Abstract
A putative endo-1,4-β-d-xylanohydrolase gene xyl10 from Aspergillus niger, encoding a 308-residue mature xylanase belonging to glycosyl hydrolase family 10, was constitutively expressed in Pichia pastoris. The recombinant Xyl10 exhibited optimal activity at pH 5.0 and 60 °C with more than 50 % of the maximum activity from 40 to 70 °C. It retained more than 90 % of the original activity after incubation at 60 °C (pH 5.0) for 30 min and more than 74 % after incubation at pH 3.0–13.0 for 2 h (25 °C). The specific activity, K m and V max values for purified Xyl10 were, respectively, 3.2 × 103 U mg−1, 3.6 mg ml−1 and 5.4 × 103 μmol min−1 mg−1 towards beechwood xylan. The enzyme degraded xylan to a series of xylooligosaccharides and xylose. The recombinant enzyme with these properties has the potential for various industrial applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Xylan, widely distributed in cell walls of plants, is the second most abundant renewable polysaccharide in nature (Giridhar and Chandra 2010). Endo-1,4-β-d-xylanohydrolase (EC 3.2.1.8, xylanase) degrades xylan to produce xylooligomers by randomly hydrolyzing 1,4-β-d-xylosidic linkages. Thus, xylanases have the great potential for various industrial applications in food production, animal feed, paper and pulp, textiles and the production of bioethanol fuel (Paës et al. 2012). Many researchers focused on the discovery of novel xylanases with excellent properties for industrial applications (Ahmed et al. 2009). A variety of xylanases occur in fungi, algae, bacteria, protozoa, gastropods and anthropods (Collins et al. 2005; Ahmed et al. 2009). Some from fungi and bacteria have broad working temperature or pH ranges, high thermotolerance, high pH stability or high catalytic efficiency. They have great advantages in industrial applications (Bai et al. 2010; Giridhar and Chandra 2010; Luo et al. 2009).
Aspergillus spp. produce many carbohydrate-active enzymes which displayed excellent properties and could be applied in various industries (Ahmed et al. 2009). Aspergillus niger produces 15 extracellular xylanases and is a good source of xylanase (Collins et al. 2005). A. niger CBS 513.88, as a mutant of A. niger NRRL 3122, produced a high activity glucoamylase, and its genome has been sequenced (Pel et al. 2007). We predicted that a putative xylanase gene (accession no. XM_001388485) from this strain might produce a protein with excellent catalytic properties. Therefore, in this research, the putative xylanase gene was optimized and synthesized by dual asymmetrical PCR and expressed in Pichia pastoris using a glyceraldehyde-3-phosphate dehydrogenase (GAP) promoter. The recombinant xylanase displaying extremely high stability over a broad pH range could have the potential to be applied in industries and provide clues to the rational protein design to improve the pH stability of xylanases.
Materials and methods
Sequence and structure analysis
The xly10 gene has been sequenced (Pel et al. 2007). The BLAST server in GenBank was used for homology searches. SignalP 4.0 was used for predicting the signal peptide sequence of the xylanase (http://www.cbs.dtu.dk/services/SignalP/). Homology analysis was performed by ClustalX and Mega 4.1. Homology modeling was implemented by MODELER 9.9 (http://salilab.org/modeller/). ProQ-Protein Quality Predictor was used for the evaluation of modeling structure (http://www.sbc.su.se/~bjornw/ProQ/ProQ.html). It was analyzed using PyMol 0.99rc6 (http://www.pymol.org/).
Gene optimization and vector construction
The cDNA of xyl10 gene was optimized using GenScript Rare Codon Analysis Tool. The optimized xyl10 gene was dissected into 40 overlapping oligonucleotides by Gene2Oligo (http://berry.engin.umich.edu/gene2oligo/). These oligonucleotide fragments were synthesized by GenScript Corporation (Nanjing, China) and assembled into full-length gene by dual asymmetrical PCR (Sandhu et al. 1992). The primers were shown in Supplementary Table 1. The full-length product was cloned into pUM-T vector and transformed into Escherichia coli TOP 10F’. Correct construction of clone plasmid (pUM-xyl10) was confirmed by PCR and sequencing (GenScript Corporation, China). After pUM-xyl10 was digested with Xba I and Xho I (ER0682 and ER0693, Thermo Fisher Scientific Inc., USA), the target product was purified and inserted into the pGAPZαA vector (V205-20, Invitrogen, USA). Correct construction of expression plasmid (pGAPZα-xyl10) was confirmed by restriction enzyme digestion and sequencing (GenScript Corporation, China).
Screening of high-level expression strains
The recombinant plasmid pGAPZα-xyl10 was linearized with BglII (Thermo Fisher Scientific Inc., USA) and then transformed into P. pastoris X33 by electroporation according to the described method in Pichia Expression Kit (Invitrogen, USA). The empty pGAPZαA vector was also transformed into P. pastoris X33 and it was used as a control. Screening of high-level expression and inheritance stability strains was implemented according to the method as previously described (Zhao et al. 2011).
Expression and purification of recombinant endo-1,4-β-d-xylanohydrolase
The colony was picked into 10 ml YPD medium (10 g yeast extract l−1, 20 g peptone l−1 and 20 g glucose l−1) and cultivated at 30 °C (250 rpm) for 24 h in 100 ml shake-flasks. One ml culture was transferred into 100 ml YPD and cultivated for 108 h at 30 °C (250 rpm) in 500 ml shake-flasks. Dry cell weight, enzyme activity, crude protein and glucose concentrations were determined by sampling 1 ml culture every 12 h. The recombinant xylanase was purified by size exclusion chromatography on a Superdex G-75 column (see Zhao et al. 2011).
Protein determination and SDS-PAGE analyses
Protein concentration was determined using a kit. Molecular mass of the recombinant enzyme was estimated by SDS-PAGE with an unstained protein molecular weight marker as the standard (Schägger 2006).
Enzymatic activity assay
The 3,5-dinitrosalicylic acid (DNS) method was used for determining xylanase activity (Bailey et al. 1992). The reaction was started by incubating 0.5 ml appropriately diluted enzyme sample with 0.5 ml 10 g oat spelt xylan l−1 in 0.1 M citric acid/Na2HPO4 buffer (pH 5.0) for 5 min at 60 °C. One ml DNS was added to stop the reaction. The mixture was boiled for 5 min and cooled to room temperature. The absorbance was measured at 540 nm. One unit of xylanase activity was defined as the amount of enzyme releasing 1 μmol xylose equivalents per minute. All experiments were done in triplicate.
Hydrolysis products analysis by thin-layer chromatography (TLC)
After incubation with 10 g substrate l−1 (10 ml) with 10 U of the recombinant enzyme at 60 °C in 0.1 M citric acid/Na2HPO4 buffer (pH 5.0), hydrolysis products were detected TLC using silica gel 60F254 plates (Zhang et al. 2010). Xylose, xylobiose, xylotriose, xylotetraose and xylopentaose were used as standards.
Nucleotide sequence accession number
The cDNA sequence of xyl10 gene deposited in the GenBank database was under accession no. AM270045 and amino acid sequence of Xyl10 was under accession no. A2QFV7 (Pel et al. 2007).
Results and discussion
Sequence analysis
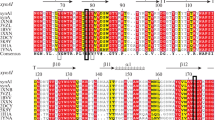
The putative xylanase gene xyl10 consisted of 984 nucleotides, encoded a 327-residue polypeptide, including a signal peptide of 19-residue and a catalytic domain of endo-1,4-β-d-xylanase belonging to glycosyl hydrolase family 10. The xylanase Xyl10 displayed 94–99 % primary sequence identity to six putative xylanases (accession nos. C5J411.2, P33559.2, JT0608, ACR83565.1, XP_001389996.2 and GAA92552.1) (Fig. 1a). However, up to date, their properties have not been reported in detail. Amino acid sequence homology and hydrophobic cluster analysis revealed that xylanases could be classified into two main glycosyl hydrolyase (GH) family 10 and 11 (Chantasingh et al. 2006), and some xylanases belonging to family 5, 7, 8, 16, 26, 43, 52 and 62 also were identified (http://www.cazy.org/Glycoside-Hydrolases.html). The amino acid sequence of the mature Xyl10 shared 86 % identity with xylanase PsXyl belonging to GH10 from Penicillium simplicissimum (PDB No. 1B30_A), whose three-dimensional structure has been reported (Schmidt et al. 1999). Homology modeling using PsXyl as the template revealed that the mature xylanase Xyl10 had the classical (α/β)8-fold. Two putative catalytic residues in Xyl10, Glu132 and Glu238 were identified based on the amino acid sequence alignment and homology modeling (Fig. 1b, c).
a Phylogenetic trees of some xylanase from GH family 10 and GH family 11. Xyl10 marked with a black uptriangle was the object of this study. The numbers at the nodes represent bootstrap values based on 1,000 replications. The lengths of the branches show the relative divergence among the reference xylanase amino acid sequences and scale bar indicates the amino acid substitutions per position. GenBank accession numbers of the xylanases were given after each species name. b Multiple amino acid sequence alignment between Xyl10 and other GH family 10 xylanases. The multiple sequence alignment between Xyl10 and other reported GH family 10 xylanases from Penicillium simplicissimum (PDB No. 1B30_A), Streptomyces sp. S27 (accession no. ACF57946.1), Streptomyces sp. SWU10 (accession no. BAK19338.1), Streptomyces megasporus DSM41476 (accession no. ADE37527.1), Saccharopolyspora sp. S582 (accession no. ADL60499.1), Streptomyces sp. S9 (accession no. ABX71815.1) and Bispora sp. MEY-1 (accession no. ACS96449.1) was done by ClastalX and Genedoc. The signal peptide was boxed and active sites were marked with black circle. c Homology modeling of Xyl10 was performed by MODELER 9.9 based on a known crystal structure of xylanase from Triticum aestivum (PDB ID:1TE1). The modeling structure was evaluated by ProQ-Protein Quality Predictor and analyzed using PyMol 0.99rc6. Two active sites Glu132 and Glu238, were indicated
Construction of recombinant strain library
The xyl10 gene was optimized by upgrading the codon adaptation index from 0.68 to 0.95 to increase the probability of high-level expression in P. pastoris. The Stem-Loop structures which impacted ribosomal binding and stability of mRNA were removed and the GC content was changed from 53 to 39.1 % to prolong the half-life of mRNA in P. pastoris (Supplementary data Optimized xyl10 gene). The full-length of optimized xyl10 gene was obtained by dual asymmetrical PCR (Fig. 2). The optimized xyl10 gene was inserted into pGAPZαA vector downstream of α-factor signal peptide from Saccharomyces cerevisiae which enabled the secretion of the recombinant xylanase into culture broth.
Identification of full-length xyl10 gene by agarose gel electrophoresis. Lanes 1, 2 products of fragments I and II by dual asymmetrical PCR; lane 3 products of full-length xyl10 gene by one-step dual asymmetrical PCR; lane 4 products of full-length xyl10 gene assembled from fragments I and II by overlap extention PCR; lane 5 DNA marker; lane 6 negative control for one-step dual asymmetrical PCR; lane 7 negative control for assembling full-length xyl10 gene by overlap extention PCR
After being linearized by Bgl II, the correct construction of pGAPZα-xyl10 plasmid was transformed into P. pastoris X33 by electrotransformation. A total of 338 positive transformants was chosen from YPDS plates containing 100–500 μg Zeocin ml−1 for screening high-level expression of xylanase in shaken flask cultures. A stable-inheritance strain named A × 10−4 selected from 338 positive transformants with high levels of xylanase activity and inheritance stability was used for further studies.
Expression and purification of the recombinant enzyme
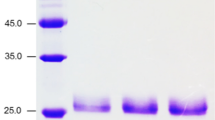
The maximum activity of strain A × 10−4 was 362 U ml−1. Crude protein in supernatant reached 145 μg ml−1 after cultivation of the transformed yeast for 108 h (Fig. 3).The recombinant Xyl10 was secreted into culture and purified using gel filtration chromatography, resulting in a purification yield of 32 % (Fig. 4a). The specific activity of recombinant Xyl10 was 2.5 × 103 U mg−1 for oat spelt xylan after 1.2-fold purification. SDS-PAGE analysis showed a protein with a size of ~33 kDa, which was identical to the predicted size (Fig. 4b).
Purification of the recombinant Xyl10. a Elution profile of recombinant Xyl10 on Sephadex™ G-75 chromatography. Relative xylanase activity in fractions (black circle); relative protein concentration in fractions (open square). b SDS-PAGE analysis of proteins present in the crude supernatant. Lane 1 purified protein by size exclusion chromatography on Sephadex G-75 chromatography; lane 2 molecular mass standard protein; lane 3 the proteins in crude supernatant of Pichia pastoris X33. 233 μg Xyl10 ml−1 and 582 U ml−1 of xylanase activity were taken as 100 % of protein concentration and xylanase activity, respectively
Effects of pH, temperature and chemical reagents on the recombinant enzyme activity
The recombinant enzyme displayed the maximal activity at pH 5.0 (Fig. 5a). Its activity declined drastically below pH 3.0 and above pH 7.0. From pH of 3 to 13 more than 74 % of its original activity was retained after incubation for 2 h at room temperature (Fig. 5b). A comprehensive literature review revealed that few GH10 xylanases with the similar pH optima possess a similar property. GH10 xylanases from Streptomyces sp. displayed high stability over a broad pH range, especially under alkaline conditions (Supplementary Table 2). Two GH10 xylanases from Bispora sp. MEY-1 and Penicillium pinophilum C1 displayed high stability under acidic conditions (Luo et al. 2009; Cai et al. 2011). Compared with these xylanases, the recombinant Xyl10 was highly stable over a broad pH range. A GH10 xylanase from Streptomyces sp. S27, similar to Xyl10, also showed excellent pH stability with about 80 % residual activity at pH 2.2–12 after incubation at 37 °C for 1 h (Li et al. 2009).
Effects of pH and temperature on the recombinant Xyl10 activity. a Effect of pH on the activity of recombinant Xyl10. The activity was determined at 60 °C in 0.1 M citric acid/Na2HPO4 buffer (pH 2.2–8.0). b pH stability of recombinant Xyl10. This was determined by measuring the residual activity after incubation at various pH values at room temperature for 2 h without substrate. pH 2.2–8.0: 0.1 M citric acid/Na2HPO4 buffer; pH 9.0–11.0: 0.05 M glycine/NaOH buffer; pH 12.0–13.0: 0.05 M KCl/NaOH buffer. c Effect of temperature on the activity of recombinant Xyl10. The activity was determined in 0.1 M citric acid/Na2HPO4 buffer (pH 5.0) at temperatures between 20 and 80 °C. d Thermostability of recombinant Xyl10. It was determined by measuring the residual activity under the optimal condition after the enzyme had been incubated at the indicated temperature 60 °C (black square), 70 °C (open circle) and 80 °C (black uptriangle) for 0–30 min. The error bars represent mean ± SD (n = 3) and 2.5 × 103 U mg−1 was taken as 100 % of xylanase activity
The optimal temperature for the recombinant enzyme was 60 °C. It displayed more than 50 % of its maximal activity over the temperature range from 40 to 70 °C (Fig. 5c). Almost no activity was detected above 85 °C. The recombinant Xyl10 retained more than 90 % relative activity after incubation at 60 °C for 30 min and its half-life time at 70 °C was less than 5 min (Fig. 5d). The optimal temperature for most GH 10 xylanases varies from 50 °C to 70 °C (Supplementary Table 2). A cold-active xylanase from Glaciecola mesophila KMM241 displayed the maximal activity at 30 °C (pH 7.0) and the most thermostable xylanase belonging to GH 10 from Acidothermus cellulolyticus 11B showed the maximal activity at 90 °C (pH 6.0) (Guo et al. 2009; Barabote et al. 2010).
Some metal ions and chemicals significantly inhibit the activity of most xylanases in industrial applications. However, among metal ions and chemicals tested in this study, only SDS greatly inhibited the recombinant enzyme (Supplementary Table 3).
Detection of the recombinant enzyme substrate specificity, kinetic parameters and hydrolysis products
The recombinant enzyme displayed the highest specific activity towards beechwood xylan and it had a negligible specific activity towards CMC-Na (Table 1). Xyl10 had the maximum k cat/K m toward beechwood xylan but the lowest K m against birchwood xylan, showing that it had the highest catalytic efficiency towards beechwood xylan but the highest affinity towards birchwood xylan. The specific activity and V max for purified Xyl10 were higher than most reported xylanases belonging to GH10 except the recombinant xylanase from Bispora sp. MEY-1, whose specific activity and V max towards oat spelt xylan were 1.9 × 104 U mg−1 and 2.4 × 104 μmol min−1 mg−1 (Luo et al. 2009). The main hydrolysis products of oat spelt xylan by Xyl10 were xylotriose and xylobiose (Supplementary Fig. 1). Xylose and xylotetraose were also detected. Xylobiose promoted the growth of intestinal Bifidobacteria, inhibited proliferation of pathogenic bacteria, increased nutrients absorption and also used in antiobesity diets because it was low in calories (Vazquez et al. 2000; Moure et al. 2006). The recombinant Xyl10 hydrolyzed xylan to produce xylobiose as the major products and thus could be used in food and health care products.
In conclusion, A GH10 xylanase Xyl10, was expressed in P. pastoris with functional activity. The recombinant Xyl10 had high stability from pH 3 to 13 and, because of its good thermotolerance and high catalytic efficiency, it has a great potential to be used in industries.
References
Ahmed S, Riaz S, Jamil A (2009) Molecular cloning of fungal xylanases: an overview. Appl Microbiol Biotechnol 84:19–35
Bailey MJ, Biely P, Poutanen K (1992) Interlaboratory testing of methods for assay of xylanase activity. J Biotechnol 23:257–270
Barabote RD, Parales JV, Guo YY et al (2010) Xyn10A, a thermostable endoxylanase from Acidothermus cellulolyticus 11B. Appl Environ Microbiol 76:7363–7366
Cai H, Shi P, Bai Y et al (2011) A novel thermoacidophilic family 10 xylanase from Penicillium pinophilum C1. Process Biochem 46:2341–2346
Chantasingh D, Pootanakit K, Champreda V et al (2006) Cloning, expression, and characterization of a xylanase 10 from Aspergillus terreus (BCC129) in Pichia pastoris. Protein Expr Purif 46:143–149
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Giridhar PV, Chandra T (2010) Production of novel halo-alkali-thermo-stable xylanase by a newly isolated moderately halophilic and alkali-tolerant Gracilibacillus sp. vTSCPVG. Process Biochem 45:1730–1737
Guo B, Chen XL, Sun CY et al (2009) Gene cloning, expression and characterization of a new cold-active and salt-tolerant endo-β-1, 4-xylanase from marine Glaciecola mesophila KMM 241. Appl Microbiol Biotechnol 84:1107–1115
Li N, Shi P, Yang P et al (2009) A xylanase with high pH stability from Streptomyces sp. S27 and its carbohydrate-binding module with/without linker-region-truncated versions. Appl Microbiol Biotechnol 83:99–107
Luo H, Li J, Yang J et al (2009) A thermophilic and acid stable family-10 xylanase from the acidophilic fungus Bispora sp. MEY-1. Extremophiles 13:849–857
Moure A, Gullón P, Domínguez H et al (2006) Advances in the manufacture, purification and applications of xylo-oligosaccharides as food additives and nutraceuticals. Process Biochem 41:1913–1923
Paës G, Berrin JG, Beaugrand J (2012) GH11 xylanases: structure/function/properties relationships and applications. Biotechnol Adv 30:564–592
Pel HJ, de Winde JH, Archer DB et al (2007) Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat Biotechnol 25:221–231
Sandhu G, Aleff R, Kline B (1992) Dual asymmetric PCR: one-step construction of synthetic genes. Biotechniques 12:14–16
Schägger H (2006) Tricine-SDS-PAGE. Nat Protoc 1:16–22
Schmidt A, Gübitz GM, Kratky C (1999) Xylan binding subsite mapping in the xylanase from Penicillium simplicissimum using xylooligosaccharides as cryo-protectant. Biochem 38:2403–2412
Vazquez MJ, Alonso JL, Dominguez H et al (2000) Xylooligosaccharides: manufacture and applications. Trends Food Sci Technol 11:387–393
Wang SY, Hu W, Lin XY et al (2012) A novel cold-active xylanase from the cellulolytic myxobacterium Sorangium cellulosum So9733-1: gene cloning, expression, and enzymatic characterization. Appl Microbiol Biotechnol 93:1503–1512
Zhang M, Jiang Z, Yang S et al (2010) Cloning and expression of a Paecilomyces thermophila xylanase gene in E. coli and characterization of the recombinant xylanase. Bioresour Technol 101:688–695
Zhao W, Zheng J, Zhou H (2011) A thermotolerant and cold-active mannan endo-1, 4-[beta]-mannosidase from Aspergillus niger CBS 513.88: constitutive overexpression and high-density fermentation in Pichia pastoris. Bioresour Technol 102:7538–7547
Acknowledgments
This work was financially supported by Genencor Innovation Grant.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jia Zheng and Ning Guo contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zheng, J., Guo, N., Wu, L. et al. Characterization and constitutive expression of a novel endo-1,4-β-d-xylanohydrolase from Aspergillus niger in Pichia pastoris . Biotechnol Lett 35, 1433–1440 (2013). https://doi.org/10.1007/s10529-013-1220-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-013-1220-8