Abstract
A novel actinobacterium, designated MM109T, was isolated from a moonmilk deposit collected from the cave ‘Grotte des Collemboles’ located in Comblain-au-Pont, Belgium. Based on a polyphasic taxonomic approach comprising chemotaxonomic, phylogenetic, morphological, and physiological characterization, the isolate has been affiliated to the genus Streptomyces. Multilocus sequence analysis based on the 16S rRNA gene and five other house-keeping genes (atpD, gyrB, rpoB, recA and trpB) showed that the MM109T isolate is sufficiently distinct from its closest relative, Streptomyces peucetius strain AS 4.1799T, as to represent a novel species. The phylogenetic distinctiveness of the taxon represented by isolate MM109T was supported by the isolation and identification of additional twelve moonmilk-derived isolates, which according to multilocus sequence analysis were clustered along with MM109T. Scanning electron microscopy observations revealed complex and diversified structures within a MM109T colony, made from branching vegetative mycelia. The spore chains of the MM109T isolate undergo complete septation at the late stages of the morphological differentiation process, leading to the formation of packs of smooth cylindrical-shaped spores. Isolate MM109T produces several intracellular and diffusible pigments, particularly an intracellular green-pigmented secondary metabolite, which was identified through UPLC-ESI–MS analysis as ferroverdin A, an iron-chelating molecule formerly extracted and characterized from Streptomyces sp. strain WK-5344. The isolate MM109T is thus considered to represent a novel species of Streptomyces, for which the name Streptomyces lunaelactis sp. nov. is proposed with the type strain MM109T (=DSM 42149T = BCCM/LMG 28326T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Streptomyces species are high-GC Gram-positive bacteria, well-known for their prolific secondary metabolism, being a source of valuable bioactive compounds with medical and biotechnological properties (Hopwood 2007). These soil-dwelling bacteria are also active in soil mineralization through their enzymatic arsenal specialized in carbohydrate utilization (Hodgson 2000; Bertram et al. 2004). Surprisingly, these active decomposers of organic matter have also been found in oligotrophic environments such as the deep sea (Gartner et al. 2011), deep subsurface soil (Zlatkin et al. 1996) and cave deposits (Barton et al. 2007; Portillo et al. 2009; Porca et al. 2012). Among the latter, a diverse and rich microbiome has been reported for a wide range of secondary structures, also known as speleothems, such as pool fingers, stalactites, stalagmites, cave pisoliths and moonmilk (Tetu et al. 2013; Ortiz et al. 2014). Moonmilk is a white carbonate deposit with a morphology—powdery, dry, pasty, wet or even liquid—that depends on its water content (Canaveras et al. 2006). Within the moonmilk structure, along with various calcite minerals, dense webs of bacterial filaments have been observed (Canaveras et al. 2006). Actinobacteria, including Streptomyces, are considered to be the major contributors to these filaments and the moonmilk microbial community (Canaveras et al. 2006; Barton and Northup 2007; Cuezva et al. 2012). In order to assess the genetic and phenotypic features of hypogene streptomycetes in comparison to their soil counterparts, we selectively isolated actinomycetes from a moonmilk speleothem collected in the cave ‘Grotte des Collemboles’ located in Comblain-au-Pont, Belgium. This work presents the first polyphasic taxonomic description of a Streptomyces species isolated from a moonmilk deposit, based on phylogenetic and chemotaxonomic analyses combined with phenotypic and physiological characterizations.
Materials and methods
Isolation of strains
Strain MM109T was isolated together with twelve other isolates, which displayed green-pigmentation of the substrate mycelium, from the moonmilk deposits collected in the cave ‘Grotte des Collemboles’ (Comblain-au-Pont, Belgium) in January 2012. Moonmilk samples were freeze-dried on a VirTis Benchtop SLC Lyophilizer (SP Scientific, Warminster, PA, USA) and around 250 mg of the lyophilisates were suspended in 0.25X strength Ringer’s solution supplemented with 0.001 % Tween 80. Serial dilutions were inoculated on International Streptomyces Project (ISP) media (Shirling and Gottlieb 1966) and starch nitrate (SN) medium (Waksman 1961) supplemented with nalidixic acid (75 µg/ml) and nystatin (50 µg/ml) to suppress the growth of Gram-negative bacteria and fungi, respectively. Strains were isolated after 1 month of incubation at 17 °C and preserved on ISP-2 slopes at 4 °C and as 20 % glycerol mycelium stock at −20 °C.
Chemotaxonomic characterization
Chemotaxonomy analyses were carried out by the Identification Service of DSMZ, Braunschweig, Germany. Isolate MM109T was cultivated in tryptone-yeast extract (ISP-1) broth for 3 days at 28 °C and 180 rpm. Cells were harvested by centrifugation (10 min at 4,000 rpm), washed twice with distilled water and freeze-dried. Fatty acid methyl esters were extracted according to the method of Miller (1982) and Kuykendall et al. (1988), separated on an Agilent 6890N Network Gas Chromatograph (Agilent Technologies, Santa Clara, CA, USA) and subsequently identified and quantified using the ACTIN6 method of the Sherlock Microbial Identification System (MIS) 6.1 software (MIDI Inc, Newark, USA). Respiratory lipoquinones and polar lipids were isolated using the two stage method described by Tindall (Tindall 1990a, b) and separated by thin layer chromatography (TLC). Respiratory lipoquinones were further analyzed by HPLC, and polar lipids by the previously described method of Tindall et al. (2007). Analysis of the isomeric forms of diaminopimelic acid (DAP) and sugars in the whole-cell hydrolysates were carried out by TLC according to Staneck and Roberts (1974).
Amplification and sequencing of house-keeping genes
Amplification of the 16S rRNA gene was performed using the bacterial universal primers 8F and 1541R (Onstott et al. 2009). In addition, amplicons of five house-keeping genes—atpD (ATP synthase subunit beta), gyrB (DNA gyrase subunit B), recA (recombinase A), rpoB (RNA polymerase subunit B) and trpB (tryptophan synthase subunit B)—were generated using primers atpDPF-atpDPR, gyrBPF-gyrBPR, recAPF-recAPR, rpoBPF-rpoBPR, and trpBPF-trpBPR, respectively, as described previously (Guo et al. 2008). Isolates were cultivated for 3 days at 28 °C in LB (Luria–Bertani) liquid medium. Genomic DNA was extracted from the mycelium with the GenElute Bacterial Genomic DNA Kit (Sigma-Aldrich, St. Louis, MO, USA) according to manufacturer’s instructions. PCR reactions contained 1.5 ng/μl template DNA, 1× Pfu buffer (Promega, Madison, WI, USA), 200 μM of each dNTP, 10 % DMSO, 1 µM of each primer, 2.5 U Pfu DNA Polymerase (Promega) and sterile water to a final volume of 50 μl. The thermal cycling conditions comprised an initial denaturation step at 96 °C for 2 min, followed by 35 cycles of denaturation (94 °C for 30 s), primer annealing (62 °C for 30 s) and extension (72 °C for 2 min). A final extension step was performed at 72 °C for 5 min. PCR products were separated by electrophoresis on a 1 % agarose gel and corresponding bands were excised. Amplicons were purified using the Gene JET Gel Extraction Kit (Thermo Scientific, Waltham, MA, USA) and sequenced on an ABI 3730 DNA Analyzer (Applied Biosystems, Foster City, CA, USA) using both forward and reverse primers. Amplification and sequencing were carried out in triplicate and the consensus sequence of each gene was deduced from multiple sequence alignment using the Multalin software (Corpet 1988). The sequences of the 16S rRNA, atpD, gyrB, recA, rpoB and trpB genes of the isolate MM109T have been deposited in GenBank with the accession numbers KM207217, KM207218, KM207219, KJ862806, KJ862819, and KJ862832, respectively. The accession numbers of 16S rRNA, recA, rpoB and trpB genes of twelve additional green-pigmented isolates deposited in the GenBank database are available in the Online Supplementary Table 1.
16S rRNA gene and multilocus sequence (MLSA) phylogenetic analyses
The almost complete 16S rRNA gene sequence of isolate MM109T was aligned along with the sequences of the most closely related Streptomyces species (>99 % of identity) found by the EzTaxon server (http://www.ezbiocloud.net/eztaxon, Kim et al. 2012), and representative Streptomyces type strains, retrieved from GenBank, using MUSCLE (Edgar 2004). The alignment was manually trimmed to 1,395 bp, and a neighbour-joining (NJ) (Saitou and Nei 1987) tree based on the Kimura two-parameter model (Kimura 1980) was constructed using the MEGA 6.0 software (Tamura et al. 2013), with Saccharopolyspora erythrea NRRL 2338T as an outgroup. The robustness of the tree topology was evaluated by bootstrap analysis (Felsenstein 1985) based on 1,000 resamplings. In addition, a maximum-likelihood (ML) tree (Felsenstein 1981) was constructed as above in order to validate the topology observed with the NJ tree.
Multilocus sequence analysis (MLSA) was carried out according to Guo et al. (2008). Briefly, the partial sequences of the six house-keeping genes obtained for isolate MM109T were trimmed to 496 bp (atpD), 386 bp (gyrB), 504 bp (recA), 540 bp (rpoB) and 571 bp (trpB), along with a 426 bp fragment of the 16S rRNA gene containing the Streptomyces-specific variable γ region (Guo et al. 2008). A BLAST search was used to identify close relatives and their representative 16S rRNA genes and other house-keeping genes were retrieved from GenBank. Trimmed sequences of the six house-keeping genes were concatenated head-to-tail in-frame in the order 16S rRNA-atpD-gyrB-recA-rpoB-trpB, generating a final sequence of 2,828 bp. Concatenated sequences were aligned using MUSCLE (Edgar 2004) and a poorly aligned region corresponding to the last 95 bp of the gyrB gene was removed to increase phylogenetic resolution. MLSA for all thirteen green-pigmented isolates was carried out based on the trimmed sequences of four house-keeping genes concatenated in the order 16S rRNA-recA-rpoB-trpB, generating a 2,040 bp final sequence. NJ and ML trees for each MLSA were constructed as described above for the 16S rRNA gene.
Phenotypic characterization
Phenotypic properties of isolate MM109T were determined on ISP media 1 to 7 (prepared as described by Shirling and Gottlieb (1966) or obtained from HiMedia Laboratories, Mumbai, India), Czapek’s agar (Waksman 1967) and nutrient agar (Waksman 1961) after 2 and 3 weeks of incubation at 28 °C. Colours of substrate mycelium and diffusible pigments were characterized using ISCC-NBS colour charts (Kelly 1964). Carbon source utilization pattern was established by inoculation of isolate MM109T on minimal media (Kieser et al. 2000) supplemented with 1 % d-(+)-glucose, sucrose, l-(+)-arabinose, d-(+)-raffinose, l-(+)-rhamnose, d-(−)-fructose, d-mannitol, maltose, inositol, N-acetylglucosamine, d-(+)-xylose, and water as negative control. Tolerance to a range of NaCl concentrations (1–10 %, at 1 % intervals) and pH (5.0–12.0, at 1.0 pH unit intervals) was quantified after 2 weeks of cultivation on ISP-1 solid medium (covered with cellophane discs). Growth at different pH values was determined using the buffer system described previously (Xu et al. 2005). The tolerance to a range of temperatures (4, 10, 12, 14, 17, 20, 28, 30 and 37 °C) was first roughly assessed after 2 weeks of cultivation on ISP-1 solid medium and next quantified after 5 days of cultivation in ISP-1 liquid medium.
Scanning electron microscopy (SEM)
The morphological characteristics of vegetative and aerial hyphae of isolate MM109T were assessed after 2 weeks and 1 month of incubation at 28 °C on ISP-4 medium by cryo-scanning electron microscopy (JSM-840A, JEOL, Japan), as previously described (Tenconi et al. 2012).
UPLC-ESI–MS analysis of ferroverdin A
A green pigment was extracted from three moonmilk-derived strains—MM28, MM37 and MM109T. Isolates were inoculated on different ISP media covered with cellophane discs (GE Osmonics Labstore, Ref K01CP09030) and incubated for 1 week at 28 °C. The green-pigmented metabolite was extracted from the fresh biomass with acetone and concentrated using a rotary evaporator (IKA RV 10 digital, VWR International, LLC, USA). The crude extract from the isolate MM109T was separated on a silica membrane by TLC (Sigma-Aldrich, St. Louis, MO, USA) for 1 h using acetone as the mobile phase. The green band was collected from the TLC plate and extracted with acetone. The purified green pigment from isolate MM109T and the crude extracts from the isolates MM28 and MM37 were analyzed with an UPLC (Acquity H-class, Waters s.a., Zellik, Belgium) coupled to a single quadrupole mass spectrometer (Waters SQD mass analyzer) on an ACQUITY UPLC BEH C18 1.7 µm column, as described previously (Lambert et al. 2014). The elution of the green pigment was carried out at 40 °C with a gradient (started 2 min after injection) of acetonitrile (ACN) ranging from 15 to 95 % in 5 min at a constant flow rate of 0.6 ml/min (ACN and water were acidified with 0.1 % formic acid). Pigment was detected in electrospray negative ion mode (scan in the mass range m/z 200–1,300) by setting SQD parameters as follows: source temperature 130 °C; desolvation temperature 80 °C; and desolvation/cone nitrogen flow: 600/50 L h−1. For optimal detection, the cone voltage was set at −80 V.
Results and discussion
Chemotaxonomic characteristics
A green-pigmented colony designated MM109T was selected in the course of an actinomycetes-specific isolation procedure from a lyophilized moonmilk sample obtained from the cave ‘Grotte des Collemboles’ (Comblain-au-Pont, Belgium). Observation under a light microscope of pellet-forming filaments in liquid culture suggested that isolate MM109T is most likely a filamentous actinomycete (data not shown). Since chemotaxonomy has been shown to be appropriate for differentiation of genera within the order Actinobacteria (Goodfellow et al. 2012), a complex chemical characterization was carried out for this isolate. The detection of ll-diaminopimelic acid in the whole-organism hydrolysate indicated that the cell-wall is of chemotype 1 (Lechevalier and Lechevalier 1970), suggesting that the isolate MM109T belongs to the genus Streptomyces (Leyh-Bouille et al. 1970). Additionally, as typically observed for streptomycetes (Manfio et al. 1995), the predominant respiratory quinones detected were the octa- and hexahydrogenated menaquinones with nine isoprene units—MK-9(H8) (63 %) and MK-9(H6) (21 %) respectively, followed by MK-10(H2) (6 %), MK-9(H4) (4 %), MK-10 (3 %) and MK-9(H2) (1 %). The major polar lipids were identified as phosphatidylinositol, diphosphatidylglycerol and a phosphoglycolipid, which are commonly observed among actinomycetes, and phosphatidylethanolamine, characteristic of the phospholipid type PII reported for members of the genus Streptomyces (Lechevalier et al. 1977). The cellular fatty acid profile was found to be a mixture of saturated straight-chain as well as iso- and anteiso- branched-chain fatty acids (fatty acid type 2c) which are also characteristic of actinomycetes (Kroppenstedt 1985). Major components (>10 %) were identified as anteiso-C15:0 (31.0 %) and iso-C16:0 (27.8 %), and minor components (<10 %) were anteiso-C17:0 (6.6 %), iso-C14:0 (5.7 %), iso-C15:0 (5.7), iso C16:1 H (4.3 %), C15:0 (4.0 %), anteiso-C17:1 (3.2 %), C16:0 (2.4 %), cis 9-C16:1 (2.1 %) and methyl-C16:0 (1.8 %). The chemotaxonomic features are thus consistent with the assignment of isolate MM109T to the genus Streptomyces (Kämpfer 2012).
Phylogenetic analysis
An EzTaxon analysis of the almost complete (1,420 bp) 16S rRNA gene sequence of MM109T revealed the highest sequence similarity with Streptomyces rubiginosohelvolus NBRC 12912T (99.37 % identity—1,411 nt out of 1,420 nt). A phylogenetic analysis including ten other related Streptomyces species (>99 % sequence identity) as well as a selection of representative Streptomyces type strains confirmed the affiliation of the isolate MM109T to the genus Streptomyces (Online Supplementary Fig. 1). The phylogenetic tree created by both tree-making algorithms (NJ and ML) showed that MM109T forms a branch distinct from its closest relatives.
However, analysis of the 16S rRNA gene by itself has been reported to be insufficient to define phylogenetic relationships among closely related Streptomyces species (Guo et al. 2008; Rong et al. 2009; Rong and Huang 2010, 2012; Labeda 2011; Han et al. 2012), mainly due to the highly conserved nature of this gene within the genus (Labeda 2011). Therefore, the phylogenetic analysis was refined by carrying out a MLSA (Guo et al. 2008). This approach increases the phylogenetic resolution (Rong and Huang 2010), as it is based on several house-keeping genes which are conserved enough to constitute a phylogenetic marker and possess more variable sites than the 16S rRNA gene (Rong and Huang 2010). Therefore, in addition to the 16S rRNA gene, partial sequences of five other house-keeping genes, namely atpD, gyrB, recA, rpoB and trpB, were obtained for the isolate MM109T. BLAST analyses of these five protein-coding genes revealed that the closest relatives of MM109T were Streptomyces peucetius strain AS 4.1799T (97.4 % atpD gene sequence similarity), Streptomyces spororaveus strain AS 4.1926T (87.7 % gyrB gene sequence similarity), Streptomyces yanii strain AS 4.1146T (93.8 % recA gene sequence similarity), Streptomyces mauvecolor strain AS 4.1997T (93.9 % rpoB gene sequence similarity) and Streptomyces hirsutus strain NRRL B-2713T (91.2 % trpB gene sequence similarity). Affiliation of MM109T to different Streptomyces species according to different house-keeping genes suggested that this isolate is phylogenetically distinct from its neighbours. Phylogenetic analysis based on the alignment of concatenated sequences showed that the isolate MM109T is most closely related to Streptomyces peucetius strain AS 4.1799T (Fig. 1). The topology of the tree was not affected by tree-making algorithms. Moreover, pairwise distances calculated for MM109T and the related species using concatenated sequences of atpD-gyrB-recA-rpoB-trpB were above 0.007, which corresponds to 70 % DNA–DNA homology, considered to be the threshold for species determination (Rong and Huang 2012; Busarakam et al. 2014; Labeda et al. 2014). Thus, MLSA analysis confirmed that MM109T should be classified as a novel species within the genus Streptomyces.
NJ tree based on MLSA analysis of the partial sequences from six house-keeping genes (16S rRNA, atpD, gyrB, recA, rpoB, trpB) of isolate MM109T (in bold), 16 closest neighbours and 8 selected representative Streptomyces type strains. Saccharopolyspora erythraea NRRL 2338T was used as an outgroup. Node numbers indicate percent bootstrap support with 1,000 resamplings. Bar 0.02 substitutions per nucleotide position
Phenotypic and physiological characterization
Isolate MM109T was able to grow in all tested media, with good growth observed on ISP-1, ISP-3, ISP-6 and nutrient agar (Table 1). The colour of the vegetative mycelium varied from yellowish grey, yellowish green to strong yellow green, while aerial hyphae were white to grey. The same diffusible pigments were produced in both commercial and non-commercial media. Melanin was produced on peptone-yeast extract-iron agar (ISP-6). The green pigmentation of the isolate, which was the phenotype that prompted us to select the MM109T colony in our original screening for moonmilk-dwelling Actinobacteria, was also observed when the strain was cultivated on ISP3 media.
Isolate MM109T was found to grow on l-(+)-arabinose, sucrose, d-(+)-xylose, inositol, d-(+)-raffinose, maltose and N-acetylglucosamine, whereas growth was observed to be drastically reduced when d-(+)-glucose, d-(−)-fructose, d-mannitol, and l-(+)-rhamnose were used as sole carbon sources. Isolate MM109T was observed to grow within the temperature range of 10 to 30 °C, between pH 6 to 8, and in the presence of 1 to 4 % NaCl, with the highest biomass accumulation observed at 18–28 °C, 3 % NaCl and pH 6 (Fig. 2). The comparison of several phenotypic and physiological properties of MM109T with the most closely related Streptomyces type strains found by MLSA analysis clearly highlighted differences between them, supporting the divergence of MM109T strain from its phylogenetic neighbours (Table 2).
Salinity, pH, and temperature tolerance of isolate MM109T. Influence of increasing concentration of NaCl (a), medium pH (b) and the range of temperatures (c) on biomass accumulation. Salinity and pH tolerance assays were performed on solid ISP-1 media while assays for tolerance to temperature were performed on liquid ISP-1 medium
Analysis of vegetative filament morphology and spore-forming aerial hyphae
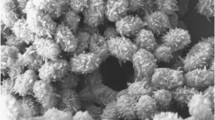
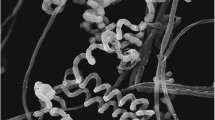
Examination of morphological features by scanning electron microscopy allowed visualization of the complex architecture of the MM109T colony (Fig. 3a). Several types of filamentous structures were observed depending on the region of the colony: filaments were observed to be either well separated (Fig. 3b) or fused together in thick root-shaped branching structures (Fig. 3c) or networking cavities (Fig. 3d); sporadically, round or rod-shaped small aggregates and sphere-like structures were detected (Fig. 3e), but they seemed to be generated from inorganic matter rather than through differentiation of vegetative filaments. Despite a clear whiE-associated grey-pigmentation typical of fully maturated spore chains (Davis and Chater 1990), examination of the whole colony surface (Fig. 3a) revealed only occasional aerial hyphae, with a higher frequency in the region of the colony detailed in Fig. 3f. Aerial hyphae were either linear or coiled, with regular, but incomplete septations every ~0.8 µm (Fig. 3f–g). Fully maturated rod-shaped or cylindrical smooth-surfaced spores were observed after prolonged incubation (~30 days). Due to complete septation, mature spores did not remain in aerial chains but collapsed into packs of about 20–40 spores (Fig. 3f–g), a phenotype that is more similar to the premature autolysis-driven spore separation observed for a Streptomyces coelicolor ssgE mutant (GSE1) (Noens et al. 2005), rather than ectopic sporulation emerging from the substrate mycelium.
Cryo-scanning electron micrographs of isolate MM109T grown on inorganic salts-starch agar (ISP-4) at 28 °C for 4 weeks (micrographs a–f) and 9 weeks (g and h). Morphological observations revealed a structurally complex colony of isolate MM109T (a) developing differential vegetative filaments—from well separated (b) or fused branching structures (c) to networking cavities (d). Round or rod-shaped small aggregates of unknown origin were observed sporadically (e). Linear or coiled aerial hyphae (f) maturated into packs of collapsed spore chains (g and h)
Identification of additional 12 moonmilk isolates belonging to the same lineage as the type strain MM109T
In order to convincingly demonstrate the novelty of the isolate MM109T and to assess if more representatives of this species are present in the different moonmilk deposits from the same cave, we screened our moonmilk actinobacterial collection for isolates displaying a phenotype similar to MM109T. Since a particularly distinctive physiological feature of the strain MM109T was the production of an intracellular green compound, the green pigmentation of the colony was used as a selection criterion for further phylogenetic analysis. The phenotype of over 100 isolates, derived from three moonmilk sampling points—COL1, COL3, COL4 (Fig. 4a), was evaluated on different media after 2 weeks of growth, which resulted in the identification of an additional twelve green-pigmented colonies: two originating from the moonmilk sampling point COL1 (MM91, MM93), eight from the sampling point COL3, like the MM109T isolate (MM15, MM25, MM28, MM29, MM31, MM32, MM37, MM40), and two from the sampling point COL4 (MM113, MM115). In order to assess the phylogenetic position of the green-pigmented isolates and verify if the green pigmentation is a common trait of a monophyletic cluster, MLSA, based on the concatenated sequences of four house-keeping loci, was carried out. This phylogenetic analysis showed that all thirteen green-pigment producing isolates, including the type strain MM109T, represent a monophyletic cluster, clearly separated from their nearest neighbour, S. mauvecolor strain AS 4.1997T (Fig. 4b). Additionally, the MLSA generated exclusively for the thirteen moonmilk isolates, demonstrated that all the green-pigment producers, although highly similar (based on the number of nucleotide substitutions per site), when compared to the closely related Streptomyces, still differ from each other, being classified into eight sub-clusters (Fig. 4c). In order to precisely establish the relationships between those thirteen isolates sequencing of the additional house-keeping genes or of the whole genome would be necessary. Nevertheless, this data strongly supports the isolation and identification of a novel Streptomyces species represented by thirteen isolates, belonging at least to eight sub-clusters, with the type strain MM109T, for which we propose the name Streptomyces lunaelactis sp. nov., as the first fully characterized Streptomyces species isolated from a moonmilk speleothem.
a Map of the cave ‘Grotte des Collemboles’ in Comblain-au-Pont in Belgium, with the spatial distribution of moonmilk sampling sites (COL1 to 5) indicated by black arrows. b, c MLSA analysis of all green-pigmented moonmilk (MM) isolates based on the partial sequences of four house-keeping genes (16S rRNA, recA, rpoB, trpB) showing their phylogenetic position compared with the closest related species and representative Streptomyces type strains, as well as displaying relatedness between moonmilk isolates within the cluster. Sampling sites from which each moonmilk isolate originates are indicated in brackets. Node numbers indicate percent bootstrap support with 1,000 resamplings. Bar number of substitutions per nucleotide position
Ferroverdin A is an intracellular green-pigmented metabolite produced by isolates belonging to Streptomyces lunaelactis sp. nov.
Selective isolation for moonmilk-specific Actinobacteria displaying green pigmentation of vegetative mycelium led to the identification of a group of isolates which shared this particular physiological property and belonged to the same phylogenetic cluster within the genus Streptomyces. In order to identify the metabolite associated with the green pigmentation, an acetone extract from the mycelium of the MM109T isolate was first subjected to TLC to separate the green pigment from the other metabolites of the complex mixture. The TLC-purified green compound was subjected to UPLC-ESI–MS analyses, which revealed a major peak containing one single compound with a molecular ion species [M]− at m/z 860 which could correspond to a coordination compound known as ferroverdin A, composed of three p-vinylphenyl-3-nitroso-4-hydroxybenzoate ligands complexed with a ferrous ion (C45H30N3O12Fe) formerly extracted and characterized from Streptomyces sp. strain WK-5344 (Tabata et al. 1999; Tomoda et al. 1999). Two other representative isolates of S. lunaelactis belonging to different phylogenetic sub-clusters —MM28, and MM37—were grown under conditions stimulating green pigment production (Online Supplementary Fig. 2) and UPLC-ESI–MS analyses of acetone extracts of each isolate also identified a compound with a molecular ion species [M]− at m/z 860 corresponding to ferroverdin A. This was unambiguously confirmed by the presence of typical fragment ions including those at m/z 592 and m/z 268 (see Fig. 5 for MM109T and Online Supplementary Fig. 2 for MM28 and MM37) resulting from the loss of one ligand and two ligands coupled to ferrous iron, respectively (Tabata et al. 1999).
Electrospray ionization (ESI) negative mass spectrum of ferroverdin A identified from the green-pigmented isolate MM109T, the phenotype and morphology of a single colony of which is displayed above the graph. The predominant peak at m/z 860 corresponds to ferroverdin A consisting of three p-vinylphenyl-3-nitroso-4-hydroxybenzoate ligands complexed with a ferrous ion, while fragment ions at m/z 592 and 268 are associated with the loss of one ligand and two ligands coupled to ferrous iron, respectively. Fragment ions at m/z 325 and 562 could not be assigned unambiguously
Description of Streptomyces lunaelactis sp. nov.
Streptomyces lunaelactis sp. nov. (lu.nae.lac’tis. L. gen. n. lunae, of the moon; L. n. lac, lactis, milk; N.L. gen. n. lunaelactis; from moonmilk, referring to the speleothem from which the organism was first isolated).
Gram-positive, aerobic actinobacterium. Grows on all media tested in this study and particularly well on ISP-1, ISP-3, ISP-6, and nutrient agar. Produces white to grey aerial mycelium and variously pigmented substrate mycelium. On ISP-7 medium synthesizes an intracellular green pigment identified as ferroverdin A. Diffusible moderate yellow, greyish yellow and deep orange yellow pigments are produced on ISP-1, ISP-7 and nutrient agar, respectively, as well as dark brown melanin on ISP-6. The substrate mycelium forms various structures from loose to tightly compacted filaments, while the linear to coiled aerial hyphae differentiate into fully septated rod-shaped/cylindrical, smooth-surfaced spores. Grows within the temperature range 10 to 30 °C, and at slightly acidic to weak alkaline pH (pH 6–8), with highest accumulation of biomass at 18–28 °C and pH 6. Tolerates up to 4 % NaCl, growing optimally at 3 % NaCl. The chemotaxonomic features are typical of the genus Streptomyces. The whole-organism hydrolysate contains l-diaminopimelic acid, indicating that the cell wall is of chemotype-I. Octa- and hexahydrogenated menaquinones with nine isoprene units [MK-9(H8), MK-9(H6)] are the major menaquinones, while branched-chain anteiso-C15:0 and iso-C16:0 are the predominant fatty acids. The polar lipids profile corresponds to the type PII phospholipid pattern being mainly composed of phosphatidylethanolamine, followed by phosphoglycolipid, phosphatidylinositol and diphosphatidylglycerol.
The type strain MM109T (=DSM 42149T = BCCM/LMG 28326T) was isolated from a moonmilk speleothem from the cave ‘Grotte des Collemboles’, Comblain-au-Pont, Belgium. The sequences of six house-keeping genes—16S rRNA, atpD, gyrB, recA, rpoB, and trpB—have been deposited in GenBank with the respective accession numbers: KM207217, KM207218, KM207219, KJ862806, KJ862819, and KJ862832.
References
Barton HA, Northup DE (2007) Geomicrobiology in cave environments: past, current and future prospectives. J Cave Karst Stud 69:163–178
Barton HA, Taylor NM, Kreate MP, Springer AC, Oehrle SA, Bertog JL (2007) The impact of host rock geochemistry on bacterial community structure in oligotrophic cave environments. Int J Speleol 36:93–104
Bertram R, Schlicht M, Mahr K, Nothaft H, Saier MH Jr, Titgemeyer F (2004) In silico and transcriptional analysis of carbohydrate uptake systems of Streptomyces coelicolor A3(2). J Bacteriol 186:1362–1373
Busarakam K, Bull AT, Girard G, Labeda DP, van Wezel GP, Goodfellow M (2014) Streptomyces leeuwenhoekii sp. nov., the producer of chaxalactins and chaxamycins, forms a distinct branch in Streptomyces gene trees. Antonie Van Leeuwenhoek 105:849–861
Canaveras JC, Cuezva S, Sanchez-Moral S, Lario J, Laiz L, Gonzalez JM, Saiz-Jimenez C (2006) On the origin of fiber calcite crystals in moonmilk deposits. Naturwissenschaften 93:27–32. doi:10.1007/s00114-005-0052-3
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16:10881–10890
Cuezva S et al (2012) The biogeochemical role of Actinobacteria in Altamira Cave, Spain. FEMS Microbiol Ecol 81:281–290. doi:10.1111/j.1574-6941.2012.01391.x
Davis NK, Chater KF (1990) Spore colour in Streptomyces coelicolor A3(2) involves the developmentally regulated synthesis of a compound biosynthetically related to polyketide antibiotics. Mol Microbiol 4:1679–1691
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. doi:10.1093/nar/gkh340
Felsenstein J (1981) Evolutionary trees from DNA sequences: a Maximum Likelihood Approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gartner A, Blumel M, Wiese J, Imhoff JF (2011) Isolation and characterisation of bacteria from the Eastern Mediterranean deep sea. Antonie Van Leeuwenhoek 100:421–435. doi:10.1007/s10482-011-9599-5
Goodfellow M, Kampfer P, Busse HJ, Trujillo ME, Suzuki K-I, Ludwig W, Whitman WB (2012) The Actinobacteria, Parts A and B vol 5. Bergey’s Manual of Systematic Bacteriology. Springer, New York
Grein A, Spalla C, Di Marco A, Canevazzi G (1963) Descrizione e classificazione di un attinomicete (Streptomyces peucetius sp. nova) produttore di una sostanza attivita antitumorale: La daunomicina. G Microbiol 11:109–118
Guo Y, Zheng W, Rong X, Huang Y (2008) A multilocus phylogeny of the Streptomyces griseus 16S rRNA gene clade: use of multilocus sequence analysis for streptomycete systematics. Int J Syst Evol Microbiol 58:149–159. doi:10.1099/ijs.0.65224-058/1/149
Han JH, Cho MH, Kim SB (2012) Ribosomal and protein coding gene based multigene phylogeny on the family Streptomycetaceae. Syst Appl Microbiol 35:1–6. doi:10.1016/j.syapm.2011.08.007
Hodgson DA (2000) Primary metabolism and its control in streptomycetes: a most unusual group of bacteria. Adv Microb Physiol 42:47–238
Hopwood DA (2007) Streptomyces in nature and medicine—the antibiotic makers. Oxford University Press, New York
Kämpfer P (2012) Genus I. Streptomyces Waksman and Henrici 1943, 339 emend. Witt and Stackebrandt 1990, 370 emend. Wellington, Stackebrandt, Sanders, Wolstrup and Jorgensen 1992, 159. In: Goodfellow M, Kämpfer P, Busse H-J, Trujillo ME, Suzuki K-I, Ludwig W, Whitman WB vol 5. The Actinobacteria, Part B. 2., Bergey’s Manual of Systematic Bacteriology edn. Springer, New York
Kelly KL (1964) Inter-Society Color Council—National Bureau of Standards Color Name Charts Illustrated with Centroid Colors. US Government Printing Office, Washington
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical Streptomyces Genetics. John Innes Foundation, Norwich
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon: a prokaryotic 16S rRNA Gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kroppenstedt RM (1985) Fatty acid and menaquinone analysis of actinomycetes and related organisms. Chem Methods Bact Syst 20:173–199
Kuykendall LD, Roy MA, O’Neill JJ, Devine TE (1988) Fatty acids, antibiotic resistance, and deoxyribonucleic acid homology groups of Bradorhizobium japonicum. Int J Syst Bacteriol 38:358–361
Labeda DP (2011) Multilocus sequence analysis of phytopathogenic species of the genus Streptomyces. Int J Syst Evol Microbiol 61:2525–2531. doi:10.1099/ijs.0.028514-0
Labeda DP, Doroghazi JR, Ju KS, Metcalf WW (2014) Taxonomic evaluation of Streptomyces albus and related species using multilocus sequence analysis and proposals to emend the description of Streptomyces albus and describe Streptomyces pathocidini sp. nov. Int J Syst Evol Microbiol 64:894–900
Lambert S et al (2014) Altered desferrioxamine-mediated iron utilization is a common trait of bald mutants of Streptomyces coelicolor. Metallomics 6:1390–1399
Lechevalier MP, Lechevalier H (1970) Chemical composition as a criterion in the classification of aerobic actinomycetes. Int J Syst Bacteriol 20:435–443
Lechevalier MP, De Bie`vre C, Lechevalier HA (1977) Chemotaxonomy of aerobic actinomycetes: phospholipid composition. Biochem Syst Ecol 5:249–260
Leyh-Bouille M, Bonaly R, Ghuysen JM, Tinelli R, Tipper D (1970) LL-Diaminopimelic acid containing peptidoglycans in walls of Streptomyces sp. and of Clostridium perfringens (type A). Biochemistry 9:2944–2952
Manfio GP, Zakrzewska-Czerwinska J, Atalan E, Goodfellow M (1995) Towards minimal standards for the description of Streptomyces species. Biotekhnologia 7–8:242–253
Miller LT (1982) Single derivatization method for routine analysis of bacterial whole-cell fatty acid methyl esters, including hydroxy acids. J Clin Microbiol 16:584–586
Murase M, Hikiji T, Nitta K, Okami Y, Takeuchi T, Umezawa H (1961) Peptimycin, a product of Streptomyces exhibiting apparentinhibition against Ehrlich carcinoma. J Antibiot Ser A 14:113–118
Noens EE, Mersinias V, Traag BA, Smith CP, Koerten HK, van Wezel GP (2005) SsgA-like proteins determine the fate of peptidoglycan during sporulation of Streptomyces coelicolor. Mol Microbiol 58:929–944
Onstott TC et al (2009) Microbial communities in subpermafrost saline fracture water at the Lupin Au mine, Nunavut, Canada. Microb Ecol 58:786–807. doi:10.1007/s00248-009-9553-5
Ortiz M, Legatzki A, Neilson JW, Fryslie B, Nelson WM, Wing RA (2014) Making a living while starving in the dark: metagenomic insights into the energy dynamics of a carbonate cave. ISME J 8:478–491
Porca E, Jurado V, Žgur-Bertok D, Saiz-Jimenez C, Pašić L (2012) Comparative analysis of yellow microbial communities growing on the walls of geographically distinct caves indicates a common core of microorganisms involved in their formation. FEMS Microbiol Ecol 81:255–266
Portillo MC, Saiz-Jimenez C, Gonzalez JM (2009) Molecular characterization of total and metabolically active bacterial communities of ‘white colonizations’ in the Altamira Cave, Spain. Res Microbiol 160:41–47
Rong X, Huang Y (2010) Taxonomic evaluation of the Streptomyces griseus clade using multilocus sequence analysis and DNA–DNA hybridization, with proposal to combine 29 species and three subspecies as 11 genomic species. Int J Syst Evol Microbiol 60:696–703
Rong X, Huang Y (2012) Taxonomic evaluation of the Streptomyces hygroscopicus clade using multilocus sequence analysis and DNA–DNA hybridization, validating the MLSA scheme for systematics of the whole genus. Elsevier 35:7–18
Rong X, Guo Y, Huang Y (2009) Proposal to reclassify the Streptomyces albidoflavus clade on the basis of multilocus sequence analysis and DNA–DNA hybridization, and taxonomic elucidation of Streptomyces griseussubsp.solvifaciens. Syst Appl Microbiol 32:314–322
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol 28:226–231
Tabata N, Tomoda H, Omura S (1999) Ferroverdins, inhibitors of cholesteryl ester transfer protein produced by Streptomyces sp. WK-5344, II. Structure elucidation. J Antibiot 52:1108–1113
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30:2725–2729
Tenconi E, Jourdan S, Motte P, Virolle MJ, Rigali S (2012) Extracellular sugar phosphates are assimilated by Streptomyces in a PhoP-dependent manner. Antonie Van Leeuwenhoek 102:425–433
Tetu SG, Breakwell K, Elbourne LD, Holmes AJ, Gillings MR, Paulsen IT (2013) Life in the dark: metagenomic evidence that a microbial slime community is driven by inorganic nitrogen metabolism. The ISME journal 7:1227–1236
Tindall BJ (1990a) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130
Tindall BJ (1990b) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Letts 66:199–202
Tindall BJ, Sikorski J, Smibert RM, Kreig NR (2007) Phenotypic characterization and the principles of comparative systematics. In methods for general and molecular microbiology, 3rd edn. ASM Press, Washington, pp 330–393
Tomoda H, Tabata N, Shinose M, Takahashi Y, Woodruff HB, Omura S (1999) Ferroverdins, inhibitors of cholesteryl ester transfer protein produced by Streptomyces sp. WK-5344. I. Production, isolation and biological properties. J Antibiot 52:1101–1107
Waksman SA (1961) The actinomycetes, classification, identification and description of genera and species, vol 2. Williams & Wilkins, Baltimore
Waksman SA (1967) The Actinomycetes. Ronald Press, A Summary of Current Knowledge. New York
Xu P et al (2005) Naxibacter alkalitolerans gen. nov., sp nov., a novel member of the family ‘Oxalobacteraceae’ isolated from China. Int J Syst Evol Microbiol 55:1149–1153
Zlatkin IV, Schneider M, de Bruijn FJ, Forney LJ (1996) Diversity among bacteria isolated from the deep subsurface. J Ind Microbiol 17:219–227
Acknowledgments
MM, and ISP work is supported by a Research Foundation for Industry and Agriculture (FRIA) grant. SR is a research associate at Belgian Funds for Scientific Research (F.R.S-FNRS). AAA work is supported by a F.R.S-FNRS grant (R.FNRS.3342). This work is supported in part by the Belgian program of Interuniversity Attraction Poles initiated by the Federal Office for Scientific Technical and Cultural Affairs (PAI no. P7/44). We are thankful to Prof Denis Baurain (ULg) and Maxime Licops for useful advices for our phylogeny analyses and technical assistance, respectively.
Conflict of Interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Maciejewska, M., Pessi, I.S., Arguelles-Arias, A. et al. Streptomyces lunaelactis sp. nov., a novel ferroverdin A-producing Streptomyces species isolated from a moonmilk speleothem. Antonie van Leeuwenhoek 107, 519–531 (2015). https://doi.org/10.1007/s10482-014-0348-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0348-4