construct allows rapid verification of successful transformation. As the T-DNA inserts into the genome as a single copy DNA fragment, the transformation method established herein is suitable for functional genomic studies as well as for genetic engineering and synthetic biology (Michielse et al. 2005; Soltani et al. 2008). Furthermore, our findings could be useful for other researchers in the fungal research community who are interested to establish a reliable and efficient transformation method for recalcitrant species.
References
Abbott EP, Ianiri G, Castoria R, Idnurm A (2013) Overcoming recalcitrant transformation and gene manipulation in Pucciniomycotina yeasts. Appl Microbiol Biotechnol 97:283–295
Bakkeren G, Kronstad JW (1996) The pheromone cell signaling components of the Ustilago α mating-type loci determine intercompatibility between species. Genetics 143:1601–1613
Bundock P, den Dulk-Ras A, Beijersbergen A, Hooykaas PJ (1995) Trans-kingdom T-DNA transfer from Agrobacterium tumefaciens to Saccharomyces cerevisiae. EMBO J 14:3206–3214
de Groot M, Bundock P, Hooykaas P, Beijersbergen A (1998) Agrobacterium tumefaciens-mediated transformation of filamentous fungi. Nat Biotechnol 16:839–842
Ferreira S, Comstock J (1989) Smut. In: Ricaud C, Egan B, Gillaspie J, Hughes C (eds) Diseases of sugar-cane. Major diseases. Elsevier, Amsterdam, pp 211–229
Ji L, Jiang ZD, Liu Y, Koh CM, Zhang LH (2010) A simplified and efficient method for transformation and gene tagging of Ustilago maydis using frozen cells. Fungal Genet Biol 47:279–287
Lazo GR, Stein PA, Ludwig RA (1991) A DNA transformation-competent Arabidopsis genomic library in Agrobacterium. Biotechnology 9:963–967
Liu Y, Koh CM, Sun L, Hlaing MM, Du M, Peng N, Ji L (2013) Characterization of glyceraldehyde-3-phosphate dehydrogenase gene RtGPD1 and development of genetic transformation method by dominant selection in oleaginous yeast Rhodosporidium toruloides. Appl Microbiol Biotechnol 97:719–729
Michielse CB, Hooykaas PJ, van den Hondel CA, Ram AF (2005) Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Curr Genet 48:1–17
Morita T, Ishibashi Y, Hirose N, Wada K, Takahashi M, Fukuoka T, Imura T, Sakai H, Abe M, Kitamoto D (2011) Production and characterization of a glycolipid biosurfactant, mannosylerythritol lipid B, from sugar-cane juice by Ustilago scitaminea NBRC 32730. Biosci Biotechnol Biochem 75:1371–1376
Mullins ED, Chen X, Romaine P, Raina R, Geiser DM, Kang S (2001) Agrobacterium-mediated transformation of Fusarium oxysporum: an efficient tool for insertional mutagenesis and gene transfer. Phytopathology 91:173–180
Shi L, Fang X, Li M, Mu D, Ren A, Tan Q, Zhao M (2012) Development of a simple and efficient transformation system for the basidiomycetous medicinal fungus Ganoderma lucidum. World J Microbiol Biotechnol 28:283–291
Soltani J, Heusden GP, Hooykaas PJ (2008) Agrobacterium-mediated transformation of non-plant organisms. In: Citovsky V, Tzfira T (eds) Agrobacterium: from biology to biotechnology. Springer, New York, pp 649–675
Sundar AR, Barnabas EL, Malathi P, Viswanathan R (2012) A mini-review on smut disease of sugarcane caused by Sporisorium scitamineum. In: Mworia J (ed) InTech, vol 5. InTech Europe, Winchester, pp 107–128
Weng LX, Deng HH, Xu JL, Li Q, Wang LH, Jiang ZD, Zhang HB, Li QW, Zhang LH (2006) Regeneration of sugar-cane elite breeding lines and engineering of stem borer resistance. Pest Manag Sci 62:178–187
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Project No. 31271993) and the Temasek Trust and the Economic Development Board, Singapore. The authors express sincere gratitude to Prof. Lian-Hui Zhang, Institute of Molecular and Cell Biology, Singapore for his valuable suggestions and revision of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, L., Yan, M., Ding, Z. et al. Improved dominant selection markers and co-culturing conditions for efficient Agrobacterium tumefaciens-mediated transformation of Ustilago scitaminea . Biotechnol Lett 36, 1309–1314 (2014). https://doi.org/10.1007/s10529-014-1486-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-014-1486-5