Abstract
A new superoxide dismutase (SOD) gene from the thermophilic fungus Chaetomium thermophilum (Ctsod) was cloned and expressed in Pichia pastoris and its gene product was characterized. The specific activity of the purified CtSOD was 2,170 U/mg protein. The enzyme was inactivated by KCN and H2O2 but not by NaN3, confirming that it belonged to the type of Cu, ZnSOD. The amino acid residues involved in coordinating copper and zinc were conserved. The recombinant CtSOD exhibited optimum activity at pH 6.5 and 60°C. The enzyme retained 65% of the maximum activity at 70°C for 60 min and the half-life was 22 and 7 min at 80 and 90°C, respectively. The recombinant yeast exhibited higher stress resistance than the control yeast cells to salt and superoxide-generating agents, such as paraquat and menadione.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The superoxide radicals (O •−2 ) are generated as normal by-products of aerobic cellular metabolism and can damage all types of biomolecules including DNA, proteins, and membrane lipids. It can lead to many pathophysiological events correlated with mutagenesis, cancer, degenerative processes, cell death and aging (Bannister et al. 1987). Superoxide dismutase (SOD) converts the O •−2 to H2O2 which, in turn, is enzymatically transformed by catalase or glutathione peroxidase into O2 and H2O.
SOD is important in the pharmaceutical, nutraceutical and cosmeceutical industries (Raimondi et al. 2008). Recombinant SODs may be useful for treating a wide variety of disorders related to oxidative stress, including prevention of oncogenesis, Alzheimer’s disease, acute radiodermatitis, inflammation in wounds, and diabetes (Angelova et al. 2001; Nishikawa et al. 2001). However, SOD is easily degraded at high temperatures, or in acid or alkali. The stability of SOD is low at room temperature and cannot be stored for long periods (Wu et al. 2010). Thermostability is one of the most important properties for commercial SODs, since thermal denaturation is a common cause of enzyme inactivation (Li et al. 2005). In addition, thermostable enzymes in industrial processes can reduce the risk of contamination and the cost of external cooling (Vieille and Zeikus 2001). Therefore, there has been increasing interest in thermophilic organisms as a source of thermostable SODs with industrial and biomedical applications.
Although a number of thermostable SODs have been isolated from hyperthermophilic archaea and thermophilic bacteria (Song et al. 2009), the applications of thermophilic fungal SODs are currently limited. Until now, only one study of Cu, ZnSOD from the thermophilic fungus Thermoascus aurantiacus var. levisporus, has been reported (Shinjin et al. 2007). Chaetomium thermophilum is a soil-borne thermophilic fungus, which grows well at 50°C. During the past decade, many enzymes isolated from this microorganism have been extensively characterized and commercialized, including glucoamylase, cellulase, endocellulase, xylanase, and laccase (Chen et al. 2007). However, to our knowledge, studies describing SOD from Chaetomium thermophilum are not available. Therefore, in the present work, the overexpression, purification and characterization of a novel thermostable Cu, ZnSOD from this thermophilic fungus are described for the first time.
Materials and methods
Strains, vectors and culture media
Chaetomium thermophilum (HSAUP072651) was used as mRNA donor. Escherichia coli DH5a and JM109 were used for cloning and nucleotide sequencing. Pichia pastoris host strain GS115 and the pPIC9K vector (Invitrogen) were used as expression system.
Cloning of Cu, ZnSOD gene
Total RNA was isolated from the mycelia of Chaetomium thermophilum using Trizol according to the manufacturer’s instructions. The first strand cDNA was synthesized with Takara RNA PCR Kit (AMV) Ver. 3.0. Degenerate primers (Supplementary Table 1) were designed based on the conserved sequence of Cu, Zn-SOD and used to amplify partial sequence of target gene with RT-PCR. RACE method was used to clone the full-length cDNA using the 5′- and 3′-RACE Kit. All the PCR products were cloned into pMD18-T easy vectors, and transformed into E.coli DH5a. Sequences were conducted on an ABI PRISM 377 DNA Sequencer with SP6 and T7 primers. The resulting SOD gene was designated as Ctsod.
Construction of expression vector
The cDNA sequence encoding the mature CtSOD was amplified by PCR from the genomic cDNA of C. thermophilum using two primers with recognition sites for EcoRI and NotI (sense primer: 5′-CCGAATTCGTCAAGGCAGTTGCTGT-3′, antisense primer: 5′-GGGCGGCCGCTTACTGGGCA ATGCCAAT-3′). PCR was performed using pre-denaturation at 94°C for 4 min, 35 cycles of 30 s at 94°C, 20 s at 56°C and 40 s at 72°C; and a final elongation step of 10 min at 72°C. PCR product purified with QIAquick gel extraction kit was digested with EcoRI and NotI, then ligated into the EcoRI–NotI site of the expression vector pPIC9K to generate pPIC9K/Ctsod.
CtSOD expression and purification
After linearization of the corresponding plasmid pPIC9K/Ctsod with SacI, the GS115 strain was transformed using the electroporation method. Transformants which grew normally on the MD and MM plates were seeded onto YPD medium plates containing 1, 2, 3 or 4 mg G418/ml and cultured at 28°C for 2 days to select the multicopy integrants. The selected multicopy transformants were grown in BMGY at 28°C until OD600 reached 1–2. The cells were harvested by centrifugation at 5,000×g for 5 min, and then cultured in BMMY. Methanol was added every day to give 1% (v/v) and to induce expression for 7 days. Cultures were sampled daily and SOD activity was analyzed in the harvested supernatants.
The recombinant CtSOD protein was purified with DEAE–Sepharose column chromatography according to the supplier’s manuals. The purity of the enzyme was checked by SDS-PAGE (12% w/v).
SOD activity assays and protein level
The supernatant protein concentrations were determined using the Bradford’s method with crystalline bovine serum albumin as a standard. SOD activity was measured following the method of Beauchamp and Fridovich (1971). Briefly, the reaction mixture in 3 ml was composed of 13 mM l-methionine, 75 μM NBT (Nitroblue Tetrazolium), 2 μM riboflavin, 10 μM EDTA–Na2 and 10 μl purified enzyme in 50 mM PBS (pH 7.8). The test tubes were exposed to a source of light. The reduction of NBT was monitored after 10 min at 560 nm. One unit SOD activity was defined as the amount of enzyme which caused 50% of maximum inhibition of NBT reduction. All experiments were performed in triplicate and average values were reported.
Results and discussion
Cloning and sequence analysis of Ctsod from C. thermophilum
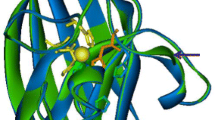
The full-length of Ctsod gene from C. thermophilum was 712 base pairs (bp) and contained a 465 bp ORF. The ORF encoded a putative protein of 154 amino acids with a calculated molecular mass of 16.1 kDa and a pI value of 6.4. The obtained nucleotide sequence was submitted to GenBank (Accession No. DQ493760). Multiple alignments of the putative CtSOD with other Cu, ZnSODs showed that the amino acid residues H-47, H-49, H-64, H-72, H-81, and H-121, which are involved in coordinating copper and zinc, were conserved. Furthermore, the arginine residue R-144, participating in enhancing the binding of O •−2 to the copper atom, was also conserved in CtSOD. The putative amino acid sequence of CtSOD analyzed by signal P 3.0 program revealed that it did not contain a hydrophobic signal peptide in the 5′-region, indicating that it might be an endocellular enzyme. This is consistent with the fact that most Cu, ZnSODs are intracellular proteins.
A phylogenetic tree of the CtSOD and other Cu, ZnSODs by DNAMAN software showed that it shared 90.26% homology with Cu, ZnSOD from Chaetomium globosum (XP_001222495), and shared moderate identities of 71.43, 64.74, 59.24% with Cu, ZnSODs from Aspergillus flavus (AAM94904), Nectri haematococca (EEU33541) and Cryptococcus liquefaciens (BAF42028), respectively. However, it shared a low similarity (37.31%) with Cu, ZnSOD from Penicillium marneffei (XP_002148587) (Supplementary Fig. 1).
Heterologous expression and purification of CtSOD
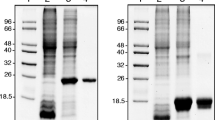
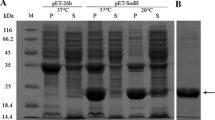
The cDNA of the Ctsod was introduced into the yeast expression system, and the heterologous expression of CtSOD was under transcriptional control of the AOX1 promoter in P. pastoris GS115. After being induced, the CtSOD was successfully expressed and excreted. Figure 1a shows SDS-PAGE of the induced recombinant CtSOD from a transformant strain CZ16. 6 days after induction, this strain had the highest SOD activity of 1,660 U/ml culture, and its expression level was 1.54 mg/ml. This high expression of protein did not affect cell growth. The expressed enzyme was easily purified with DEAE–Sepharose column chromatography (Table 1). The purified enzyme gave a single band on SDS-PAGE with a molecular mass of around 17 kDa, which was close to the predicted molecular weight for this protein (Fig. 1b). The specific activity of the purified enzyme was 2,170 U/mg, similar to commercially available products (2,000–4,000 U/mg). The purified CtSOD was inhibited by 5 mM KCN and 5 mM H2O2, but not by 10 mM NaN3 (Fig. 1c), further confirming that the CtSOD from C. thermophilum belonged to Cu, ZnSOD classification.
SDS-PAGE and PAGE analysis of recombinant CtSOD from P. pastoris GS115. a SDS-PAGE analysis of expressed CtSOD protein. Lane 1 Protein marker, low (14.3–97.2 kDa); Lane 2 P.pastoris GS115 control; Lanes 3–8 1, 2, 3, 4, 5, and 6 days after induction with 1% (v/v) methanol, respectively. b The purified protein. Lane 1 Protein marker; Lane 2 Purified CtSOD visualized by Coomassie Brilliant Blue R250 staining. c The effect of inhibitors on the purified CtSOD. Activity staining with NBT was done according to Beauchamp and Fridovich (1971). The samples were pre-incubated with the inhibitors of KCN (5 mM), NaN3 (10 mM) or H2O2 (5 mM) at pH 7.0, 37°C for 40 min. For the detection of SOD activity, the treated samples were electrophoresed on a 12% (w/v) native PAGE. Then the gel was soaked in 750 μM NBT solution for 15 min and 2 μM riboflavin for 5 min in the dark, respectively. Followed by immersion with illumination in 1 mM EDTA–Na2. Lane 1 as the control, no treatment; Lane 2 5 mM KCN as inhibitor; Lane 3 5 mM H2O2 as inhibitor; Lane 4 10 mM NaN3 as inhibitor
The P. pastoris expression system uses only pure reagents, free of toxins, while bacterial contamination is prevented by the use of methanol (Eckart and Bussineau 1996). Therefore, the expressed thermostable CtSOD is very promising for commercialization especially in the food and pharmaceutical industries.
Properties of the recombinant CtSOD
The purified recombinant CtSOD was optimally active at pH 6–7, with maximum activity at pH 6.5 (Fig. 2a). The enzyme activity was relatively stable at pH 5–9, and more than 70% of full activity retained at pH 5.5–8 (Fig. 2b). The optimal enzyme activity was established at 60°C. The enzyme retained more than 65% of full activity after 60 min incubation at 70°C. The purified Cu, ZnSOD from Thermoascus aurantiacus var. levisporus was stable at 50 and 60°C, and retained 53% activity after 60 min at 70°C (Shinjin et al. 2007). The purified MnSOD from Thermomyces lanuginosus had maximum activity at 55°C and retained 55% activity after 60 min at 70°C (Li et al. 2005). In addition, the recombinant CtSOD exhibited remarkably thermal stability, the half-life of the enzyme was 22 and 7 min under incubation at 80 and 90°C, respectively (Fig. 2c). Though not as thermostable as those exhibited by hyperthermophilic archaea such as Aquifex pyrophilus, Sulfolobus solfataricus and Pyrobaculum calidifontis (Song et al. 2009), eukaryotic organisms have been suggested as a better source of enzymes that are more similar to those from humans than prokaryotic organisms (Shin et al. 2009), and the major enzymatic action occurs at 40–60°C in most operational situations. Therefore, thermostable enzymes from thermophilic fungi may be better suited than enzymes from hyperthermophiles (Maheshwari et al. 2000).
Effect of temperature and pH on enzyme activity and stability of purified CtSOD from P. pastoris GS115. a The optimum pH value for CtSOD activity. SOD activity was determined at pH range of 5–10 at 60°C. The following buffers were used: CH3COOH/CH3COONa (pH 4–6), NaH2PO4/Na2HPO4 (pH 6–7) and Tris/HCl (pH 7–10). b The pH stability of CtSOD. After incubating the enzyme in buffers ranging from pH 3 to 10 at 37°C for 1 h, enzyme activity was determined in pH 6.5 at 60°C. c The thermostability of CtSOD. For each sample, SOD activity was measured after incubating at 60, 70, 80 and 90°C without substrate for 10, 20, 30, 40, 50, and 60 min, separately. The remaining activity was detected under standard conditions and calculated as the percentage of the maximum SOD activity. The absolute enzyme activity corresponding to 100% was 2,170 U mg−1. The values represent the means of three independent experiments (Mean ± standard error)
Viability under oxidative stress and salt
To assess a protective role of the Ctsod, the same numbers of the recombinant yeast harboring pPIC9K/Ctsod and the control yeast harboring pPIC9K were spotted on the YPD plates with 50–100 μM menadione, 300–500 μM Paraquat or 0.3–1 M NaCl, which cause oxidative stress. The recombinant yeast over-expressing CtSOD appeared to be more resistant to oxidative stress than the control yeast cells (Fig. 3). The recombinant yeast and control yeast cells grew at similar rates on the YPD plates without any stressful agents. However, concentrations of menadione above 50 μM and Paraquat over 300 μM substantially reduced the growth of the control yeast cells but the Ctsod transformed yeasts showed better growth.
Antioxidant activity tests. The transformant strains carrying pPIC9K/Ctsod vector and the control strains carrying empty pPIC9K were grown on YPD medium at 28°C for 20 h, adjusted to the same absorbance (OD600) and then serially ten fold diluted to 10−1, 10−2, 10−3, 10−4, and 10−5 in sterile water. Samples, 3 μl, of each dilutions were spotted on plates supplemented with various concentrations of paraquat, menadione and salt, and incubated for 3 days at 28°C. The control is plate without any stressful agents
These results were in good agreement with previous studies which showed that recombinant yeast cells bearing the human sod gene were more resistant to Paraquat and menadione-mediated oxidative stress (Yoo et al. 1999), and mammalian cells transfected with Cu, Zn-SOD gene were also better protected against paraquat toxicity (Krall et al. 1988). In addition, our results demonstrated that the Ctsod from C. thermophilum confers salt resistance. The control yeast cells were very sensitive to elevated salt concentrations as severe growth reduction was observed at both 0.3 and 0.5 M NaCl, and growth was completely arrested at higher levels. In contrast, the Ctsod transformed yeast cells had better growth than the control yeast cells at NaCl levels exceeding 0.3 M, and they could grow even at 1.5 M NaCl (data not shown). These results showed that Ctsod is an excellent salt tolerance gene.
To our knowledge, this is the first report describing an sod gene from thermophilic fungi that is able to confer resistance to salt stress. Therefore, the Ctsod gene may have great potential for the genetic improvement of salt tolerance in plants. Soil salinity is one of the most important environmental factors limiting crop productivity. Today, approximately 20% of the world’s irrigated land is affected by salt accumulation (Munns 2005). A variety of genes such as barley oxalate oxidase and Arabidopsis tocopherol cyclase have been utilized to develop transgenic plants with enhanced tolerance to salt and oxidative stress. Also, we are currently trying to transfer the Ctsod gene to tobacco and rice to investigate its ability to improve plant tolerance not only to high salinity but also to other abiotic stress.
References
Angelova MP, Dolashka-Angelova E, Ivanova J, Serkedjieva et al (2001) A novel glycosylated Cu/Zn-containing superoxide dismutase: production and potential therapeutic effect. Microbiol-SGM 147:1641–1650
Bannister JV, Bannister WH, Rotilio G (1987) Aspects of the structure, function and applications of superoxide dismutase. Crc Crit Rev Biochem 22:111–180
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44:276–278
Chen J, Zhang YQ, Zhao CQ, Li AN, Zhou QX, Li DC (2007) Cloning of a gene encoding thermostable glucoamylase from Chaetomium thermophilum and its expression in pichia pastoris. J Appl Microbiol 103:2277–2284
Eckart MR, Bussineau CM (1996) Quality and authenticity of heterologous proteins synthesized in yeast. Curr Opin Biotechnol 7:525–530
Krall J, Bagley A, Mullenbach G, Hallewell R, Lynch R (1988) Superoxide mediates the toxicity of paraquat for cultured mammalian cells. J Biol Chem 263:1910–1914
Li DC, Gao J, Li YL, Lu J (2005) A thermostable manganese-containing superoxide dismutase from the thermophilic fungus Thermomyces lanuginosus. Extremophiles 9:1–6
Maheshwari R, Bharadwaj G, Bhat MK (2000) Thermophilic fungi: their physiology and enzymes. Microbiol Mol Biol Rev 64:461–488
Munns R (2005) Genes and salt tolerance: bringing them together. New Phytol 167:645–663
Nishikawa M, Nagatomi H, Nishijima M, Ohira G, Chang BJ, Sato E, Inoue M (2001) Targeting superoxide dismutase to renal proximal tubule cells inhibits nephrotoxicity of cisplatin and increases the survival of cancer-bearing mice. Cancer Lett 171:133–138
Raimondi S, Uccelletti D, Matteuzzi D, Pagnoni U, Rossi M, Palleschi C (2008) Characterization of the superoxide dismutase SOD1 gene of Kluyveromyces marxianus L3 and improved production of SOD activity. Appl Microbiol Biotechnol 77:1269–1277
Shin DS, Didonato M, Barondeau DP, Hura GL, Hitomi C, Berglund JA, Getzoff ED, Cary SC, Tainer JA (2009) Superoxide dismutase from the eukaryotic thermophile Alvinella pompejana: structures, stability, mechanism, and insights into amyotrophic lateral sclerosis. J Mol Biol 385:1534–1555
Shinjin E, Guo FX, Liu SA, Chen J, Wang YJ, Li DC (2007) Purification, characterization, and molecular cloning of a thermostable superoxide dismutase from Thermoascus aurantiacus. Biosci Biotechnol Biochem 71:1090–1093
Song NN, Zheng Y, SJ E, Li DC (2009) Cloning, expression, and characterization of thermostable manganese superoxide dismutase from Thermoascus aurantiacus var. levisporus. J Microbiol 47:123–130
Vieille C, Zeikus GJ (2001) Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 65:1–43
Wu JR, Lin Y, Zheng ZY, Lin CC, Zhan XB, Shen YQ (2010) Improvement of the CuZn-superoxide dismutase enzyme activity and stability as a therapeutic agent by modification with polysialic acids. Biotechnol Lett. doi:10.1007/s10529-010-0382-x
Yoo HY, Kim SS, Rho HM (1999) Over-expression and simple purification of human superoxide dismutase (SOD1) in yeast and its resistance to oxidative stress. J Biotechnol 68:29–35
Acknowledgments
This work was supported by the Chinese National Nature Science Foundation (31071723), the Chinese National Programs for High Technology Research and Development (2006AA10Z304), and the Chinese Project of Transgenic Organisms (2008ZX08001-002). We thank Dr Jiarui Li from Kansas State University for his suggested revisions of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10529_2011_543_MOESM2_ESM.tif
Supplementary Fig. 1 Phylogenetic tree of Chaetomium thermophilum Cu, ZnSOD and other fungal Cu, ZnSODs. C.T: Chaetomium thermophilum (DQ493760); C.G: Chaetomium globosum (XP_001222495); K.M: Kluyveromyces marxianus (CAO02396); A.F: Aspergillus flavus (AAM94904); A.C: Aspergillus clavatus NRRL (XP_001274132); C.P: Coccidioides posadasii (ABF73315); N.H: Nectria haematococca mpVI (EEU33541); S.P: Schizosaccharomyces pombe (NP593163); C.L: Cryptococcus liquefaciens (BAF42028); P.M: Penicillium marneffei ATCC (XP_002148587). (TIFF 199 kb)
Rights and permissions
About this article
Cite this article
Zhang, LQ., Guo, FX., Xian, HQ. et al. Expression of a novel thermostable Cu, Zn-superoxide dismutase from Chaetomium thermophilum in Pichia pastoris and its antioxidant properties. Biotechnol Lett 33, 1127–1132 (2011). https://doi.org/10.1007/s10529-011-0543-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-011-0543-6