Abstract
Vascular endothelial growth factors (VEGFs) regulate blood and lymph vessel development upon activation of three receptor tyrosine kinases (VEGFRs). The extracellular domain of VEGFRs consists of seven Ig-homology domains, of which D2–3 form the ligand-binding site, while the membrane proximal domains D4–7 are involved in homotypic interactions in ligand-bound receptor dimers. Based on low-resolution structures, we identified allosteric sites in D4–5 and D7 of vascular endothelial growth factor receptor 2 (VEGFR-2) accomplishing regulatory functions. Allosteric inhibition of VEGFR-2 signaling represents an attractive option for the treatment of neovascular diseases. We showed earlier that DARPin® binders to domains D4 or D7 are potent VEGFR-2 inhibitors. Here we investigated in detail the allosteric inhibition mechanism of the domain D4 binding inhibitor D4b. The 2.38 Å crystal structure of D4b in complex with VEGFR-2 D4–5, the first high-resolution structure of this VEGFR-2 segment, indicates steric hindrance by D4b as the mechanism of inhibition of receptor activation. At the cellular level, D4b triggered quantitative internalization of VEGFR-2 in the absence of ligand and thus clearance of VEGFR-2 from the surface of endothelial cells. The allosteric VEGFR-2 inhibition was sufficiently strong to efficiently inhibit the growth of human endothelial cells at suboptimal dose in a mouse xenograft model in vivo, underlining the therapeutic potential of the approach.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The formation of functional blood and lymphatic vessels is a keystone during embryo development and is essential for supplying all organs with oxygen and nutrients and for disposal of catabolites. Vascular endothelial growth factors (VEGFs) activate the type V subfamily of receptor tyrosine kinases (RTKs) in the human kinome representing the main drivers of vasculogenesis, the de novo formation of vessels, and angiogenesis, the formation of new vessels from preexisting vasculature [1,2,3]. Distinct VEGF subfamily members activate either VEGFR-1, -2 or -3, or combinations thereof [4]. Mutation or ablation of single VEGFRs gave rise to highly specific disease profiles documenting signal diversity among the three receptors [2]. Ablation of VEGFR-1 was embryonic lethal due to severe disorganization of blood vessels [5], while VEGFR-2 knockout led to the complete absence of endothelial cells and early embryonal death [6]. In addition, VEGFR-3 was shown to be indispensable in early embryonic vessel development and in lymphangiogenesis [7]. Similar to other RTKs, VEGFR activation is initiated by ligand-mediated receptor dimerization, thereby instigating transmembrane signaling and activation of the intracellular kinase domain [8].
Low-resolution structures of a VEGFR-2 ECD/ligand complex suggested that domains D2–3 of the seven immunoglobulin-homology domains (Ig-domains) comprising the receptor extracellular domain, ECD, form the ligand-binding site, while domains D4–7 are involved in homotypic receptor contacts, presumably regulating the exact orientation of the membrane proximal receptor domains and thus kinase activation [9]. X-ray structures confirmed this and established the details of ligand binding to Ig-domains D2–3 of VEGFR-2 and D1–2 of VEGFR-3, respectively [10,11,12]. We recently published a composite model for the entire ECD encompassing D1–7 of VEGFR-1 in complex with VEGF-A based on X-ray crystallographic data [13]. This structure agreed with earlier data for D2 defining this domain as the major binding site for VEGF family ligands [14,15,16] and documented the additional role of D3 and the interdomain linker between D2 and D3 in ligand binding. Taken together, these structures show how the membrane distal Ig-domains mediate ligand binding with VEGF, while the membrane proximal domains are involved in homotypic contacts presumably required for proper receptor orientation in active dimers.
Classical anti-angiogenic therapy relies on several approaches; the most prominent example applied in the clinic is sequestration of VEGF using antibodies [17, 18]. The VEGF-A-specific antibody bevacizumab applied in anti-angiogenic cancer therapy, for example, neutralizes all VEGF-A isoforms and thus blocks VEGF receptor activation. As a drawback systemic destruction of the normal vasculature was observed [19, 20]. On the other hand, domain D2–3-specific antibodies inhibiting VEGF binding to VEGFR-2 must be administered in large amounts to successfully compete with VEGF for blocking receptor activity [21]. In addition, kinase inhibitors such as sunitinib acting intracellularly block VEGFR kinase activation and signaling. The lack of receptor specificity of such kinase inhibitors is known to lead to a suboptimal safety profile [22].
Based on our discovery of novel allosteric regulatory sites in the VEGFR-2 ECD, we previously set out to develop a new class of highly specific designed ankyrin repeat protein inhibitors (DARPins). We isolated DARPin® domains interacting with either D2–3, D4 or D7 [23]. DARPin® domains interacting with D2–3 inhibited ligand binding, receptor dimerization, and receptor kinase activation similar to bevacizumab. DARPin® domains specific for D4 or D7, on the other hand, did not prevent ligand-mediated receptor dimerization, nevertheless efficiently blocked functional receptor output.
Here we analyzed the role of VEGFR-2 D4 as a drug-targetable site in more detail. We determined the crystal structure of DARPin® domain D4b in complex with the VEGFR-2 ECD enabling the in-detail analysis of the mechanism of allosteric inhibition. Based on cellular analyses, we propose a possible mechanism for DARPin® domain-mediated receptor inhibition. Finally, we show the efficacy of a pharmacokinetically engineered variant of D4b, HD4b, in inhibiting human umbilical vein endothelial (HUVE) cell growth in a mouse xenograft angiogenesis system.
Methods
Cloning of the VEGFR-2 D4–5 construct
The gene sequence encoding the extracellular domain D4–5 fragment of the human VEGFR-2 (residues 329–549) with an N-terminal SPARC secretion signal sequence and a C-terminal 6× histidine tag was cloned into the pCepPu vector for constitutive expression in mammalian cells.
VEGFR-2 protein production and purification
For expression of VEGFR-2 D4–5, the pCepPu vector was transfected into HEK293 EBNA cells using polyethylenimine, PEI. Cells were maintained in adherent cultures with DMEM (BioConcept, Basel, Switzerland) containing 10% fetal bovine serum (FBS) and 25 µg/mL G418 in an incubator at 37 °C and 5% CO2. Stable cell lines were generated by the addition of 1 µg/mL puromycin to select for cells containing the pCepPu vector, followed by clonal selection and expansion. For scale-up production of VEGFR-2 D4–5, adherent cultures were expanded and transferred to Erlenmeyer shaking flasks with protein expression media (PEM, Thermo Fischer Scientific), 1% FBS, 1× GlutaMax, 25 µg/mL G418, 1 µg/mL puromycin), at an initial cell density of approximately 0.5 million cells/mL. Suspension cultures were grown at 37 °C, 5% CO2, with agitation at 100 rpm. The culture was scaled-up keeping the cell density below 3 million cells/mL. Puromycin and G418 were not used past the initial 70 mL starter suspension culture. Suspension cultures were expanded until an approximately 2 L volume was reached. The culture was then centrifuged in sterile 1 L centrifuge bottles using a fixed-angle rotor (1500×g, 5 min), and the cells resuspened in two 1 L volumes of expression media (PEM, 1× GlutaMax, 0.5 µg/mL kifunensine, 5 mM sodium butyrate, 1% penicillin and streptomycin). The culture was allowed to incubate for a further 6 days before harvesting the secreted protein by centrifuging the culture to remove the cells (4000×g, 15 min) and storing the supernatant at − 20 °C. 2 L of protein-containing culture supernatant was passed through a 0.45-µm filter, concentrated with a 3 kDa Mr tangential-flow concentrator to an approximate volume of 500 mL. The sample was buffer exchanged twice with 500 mL dilution buffer (50 mM HEPES pH 8.0, 300 mM NaCl), concentrated to 250 mL and loaded on a 5 mL HisTrap HP column. After washing with 50 mM HEPES pH 8.0, 300 mM NaCl, 20 mM imidazole, the protein was step eluted with 40% elution buffer (50 mM HEPES pH 7.5, 300 mM NaCl, 400 mM imidazole) and concentrated to 5 mL using a 10 kDa Mr centrifugal concentrator. The sample was filtered with a 0.45-µm syringe filter and passed over a Superdex 75 16/60 column in 50 mM HEPES pH 7.5, 150 mM NaCl. Peak fractions were concentrated to 10 mg/mL.
DARPin® domain production
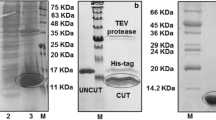
DARPin® domain D4b [23] and the construct HD4b, consisting of D4b additionally carrying a serum albumin-binding DARPin® domain at its aminoterminus to improve pharmacokinetic properties, were expressed in E. coli BL21 and purified to homogeneity using IMAC and size-exclusion chromatography as described previously [24].
Crystallization, data collection, and structure determination
The purified VEGFR-2 domain D4–5 protein was mixed in a 1:1 molar ratio with DARPin® domain D4b to achieve a final protein concentration of 7.7 mg/mL. Crystallization trials were set up in 24-well hanging-drop plates, with three drop ratios per reservoir (1:1, 1:2, and 2:1 protein:reservoir volume). Successful crystal growth was achieved with a reservoir solution of 0.2 M calcium acetate monohydrate, 15% PEG 4000, 0.1 M sodium cacodylate pH 6.4, and streak-seeding drops using a mixture of initial crystal hits. Crystals were grown at 20 °C and harvested after 2 weeks, cryoprotected by quick soaking in 15% trehalose, and flash-frozen in liquid nitrogen.
Diffraction data were collected at the Swiss Light Source beamline X06SA (PXI). A complete dataset was recorded from a single crystal at 100 K. Data were measured with a DECTRIS EIGER X 16 M detector in a 360° sweep at 0.1° per image. Diffraction images were indexed, integrated and scaled using Xia2 [25], DIALS [26], and Aimless [27, 28], respectively. Correct space group determination and re-indexing was accomplished using Pointless [27]. The structure was solved by molecular replacement using Phaser [29] as implemented in Phenix [30] with search models PDB ID 4BSJ (VEGFR-3 extracellular domain D4–5 structure [10] and PDB ID 2QYJ consensus DARPin® domain structure [31]). The resulting starting model contained two copies of the VEGFR-2 domain D4–5 and two copies of the DARPin® domain D4b. Phases were improved by iterative manual rebuilding and refinement using Coot [32] and phenix.refine [30], with non-crystallographic symmetry restraints across the two VEGFR-2 domain D4–5 chains and across the two DARPin® domain D4b chains. TLS grouping was used to refine anisotropic B-factor parameters. See Table 1 for data collection and refinement statistics.
Receptor expression and internalization in cell culture
Cell culture conditions
Porcine aortic endothelial cells expressing VEGFR-2 (PAE-KDR) were grown in DMEM supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. Cells were propagated in a humidified atmosphere at 37 °C with 5% CO2.
Immunofluorescence microscopy
Cells were grown on glass coverslips coated with poly-l-lysine (P4707, Sigma) to 60% confluency. Cells were treated either with 1 µM D4b or 1.5 nM VEGF-A for 10 min in starvation media (DMEM supplemented with 0.1% FBS). Cells were fixed with 3.7% formaldehyde in phosphate-buffered saline (PBS) for 20 min at 37 °C, permeabilized for 10 min with 0.1% NP-40 in PBS, and blocked for 20 min in 5% FBS in PBS at room temperature. Samples were exposed to primary VEGFR-2-specific antibody 55B11 (Cell Signaling Technology, BioConcept, Allschwil, Switzerland, diluted 1:1000) and fluorescently labeled secondary antibodies in PBS containing 5% bovine albumin and embedded. Images were acquired with a Leica SP5 laser scanning confocal microscope.
Squassh analysis of VEGFR-2 internalization
Analysis of 25 images per condition was performed using the ImageJ plugin Squassh [33]. Squassh combines segmentation and deconvolution of images in a single step, yielding better results for small objects close to the diffraction limit of the microscope. Squassh was used to segment intracellular vesicles and cells. Squassh Analyst was used for data analysis and normalization of the VEGFR-2 positive vesicle area relative to the total cell area.
Trypsin digestion of cell surface exposed receptors
Protection of VEGFR-2 from extracellular trypsin treatment was exploited in the quantification of the internalized VEGFR-2. Following incubation with D4b or VEGF-A in starvation media, PAE-KDR cells were washed three times with ice-cold PBS, then incubated on ice with freshly prepared trypsin (1 mg/mL, Sigma) for 30 min. The enzymatic reaction was quenched by the addition of soybean trypsin inhibitor (50 mg/mL, Sigma). Cells were scraped off the plate and centrifuged at 500 rpm at 4 °C for 5 min. The cell pellet was lysed in gel electrophoresis sample buffer, heated to 95 °C for 5 min, and analyzed on Western blots using VEGFR-2 antibody 55B11(Cell Signaling Technology) as described by Hyde et al. [23].
Determination of VEGFR-2 activity
Cells were treated with D4b, VEGF-A or both and processed for Western analysis as described by Hyde et al. [23]. Phosphorylation level of VEGFR-2 was determined using the p1175-specific antibody 19A10 (Cell Signaling Technology); total receptor level was determined with VEGFR-2 antibody 55B11.
Statistical analysis
For statistical analysis of experimental data, multiple comparisons were investigated for significant differences by 1- and 2-way ANOVA. Individual comparisons were subsequently performed by using the unpaired Student’s t test. Data are expressed as mean ± SD, where n is the number of individual experiments; p values ≤ 0.05 were considered statistically significant in two-sided tests. Numerical analyses were performed in GraphPad Prism version 7.0 (GraphPad Software, USA).
In vivo HUVE cell angiogenesis model
An in vivo angiogenesis model described earlier [34] was performed at ProQinase GmbH (Freiburg, Germany) as follows. HUVE cells passage 3–4 (PromoCell, Heidelberg, Germany) were cultured in ECBM/ECGM media and transduced by a lentivirus encoding a luciferase–neomycin fusion protein (ECsLuc+). HUVE cell spheroids (ECsLuc+) were prepared as described in Lichtenbeld et al. [35] by pipetting ECs to a dish coated with 0.5% agarose. The following day the EC spheroids were harvested and mixed in a Matrigel/fibrin solution to reach a final number of 300,000 ECs as spheroids per injected plug. VEGF-A and FGF-2 were added at a final concentration of 1000 ng/mL.
SCID mice were subcutaneously injected into the right flank with 500 µL of the cell/matrix suspension that quickly solidified. The study consisted of three study arms containing 10 female SCID mice each. At day 0, ECsLuc+ spheroids were injected subcutaneously in a Matrigel/fibrin matrix on the right flank of the mice. ECsLuc+ spheroids were additionally injected on the left flank of four mice of each group. The first perfused vessels were detectable between day 4 and day 6. After 20 days of in vivo growth a well-established vasculature with around 50–60% pericyte-covered and perfused vessels was established [36]. Group 1 was treated with vehicle control, group 2 with sunitinib (40 mg/kg), and group 3 with HD4b (1 mg/kg).
The development of the ECsLuc+ vasculature was investigated weekly by bioluminescence imaging (BLI signal). BLI signal at day 1 was used to randomize the animals. Animals were randomized into 3 groups of 10 animals each according to the BLI signal. Randomization was performed with 12 animals that received two Matrigel/fibrin matrix plugs and 18 animals that received a single Matrigel/fibrin matrix plug. On the same day, treatment with Sunitinib or DARPin® candidate HD4b or PBS was started. At study end (day 21) the right plugs containing ECsLuc+ were removed, homogenized and analyzed for ex vivo luciferase activity.
The four mice containing two ECsLuc+ plugs were injected prior to necropsy with 100 µL of Ulex europaeus agglutinin-I:biotin (UEA-I; 1 mg/mL; Vector laboratories, Peterborough, UK, B-1065, lot #ZC0331) into the tail vein of each mouse for 10 min before being anaesthetized by isoflurane and euthanization by cervical dislocation. The Matrigel plugs from the right flank were collected in luciferase assay buffer for ex vivo luciferase activity measurement. The plugs from the left flank were fixed in 4% formalin at room temperature for 8–12 h and paraffin embedded using a Leica TP1020 processor. The ECsLuc+ Matrigel plugs were collected, homogenized, and assayed for luciferase activity using a Promega kit (E1501).
Immunohistochemistry of Matrigel plugs
For histological examination of the human vasculature in the Matrigel plug paraffin sections (thickness 8–10 µm) were prepared. Blood vessel formation and coverage by pericytes was detected by staining for human CD34 (green fluorescence FITC, NCL-END, Menarini, Berlin, Germany) and anti-smooth muscle actin (SMA, red fluorescence Cy3; clone 1A4, Sigma, Taufkirchen, Germany). Three sections per plug were analyzed, and three images were taken from each section at a magnification of 200× using an Eclipse TE2000-U microscope (Nikon, Kanagawa, Japan). The area analyzed per section was 0.44 mm2. The vessel number (CD34 positive) and coverage (SMA positive) was manually determined using the NIS-elements basic research software (Nikon, Kanagawa, Japan). Irrespective of size, each green fluorescent spot was counted as a vessel. Each branching was counted separately and larger vessels received up to 3 counts. The marked vessels were inspected for red fluorescence and counted irrespective of strength of appearance.
Results
Structure of the DARPin® domain D4b/VEGFR-2 D4–5 ECD complex
We crystallized a complex of recombinantly produced D4–5 protein derived from the VEGFR-2 ECD bound to DARPin® domain D4b. The proteins were produced and purified separately by affinity and size-exclusion chromatography, followed by mixing of the two components prior to setting up crystallization screens. Crystals of the complex were obtained by streak seeding from initial hits into grid screens set in hanging-drop format. Single crystals were obtained with rhomboid morphology measuring approximately 150–200 µm in length and 20–30 µm in thickness. A complete crystallographic dataset was measured to 2.38 Å resolution, enabling structure determination by molecular replacement and refinement to final Rwork/Rfree values of 0.178/0.229 (Table 1; PDB ID: 5OYJ).
The crystal structure of the DARPin® domain D4b/VEGFR-2 D4–5 complex shows that this DARPin® domain specifically binds to D4 of the receptor. The total interface area between D4b and VEGFR-2 D4 is approximately 1200 Å2. The complex is stabilized by a number of hydrogen bonds and salt bridges between VEGFR-2 D4 and DARPin® domain D4b, including in particular salt bridges between R347 and D77 and R349 and E89, respectively (Fig. 1a, Table S1). Interacting residues in the DARPin® domain D4b are located in the beta-turn as well as the alpha helices of the ankyrin repeats. For the full list of interfacing residues see supplementary Table S1. In order to provide a structural basis for the inhibitory effect of D4b, we superimposed the D4b/VEGFR-2 D4–5 structure onto the published VEGFR-1 ECD/VEGF-A composite model [13] as shown in Fig. 1b. The superposition shows that D4b clashes with the adjacent monomer in receptor ECD dimers and thus blocks close apposition of two receptor chains in the membrane proximal part of the dimeric receptor. This finding establishes that inhibition of VEGFR-2 signaling by DARPin® domain D4b results from steric hindrance of receptor alignment in domain D4, thereby preventing transmembrane signaling.
Structure of VEGFR-2/DARPin® domain complex. a Structural model of VEGFR-2 D4–5 in complex with DARPin® domain D4b. Left, representation of the full structure, right, details showing the interacting interface between D4–5 and D4b. b Superposition of model shown in a with VEGFR-1 ECD/ligand complex structural model [13], left ‘side view’, right ‘top view’ of structural model
VEGF-A and DARPin® domain D4b promote internalization of VEGFR-2
Based on work published on other receptor tyrosine kinases such as HER2 and the EGF receptor, we studied internalization of VEGFR-2 in the presence and absence of VEGF-A or DARPin® domain D4b. We used two approaches to determine receptor internalization. First, we followed receptor uptake into intracellular vesicles in immunostained VEGFR-2-expressing cells and monitored VEGFR-2 positive vesicles area after addition of either VEGF-A or D4b or both (Fig. 2a). Incubation with VEGF-A or D4b led to a significant increase in the number of internalized vesicles. The same effect was observed when VEGF-A and D4b were added simultaneously. To quantify vesicle accumulation in cells we performed a Squassh analysis [33] and analyzed 25 individual microscopic images per condition (Fig. 2b). The data show that VEGFR-2 accumulation in intracellular vesicles is significantly increased upon treatment with VEGF-A or D4b or both.
DARPin® domain D4b promotes internalization of VEGFR-2. VEGFR-2 internalization. a Immunostaining of VEGFR-2 receptors expressed in PAE-KDR cells showing receptor internalization following VEGF-A or DARPin® domain D4b administration. PAE-KDR cells were fixed and stained 10 min after addition of VEGF-A, D4b or VEGF-A together with D4b. b Area of VEGFR-2 positive vesicles relative to total cell area was analyzed by Squassh. Statistical data analysis was performed using GraphPad Prism 7. Data show a representative experiment of 25 independent cell samples. Error bars represent ± SD. The statistical significance based on a Student’s t test is indicated by * representing p < 0.05. c Top gel shows Western analysis of control and trypsin-treated cells using VEGFR-2-specific antibody. Cells were treated with VEGF-A or D4b for 10 min. Top two arrows on the left of the blot indicate bands representing undegraded receptor, bottom arrow points to band of degraded receptor. Lower gel shows kinase activity of VEGFR-2 determined with phospho-tyrosine-specific antibody in PAE-KDR cells. Shown are control, VEGF-A, D4b, or VEGF-A plus D4b treated cells. Bottom row shows total level of VEGFR-2 of the same gel
To further confirm that VEGFR-2 is cleared from the cell surface upon VEGF-A or D4b binding, a method that distinguishes between internalized and cell surface exposed protein was used. Intact cells were treated with trypsin resulting in digestion of membrane-bound extracellular VEGFR-2, while internalized protein was protected and remained intact. VEGFR-2 protein was examined by SDS–PAGE and Western blot analysis (Fig. 2c upper gel). In untreated control cells, only full-length VEGFR-2 was observed (lane 1), while trypsin treatment led to the degradation of the receptor in control cells as indicated by the two lower bands (Fig. 2c upper gel, lane 2). In the presence of VEGF-A or DARPin® domain D4b, on the other hand, VEGFR-2 was protected from trypsin degradation and predominantly the bands corresponding to intact VEGFR-2 were observed (Fig. 2c, upper gel, lanes 3 and 4).
DARPin® domain D4b, although stimulating receptor internalization and degradation, did not induce kinase activation of VEGFR-2 (Fig. 2c lower gel, see also Hyde et al. [23]).
Efficacy of vessel growth inhibition by DARPin® domain HD4b in an in vivo angiogenesis model
A pharmacokinetically optimized variant of D4b, called HD4b, was engineered for in vivo application. To test the efficacy of DARPin® domain HD4b in vivo, we used an angiogenesis model in xenograft mice. In vivo bioluminescence imaging of implanted luciferase transfected HUVE cells demonstrated that both group 3 animals, treated with HD4b (1 mg/kg), and group 2 animals, treated with the reference compound sunitinib (40 mg/kg), showed prominent and significant luciferase signal reduction starting on day 8 until study end at day 21 (Fig. 3a, for details see also Fig. S1). The ex vivo luciferase assay confirmed the result of in vivo bioluminescence imaging. The HD4b-treated group showed a significant reduction (p = 0.0015) and the sunitinib reference control group (p = 0.0006) a highly significant reduction in luciferase signal compared to the vehicle control group (Fig. 3b).
DARPin® domain HD4b-mediated inhibition of endothelial cell growth in xenograft mice. a In vivo luciferase activity (photons/s). Light emission was measured 10 min post-injection of 150 mg/kg D-luciferin with a CCD-camera for 8 min using a NightOWL LB 981 bioluminescence imaging system. Light emission overlay images of day 1, 8, 15, and 20 displaying all animals in the groups as indicated in the legend. Individual primary animal data are depicted in Figure S1. b The plugs with the ECsLuc+ were collected during necropsy and homogenized. The luciferase activity of the homogenates was measured to determine the content of luciferase positive endothelial cells. c New vessel formation within matrix plugs was analyzed in three sections per matrix plug for human CD34 staining. Mean vessel number of each plug is shown in dots and the median value from each group as a horizontal bar. The animals in the different groups were treated according to the legend
Determining the human vessel number in a subgroup of four animals implanted with Matrigel plugs revealed an apparent reduction in mean vessel number of both the HD4b and sunitinib-treated animals compared to the vehicle control group (Fig. 3c). The low vessel number in groups 2 and 3 did not allow detailed quantitative analysis of the coverage of the human vessels with mural cells (data not shown).
Discussion
Structure of DARPin® domain D4b/VEGFR-2 complex
To unravel the mechanism of DARPin® domain D4b inhibition of angiogenesis, we solved the crystal structure of a DARPin® domain D4b/VEGFR-2 D4–5 complex. Assuming that the dimeric arrangement of the VEGFR-2 ECD in complex with VEGF-A corresponds to the VEGFR-1 ECD/VEGF-A structure published earlier, we propose a structural model of DARPin® domain D4b bound to the full-length VEGFR-2 dimer (Fig. 1b) [13]. According to the model DARPin® domain D4b binding results in steric hindrance preventing close association of two receptor monomers in the membrane proximal part of the receptor ECD. This model is in line with the previous notion that the homotypic contacts in VEGFR-2 are essential for receptor activation and downstream signaling [23]. The structural model also agrees with our earlier published data where we identified conserved residues in homotypic contacts in D5 of VEGFR-2 that, upon mutation, reduced ligand-induced receptor phosphorylation [13].
An additional key value of the DARPin® domain D4b VEGFR-2 D4–5 structure is the high-resolution structural information now available for D4 and D5 of VEGFR-2. This leaves only the structural details of D1 and D6 of VEGFR-2 unknown.
Our structure shows D4 in the monomeric state where the flexible A–A′ strand has a different conformation than in dimeric VEGFR-1 [13] and VEGFR-3 [10], with high B-factors illustrating flexibility of this region. The B-factors of the D4 structure in our model are, however, low and thus differ from D4 of VEGFR-3 (PDB ID: 4bsj) where they were observed to be high due to the lack of disulfide bridging between the beta-sheets. It is likely that binding of D4b further stabilizes D4 and makes it more rigid and therefore unable to trigger the intertwining of the membrane proximal domains of receptor monomers required for receptor activation. DARPin® domain D4b binding to D4 may therefore block transmission of signaling in the receptor ECD from the ligand-binding domain toward the membrane by interfering with the alignment of receptor monomers. This would be in agreement with the earlier finding that close proximity of the D7 domains is required for transmembrane signaling to the intracellular kinase domain [37].
Taken together, our data provide a structural rationale for allosteric inhibition of VEGFR-2 by DARPin® domain D4b. D4b prevents receptor dimers from adopting the active conformation obligatory for VEGF signaling by preventing the correct orientation of D4–7 in ligand-bound receptor dimers.
Mechanism of VEGFR-2 inhibition
Our internalization studies performed on cultured endothelial cells revealed a possible cellular mechanism of D4b-mediated receptor inactivation. D4b binding to VEGFR-2 led to receptor internalization as demonstrated by an increase in the total area of intracellular receptor-positive vesicles and the clearance of the receptor from the plasma membrane (Fig. 2). In agreement with our earlier publication by Hyde et al. [23], internalization stimulated by D4b was not accompanied by kinase activation (Fig. 2c, lower gel). Whether the receptor is internalized as a dimer or monomer remains at this point unclear; however, based on our earlier work showing that VEGFR-2 exists in live cells in an equilibrium between monomer and dimer even in the absence of ligand, D4b might downregulate both monomeric and dimeric receptor [38]. D4b apparently shares the mechanism of downregulation with antibodies specific for HER2 or EGFR [39,40,41]. Drebin et al. [42] and Hudziak et al. [43] have demonstrated that receptor-specific antibodies reduced the amount of HER2 expressed on the surface of cancer cells. Analogous to therapeutic antibodies specific for HER2 [44, 45], the allosteric properties of D4b might provoke structural changes in the receptor ECD resulting in non-productive VEGFR-2 internalization and clearing from the plasma membrane. Recently, receptor clustering followed by unspecific internalization was also demonstrated for another HER2-specific antibody Pertuzumab [46].
In vivo angiogenesis inhibition in mice
Our earlier study using several cell culture systems showed that DARPin® domain D4b specific for subdomain D4 of the ECD blocked signaling by VEGFR-2 [23]. Interestingly, this DARPin® domain did not block ligand binding or receptor dimerization and thus behaved like an allosteric inhibitor of VEGFR-2.
Here, we set out to confirm the inhibitory activity of DARPin® domain D4b in vivo in an animal angiogenesis model. D4b is specific for human VEGFR-2 and could not directly be tested in mouse angiogenesis models. We therefore investigated the inhibitory effect of the pharmacokinetically engineered DARPin® domain HD4b using human endothelial cells implanted in Matrigel into female SCID mice. All animals implanted with Matrigel plugs were healthy throughout the study with no weight loss. In the absence of inhibitor, the implanted endothelial cells divided and anostomosed with the mouse vasculature upon VEGF-A exposure resulting in a well-vascularized Matrigel plug. The dosing of HD4b was about 3000-fold lower on a molar basis than for suntinib and has not been optimized for maximal efficacy in our study. Using this protocol DARPin® domain HD4b inhibited the growth of implanted endothelial cells as efficiently as the classical VEGFR-2 receptor kinase inhibitor sunitinib. DARPin® domain HD4b and sunitinib also blocked human endothelial cells from connecting to the mouse vasculature, preventing the formation of contiguous vascular structures.
The data presented here show that systemically applied DARPin® domain HD4b, a variant of D4b which was engineered for extended half-life in circulation [24], inhibits angiogenesis in vivo to a similar extent as the clinically established VEGFR-2 inhibitor sunitinib. With its allosteric inhibition mode and high VEGFR-2 selectivity, DARPin® domain HD4b promises to be an attractive candidate for anti-angiogenic therapy. Together with published work using similar reagents called Affimers [47] our data document the usefulness of novel alternative binding scaffolds for constructing so-called molecular recognition reagents. These reagents can be produced in large quantities and chemically modified in multiple ways to generate complementary affinity reagents for molecular and cell biology applications and as targeting devices for therapeutic applications in vivo.
References
Smith GA, Fearnley GW, Harrison MA, Tomlinson DC, Wheatcroft SB, Ponnambalam S (2015) Vascular endothelial growth factors: multitasking functionality in metabolism, health and disease. J Inherit Metab Dis 38(4):753–763
Shibuya M (2014) VEGF-VEGFR Signals in Health and Disease. Biomol Ther 22(1):1–9
Moens S, Goveia J, Stapor PC, Cantelmo AR, Carmeliet P (2014) The multifaceted activity of VEGF in angiogenesis—implications for therapy responses. Cytokine Growth Factor Rev 25(4):473–482
Shibuya M (2013) VEGFR and type-V RTK activation and signaling. Cold Spring Harb Perspect Biol 5(10):a009092
Fong GH, Rossant J, Gertsenstein M, Breitman ML (1995) Role of the Flt-1 receptor tyrosine kinase in regulating the assembly of vascular endothelium. Nature 376(6535):66–70
Shalaby F, Rossant J, Yamaguchi TP, Gertsenstein M, Wu XF, Breitman ML, Schuh AC (1995) Failure of blood-island formation and vasculogenesis in Flk-1-deficient mice. Nature 376(6535):62–66
Veikkola T, Jussila L, Makinen T, Karpanen T, Jeltsch M, Petrova TV, Kubo H, Thurston G, McDonald DM, Achen MG, Stacker SA, Alitalo K (2001) Signalling via vascular endothelial growth factor receptor-3 is sufficient for lymphangiogenesis in transgenic mice. EMBO J 20:1223–1231
Lemmon MA, Schlessinger J (2010) Cell signaling by receptor tyrosine kinases. Cell 141(7):1117–1134
Ruch C, Skiniotis G, Steinmetz MO, Walz T, Ballmer-Hofer K (2007) Structure of a VEGF-VEGF receptor complex determined by electron microscopy. Nat Struct Mol Biol 14(3):249–250
Leppänen VM, Tvorogov D, Kisko K, Prota AE, Jeltsch M, Anisimov A, Markovic-Mueller S, Stuttfeld E, Goldie KN, Ballmer-Hofer K, Alitalo K (2013) Structural and mechanistic insights into VEGF receptor 3 ligand binding and activation. Proc Natl Acad Sci USA 110(32):12960–12965
Brozzo MS, Bjelic S, Kisko K, Schleier T, Leppänen VM, Alitalo K, Winkler FK, Ballmer-Hofer K (2012) Thermodynamic and structural description of allosterically regulated VEGF receptor 2 dimerization. Blood 119(7):1781–1788
Leppänen VM, Prota AE, Jeltsch M, Anisimov A, Kalkkinen N, Strandin T, Lankinen H, Goldman A, Ballmer-Hofer K, Alitalo K (2010) Structural determinants of growth factor binding and specificity by VEGF receptor 2. Proc Natl Acad Sci USA 107(6):2425–2430
Markovic-Mueller S, Stuttfeld E, Asthana M, Weinert T, Bliven S, Goldie KN, Kisko K, Capitani G, Ballmer-Hofer K (2017) Structure of the full-length VEGFR-1 extracellular domain in complex with VEGF-A. Structure 25(2):341–352
Christinger HW, Fuh G, de Vos AM, Wiesmann C (2004) The crystal structure of PlGF in complex with domain 2 of VEGFR1. J Biol Chem 279(11):10382–10388
Wiesmann C, Fuh G, Christinger HW, Eigenbrot C, Wells JA, de Vos AM (1997) Crystal structure at 1.7 Å resolution of VEGF in complex with domain 2 of the Flt-1 receptor. Cell 91(5):695–704
Iyer S, Darley PI, Acharya KR (2010) Structural insights into the binding of VEGF-B by VEGFR-1D2: recognition and specificity. J Biol Chem 285(31):23779–23789
Ellis LM (2005) Bevacizumab. Nat Rev Drug Discov 4(Supplement):S8–S9
Ferrara N, Hillan KJ, Gerber HP, Novotny W (2004) Case history: discovery and development of bevacizumab, an anti-VEGF antibody for treating cancer. Nat Rev Drug Discov 3(5):391–400
Yang Y, Zhang Y, Cao Z, Ji H, Yang X, Iwamoto H, Wahlberg E, Lanne T, Sun B, Cao Y (2013) Anti-VEGF- and anti-VEGF receptor-induced vascular alteration in mouse healthy tissues. Proc Natl Acad Sci USA 110(29):12018–12023
Inai T, Mancuso M, Hashizume H, Baffert F, Haskell A, Baluk P, Hu-Lowe DD, Shalinsky DR, Thurston G, Yancopoulos GD, McDonald DM (2004) Inhibition of vascular endothelial growth factor (VEGF) signaling in cancer causes loss of endothelial fenestrations, regression of tumor vessels, and appearance of basement membrane ghosts. Am J Pathol 165(1):35–52
Krupitskaya Y, Wakelee HA (2009) Ramucirumab, a fully human mAb to the transmembrane signaling tyrosine kinase VEGFR-2 for the potential treatment of cancer. Curr Opin Investig Drugs 10(6):597–605
Mendel DB, Laird AD, Xin X, Louie SG, Christensen JG, Li G, Schreck RE, Abrams TJ, Ngai TJ, Lee LB, Murray LJ, Carver J, Chan E, Moss KG, Haznedar JO, Sukbuntherng J, Blake RA, Sun L, Tang C, Miller T, Shirazian S, McMahon G, Cherrington JM (2003) In vivo antitumor activity of SU11248, a novel tyrosine kinase inhibitor targeting vascular endothelial growth factor and platelet-derived growth factor receptors: determination of a pharmacokinetic/pharmacodynamic relationship. Clin Cancer Res 9(1):327–337
Hyde CA, Giese A, Stuttfeld E, Abram SJ, Villemagne D, Schleier T, Binz HK, Ballmer-Hofer K (2012) Targeting the extracellular domains D4 and D7 of VEGFR-2 reveals allosteric receptor regulatory sites. Mol Cell Biol 32(19):3802–3813
Steiner D, Merz FW, Sonderegger I, Gulotti-Georgieva M, Villemagne D, Phillips DJ, Forrer P, Stumpp MT, Zitt C, Binz HK (2017) Half-life extension using serum albumin-binding DARPin(R) domains. Protein Eng Des Sel 30(9):583–591
Winter G (2010) xia2: an expert system for macromolecular crystallography data reduction. J Appl Cryst 43(1):186–190
Waterman DG, Winter G, Parkhurst JM, Fuentes-Montero L, Hattne J, Brewster A, Sauter NK, Evans G (2013) The DIALS framework for integration software. CCP4 Newsl Protein Crystallogr 49:16–19
Evans P (2006) Scaling and assessment of data quality. Acta Crystallogr D Biol Crystallogr 62(Pt 1):72–82
Evans PR, Murshudov GN (2013) How good are my data and what is the resolution? Acta Crystallogr Sect D 69(7):1204–1214
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ (2007) Phaser crystallographic software. J Appl Crystallogr 40(Pt 4):658–674
Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66(Pt 2):213–221
Merz T, Wetzel SK, Firbank S, Pluckthun A, Grutter MG, Mittl PR (2008) Stabilizing ionic interactions in a full-consensus ankyrin repeat protein. J Mol Biol 376(1):232–240
Emsley P, Cowtan K (2004) Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60(Pt 12 Pt 1):2126–2132
Rizk A, Mansouri M, Ballmer-Hofer K, Berger P (2015) Subcellular object quantification with Squassh3C and SquasshAnalyst. Biotechniques 59(5):309–312
Laib AM, Bartol A, Alajati A, Korff T, Weber H, Augustin HG (2009) Spheroid-based human endothelial cell microvessel formation in vivo. Nat Protoc 4(8):1202–1215
Lichtenbeld HH, Muller AD, van Dam-Mieras MC, Blijham GH (1993) Tumor spheroid-induced vesicle formation on endothelial cells is associated with procoagulant properties. J Cell Sci 106(Pt 2):657–662
Alajati A, Laib AM, Weber H, Boos AM, Bartol A, Ikenberg K, Korff T, Zentgraf H, Obodozie C, Graeser R, Christian S, Finkenzeller G, Stark GB, Heroult M, Augustin HG (2008) Spheroid-based engineering of a human vasculature in mice. Nat Methods 5(5):439–445
Yang Y, Xie P, Opatowsky Y, Schlessinger J (2010) Direct contacts between extracellular membrane-proximal domains are required for VEGF receptor activation and cell signaling. Proc Natl Acad Sci USA 107(5):1906–1911
Sarabipour S, Ballmer-Hofer K, Hristova K (2016) VEGFR-2 conformational switch in response to ligand binding. Elife 5:e13876
Friedman LM, Rinon A, Schechter B, Lyass L, Lavi S, Bacus SS, Sela M, Yarden Y (2005) Synergistic down-regulation of receptor tyrosine kinases by combinations of mAbs: implications for cancer immunotherapy. Proc Natl Acad Sci USA 102(6):1915–1920
Baselga J, Albanell J (2001) Mechanism of action of anti-HER2 monoclonal antibodies. Ann Oncol 12(Suppl 1):S35–S41
Patel D, Lahiji A, Patel S, Franklin M, Jimenez X, Hicklin DJ, Kang X (2007) Monoclonal antibody cetuximab binds to and down-regulates constitutively activated epidermal growth factor receptor vIII on the cell surface. Anticancer Res 27(5A):3355–3366
Drebin JA, Link VC, Stern DF, Weinberg RA, Greene MI (1985) Down-modulation of an oncogene protein product and reversion of the transformed phenotype by monoclonal antibodies. Cell 41(3):697–706
Hudziak RM, Lewis GD, Winget M, Fendly BM, Shepard HM, Ullrich A (1989) p185HER2 monoclonal antibody has antiproliferative effects in vitro and sensitizes human breast tumor cells to tumor necrosis factor. Mol Cell Biol 9(3):1165–1172
Fan Z, Mendelsohn J (1998) Therapeutic application of anti-growth factor receptor antibodies. Curr Opin Oncol 10(1):67–73
Zhu W, Okollie B, Artemov D (2007) Controlled internalization of Her-2/neu receptors by cross-linking for targeted delivery. Cancer Biol Ther 6(12):1960–1966
Bertelsen V, Stang E (2014) The Mysterious Ways of ErbB2/HER2 Trafficking. Membranes (Basel) 4(3):424–446
Tiede C, Bedford R, Heseltine SJ, Smith G, Wijetunga I, Ross R, AlQallaf D, Roberts AP, Balls A, Curd A, Hughes RE, Martin H, Needham SR, Zanetti-Domingues LC, Sadigh Y, Peacock TP, Tang AA, Gibson N, Kyle H, Platt GW, Ingram N, Taylor T, Coletta LP, Manfield I, Knowles M, Bell S, Esteves F, Maqbool A, Prasad RK, Drinkhill M, Bon RS, Patel V, Goodchild SA, Martin-Fernandez M, Owens RJ, Nettleship JE, Webb ME, Harrison M, Lippiat JD, Ponnambalam S, Peckham M, Smith A, Ferrigno PK, Johnson M, McPherson MJ, Tomlinson DC (2017) Affimer proteins are versatile and renewable affinity reagents. Elife. https://doi.org/10.7554/eLife.24903
Acknowledgements
Andreas Cornelius, Johan Abram and Nicole Bassler of Molecular Partners are acknowledged for production and characterization of D4b and HD4b. We thank Drs. Richard Kammerer and Roger Benoit for designing the receptor D4–5 construct, Dr. Aurélien Rizk for advice with Squassh analysis, and Mayanka Asthana for assistance in protein purification. We also thank staff at beamline X06SA at the Swiss Light Source in Villigen Switzerland for help with X-ray data acquisition. This work was supported by grants from Schweizerischer Nationalfonds zur Förderung der Wissenschaftlichen Forschung (31003A_152908 to KB-H) and Oncosuisse (KLS-3569-02-2015 to KB-H).
Author information
Authors and Affiliations
Contributions
Experiments were conceived, designed and interpreted by HKB, SMM, CLP, DA and KB-H. Experiments were performed by KMT, CLP, DA and SMM. The manuscript was written by KMT, CLP, DA, SMM, HKB and KB-H.
Corresponding authors
Ethics declarations
Conflict of interest
HKB is shareholder of Molecular Partners AG commercializing the DARPin® Technology. The other authors do not declare competing financial interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Thieltges, K.M., Avramovic, D., Piscitelli, C.L. et al. Characterization of a drug-targetable allosteric site regulating vascular endothelial growth factor signaling. Angiogenesis 21, 533–543 (2018). https://doi.org/10.1007/s10456-018-9606-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10456-018-9606-9