Abstract
Phylogenetic relationships of Indian Citron (Citrus medica L.) with other important Citrus species have been inferred through sequence analyses of rbcL and matK gene region of chloroplast DNA. The study was based on 23 accessions of Citrus genotypes representing 15 taxa of Indian Citrus, collected from wild, semi-wild, and domesticated stocks. The phylogeny was inferred using the maximum parsimony (MP) and neighbor-joining (NJ) methods. Both MP and NJ trees separated all the 23 accessions of Citrus into five distinct clusters. The chloroplast DNA (cpDNA) analysis based on rbcL and matK sequence data carried out in Indian taxa of Citrus was useful in differentiating all the true species and species/varieties of probable hybrid origin in distinct clusters or groups. Sequence analysis based on rbcL and matK gene provided unambiguous identification and disposition of true species like C. maxima, C. medica, C. reticulata, and related hybrids/cultivars. The separation of C. maxima, C. medica, and C. reticulata in distinct clusters or sub-clusters supports their distinctiveness as the basic species of edible Citrus. However, the cpDNA sequence analysis of rbcL and matK gene could not find any clear cut differentiation between subgenera Citrus and Papeda as proposed in Swingle’s system of classification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Horticulture concerned with plants that are used by people for food, either as edible products, or for culinary ingredients, for medicinal use or ornamental and aesthetic purposes. They are genetically very diverse group and play a major role in modern society end economy. Fruits and vegetables are an important component of traditional food, but are also central to healthy diets of modern urban population (Bajpai et al. 2014; Feng et al. 2014; Ruttanaprasert et al. 2014; Mlcek et al. 2015). The genus Citrus L. belongs to the family Rutaceae of the subfamily Aurantioideae and is assumed to have originated in South and Southeast Asia particularly in the region extending from northeast India, eastward through the Malayan Archipelago to China and Japan, and southward to Australia (Swingle and Reece 1967). In Citrus fruits production, India is among the top five countries with its production of 11.15 million tons in a total area of 1.1 million hectare (Anonymous 2015).
Citrus taxonomy and phylogeny are very complex and ambiguous owing to sexual compatibility between citrus and related genera, long history of cultivation, adventive type of apomixis (nucellar polyembryony), high frequency of somatic bud mutation, etc. Sexual compatibility even between Citrus and related genera like Fortunella, Poncirus, etc., has contributed to the taxonomic misunderstanding (Frost and Soost 1968; Nicolosi et al. 2000). Citrus classification systems have been invented from time to time because taxonomy was based mainly on morphological and geographical data. Currently, two usually Citrus classification systems suggested by Swingle and Reece (1967) and Tanaka (1977) have been most widely used. The incongruity between them is that Swingle’s system distinguishes just 16 species, while Tanaka’s system identifies 162 species. Various studies undertaken by modern citrus taxonomists considered citron (Citrus medica), mandarin (C. reticulata), and pummelo (C. maxima) as the true Citrus species within the subgenus Citrus or the most similar ancestors to the most of the modern day cultivated Citrus species, and other species within this subgenus are hybrids derived from these true species, species of subgenus Papeda or closely related genera (Scora 1975; Barrett and Rhodes 1976; Gmitter and Hu 1990).
Taxonomic characterization leading to unequivocal identification of Citrus species and their genetic resources are essential fundamentals for Citrus breeding, Citriculture, and Citrus industry. Nair and Nayar (1997) followed primarily the scheme of Swingle and Reece (1967) and moderately that of Tanaka (1977) including 18 taxa, covering of eight species under subgenus Citrus, three under subgenus Papeda, and seven other indigenous Citrus varieties with a suspected hybrid origin and uncertain taxonomic affinities in a systematic explanation on Indian Citrus.
India takes pleasure in an outstanding situation in the “Citrus belt of the world” owing to her affluent prosperity of both wild and cultivated Citrus genetic resources (Nair and Nayar 1997; Malik et al. 2013). The north-eastern region of India is a rich wealth of various Citrus species, and this area might be the center of origin of several Citrus species including 17 Citrus species with their 52 cultivars, and 7 probable natural hybrids are originated in the North-eastern province of India (Bhattacharya and Dutta 1956) due to undisturbed deep forests by abiotic factors have also been reported from the region, thus bestowing this area with a special status of “wealth domicile” of Citrus germplasm (Sharma et al. 2004).
C. medica commonly known as citron is indigenous to India. It is monoembryonic in nature and is considered as one of the three basic species of Citrus (Scora 1975; Barrett and Rhodes 1976). The importance of C. medica in the ancestry of Citrus has been put into question mark. Such a geographically diverse species functioning as the other parent of these cultigens suggests that the natural distribution of C. medica is not restricted to India (Bayer et al. 2009). Due to this confusion of the role of C. medica as the basic species, there is an urgent need for studying the extent of diversity occurring in citron in different parts of India and their phylogenetic relationships with other closely related Citrus species and genera.
Recent progress in DNA sequencing techniques has allowed the extensive use of short DNA fragments, especially those of the chloroplast genome, in the study of phylogenetic relationships. Phylogenetic analyses based on the various regions of the chloroplast genome have been conducted in the family Rutaceae and the subfamily Aurantioideae (Jung et al. 2005; Li et al. 2007; Groppo et al. 2008; Bayer et al. 2009; Salvo et al. 2010; Lu et al. 2011; Hynniewta et al. 2014). The rbcL gene, located on the chloroplast DNA (cpDNA), encodes the large subunit of ribulose 1, 5-bisphosphate carboxylase/oxygenase, an enzyme that catalyzes carbon fixation in photosynthesis. Compared to most genes encoded in the cpDNA, the rbcL gene has a relatively slow nucleotide substitution rate (Tshering et al. 2010). The matK gene is also located on the cpDNA and encodes a maturase involved in splicing type II introns from RNA transcripts. The matK gene is encoded by the chloroplast trnK intron. Since matK has a relatively fast mutation rate, it evolves faster than the rbcL gene (Olmstead and Palmer 1994; Hilu and Liang 1997; Hilu et al. 2003; Tshering et al. 2010, 2013).
In the present study, the phylogenetic relationship of Indian Citron (C. medica) with other important Citrus species was analyzed using rbcL and matK gene sequence of the chloroplast DNA (cpDNA).
Materials and Methods
Plant Samples
Twenty three accessions representing 15 Citrus taxa and one out-group taxon Poncirus trifoliata were collected from wild as well as domesticated stocks from different parts of India. Young fresh leaf tissues from all the sample materials were collected and stored in silica gel (20–60 mesh) and were used subsequently for genomic DNA isolation. Details of accessions used for rbcL and matK sequence analyses are given in Table 1.
DNA Extraction
Total genomic DNA was isolated from a final set of 23 representative accessions through cetyl trimethyl ammonium bromide (CTAB) method (Rogstad 1993). Quantitation of isolated DNA was done spectrophotometrically, and its quality was checked by electrophoresis on 1.0% agarose gel.
PCR Amplification
Two regions of cpDNA (rbcL and matK) were amplified from each of the 23 accessions via PCR. The primers, used for polymerase chain reaction amplification of the rbcL gene, were rbcL 1-1-F (aF) (5′-ATGTCACCACAAACAGAGACTAAAGC-3′) and rbcL NN3-2 R (cR) (5′-GCAGCAGCTAGTTCCGGGCTCCA-3′) (Bayer et al. 2009). For matK gene, the primers used were matK-5′ trnK spacer matK 6 (5′-TGGGTTGCTAACTCAATGG-3′) and matK-5′ trnK spacer matK 5′ R (5′-GCATAAATATAYTCCYGAAARATAAGTGG-3′) (Bayer et al. 2009). DNA amplification was carried out in a BioeR Xp thermocycler, and the concentration of PCR components was optimized for amplification: 10 mMTris (pH 8.3), 50 mM KCl, 0.2 mM dNTP each, 2.5 mM MgCl2, 1 U Taq DNA polymerase, 10 pmol primer each, and 50 ng genomic DNA in 50 µl final reaction volume. The PCR was programmed as pre-denaturation at 94°C for 3 min, 35 cycles of denaturation at 94°C for 1 min, annealing at 57°C for 1 min, extension at 72°C for 1 min, and final extension at 72°C for 7 min. The total PCR products were electrophoresed in 0.8% low melting agarose gel (G Biosciences) at 120 V for 3 h, and bands were visualized in UV gel doc system (Mega Biosystematica, U.K.).
Sequence Analysis of rbcL and matK Gene
Twenty three representatives, including one accession of P. trifoliata as out-group taxon, were used for comparison of rbcL and matK gene sequences. PCR products were excised and purified using Clean HiMedia kit. The yield of purified DNA was quantified using UV spectrophotometer. Eluted PCR products were sequenced using an Applied Biosystems Automated Sequencer (Model 3730, version 3.1) using both forward and reverse primer. Sequences of 23 accessions of citrus including one out-group were annotated.
The identity of sequences was confirmed through a BLASTn search in NCBI database (Altschul et al. 1997). The sequences were aligned using Clustal-W program (Higgins et al. 1994) with the default settings. Phylogenetic analysis was carried out in MEGA 5 software (Tamura et al. 2011). Pair-wise sequence divergence rates between accessions were calculated using Maximum Composite Likelihood method (Tamura et al. 2004). Phylogeny reconstruction was carried out using maximum parsimony (MP) and neighbor joining (NJ) methods. MP tree was constructed using the close-neighbor-interchange algorithm, while NJ tree was obtained using the maximum composite likelihood criterion. Support values of the internal branches of MP and NJ trees were evaluated through boot strap method (500 replicates) (Felsenstein 1985).
Results
Sequence Analysis of matK Gene
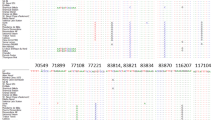
The BLASTn search helped determine that the new sequences were from sequence region and maximum homology was obtained from the sequences of Citrus. Sequence length of matK in the 23 Citrus accessions ranged from 736 to 862 bp (avg. Sequence length 824 bp). The dataset including alignment gaps and missing data comprised 887 bp aligned nucleotide positions, which included 786 conserved sites, 100 variable sites, and 65 parsimony informative sites. In the matK sequences, G+C content ranged from 32.9 to 35.2% with an average of 34.2%. Transition/Transversion bias (R) is 0.40. Substitution pattern and rates were estimated under the Kimura (1980) 2-parameter model (+G+I). A discrete Gamma distribution was used to model evolutionary rate differences among sites (5 categories, [+G], parameter = 0.6399). The rate variation model allowed for some sites to be evolutionarily invariable ([+I], 65.4397% sites). The nucleotide frequencies are A = 25.00%, T/U = 25.00%, C = 25.00%, and G = 25.00%. All positions containing gaps and missing data were eliminated. There were a total of 726 positions in the final dataset. Summary of matK sequence data is given in Table 2. The matK sequence analysis showed moderate rate of nucleotide divergence within the Citrus taxa (Table 3). Genetic divergence within Citrus group ranged from 0 (among citron accessions) to 0.021 (C. ichangensis and C. jambhiri) with an average of 0.007.
The phylogeny among Citrus genotypes was constructed through NJ method. In the NJ bootstrap consensus tree (Fig. 1), all the Citrus accessions were grouped into five distinct clusters:
Cluster I: C. ichangensis, C. maxima, and C. latipes
Cluster II: C. sinensis, C. karna, and C. limmettoides
Cluster III: C. macroptera, C. aurantiifolia, and C. limon
Cluster IV: C. reticulata, C. jambhiri, C. aurantium, and C. limonia
Cluster V: C. medica and C. indica
P. trifoliata was separately attached at the base of the tree as the diverging Citrus relative’s lineage. The phylogeny was also inferred through the maximum parsimony method which separated all the 23 accessions into five clusters as similar to NJ tree (Fig. 2).
Sequence Analysis of rbcL Gene
Sequence length of rbcL in the 23 Citrus accessions ranged from 1207 to 1297 bp (avg. Sequence length 1245 bp). The dataset including alignment gaps and missing data comprised 1307 bp aligned nucleotide positions, which included 1186 conserved sites, 115 variable sites, and 56 parsimony informative sites. In the rbcL sequences, G+C content ranged from 45.0% to 45.8% with an average of 45.4%. The estimated Transition/Transversion bias (R) is 0.82. Substitution pattern and rates were estimated under the Kimura (1980) 2-parameter model (+G+I). A discrete Gamma distribution was used to model evolutionary rate differences among sites (5 categories, [+G], parameter = 0.1230). The nucleotide frequencies are A = 25.00%, T/U = 25.00%, C = 25.00%, and G = 25.00%. All positions containing gaps and missing data were eliminated. There were a total of 1200 positions in the final dataset. Summary of rbcL sequence data is given in Table 4. The rbcL sequence analysis showed moderate rate of nucleotide divergence within the Citrus taxa (Table 5). Genetic divergence within Citrus group ranged from 0.00 (among the citron accessions) to 0.034 (C. karna, C. medica -MD99; C. karna, C. indica) (avg. 0.017).
The phylogeny constructed based on NJ bootstrap consensus tree divided all the 23 Citrus accessions into five distinct clusters as shown in Fig. 3.
Cluster I: C. macroptera, C. sinensis, C. jambhiri, C. maxima, C. reticulata, and C. karna Cluster II: C. indica
Cluster III: C. medica
Cluster IV: C. limonia, C. limon, and C. aurantiifolia
Cluster V: C. latipes, C. ichangensis, C. limmettoides, and C. aurantium
In rbcL sequence analysis also, P. trifoliata was found to be separately attached at the base of the tree as the diverging Citrus relative’s lineage. The phylogeny inferred through the maximum parsimony method also separated all the 23 accessions into five distinct clusters as similar to NJ tree (Fig. 4).
Discussion
Citrus classification is confusing and highly notorious. Various taxonomists have recognized 16 to 162 species in the genus Citrus (Swingle 1943). Most of the puzzlement is due to free hybridization of different species and incidences of intermediate forms. The cpDNA sequences are the primary source of characters for phylogenetic studies in plants (Bayer et al. 2009; Small et al. 2005). Protein-coding gene sequences such as rbcL and matK have been used to elucidate phylogenetic relationships among higher-level taxa (Chase et al. 1993; Tshering et al. 2010, 2013). Subsequently, the potential utility of non-coding regions of the chloroplast genome was recognized for lower-level studies (Taberlet et al. 1991). Recently, Lu et al. (2011) investigated the molecular phylogeny of 30 genotypes from six genera of the true citrus fruit trees by conducting research on three cpDNA regions. In another report, Morton et al. (2003) and Jena et al. (2009) carried out molecular phylogeny in Indian Citrus L. (Rutaceae) based on trnL-trnF sequence data of chloroplast DNA. Wali et al. (2013) studied the phylogenetic relationships on selected Citrus species based on Chloroplast Gene, rps14. Earlier, with the help of cpSSR, Cheng et al. (2005) and Deng et al. (2007) have reported the molecular phylogeny of Citrus.
In the present study, rbcL and matK gene sequence analyses of the cpDNA were used to investigate the phylogenetic relationship of Indian citron with other important commercial Citrus Spp. In our studies, C. maxima, C. medica, and C. reticulata were separated in distinct groups or sub-clusters which supports their distinctiveness as the true basal species of edible Citrus. This concept has gained much acceptance and support through recent morphological, biochemical, and molecular studies conducted by different citrus taxonomists (Scora 1975; Barrett and Rhodes 1976; Nicolosi et al. 2000; Arau´jo et al. 2003; Mabberley 2004; Liang et al. 2007; Jena et al. 2009).
C. medica (Citron) is one of the basal species of Indian origin and is believed to have acted as male parent in the origin of several hybrids/cultivars of Citrus such as all true lemons and rough lemon (Barrett and Rhodes 1976; Federici et al. 1998; Nicolosi et al. 2000; Gulsen and Roose 2001; Moore 2001; Mabberley 2004). Our rbcL and matK sequence data recognized C. medica as a true basic species as both wild and domesticated accessions of the species grouped in the cluster I with a very high bootstrap value of 87% (NJ tree) and 88% (MP tree). Citron, as an important true species, took part in the origin of many Citrus species, but our cpDNA data analysis indicates that citron has always acted as the male parent (Nicolosi et al. 2000).
C. maxima (Pummelo) and C. reticulata (Mandarin) are believed to have contributed to the development of several commercial citrus fruits, such as sour orange (C. aurantium, a cross between mandarin and pummelo), sweet orange (C. sinensis) (L.) Osbeck (a backcross between pummelo and mandarin), grapefruit (C. paradisi) (a backcross between pummelo and sweet orange) (Moore 2001; Mabberley 2004). Pummelo was reported as one of the three true Citrus species by Barrett and Rhodes (1976), and most of subsequent studies were in agreement with this statement (Federici et al. 1998; Nicolosi et al. 2000; Barkley et al. 2006; Uzun et al. 2009). Pummelo has played an important role as a parent of many citrus fruits, such as lemons, oranges, and grapefruits. C. maxima clustered with papedas, particularly with C. latipes. In UPGMA tree, the sour and sweet oranges were grouped together in a separate cluster along with the Khasi papeda and Melanesian papeda, while in the MP and NJ trees, the grapefruit and sour orange formed a separate cluster along with C. reticulata, and the sweet orange grouped with C. maxima along with the Khasi papeda and Melanesian papeda. The consistent grouping of sweet orange with C. maxima in the rbcL and matK derived trees indicates the role of C. maxima as a male parent in the origin of sweet oranges.
C. indica (Indian Wild Orange) is a true wild species endemic to the Garo hills in Meghalaya. Tanaka (1928) was the first to describe it as a new species. Swingle and Reece (1967), however, suspected C. indica to be of hybrid origin involving a wild species of Citrus (C. latipes) and one of the cultivated species of Citrus as putative parents. Therefore, elucidating its special taxonomic position as a true species or progenitor species of cultivated Citrus taxa. C. medica (citron), C. reticulata (mandarin), and C. maxima (pummelo) are defined as basic true species by Swingle and Reece (1967) a phylogenetic truth which was later supported by a number of workers (Barrett and Rhodes 1976; Jena et al. 2009; Kyndt et al. 2010; Kumar et al. 2012). C. indica accessions clustered with C. medica in both rbcL and matK sequence data based on NJ and MP trees. Similar clustering pattern was reported earlier by Nicolosi et al. (2000), Federici et al. (1998), and Jena et al. (2009) based on PCR–RFLP of cpDNA. Mabberley (2004) also subscribed Swingle’s view in treating C. indica as a species of suspected hybrid origin. Based on RAPD and PCR–RFLP data, Federici et al. (1998) argued against the hybrid origin of C. indica. Our studies also do not support the hybrid origin of C. indica as it consistently separated out as a distinct group along with C. medica.
C. aurantium (sour orange) is considered as a hybrid, a cross between C. reticulata (Mandarin) and C. maxima (Pummelo). In our study, we found that C. aurantium clustered with C. reticulata. Thus, our data support the role of Mandarin as one of the maternal parents in the hybrid origin of C. aurantium (Jena et al. 2009; Kumar et al. 2012). C. jambhiri (rough lemon) is considered as a hybrid originated from C. medica and C. reticulata (Scora 1975; Barrett and Rhodes 1976; Nicolosi et al. 2000; Mabberley 2004). Based on the UPGMA obtained through NJ tree and MP tree, C. jambhiri was found to be clustered with C. reticulata. Our data thus show a close relationship between C. reticulata and C. jambhiri and support the role of C. reticulata as maternal parent in the hybrid origin of C. jambhiri (Federici et al. 1998; Nicolosi et al. 2000; Barkley et al. 2006).
C. sinensis loosely clustered with C. maxima and C. reticulata in our rbcL sequence data. Its clustering with C. maxima in the rbcL NJ tree indicates the hybrid origin of C. sinensis involving C. maxima as one of the putative parents, thereby supporting the views of Barrett and Rhodes (1976), Luro et al. (1995), and Nicolosi et al. (2000). Several earlier workers hypothesized C. limon to be of complex hybrid origin involving two parents: citron and lime (Swingle 1943; Malik et al. 1974; Scora 1975) or citron and sour orange (Nicolosi et al. 2000; Gulsen and Roose 2001) or sour orange and lime (Hirai and Kozaki 1981; Torres et al. 1978a, b). Most lemons have highly similar morphological and biochemical characters, and some are reported to have originated by mutation from a single parental lemon tree. In our study, C. limon grouped with C. limonia, C. aurantifolia and C. macroptera based on cpDNA data. This study showed that C. aurantiifolia (sour lime) is involved as one of the parents in the origin of C. limon.
C. aurantiifolia was proposed as a trihybrid origin, involving citron, pummelo, and a species of Microcitrus in the parentage (Barrett and Rhodes 1976)). RFLP data of Federici et al. (1998) supported citron as one of the parents involved in the origin of C. aurantiifolia. In our data based on rbcL NJ tree, C. aurantiifolia was found to be closely related to C. limon and C. limonia. It was loosely clustered with C. medica, suggesting the role of Citron as one of the maternal parents involved in the origin of C. aurantiifolia. C. karna (Karna orange or Karna khatta) has long been known in India and exploited as a root stock for grafting commercial Citrus varieties. Fruit characters of C. karna show resemblances with C. aurantium, C. medica, and C. maxima. In our cpDNA analysis, C. karna consistently found a place along with other taxa of suspected hybrid origin. In our study based on rbcL sequence data, C. karna was found to be closely related with C. reticulata and C. maxima, which suggests the involvement of either C. reticulata or C. maxima as one of the maternal parents in the origin of C. karna. However, there is no conclusive evidence to elucidate the mode and actual parentage involved in the origin of C. karna.
C. limmettoides was supposed to have originated as a hybrid of C. aurantiifolia with C. limetta Risso or with a sweet citron (C. medica var. dulcis Risso et Poit) (Webber 1943) or a cross between C. aurantiifolia with C. sinensis (Barrett and Rhodes 1976). cpDNA profiling by Nicolosi et al. (2000) could not trace the parents involved in the origin of C. limmettoides, although a SCAR analysis by the same authors indicated citron and sweet orange as putative male and female parents, respectively, of the Indian sweet lime. In our study, C. limmettoides was found to be closely related with C. sinensis based on matK NJ and MP tree. This suggests that C. sinensis may be one of the possible parents of C. limmettoides (Barrett and Rhodes 1976). C. macroptera commonly called as Melanesian papeda has wide spread distribution in India especially in the northeastern part of India as compared to other endangered Citrus species. The fruits are being very juicy and vesicles very small resembling that of the lime. Swingle and Reece (1967) considered it as a promising rootstock and useful for breeding new rootstocks. In our study, C. macroptera clustered together with C. sinensis on the basis of rbcL sequence data, and it clustered with C. aurantiifolia and C. limon based on matK gene sequence in both NJ and MP trees. Our data infer close genetic relationship between this species and their probable origin from the same genetic lineage.
C. latipes (Khasi Papeda) is known to have originated in India probably in the North-Eastern part of India (Bhattacharya and Dutta 1956). The fruit being inedible have little commercial value. It has been tried as a root-stock for the Khasi orange (C. reticulata) and is found to be incompatible. Tanaka (1977) hypothesizes that C. latipes may have originated from C. maxima. The cpDNA data in our studies support this hypothesis. The presence of C. latipes in the pummelo cluster might indicate that the ancient maternal relationship is in the cluster. The cpDNA profiling by Nicolosi et al. (2000) also supported Pummelo as the maternal parent of C. latipes. C. ichangensis (Ichang papeda) is not a cultivated fruit and is absolutely inedible. It is reported to be very much cold-resistant. The fruits practically contain no juice and have no commercial importance. Its value as a root-stock has not yet been ascertained. Major differences exist between Swingle’s (1943) and Tanaka (1977) systems regarding the taxonomy of C. ichangensis. Swingle placed it in the subgenus Metacitrus, which contained all Mandarin species and some hybrids of C. ichangensis, but no other Papeda species at all. Zhu (1988) showed that C. ichangensis was a primitive Citrus species. Herrero et al. (1996) found that isozyme data clustered C. ichangensis with C. karna and C. meyeri, which are lemon types. The analysis of Fraction I protein conducted by Handa et al. (1986) showed that C. ichangensis obviously differs from the other Papeda species which originated in tropical or subtropical regions by its cold hardiness and having single flowers. The present studies based on sequence data of cpDNA results show that C. ichangensis is a distinct species very different from most other Citrus species (Federici et al. 1998; Nicolosi et al. 2000).
Swingle and Reece (1967) had divided citrus into two subgenera: Citrus and Papeda. The members of subgenus Papeda are distinguishable from subgenus Citrus in having large sized fruits containing acrid oil droplets in their pulp-vesicles; leaflets with broadly winged petioles that are usually as long or longer than the leaflet blades; free stamens; presence of purplish tinged on new shoots and flowers; and a epigeous mode of seed germination. However, our cpDNA analysis, based on rbcL and matK gene sequence, could not find any clear cut differentiation between subgenera Citrus and Papeda. This supports the earlier findings of earlier workers (Nicolosi et al. 2000; Jena et al. 2009; Hynniewta et al. 2014).
To conclude that the chloroplast DNA (cpDNA) analysis based on rbcL and matK sequence data carried out in Indian taxa of Citrus was useful in differentiating all the true species and species/varieties of probable hybrid origin in distinct clusters or groups. Sequence analysis based on rbcL and matK gene was able to provide unambiguous identification and disposition of true species like C. maxima, C. medica, C. reticulata, and related hybrids/cultivars. The separation of C. maxima, C. medica, and C. reticulata in distinct clusters or sub-clusters supports their distinctiveness as the basic species of edible citrus. The cpDNA sequence analysis of rbcL and matK gene could not find any clear cut differentiation between subgenera Citrus and Papeda according to Swingle’s system. However, this study was helpful in supporting the distinctiveness of C. indica, C. latipes and C. ichangensis as true species, besides elucidating the hybrid origin and relationships among the cultivated species/biotypes, such as C. aurantiifolia, C. limon, C. limmettoides, C. aurantium, C. sinensis, C. karna, and C. macroptera. The outcomes of this study will be further helpful in elucidating correct taxonomic identification, documentation, characterization and evaluation of Indian citron and its genetic resources to be used in future crop improvement programs.
Change history
24 October 2019
The Editor-in-Chief and the publisher have retracted this article [1] because of significant overlap with previously published articles [2, 3, 4, 5]. Ajit Uchoi, Surendra Kumar Malik, Ravish Chaudhary, Susheel Kumar, M.R. Rohini, Digvender Pal, and Sezai Ercisli disagree with the retraction. The publisher was not able to get in contact with Rekha Chaudhury, she did not respond to any correspondence about this retraction.
24 October 2019
The Editor-in-Chief and the publisher have retracted this article [1] because of significant overlap with previously published articles [2���5]. Ajit Uchoi, Surendra Kumar Malik, Ravish Chaudhary, Susheel Kumar, M.R. Rohini, Digvender Pal, and Sezai Ercisli disagree with the retraction. The publisher was not able to get in contact with Rekha Chaudhury, she did not respond to any correspondence about this retraction.
References
Altschul SF, Thomas LM, Alejandro AS, Jinghui Z, Zheng Z, Webb M, David JL (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res 25:3389–3402
Anonymous (2015) Indian Horticulture Database 2014 (National Horticulture Board, Ministry of Agriculture, Gurgaon, Haryana), pp 1–286
Arau´jo EF, Queiroz LP, Machado MA (2003) What is citrus? Taxonomic implications from a study of cp-DNA evolution in the tribe Citreae (Rutaceae subfamily Aurantioideae). Org Div Evol 3:55–62
Bajpai PK, Warghat AR, Sharma RK, Yadav A, Thakur AK, Srivastava RB, Stobdan T (2014) Structure and genetic diversity of natural populations of Morus alba in the Trans-Himalayan Ladakh Region. Biochem Genet 52(3):137–152
Barkley NA, Roose ML, Krueger RR, Federici CT (2006) Assessing genetic diversity and population structure in a citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112:1519–1531
Barrett HC, Rhodes AM (1976) A numerical taxonomy study of affinity relationships in cultivated Citrus and its close relatives. Syst Bot 1:105–136
Bayer RJ, Mabberley DJ, Morton CM, Cathy H, Sharma IK, Pfeil BE, Rich S, Hitchcock R, Sykes S (2009) A molecular phylogeny of the orange subfamily (Rutaceae: Aurantioideae) using nine cpDNA sequences. Am J Bot 96:668–685
Bhattacharya SC, Dutta S (1956) Classification of Citrus fruits of Assam, Sc. Monogr. 20, ICAR, New Delhi, p. 110
Chase MW, Soltis DE, Olmstead RG, Morgan D, Les DH, Mishler BD, Duvall MR, Price RA, Hills HG, Qui YL (1993) Phylogenetics of seed plants: an analysis of nucleotide sequences from the plastid gene rbcL. Ann Missouri Bot Gard 80:528–580
Cheng YJ, Carmen M, Meng HJ, Guo WW, Tao NG, Deng XX (2005) A set of primers for analyzing chloroplast DNA diversity in Citrus and related genera. Tree Physiol 25:661–672
Deng ZN, La-Malfa S, Xie XM, Xiong XG, Gentile A (2007) Identification and evaluation of chloroplast uni- and trinucleotide sequence repeats in citrus. Sci Hortic 111:186–192
FAO (2009) Food and Agricultural Organization of the United Nations. http://faostat.fao.org
Federici CT, Fang DQ, Scora RW, Roose ML (1998) Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor Appl Genet 96:812–822
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Feng SG, Lu JJ, Gao L, Liu JJ, Wang HZ (2014) Molecular phylogeny analysis and species identification of Dendrobium (Orchidaceae) in China. Biochem Genet 52(3):127–136
Frost HB, Soost RK (1968) Seed reproduction: development of gametes and embryos. In: Reuther W, Batchelor LD, Webber HB (eds) The Citrus industry, vol 2. California University Press, California, pp 290–324
Gmitter FG, Hu X (1990) The possible role of Yunnan, China, in the origin of contemporary Citrus species (Rutaceae). Econ Bot 44:237–277
Groppo MG, Pirani JR, Salatino MLF, Blanco SR, Kallunki JA (2008) Phylogeny of Rutaceae based on two noncoding regions from cpDNA. Am J Bot 95:985–1005
Gulsen O, Roose ML (2001) Chloroplast and nuclear genome analysis of parentage of lemons. J Am Soc Hortic Sci 126:210–215
Handa T, Ishizawa Y, Oogaki C (1986) Phylogenetic study of fraction I protein in the genus Citrus and its close related genera. Jp J Genet 61:15–42
Herrero R, Asins MJ, Carbonell EA, Navarro L (1996) Genetic diversity in the orange subfamily Aurantoideae. I. Interspecies and intragenus genetic variability. Theor Appl Genet 92:599–609
Higgins D, Thompson J, Gibson T (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Hilu KW, Liang H (1997) The matK gene: sequence variation and application in plant systematics. Am J Bot 84:830–839
Hilu KW, Borsch T, Muller K, Soltis DE, Soltis PS (2003) Angiosperm phylogeny based on matK sequence information. Am J Bot 90:1758–1776
Hirai M, Kozaki I (1981) Isozymes of Citrus leaves. Proc Int Soc Citric 1:10–13
Hynniewta M, Malik SK, Rao SR (2014) Genetic diversity and phylogenetic analysis of Citrus (L) from north-east India as revealed by meiosis, and molecular analysis of internal transcribed spacer region of rDNA. Meta Gene 2:237–251
Jena SN, Kumar S, Nair NK (2009) Molecular phylogeny in Indian Citrus L. (Rutaceae) inferred through PCR-RFLP and trnL-trnF sequence data of chloroplast DNA. Sci Hortic 119:403–416
Jung YH, Kwon HM, Kang SH, Kang JH, Kim SC (2005) Investigation of the phylogenetic relationships within the genus Citrus (Rutaceae) and related species in Korea using plastid trnL-trnF sequences. Sci Hortic 104:179–188
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar S, Nair KN, Jena SN (2012) Molecular differentiation in Indian Citrus L. (Rutaceae) inferred from nrDNA ITS sequence analysis. Genet Resour Crop Evol 60:59–75. doi:10.1007/s10722-012-9814-x
Kyndt T, Dung TN, Goetghebeur P, Toan HT, Gheysen G (2010) Analysis of ITS of the rDNA to infer phylogenetic relationship among Vietnamese Citrus accessions. Genet Resour Crop Evol 57:183–192
Li YZ, Cheng YJ, Tao NG, Deng XX (2007) Phylogenetic analysis of mandarin landraces, wild mandarins, and related species in China using nuclear LEAFY second intron and plastid trnL-trnF sequence. J Am Soc Hortic Sci 132:796–806
Liang G, Xiong G, Guo Q, He Q, Li X (2007) AFLP analysis and the taxonomy of Citrus. Acta Hortic 760:137–142
Lu ZH, Zhou ZQ, Xie RJ (2011) Molecular phylogeny of the ‘‘true citrus fruit trees’’ group (Aurantioideae, Rutaceae) as inferred from chloroplast DNA sequence. Agric Sci China 10:49–57
Luro F, Laigret F, Bove JM, Ollitrault P (1995) DNA amplified fingerprinting, a useful tool for determination of genetic origin and diversity analysis in Citrus. Hortic Sci 30:1063–1067
Mabberley DJ (2004) Citrus (Rutaceae): a review of recent advances in etymology, systematics and medical applications. Blumea 49:481–498
Malik MN, Scora RW, Soost RK (1974) Studies on the origin of lemon. Hilgardia 42:361–382
Malik SK, Kumar S, Singh IP, Dhariwal OP, Chaudhury R (2013) Socio-economic importance, domestication trends and in situ conservation of wild Citrus species of Northeast India. Genet Resour Crop Evol. doi:10.1007/s10722-012-9948-x
Mlcek J, Valsikova M, Druzbikova H, Ryant P, Jurikova T, Sochor J, Borkovcova M (2015) The antioxidant capacity and macroelement content of several onion cultivars. Turk J Agri For 39:999–1004
Moore GA (2001) Oranges and lemons: clues to the taxonomy of Citrus from molecular markers. Trends Genet 17:536–540
Morton CM, Grant M, Blackmore S (2003) Phylogenetic relationships of the Aurantioideae inferred from chloroplast DNA sequence data. Am J Bot 90:1463–1469
Nair KN, Nayar MP (1997) Rutaceae. In: Hajra PK, Nair VJ, Daniel P (eds) Flora of India, vol IV. Botanical Survey of India, Calcutta, pp 229–407
Nicolosi E, Deng ZN, Gentile A, La Malfa S, Continella G, Tribulato E (2000) Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor Appl Genet 100:1155–1166
Olmstead RG, Palmer JD (1994) Chloroplast DNA systematics: a review of methods and data analysis. Am J Bot 81:1205–1224
Rogstad SH (1993) Saturated NaCl-CTAB solution as a means of field preservation leaves for DNA analysis. Taxon 41:701–708
Ruttanaprasert R, Banterng P, Jogloy S, Vorasoot N, Kesmala T, Kanwar RS, Holbrook CC, Patanothai A (2014) Genotypic variability for tuber yield, biomass, and drought tolerance in Jerusalem artichoke germplasm. Turk J Agri For 38:570–580
Salvo G, Ho SY, Rosenbaum G, Ree R, Conti E (2010) Tracing the temporal and spatial origins of island endemics in the Mediterranean region: a case study from the citrus family (Ruta L., Rutaceae). Syst Biol 59:705–722
Scora RW (1975) On the history and origin of citrus. Bull Torrey Bot Club 102:369–375
Sharma BD, Hore DK, Gupta SG (2004) Genetic resources of Citrus of north-eastern India and their potential use. Genet Resour Crop Evol 51:411–418
Small RL, Lickey EB, Shaw J, Hauk WD (2005) Amplification of non-coding chloroplast DNA for phylogenetic studies in lycophytes and monilophytes with comparative example of relative phylogenetic utility from ophioglossaceae. Mol Phylo Evol 36(507):522
Swingle WT (1943) The botany of Citrus and its wild relatives. In: Webber HJ, Batchelor DL (eds) The citrus industry, vol 1. University of California, Berkeley, pp 128–474
Swingle WT, Reece PC (1967) The botany of citrus and its wild relatives. In: Reuther W, Webber HJ, Batchelor LD (eds) The citrus industry, vol 1. University of California Press, Berkeley, CA, USA, pp 389–390
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenetics by using the NJ methods. Proc Nat Acad Sci (USA) 101:11030–11035
Tamura K, Dudley J, Nei M et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony method. Mol Biol Evol 28:2731–2739
Tanaka T (1928) On certain new species of Citrus. Studia Citrol 2:155–164 Japanese with English, Latin resume
Tanaka T (1977) Fundamental discussion of citrus classification. Studia Citrologia 14:1–6
Torres AM, Soost RK, Diedenhofen U (1978) Leaf isozymes as genetic markers in Citrus. Am J Bot 65:869–881
Tshering P, Anai T, Nagano Y, Matsumoto R, Yamamoto M (2010) Phylogenetic relationships of Citrus and its relatives based on rbcL gene sequences. Tree Genet Genomes 6:931–939
Tshering P, Yamamoto M, Ide MUM, Matsumoto N, Matsumoto R, Nagano Y (2013) Phylogenetic relationships of citrus and its relatives based on matK gene sequences. PLoS ONE 8(4):e62574
Uzun A, Yesiloglu T, Aka-Kacar Y, Tuzcu O, Gulsen O (2009) Genetic diversity and relationships within Citrus and related genera based on sequence related amplified polymorphism markers (SRAPs). Sci Hort 121:306–312
Wali S, Munir F, Mahmood T (2013) Phylogenetic studies of selected Citrus species based on chloroplast gene, rps14. Int J Agric Biol 15:357–361
Webber HJ (1943) Cultivated varieties of Citrus. In: Webber HJ, Batchelor DL (eds) The Citrus Industry, vol 1. University of California, Berkeley, pp 475–668
Zhu LW (1988) Numerical taxonomy of Citrus species in China (in Chinese with English abstract). Acta Phytotaxon Sin 26:353–361
Acknowledgments
The authors are grateful to Director, NBPGR and Head, Exploration and Collection Division, NBPGR, New Delhi for encouragement and support. The authors thank the officers and staff of forest departments and ICAR institutes/centres in Northeast India for assistance and support during exploration and survey trips. The authors are also thankful to Post Graduate School, IARI and University Grant Commission for awarding Doctorate Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
The Editor-in-Chief and the publisher have retracted this article because of significant overlap with previously published articles. Ajit Uchoi, Surendra Kumar Malik, Ravish Chaudhary, Susheel Kumar, M.R. Rohini, Digvender Pal, and Sezai Ercisli disagree with the retraction. The publisher was not able to get in contact with Rekha Chaudhury. She did not respond to any correspondence about this retraction.
About this article
Cite this article
Uchoi, A., Malik, S.K., Choudhary, R. et al. RETRACTED ARTICLE: Inferring Phylogenetic Relationships of Indian Citron (Citrus medica L.) based on rbcL and matK Sequences of Chloroplast DNA. Biochem Genet 54, 249–269 (2016). https://doi.org/10.1007/s10528-016-9716-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-016-9716-2