Abstract
Medium-chain-length polyhydroxyalkanoates (mcl-PHAs) are a large class of biopolymers that have attracted extensive attention as renewable and biodegradable bio-plastics. They are naturally synthesized via fatty acid de novo biosynthesis pathway or β-oxidation pathway from Pseudomonads. The unconventional yeast Yarrowia lipolytica has excellent lipid/fatty acid catabolism and anabolism capacity depending of the mode of culture. Nevertheless, it cannot naturally synthesize PHA, as it does not express an intrinsic PHA synthase. Here, we constructed a genetically modified strain of Y. lipolytica by heterologously expressing PhaC1 gene from P. aeruginosa PAO1 with a PTS1 peroxisomal signal. When in single copy, the codon optimized PhaC1 allowed the synthesis of 0.205 % DCW of PHA after 72 h cultivation in YNBD medium containing 0.1 % oleic acid. By using a multi-copy integration strategy, PHA content increased to 2.84 % DCW when the concentration of oleic acid in YNBD was 1.0 %. Furthermore, when the recombinant yeast was grown in the medium containing triolein, PHA accumulated up to 5.0 % DCW with as high as 21.9 g/L DCW, which represented 1.11 g/L in the culture. Our results demonstrated the potential use of Y. lipolytica as a promising microbial cell factory for PHA production using food waste, which contains lipids and other essential nutrients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyhydroxyalkanoates (PHAs) are a large class of natural biopolymers that is accumulated by microorganisms as a kind of carbon and energy reserves in the presence of an excess carbon source [1, 18]. These polymers have attracted extensive attention because of their thermoplastic and elastomeric properties, making them good candidates for environmentally friendly and biodegradable plastics. There are more than 150 different confirmed types of PHA monomer subunits, each containing varied monomers with different chain lengths/structures, resulting in a wide diversity of material properties [12]. They can be divided into three main types: short-chain-length PHAs (scl-PHAs) which contain repeating units of 3–5 carbon atoms, medium-chain-length PHAs (mcl-PHAs) which contain repeating units of 6–14 carbon atoms, and scl-co-mcl PHAs which consist of both SCL and MCL repeating units of 3–14 carbons [6].
Mcl-PHAs are naturally synthesized via fatty acid de novo biosynthesis pathway or β-oxidation pathway from Pseudomonads [28, 29]. Heterologous expression of the key enzymes for mcl-PHA synthesis in a proper host provides a main solution for mcl-PHA synthesis by changing the flux of intermediates of fatty acid metabolism [11, 13, 14, 26, 28, 35]. Heterologous production of mcl-PHA in recombinant Escherichia coli and Saccharomyces cerevisiae were studied extensively [17, 25, 26, 32].
In the last decades, the unconventional yeast Y. lipolytica (originally classified as Candida lipolytica) has attracted much interests, notably due to its excellent lipid/fatty acid catabolism and anabolism capacity depending on the cultivation mode [4, 9]. Therefore, it is an interesting microorganism for lipid synthesis or utilization to produce high-value bioproducts [7]. For example, metabolic engineering of Y. lipolytica resulted in a strain that produced omega-3 fatty acids eicosapentaenoic acid (EPA) at 15 % of dry cell weight by introducing several heterologous genes, encoding ∆9-elongase, ∆8-desaturase, ∆5-desaturase and ∆17-desaturase, making a breakthrough to replace an animal-derived product [30]. Lipid metabolism was rerouted in this yeast to develop a cell factory for ricinoleic acid (RA) production, capable of accumulating RA to 43 % of its total lipids and over 60 mg/g of cell dry weight [2]. Another effectual work was the synthesis of trans-10, cis-12 conjugated linoleic acid (CLA) through heterologously expressing linoleic acid isomerase gene from Propionibacterium acnes, which was further enhanced by a modified promoter and co-expression of a Δ12-desaturase gene [33, 34]. Recently, Haddouche et al. examined the feasibility of using Y. lipolytica for PHA biosynthesis and investigated the roles of multiple acyl-CoA oxidases in the routing of carbon flow towards β-oxidation [10].
Yarrowia lipolytica possesses 16 paralogs of genes coding for lipases which can catalyze the hydrolysis of the ester bond of tri-, di- and mono-glycerides of long-chain fatty acids into fatty acids and glycerol. It has six genes (POX1–POX6) encoding isoenzymes of acyl-CoA oxidases (Aoxs) with different substrate specificities that catalyze the first step and also the limiting step of β-oxidation of fatty acids [5, 10]. In contrast, S. cerevisiae has only one Aox-encoding gene. Like in S. cerevisiae, the β-oxidation cycle of fatty acid of Y. lipolytica also provides different length of (R)-specific 3-hydroxy-acyl-CoAs (catalyzed by MFE2 type enzyme), which can be used as the direct precursors for PHA synthesis. Hence, it exhibits excellent potential for PHA production. In this study, Mcl-PHA synthase (encoded by PhaC1) of Pseudomonas aeruginosa PAO1 was heterologously expressed under the recombinant hp4d promoter [21], with a PTS1 peroxisomal signal (SKL). Mcl-PHA production profiles were evaluated, using fatty acid and triolein as substrates.
Materials and methods
Strains and culture conditions
Yarrowia lipolytica strain Po1h and plasmids pINA1312 and pINA1292 have been described previously [21, 23]. Escherichia coli DH5α was used for routine subcloning and plasmid propagation. It was grown in Luria–Bertani broth (LB) containing kanamycin (25 mg/L) for plasmid selection. YPD and YNBD media were used for Y. lipolytica cultivation and transformants selection, respectively. YPD contained 1 % (w/v) yeast extract, 2 % (w/v) peptone and 2 % (w/v) dextrose. YNBD contained 0.67 % (w/v) yeast nitrogen base (without amino acids and with ammonium sulphate), 0.2 % casamino acids, and 2 % (w/v) dextrose. YNBD containing different concentration of oleic acid (0.1–1.0 %) was used as fermentation substrate by the constructed parallel strains. YPTo and YNBTo (containing 2 % (w/v) triolein instead of dextrose) were also used in mcl-PHA fermentation. Stock solutions of oleic acid (10 % oleic acid, 0.5 % Tween 80) and triolein (25 % triolein, 1.5 % Tween 80) were subjected to sonication three times for 1 min each on ice and heat sterilized separately.
For mcl-PHA production, yeast cells were cultured on YPD plates at 28 °C overnight as the first pre-cultivation step. Then a single colony was picked using a wire loop and aseptically inoculated into a 300 ml flask containing 50 ml of YPD medium and grown at 28 °C for 16–20 h. The cells were harvested by centrifugation and washed once in water and re-suspended in fermentation media with an initial OD600 of 0.5. The cultures were then grown at 28 °C, 200 rpm for 72 h before being harvested for mcl-PHA analysis. All the fermentations were done in duplicate.
Plasmids construction
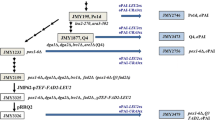
The native and codon-optimized nucleotide sequences of PhaC1 were cloned and expressed in Y. lipolytica. The native coding sequence (GeneBank AE_004091), flanked with BamHI and KpnI sites, without and with Kozak sequence, was amplified by PCR using the primers P1 (CCGGGATCCATGAGTCAGAAGAACAATAAC or CCGGGATCC GCCACAATGAGTCAGAAGAACAATAAC) and P2 (CGGGGTACCCTACAGCTTGGATCGTTCATGCACGTAGGT) on template genome of P. aeruginosa PAO1. Italic nucleotides depict the restriction sites and underlined nucleotides are Kozak element. The codon-optimized DNA sequence flanked with BamHI and KpnI, with Kozak sequence, was synthesized (Genscript, Nanjing, China). Standard PCR consisted of 15 s at 95 °C, 30 s at 56 °C, and 90 s at 68 °C for 30 cycles. The three versions of DNA fragments digested with the appropriate restriction enzymes were ligated to the same digested pINA1312 and pINA1292 to form the mono-copy and multi-copy plasmids, respectively. All newly constructed plasmids were screened by restriction enzyme digestion and PCR, and then confirmed by DNA sequencing.
Strain engineering
The expression cassettes based on newly constructed plasmids were linearized using NotI enzyme, and introduced into Y. lipolytica Po1h to obtain recombinant yeast strains (Table 1). Transformation was performed by the Lithium Acetate Method [3], and transformants were selected by plating on YNBD plates and screened by diagnostic PCR using yeast genome extracted by TIANamp Yeast DNA Kit (TIANGEN, Beijing, China).
Mcl-PHA analysis
Mcl-PHA biosynthesis was confirmed by gas chromatography-mass spectrometry (GC–MS) analysis as described by Zhuang et al. [35]. The ion source temperature was adjusted to 200 °C. The GC analyses were performed as follows: the oven temperature was initially maintained at 60 °C for 3 min, the temperature was gradually increased at a rate of 10 °C per minute up to 250 °C and then held for 6 min, one μl of sample solution was injected. Methyl esters corresponding to mcl-PHA repeating monomers were determined based on retention time and fragmentation patterns of known standards and resulting mass spectra available compared with GC–MS library database (NIST08s). The MS was operated in scanning mode between 20 and 350 m/z.
The content and monomer compositions of intracellular accumulated mcl-PHA were analyzed by gas chromatography (GC). Its content was defined as the percentage ratio of mcl-PHA concentration to dried cell weight (DCW). Liquid culture was harvested by centrifugation, washed twice in water and lyophilized. Lyophilized cells were extracted four or five times with warm methanol (65 °C) to remove lipids, free fatty acids and acyl-CoA, including 3-hydroxyacyl-CoA, while PHA (insoluble in methanol) remained associated with the cells [11]. After centrifugation and removal of the residual methanol, the material (15 mg) was subjected to methanolysis in the presence of 1 ml of chloroform and 1 ml of 3 % (v/v) sulfuric acid in methanol for 1 h at 100 °C. The samples were cooled to room temperature and then 1 ml of distilled water was added in order to extract the cell debris, soluble in the aqueous phase. The mixture was vortexed and centrifuged at 10,000 rpm for 10 min. After layer separation, the organic (chloroform) phase (500 μl) was transferred to another new vial and analyzed using a Shimadzu GC2010 gas chromatograph (Kyoto, Japan) equipped with an AOC-20i auto-injector and a RestekRxi®-5 column. PHA standard samples (Sigma-Aldrich) were also analyzed by GC according to the method above. The temperature program used was as follows: 60 °C held for 3 min, ramped from 60 to 260 °C at 10 °C min−1 and then held for 6 min.
Results
Engineering of Y. lipolytica for mcl-PHA biosynthesis
PhaC1, encoding a mcl-PHA polymerase from P. aeruginosa PAO1, was heterologously expressed with a PTS1 peroxisomal signal to ensure its proper expression in peroxisomes of Y. lipolytica. Since the codon usage of native PhaC1 sequence differs from that preferred by Y. lipolytica, the gene was codon-optimized as well. To test the effectiveness of codon optimization in the expression of PhaC1, both the native gene (PhaC1) and the codon-optimized gene (oPhaC1) were inserted into plasmid-borne expression cassettes driven by a strong promoter hp4d. Additionally, the Kozak element was added before the first AUG codon to prevent the leaky scanning of the ribosome and increase the efficiency of translation [8]. Expression cassettes were derived from single-copy vector pINA1312 which contains the non-defective ura3d1 gene for mono-copy expression [23]. Three expression cassettes were constructed and used to transform Y. lipolytica after being linearized by NotI, resulting in three parallel mcl-PHA producing strains: PSNC, PSC and PSOC (Table 1). Recombinant yeast strains harboring these variant versions of PhaC1 expression cassette were fermented in YNBD media containing 0.1 % oleic acid. Three transformants of each recombinant strain were picked randomly for PHA fermentation. GC–MS results showed that the engineered strain harboring mcl-PHA synthase accumulated mcl-PHA compared with the control strain of Po1h (Fig. S1). PHA yields in the different recombinant Y. lipolytica strains are shown in Fig. 1. All engineered strains grew to a similar biomass (approximately 7 g/L DCW) and produced mcl-PHA in yields that were significantly different between the constructs, but similar between the 3 transformants for the same construct. The 3 PSNC clones, without Kozak element before the initial AUG codon of native phaC1 gene, produced the least amount of PHA (mean yield of 0.044 % of dried cell weight (DCW)). In contrast, the 3 PSC clones, with the Kozak element added upstream of native phaC1 gene, produced a mean yield of 0.157 % of PHA, which is 3.6-fold that produced by PSNC. Furthermore, the combined effect of codon-optimization and addition of Kozak sequence in the 3 PSOC clones resulted in an average PHA yield of 0.205 %, 4.7-fold more than that of PSNC.
Integration of PhaC1 in multiple copies
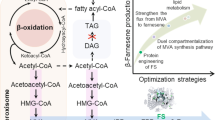
As shown in Figs. 1 and 2, exogenous PhaC introduced into peroxisomes of Y. lipolytica polymerized 3-hydroacyl-CoA, which is an intermediate of MFE2 of β-oxidation, to form mcl-PHA. Using codon-optimized gene (oPhaC1) and Kozak sequence, a relatively high PHA yield was obtained. Accordingly, enhanced activity of PhaC can seize more intermediates from β-oxidation and yield higher PHA accumulation. This allows further strengthening of the PhaC1 expression using a multiple integration strategy. The multi-copy vector pINA1292, which contains the defective ura3d4 gene and is required in multiple copies to alleviate the uracil auxotrophy of the host, was adopted to develop a more efficient PHA-producing strain [23]. Three individual transformants were picked randomly to test their abilities for mcl-PHA production, using PSOC as a control strain. As shown in Fig. 3, both DCW and PHA yields were improved. Accordingly, for the multi-copy strain, PHA yields appeared to be slightly but significantly different between PMOC1 and PMOC3 transformants (Fig. 3). PMOC1 strain was the best PHA producer, with a yield of 1.46 %, sevenfold more than that of PSOC strain. The average mcl-PHA contents and DCW of PMOCs reached 1.28 % and 8.3 g/L respectively after 72 h cultivation in a shake flask with YNBD medium containing 0.1 % oleic acid.
Schematic pathway involved in the key routes of modified catabolism of lipids, fatty acid, glycerol and glucose. LIPs, lipases; FAA, acyl-CoA synthetase; POXAOX, acyl-CoA oxidase; MFE2, a multifunctional enzyme with one 2-enoyl-CoA hydratase domain and two R-3-hydroxyacylCoA dehydrogenase domains; YLFOX3, 3-ketoacyl CoA thiolase
Optimizing cultivation for mcl-PHA production
Increasing mcl-PHA content by raising the concentration of oleic acid
In order to explore whether the ability of PHA biosynthesis can be further enhanced, fermentation using YNBD medium contained various concentrations of oleic acid from 0.1 to 1.0 % were carried out using the best-performing strain of PMOC. The biomass and mcl-PHA accumulation are presented in Table 2. As we expected, more oleic acid in the culture can lead to higher mcl-PHA accumulation, although its increase is not completely linear, especially when the levels of oleic acid increased from 0.5 to 1.0 %. The highest mcl-PHA was 0.43 g/L in YNBD with 1 % oleic acid. The composition of mcl-PHA monomers is listed in Table 2. In all conditions, (R)-3-hydroxyoctanoate (3HO) was the most abundant monomer in the final polyester, accounting for 41-48.4 mol% of the monomer composition. It indicates a specificity of the P. aeruginosa PHA synthase for (R)-3-hydroxy-octanoyl-CoA.
Increased mcl-PHA content by utilizing triolein as substrate
Since fatty acids are usually expensive, toxic to cells and insoluble (poorly miscible) in aqueous solution, glucose needs to be added as a supplementary carbon source to ensure better growth of yeast cells. In this study, triolein, as a representative triacylglycerol, was employed as single substrate for mcl-PHA synthesis in YPTo and YNBTo medium. As shown in Table 3, the PHA composition using triolein as substrate is consistent with that in YNBD medium containing oleic acid, while the PHA content increased to over 5.0 % DCW. Moreover, rich medium (YPTo) can yield higher biomass (21.9 g/L), which resulted in an accumulation of 1.11 g/L mcl-PHA. Thus, triolein constitutes an alternative and cheaper carbon source for PHA production.
Discussion
Mcl-PHAs are a large class of biopolymers that are attractive for a wide range of potential applications. The aim of this study was to apply metabolic engineering to the oleaginous yeast Y. lipolytica for the production of mcl-PHA. Yeast does not naturally synthesize PHAs, as it does not express an intrinsic PHA synthase. However, it can provide the direct precursors, different length of (R)-specific 3-hydroxy-acyl-CoAs, by β-oxidation of fatty acids that can be used as substrates for mcl-PHA biosynthesis. Mcl-PHA was previously synthesized in S. cerevisiae transformed with heterologous PHA synthase targeted to peroxisomes, using even-chain length and odd-chain length fatty acids [25, 32]. The maximum amount of PHA accumulated was 0.45 % DCW [25]. Thereafter, mcl-PHA was similarly synthesized in peroxisomes of Pichia pastoris, and reached 1 % DCW using oleic acid as substrate [24]. Recently, Y. lipolytica was engineered for synthesis of mcl-PHA, which was further improved by inactivating the R-3-hydroxyacyl-CoA dehydrogenase domain of MFE2 and blocking the neutral lipid synthesis pathway using 0.2 % tridecanoic acid in YNB minimal medium [10, 11]. These studies indicated that Y. lipolytica, which exhibits efficient lipid/fatty acid catabolism and anabolism capacities, owns a tremendous potential for mcl-PHA synthesis in comparison with other yeasts.
In this study, a series of recombinant Y. lipolytica strains was constructed for mcl-PHA production. PHA compositions remained consistent in the different strains constructed and with different carbon sources (Tables 2, 3). In all conditions, (R)-3-hydroxyoctanoate (3HO) was the most abundant monomer in the final polyester, accounting for 41–48.4 mol% of the monomer composition. This reflects the specificity of the P. aeruginosa PHA synthase for R-3-hydroxy-octanoyl-CoA, as proposed previously [22]. PHA content of PSC is 3.6-fold that of PSNC (Fig. 1), indicating that the inclusion of Kozak element increased gene expression in this study. Strain PSOC with codon optimized PhaC1 gene further increased mcl-PHA accumulation. With multiple integration of codon-optimized PhaC1, PMOC output was of 5.0 % and 1.11 g/L of PHA in the culture using triolein as the sole substrate in rich medium (YPTo), which ensured an excellent cell growth and a high resultant cell dry weight (22.0 g/L DCW).
To synthesize chemicals heterologously through strategies of metabolic engineering tends to encounter the issue of the competition between product titers and yields. To this end, improved PHA production is usually accompanied by a decrease in cell growth. In the studies of PHA synthesis, fatty acids were commonly used as substrates. However, fatty acids are insoluble and toxic to cells in aqueous solution, which led to poor cell growth. A solution for that is adding other carbon sources, such as glucose or glycerol, as co-substrates to ensure proper cell growth and biomass production. Therefore, a more balanced genetic method (evolutionary approach, for example) should be taken further to improve the ability of engineered strains. Otherwise, strategies like culture optimization using different sources of feedstocks can be applied. Herein, triolein turned out to be an excellent carbon source for mcl-PHA production, which yielded both high product concentration and biomass. Besides, Y. lipolytica used in our study was not modified for its lipid and fatty acid metabolism at all. Future works, including deletion of the neutral lipid synthesis pathway and overexpression of AOXs charging rate-limited step of β-oxidation, which might further improve the yield of mcl-PHA, are currently planned for investigation in our laboratory.
Sustainable valorization of renewable feedstock can produce value-added chemicals and biopolymers via microbial bioconversion [15, 19]. As described in our recent study, food waste is a serious global issue [27]. It contains various nutrients such as sugars, free amino nitrogen (FAN) and lipids which can be released after hydrolysis and separation. Food waste-based biorefinery, as a novel concept, received a significant attention in recent years [16, 20]. In comparison with other yeast, Y. lipolytica has excellent potential for utilising hydrophobic substrates efficiently. Our recent studies successfully demonstrated the use of glucose and FAN as major nutrients from food waste for microbial production of high value-added products such as succinate [27, 31]. Our future studies will target the full exploration of lipid fraction from food waste to produce PHA using engineered Y. lipolytica, as this study demonstrated the novel use of triolein for PHA synthesis.
References
Anderson AJ, Dawes EA (1990) Occurrence, metabolism, metabolic role, and industrial uses of bacterial polyhydroxyalkanoates. Microbiol Rev 54(4):450–472
Beopoulos A, Verbeke J, Bordes F, Guicherd M, Bressy M, Marty A, Nicaud JM (2014) Metabolic engineering for ricinoleic acid production in the oleaginous yeast Yarrowia lipolytica. Appl Microbiol Biotechnol 98(1):251–262
Chen DC, Beckerich JM, Gaillardin C (1997) One-step transformation of the dimorphic yeast Yarrowia lipolytica. Appl Microbiol Biotechnol 48(2):232–235
Dulermo T, Nicaud JM (2011) Involvement of the G3P shuttle and beta-oxidation pathway in the control of TAG synthesis and lipid accumulation in Yarrowia lipolytica. Metab Eng 13(5):482–491
Fickers P, Marty A, Nicaud JM (2011) The lipases from Yarrowia lipolytica: genetics, production, regulation, biochemical characterization and biotechnological applications. Biotechnol Adv 29(6):632–644
Gao X, Chen JC, Wu Q, Chen GQ (2011) Polyhydroxyalkanoates as a source of chemicals, polymers, and biofuels. Curr Opin Biotechnol 22(6):768–774
Gardini F, Suzzi G, Lombardi A, Galgano F, Crudele MA, Andrighetto C, Schirone M, Tofalo R (2001) A survey of yeasts in traditional sausages of southern Italy. FEMS Yeast Res 1(2):161–167
Gasmi N, Fudalej F, Kallel H, Nicaud JM (2011) A molecular approach to optimize hIFN α2b expression and secretion in Yarrowia lipolytica. Appl Microbiol Biotechnol 89(1):109–119
Groguenin A (2004) Genetic engineering of the β-oxidation pathway in the yeast Yarrowia lipolytica to increase the production of aroma compounds. J Mol Catal B-Enzym 28(2–3):75–79
Haddouche R, Delessert S, Sabirova J, Neuveglise C, Poirier Y, Nicaud JM (2010) Roles of multiple acyl-CoA oxidases in the routing of carbon flow towards beta-oxidation and polyhydroxyalkanoate biosynthesis in Yarrowia lipolytica. FEMS Yeast Res 10(7):917–927
Haddouche R, Poirier Y, Delessert S, Sabirova J, Pagot Y, Neuveglise C, Nicaud JM (2011) Engineering polyhydroxyalkanoate content and monomer composition in the oleaginous yeast Yarrowia lipolytica by modifying the ss-oxidation multifunctional protein. Appl Microbiol Biotechnol 91(5):1327–1340
Hazer B, Steinbuchel A (2007) Increased diversification of polyhydroxyalkanoates by modification reactions for industrial and medical applications. Appl Microbiol Biotechnol 74(1):1–12
Huijberts GN, Eggink G, de Waard P, Huisman GW, Witholt B (1992) Pseudomonas putida KT2442 cultivated on glucose accumulates poly(3-hydroxyalkanoates) consisting of saturated and unsaturated monomers. Appl Environ Microbiol 58(2):536–544
Jenkins LS, Nunn WD (1987) Genetic and molecular characterization of the genes involved in short-chain fatty acid degradation in Escherichia coli: the ato system. J Bacteriol 169(1):42–52
Kang Z, Du L, Kang J, Wang Y, Wang Q, Liang Q, Qi Q (2011) Production of succinate and polyhydroxyalkanoate from substrate mixture by metabolically engineered Escherichia coli. Bioresource Technol 102(11):6600–6604
Koutinas AA, Vlysidis A, Pleissner D, Kopsahelis N, Lopez Garcia I, Kookos IK, Papanikolaou S, Kwan TH, Lin CS (2014) Valorization of industrial waste and by-product streams via fermentation for the production of chemicals and biopolymers. Chem Soc Rev 43(8):2587–2627
Langenbach S, Rehm BH, Steinbüchel A (1997) Functional expression of the PHA synthase gene phaC1 from Pseudomonas aeruginosa in Escherichia coli results in poly(3-hydroxyalkanoate) synthesis. FEMS Microbiol Lett 150(2):303–309
Lee SY (1996) Bacterial polyhydroxyalkanoates. Biotechnol Bioeng 49(1):1–14
Liang Q, Qi Q (2014) From a co-production design to an integrated single-cell biorefinery. Biotechnol Adv 32(7):1328–1335
Lin CSK, Pfaltzgraff LA, Herrero-Davila L, Mubofu EB, Abderrahim S, Clark JH, Koutinas AA, Kopsahelis N, Stamatelatou K, Dickson F, Thankappan S, Mohamed Z, Brocklesby R, Luque R (2013) Food waste as a valuable resource for the production of chemicals, materials and fuels. Current situation and global perspective. Energy. Environ Sci 6(2):426–465
Madzak C, Gaillardin C, Beckerich JM (2004) Heterologous protein expression and secretion in the non-conventional yeast Yarrowia lipolytica: a review. J Biotechnol 109(1–2):63–81
Marchesini S, Poirier Y (2003) Futile cycling of intermediates of fatty acid biosynthesis toward peroxisomal beta-oxidation in Saccharomyces cerevisiae. J Biol Chem 278(35):32596–32601
Nicaud JM, Madzak C, van den Broek P, Gysler C, Duboc P, Niederberger P, Gaillardin C (2002) Protein expression and secretion in the yeast Yarrowia lipolytica. FEMS Yeast Res 2(3):371–379
Poirier Y, Erard N, MacDonald-Comber Petétot J (2002) Synthesis of polyhydroxyalkanoate in the peroxisome of Pichia pastoris. FEMS Microbiol Lett 207(1):97–102
Poirier Y, Erard N, Petetot JM (2001) Synthesis of polyhydroxyalkanoate in the peroxisome of Saccharomyces cerevisiae by using intermediates of fatty acid beta-oxidation. Appl Environ Microbiol 67(11):5254–5260
Qi Q, Rehm BH, Steinbüchel A (1997) Synthesis of poly(3-hydroxyalkanoates) in Escherichia coli expressing the PHA synthase gene phaC2 from Pseudomonas aeruginosa: comparison of PhaC1 and PhaC2. FEMS Microbiol Lett 157(1):155–162
Sun Z, Li M, Qi Q, Gao C, Lin CS (2014) Mixed food waste as renewable feedstock in succinic acid fermentation. Appl Biochem Biotechnol 174(5):1822–1833
Wang Q, Tappel RC, Zhu C, Nomura CT (2012) Development of a new strategy for production of medium-chain-length polyhydroxyalkanoates by recombinant Escherichia coli via inexpensive non-fatty acid feedstocks. Appl Environ microb 78(2):519–527
Wang Q, Zhuang Q, Liang Q, Qi Q (2013) Polyhydroxyalkanoic acids from structurally-unrelated carbon sources in Escherichia coli. Appl Microbiol Biotechnol 97(8):3301–3307
Xue Z, Sharpe PL, Hong SP, Yadav NS, Xie D, Short DR, Damude HG, Rupert RA, Seip JE, Wang J, Pollak DW, Bostick MW, Bosak MD, Macool DJ, Hollerbach DH, Zhang H, Arcilla DM, Bledsoe SA, Croker K, McCord EF, Tyreus BD, Jackson EN, Zhu Q (2013) Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat Biotechnol 31(8):734–740
Zhang AY-z, Sun Z, Leung CCJ, Han W, Lau KY, Li M, Lin CSK (2013) Valorisation of bakery waste for succinic acid production. Green Chem 15(3):690–695
Zhang B, Carlson R, Srienc F (2006) Engineering the monomer composition of polyhydroxyalkanoates synthesized in Saccharomyces cerevisiae. Appl Environ Microbiol 72(1):536–543
Zhang B, Chen H, Li M, Gu Z, Song Y, Ratledge C, Chen YQ, Zhang H, Chen W (2013) Genetic engineering of Yarrowia lipolytica for enhanced production of trans-10, cis-12 conjugated linoleic acid. Microb Cell Fact 12:70–77
Zhang B, Rong C, Chen H, Song Y, Zhang H, Chen W (2012) De novo synthesis of trans-10, cis-12 conjugated linoleic acid in oleaginous yeast Yarrowia lipolytica. Microb Cell Fact 11:51–58
Zhuang Q, Wang Q, Liang Q, Qi Q (2014) Synthesis of polyhydroxyalkanoates from glucose that contain medium-chain-length monomers via the reversed fatty acid beta-oxidation cycle in Escherichia coli. Metab Eng 24:78–86
Acknowledgments
The work described in this paper was fully supported by grants from the Research Grants Council of the Hong Kong Special Administrative Region, China (Project No. CityU189713) and the State Key Lab of Microbial Technology in Shandong University, China (M2014-03).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, C., Qi, Q., Madzak, C. et al. Exploring medium-chain-length polyhydroxyalkanoates production in the engineered yeast Yarrowia lipolytica . J Ind Microbiol Biotechnol 42, 1255–1262 (2015). https://doi.org/10.1007/s10295-015-1649-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-015-1649-y