Abstract
RNA processing, RNA editing, RNA splicing and translational activation of RNAs are essential post-transcriptional steps in chloroplast gene expression. Typically, the factors mediating those processes are nuclear encoded and post-translationally imported into the chloroplasts. In land plants, members of the large pentatricopeptide repeat (PPR) protein family are required for individual steps in chloroplast RNA processing. Interestingly, a subgroup of PPR proteins carries a C-terminal small MutS related (SMR) domain. Here we analyzed the consequences of mutations in the SVR7 gene, which encodes a PPR-SMR protein, in Arabidopsis thaliana. We demonstrate that SVR7 mutations lead to a specific reduction in chloroplast ATP synthase levels. Furthermore, we found aberrant transcript patterns for ATP synthase coding mRNAs in svr7 mutants. Finally, a reduced ribosome association of atpB/E and rbcL mRNAs in svr7 mutants suggests the involvement of the PPR-SMR protein SVR7 in translational activation of these mRNAs. We describe that the function of SVR7 in translation has expanded relative to its maize ortholog ATP4. The results provide evidence for a relaxed functional conservation of this PPR-SMR protein in eudicotyledonous and monocotyledonous plants, thus adding to the knowledge about the function and evolution of PPR-SMR proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloroplasts are endosymbiotic descendants of previously free living cyanobacteria (Margulis 1970). During co-evolution of host and endosymbiont, chloroplasts lost most of their genetic autonomy by gene transfer into the nuclear genome (Bock and Timmis 2008; Kleine et al. 2009; Martin 2003).The remaining chloroplast genes essentially code for parts of a bacterial type gene expression machinery and proteins relevant for photosynthesis. However, the overwhelming majority of proteins found in chloroplasts is nuclear encoded and post-translationally imported necessitating coordinated gene expression in both the nuclear and the plastid compartments.
Compared to cyanobacteria, gene expression in chloroplasts exhibits a surprisingly complex RNA metabolism. Group I and -II intron splicing, RNA editing, 5′-, 3′-end trimming and intracistronic RNA processing are common events in chloroplast RNA maturation (Barkan 2011; Stern et al. 2010). Additionally, chloroplast gene expression is translationally regulated (Barkan 2011; Wobbe et al. 2008). Chloroplast RNA metabolism and translational regulation are realized by nuclear encoded factors (Barkan 2011). Among those factors, the pentatricopeptide repeat (PPR) protein family is outstanding in land plants because of the large number of its members (~450 in Arabidopsis) and since PPR proteins fulfill specific functions in several steps of chloroplast RNA maturation and translation processes (Schmitz-Linneweber and Small 2008). Most PPR proteins are targeted to either chloroplasts or mitochondria and their sequences are highly conserved among land plants (Lurin et al. 2004; Schmitz-Linneweber and Small 2008). PPR proteins are composed of tandem arrays of degenerated repeating units of ~35 amino acids. These tandem arrays are predicted to form helical hairpin structures that may assemble into an alpha solenoid superstructure (Delannoy et al. 2007; Small and Peeters 2000). PPR proteins bind RNA in vitro and associate with RNA in vivo (Beick et al. 2008; Gillman et al. 2007; Hammani et al. 2011; Hattori et al. 2007; Okuda et al. 2006; Pfalz et al. 2009; Prikryl et al. 2011; Schmitz-Linneweber et al. 2005; Tasaki et al. 2010; Williams-Carrier et al. 2008; Zoschke et al. 2012). PPR domains themselves are not catalytically active but guide enzymatically active domains, enzymes or ribonucleoprotein complexes to specific sequences where they perform their essential functions in organellar RNA metabolism (Delannoy et al. 2007; Schmitz-Linneweber and Small 2008).
PPR proteins can carry additional protein domains: A large subclass of PPR proteins contains a C-terminal “E” domain that is linked to organellar RNA editing (Zehrmann et al. 2011). PPR4, which contains an RNA recognition motif, and OTP51, a PPR protein with a LAGLIDAG domain typically found in homing endonucleases are involved in chloroplast splicing (de Longevialle et al. 2008; Schmitz-Linneweber et al. 2006). Furthermore several PPR proteins carry a small MutS related domain (SMR) at the C-terminus (Liu et al. 2010).
The SMR domain was originally identified as the C-terminal part of bacterial MutS2 proteins (Moreira and Philippe 1999). Bacterial SMR domains have DNA binding and endonuclease activity and act in DNA recombination and repair processes (reviewed in Fukui and Kuramitsu 2011). It was also speculated that SMR domains could function in transcription (Fukui and Kuramitsu 2011). The function of SMR domains in PPR proteins is unknown.
Out of the eight Arabidopsis PPR-SMR proteins three were analyzed experimentally so far. pTAC2 and its tobacco ortholog were found in transcriptionally active chloroplast extracts (Pfalz et al. 2006; Suzuki et al. 2004). Arabidopsis ptac2 mutants show a strongly reduced accumulation of transcripts generated by the plastid encoded RNA polymerase (PEP) suggesting a role of pTAC2 in PEP transcription (Pfalz et al. 2006). GUN1, another PPR-SMR protein was identified in a screen for mutants with impaired retrograde chloroplast to nucleus signaling (Koussevitzky et al. 2007). Retrograde signals are elicited by changes in chloroplast physiological parameters like the photosynthetic redox status, or the accumulation of reactive oxygen species, but also by changes in chloroplast gene expression. The nucleus responds by adapting its gene expression to the requirements of the signaled plastid state. This assures a concerted gene expression in both compartments (Woodson and Chory 2008). The mutation of GUN1 disrupts several of the retrograde signaling pathways indicating a central function of this PPR-SMR protein in retrograde signaling (Koussevitzky et al. 2007). The third experimentally analyzed PPR-SMR protein is the Arabidopsis SUPPRESSOR OF VARIEGAION7 (SVR7) protein and its radish and maize orthologs P67 and ATP4, respectively (Echeverria and Lahmy 1995; Lahmy et al. 2000; Liu et al. 2010; Zoschke et al. 2012). The SVR7 mutation suppresses the variegated phenotype of var2 mutants by an unknown mechanism (Liu et al. 2010). The knockout of its maize ortholog ATP4 leads to a specific reduction of atpB/E and atpA translation causing the loss of the chloroplast ATP synthase (Zoschke et al. 2012). A more general reduction in chloroplast protein accumulation and a less severe effect on ATP synthase accumulation in the Arabidopsis svr7 mutant (Liu et al. 2010) suggested that the functions of the putative orthologs in Arabidopsis and maize differ.
To get a more detailed insight in the potential functional diversification of PPR proteins during evolution, we analyzed in the current study the effects of mutations of the PPR-SMR gene SVR7 on chloroplast gene expression in two Arabidopsis thaliana ecotypes. Here we show that SVR7 mutations in Arabidopsis cause a specific, albeit less drastic reduction in chloroplast ATP synthase accumulation compared to atp4 null mutants in maize. Additionally, a reduced ribosome association of atpB/E mRNAs was found in svr7 mutants. Different from atp4 mutants, SVR7 mutations lead in addition to an impaired rbcL translation. Further commonalities and differences found in the accumulation of other proteins of the thylakoid membrane and in chloroplast transcription and transcript accumulation are discussed in respect to findings in previously described mutants of the mono- and eudicotyledonous ATP4/SVR7/P67 orthologs (Lahmy et al. 2000; Liu et al. 2010; Zoschke et al. 2012).
Materials and methods
Plant material
SVR7 T-DNA insertion and transposon lines were ordered from the Nottingham Arabidopsis Stock Center and the Cold Spring Harbor Laboratory, respectively (seed stocks: CS819547 and GT20858). The locations of insertions in SVR7 were determined by sequencing. Plants homozygous for the insertions were identified by PCR based genotyping (primers see Table 1). Wild-type tissue was obtained from the genotyped wild-type offspring of heterozygous SVR7 mutant lines.
Plants were grown in a soil vermiculite mixture (4:1) in a growth chamber with long-day regime (16 h light [130 μmol m−2 s−1], 23 °C/8 h dark, 23 °C). Leaf tissue was harvested 30 days after sowing.
Norflurazon treatment was essentially carried out as described by Koussevitzky et al. (2007). Plants were grown on Murashige and Skoog medium supplemented with sucrose (3 %) and with or without Norflurazon (5 μM) for 8 days with long day regime (see above) before harvesting (Murashige and Skoog 1962).
Protein analyses
Proteins were isolated as described by Barkan (1998). Protein concentrations of leaf extracts were determined by protein assay measurements following manufacturer’s instructions (Bio-Rad). Proteins were size fractionated by SDS-PAGE (12 % gels), blotted onto mixed ester nitrocellulose membranes, and immuno-probed (Sambrook and Russell 2001). Antibodies used in this study were described by Roy and Barkan (1998). The quantification of immunological analyses was described previously (Zoschke et al. 2010).
Polysome analyses were carried out as described by Barkan (1993). PCR probes for polysome RNA gel blot analyses were amplified with primers listed in Table 1.
RNA analyses
RNA was extracted from leaf tissue with TRIzol reagent (Invitrogen) following manufacturer’s instructions. RNA gel blot analyses were achieved as described previously (Beick et al. 2008). PCR probes were amplified with primers given in Table 1. Labeling of PCR probes was performed with [α-32P]dCTP using the DecaLabel DNA labeling Kit (Fermentas) following manufacturer’s instructions. Run-on analyses were carried out as described (Zoschke et al. 2007; Zubo et al. 2011). 5 × 108 chloroplasts were lysed and used for in vitro transcription reactions at 25 °C for 10 min. Primers used for macro-array probe amplification are shown in Table 1. For reverse transcription reactions, the QuantiTect Rev. Transcription Kit (Qiagen) was used following manufacturer’s instructions.
Results and discussion
SVR7 mutation impairs chloroplast biogenesis and causes reduced ATP synthase levels in the Arabidopsis thaliana ecotypes Columbia and Landsberg erecta
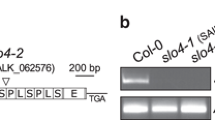
Arabidopsis Ds-transposon and T-DNA lines (generated in Landsberg erecta and Columbia accessions, respectively) with insertions in the SVR7 coding region were identified in public seed banks by database searches. The analyzed svr7 mutant alleles carry insertions upstream of the region coding for the ten PPR domains and within the sequence of the C-terminal SMR domain and are named svr7-2 and svr7-3 (Fig. 1a), respectively. svr7-1 represents a previously described mutant allele of SVR7 that leads to truncation of the reading frame within the PPR tract (Liu et al. 2010). Inbred offspring of lines carrying the mutant alleles svr7-2 and svr7-3, respectively, were PCR-genotyped and the locations of the insertions were determined by sequencing (data not shown, Fig. 1a). svr7 mutants homozygous for the alleles svr7-2 and svr7-3 are slightly pale green and developmentally retarded (Fig. 1b). However, these svr7 mutants are viable and fertile. Loss of SVR7 expression was confirmed by random primed reverse transcription and subsequent PCR amplification of the SVR7 mRNA (Fig. 2). SVR7 cDNA amplifications with primers spanning the insertions yielded a product of the expected size in wild-type derived material. No product was detected with cDNAs derived from homozygous mutants. A control amplification with actin-mRNA specific primers demonstrated successful cDNA synthesis (Fig. 2). Contamination with genomic DNA was excluded by control reactions without reverse transcriptase (Fig. 2). Weak amplification signals were observed in the svr7 mutants using amplification primers up- or downstream of the insertion, which could represent transcripts originating from promoters within the insertions. In any case, both insertions disrupt the SVR7 reading frame and generate premature stop codons at protein positions 144 and 632 downstream of the start codon, respectively.
Overview of svr7 mutant alleles. a Map of the SVR7 gene. The positions of the Ds transposon and T-DNA insertion in each of the two analyzed mutant alleles are indicated above the map (2 and 3, respectively). The DNA sequences flanking the insertions are given below. The 9-bp target site duplication generated by the Ds-insertion is underlined. Residue numbers indicate the positions relative to the start codon of SVR7. svr7-3 contains two inversely oriented T-DNAs at the shown insertion site. b Phenotype of plants of the indicated genotypes that were grown in soil in a growth chamber for 3 weeks (WT = wild-type). Note that Ds transposon lines are generated in an Arabidopsis thaliana Landsberg erecta background (Ler) whereas T-DNA insertion lines are generated in an Arabidopsis thaliana Columbia background (Col). Homozygous svr7 mutants of both alleles show a slightly pale green and developmentally retarded phenotype in comparison to wild-type plants obtained from the offspring of heterozygous mutants
Reverse-Transcription PCR analysis of SVR7 mRNA accumulation in svr7 mutant alleles. 2 μg total leaf RNA from 30 days old plants of the indicated genotypes were reverse transcribed with a random primed Reverse Transcriptase (RT +). The resulting cDNA was PCR amplified (30 cycles) with primers located within the SVR7 cDNA (location as indicated, for primer sequences see Table 1) and size fractionated by agarose gel electrophoresis. The Actin8 cDNA was amplified to control for the efficiency of the reverse transcription reaction. To check PCR reactions for genomic DNA contaminations, mock controls without Reverse Transcriptase (RT −) were carried out as well. All lanes come from the same gel; irrelevant lanes are removed from the image (vertical lines). For all analyzed regions within the SVR7 mRNA svr7 mutants accumulate lower amounts of the SVR7 transcript compared to the respective wild-type plants (WT = wild type, Ler = Landsberg erecta, Col = Columbia). No amplification products could be obtained downstream of the svr7-3 T-DNA insertion site with cDNA derived from wild-type and mutant tissue
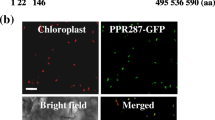
SVR7 and its radish and maize orthologs P67 and ATP4 were previously shown to be chloroplast localized (Lahmy et al. 2000; Liu et al. 2010; Lurin et al. 2004; Zoschke et al. 2012). ATP4 is involved in the expression of chloroplast encoded ATP synthase subunits (Zoschke et al. 2012). Thus the accumulation of the ATP synthase and the other photosystem complexes in svr7 mutants was analyzed by quantitative immunoblot analysis of the representative chloroplast encoded subunits AtpA/-B/-E/-F (ATP synthase), PsaD (photosystem I, PSI), D1 (photosystem II, PSII), and PetD (cytochrome b 6 f complex, Cyt b 6 f). In svr7 mutants a distinct and strong reduction in the accumulation of ATP synthase subunits A, B, F, and E to ~50 to ~10 % of wild-type levels was found (Fig. 3). In contrast PsaD and D1, representative subunits of PS I and II, were only slightly reduced in svr7 mutants (to ~80–90 % of wild-type levels) and the accumulation of PetD, a component of the Cyt b 6 f complex was not affected by SVR7 mutation. A slight decrease of the large subunit of Rubisco is indicated by Ponceau-stains of Western blots (Fig. 3). The specific reduction of the ATP synthase accompanied by minor or no effects on the other photosystem complexes (PSII, PSI, Cyt b 6 f) was also observed in knockouts of the SVR7 ortholog ATP4 in maize (Zoschke et al. 2012). The ATP synthase reduction accounts for the defect in chloroplast biogenesis in maize atp4 mutants and could also cause the observed slightly impaired chloroplast biogenesis in Arabidopsis svr7 mutants. These data are at variance with the semi-quantitative x-ray film measurements presented previously for the svr7-1 mutant allele, which show a reduced accumulation of AtpA, D1, PsaF, and RbcL to ~50 % of wild-type levels, without a specific effect on the ATP synthase (Liu et al. 2010). The mutant allele svr7-1 is chemically induced and contains amino acid exchanges and a premature stop codon within the third PPR domain (Liu et al. 2010), which should cause a knock out of SVR7 function, similar to mutations in svr7-2 and -3 alleles. Arabidopsis plants used for analysis of the svr7-1 mutant allele were grown in constant light (100 μmol m−2 s−1) whereas plants used in this study were grown under a long day regime [16 h light (130 μmol m−2 s−1)/8 h dark]. It is known that an impaired chloroplast ATP synthase activity gives rise to acidification of the thylakoid lumen since the proton gradient generated by PSII and Cyt b 6 f is not used (Maiwald et al. 2003; Rott et al. 2011). This acidification provokes the down-regulation of the expression of nuclear and plastid encoded photosystem components (Maiwald et al. 2003; Rott et al. 2011). Thus, a primary effect of SVR7 mutation on chloroplast ATP synthase accumulation could entail secondary effects on the accumulation of other photosystem complexes under conditions with continuous illumination as used for growing svr7-1 mutants (Liu et al. 2010) and might thus explain the observed differences in the accumulation of chloroplast proteins between the previous and the current study. Future comparative studies analyzing the available svr7 mutants under different light conditions may help to further clarify this question.
Quantitative immunoblot analysis of subunits of photosynthetic enzyme complexes. a 10 μg of total leaf protein extracts from plants of the indicated genotypes were analyzed by immunoblot assays (WT = wild type, Ler = Landsberg erecta, Col = Columbia). The blots were probed with antisera to four subunits of the chloroplast ATP synthase (AtpA, -B, -E, -F) and also with antisera to representative subunits of photosystem I (PsaD), photosystem II (D1), and the cytochrome b 6 f complex (PetD). Bottom: Ponceau S stains of the blots served as loading controls (RbcL marked by asterisks). Note that the Ponceau stained blots here and in c show a reduction of the large subunit of Rubisco (RbcL) in svr7 mutants. b The chemiluminescence signals from the analysis shown in a and from two biological replicate experiments were recorded using a Lumi-Imager (Roche) and quantified using the Quantity One software (Bio-Rad). The signals for wild-type samples were set to one and the means of the mutant signals were calculated and plotted relative to this wild-type signal. Error bars are based on three biological replicates. c Dilution series of wild-type and svr7 mutant protein extracts were analyzed side-by-side with an AtpB specific antiserum. Signals were quantified as described in a to verify the quantitative protein gel blot analyses shown in a and b. Upper panel Quantification of the chemiluminogram shown below. The undiluted wild-type samples were set to 1.0. Quantified signals from diluted samples correspond well with the dilution factor. Only at low sample concentrations (<10 %) is appreciable variation from the calculated values observed. Bottom: Ponceau S stain (RbcL marked by asterisks)
Mild effects of SVR7 mutation on the accumulation of chloroplast atpF, atpH, and psaJ transcripts
The SVR7 maize ortholog ATP4 is involved in processing of atpF and psaJ transcripts (Zoschke et al. 2012). To check for a similar function of SVR7 and find the reason for the reduction of ATP synthase subunits in svr7 mutants, the transcript accumulations of atpB/E, atpH, atpF, and psaJ were determined by RNA gel blot analysis (Fig. 4). In Arabidopsis the dicistronic atpB/E transcript originates from two promoters upstream of the atpB coding region, and additionally, a monocistronic atpE mRNA is transcribed from a promoter within atpB (Malik Ghulam et al. 2012; Schweer et al. 2006). In RNA gel blot hybridization experiments, a slight over-accumulation of the large dicistronic atpB/E transcript over other transcripts in the same sample was found in svr7 mutants (asterisk in Fig. 4). This resembles the atpB/E over-accumulation detected in maize mutants of the SVR7 ortholog ATP4 (Zoschke et al. 2012).
RNA gel blot analysis of plastid RNAs encoding ATP synthase subunits and PsaJ. Total leaf RNA (5 μg/lane) from plants of the indicated genotypes was size fractionated on denaturing agarose gels, blotted onto Nylon membranes and analyzed by hybridization to radiolabeled DNA probes corresponding to plastid atpB/E, atpH, atpF, and psaJ genes (primers see Table 1). Irrelevant lanes were removed (shown by vertical lines). Transcripts over- or under-accumulating in svr7 mutants are marked with asterisks or waves, respectively. The positions of RNA markers (knt = kilonucleotides) are shown. Methylene blue staining of rRNAs is shown below the blots and serves as loading control (bottom panels). Note the over-accumulation of chloroplast rRNA precursors (~2.9 and ~2.4 knt, marked with black triangles) and the slightly reduced accumulation of chloroplast 23S-rRNA breakdown products (23S#) compared to the normal accumulation of chloroplast 16S-rRNA and cytoplasmic 18S- and 25S-rRNAs in svr7 mutants
In plant chloroplasts, atpI, -H, -F, and -A are encoded in one operon, which is transcribed from several promoters and the derived polycistronic transcripts undergo an extensive processing to shorter units (Malik Ghulam et al. 2012; Miyagi et al. 1998; Pfalz et al. 2009; Stahl et al. 1993; Swiatecka-Hagenbruch et al. 2007; Zhelyazkova et al. 2012). RNA gel blot analyses with atpH- and atpF-specific probes revealed changes in steady-state levels of these transcripts in svr7 mutants (Fig. 4). A reduction of a ~1.5 kilonucleotide (knt) transcript representing the monocistronic unspliced atpF mRNA was detected with atpF exon- and intron-specific probes. The knockout of the SVR7 ortholog ATP4 in maize affects also atpF transcript accumulation. In atp4 mutants all transcripts with 3′ ends in the atpF/-A intergenic region accumulate to lower levels (Zoschke et al. 2012). By contrast, svr7 mutants display an increase in several atpH and atpF containing transcripts (Fig. 4).
PsaJ is a small non essential subunit of PSI which is encoded in an operon together with petL, petG, rpl33, and rpl18 (Pfalz et al. 2009; Schöttler et al. 2007). RNA gel blot analyses showed an over-accumulation of different psaJ containing transcripts in svr7 mutants (Fig. 4). Interestingly, the processed monocistronic psaJ mRNA, which is strongly reduced in maize atp4 mutants (Zoschke et al. 2012) is increased in svr7 mutants.
Altogether, the detected processing defects caused by SVR7 mutation are mild and most likely not the only reason for the determined minimum 50 % reduction of ATP synthase subunits, especially because most observed abnormalities are rather increases than reductions of transcript steady-state levels. Whether the detected dissimilarities between svr7 and atp4 mutants are caused by evolutionary diverged functions of the orthologous PPR-SMR proteins or by general processing differences of the analyzed operons in Arabidopsis and maize remains to be answered, e.g., by analyzing mutants of SVR7/ATP4 orthologs in other mono- and eudicotyledonous model plants such as rice (Oryza sativa) and tobacco (Nicotiana tabacum).
SVR7 mutation impairs the translation of atpB/E and rbcL transcripts
Some PPR proteins are known to act as translational activators of chloroplast mRNAs (Barkan et al. 1994; Boulouis et al. 2011; Cai et al. 2011; Pfalz et al. 2009; Zoschke et al. 2012). So far, the mechanism of translational activation has been elucidated only for PPR10. PPR10 binds a region upstream of the atpH start codon and thereby resolves a secondary structure, which constrains ribosome entry by masking the shine dalgarno sequence (Prikryl et al. 2011). The SVR7 ortholog ATP4 activates translation of atpB/E and atpA mRNAs causing a drastic ATP synthase reduction in atp4 mutants (Zoschke et al. 2012). A possible function of SVR7 in translation was examined by polysome analyses. Leaf extracts were fractionated on sucrose gradients under translational elongation inhibiting conditions (Barkan 1993). The more ribosomes are associated with an mRNA in polysomes the deeper the transcript migrates into the gradient. The distribution of atpB/E and rbcL mRNAs among gradient fractions was analyzed by RNA gel blot analysis (Fig. 5a). Methylene blue stained chloroplast rRNAs were found to be similarly distributed in svr7 mutants and wild-type plants, excluding a general translation effect in svr7 chloroplasts. By contrast, a specific shift to fractions of lower sucrose density was detected for the atpB/E and rbcL mRNAs in svr7 mutants suggesting a reduced ribosome loading of these transcripts in the mutants (Fig. 5). Thus, svr7 mutants exhibit a broader translation defect compared to atp4 mutants, which were affected in translation of ATP synthase coding mRNAs but not in that of rbcL (Zoschke et al. 2012).
Polysome-analysis of ribosome association of chloroplast atpB/E, and rbcL mRNAs. a Total leaf extracts from plants of the indicated genotypes were fractionated in sucrose gradients under polysome maintaining conditions. The more ribosomes are associated with an mRNA, the deeper it migrates into the gradient. RNAs were isolated from 12 gradient fractions (1–12, increasing sucrose concentration). Equal proportions of the RNAs from each fraction were size fractionated on denaturing agarose gels, blotted onto Nylon membranes and analyzed by subsequent hybridization to DNA probes detecting atpB/E, and rbcL mRNAs (primers see Table 1). The two dicistronic atpB/E transcripts transcribed from different promoters are labeled by waves or asterisks, respectively (Malik Ghulam et al. 2012; Schweer et al. 2006). A representative blot is shown for each mRNA and genotype. The blots were stained with methylene blue to show distribution of rRNAs (bottom panels), and then probed sequentially to detect the indicated mRNAs. Note the abberant patterns of chloroplast rRNAs in svr7 mutants as described in Fig. 4. b Quantification of the mRNA distribution within the sucrose fractions. The signals from the RNA gel blot analyses shown in a and from one biological replicate were recorded using a Phosphor-Imager (Bio-Rad) and quantified using the Quantity One software (Bio-Rad). For each transcript the percentage of the added signals in all lanes was calculated and plotted against the fractions comparing mutant and wild-type analysis (wild-type data points, blue diamonds, mutant data points, red rectangles). Error bars are based on two biological replicates. The two dicistronic atpB/E transcripts were analyzed separately and are shown in distinct diagrams labeled with waves or asterisks as the transcripts in a. The observed shifts of mRNAs to fractions of lower sucrose density in svr7 mutants suggest a reduced ribosome association of those mRNAs
It was previously proposed that a general translation defect caused by the svr7-1 mutant allele is the reason for suppression of the variegated phenotype of var2 mutants (Liu et al. 2010). The authors speculated that the detected chloroplast rRNA processing defects in svr7-1 mutants lead to impaired chloroplast translation. Indeed, also svr7-2 and -3 mutant alleles cause equal rRNA processing defects as seen by aberrant patterns of methylene blue stained chloroplast rRNAs in RNA gel blot analyses (Figs. 4, 5a). However, an over-accumulation of unprocessed chloroplast rRNA precursors was shown for several mutants with impaired chloroplast biogenesis (e.g. Barkan 1993; Beick et al. 2008, Schmitz-Linneweber et al. 2006, Zybailov et al. 2009). Moreover, the rRNA processing defects determined in mutants of the SVR7 ortholog ATP4 did not affect overall translation. These defects were found also in other mutants and consequently considered as unspecific effects linked to impaired chloroplast biogenesis (Zoschke et al. 2012). Similarly, compared to the reduction of ATP synthase subunits, the accumulation of D1, PsaD, and PetD proteins is almost unaffected by svr7-2 and -3 mutations, making a general defect in chloroplast translation in svr7 mutants unlikely (Fig. 3a, b).
To precisely unravel the role of SVR7 in chloroplast translation, a detailed analysis of the events leading to translation initiation on the atpB/E mRNAs is needed. For this, the determination of the exact binding site of SVR7 would be helpful; however, the available ATP4 antiserum did not detect SVR7 (data not shown) and thus, identification of the RNA targets of SVR7 by RIP-Chip or related techniques is not feasible at present.
Unaltered chloroplast transcription activity in svr7 mutants
The PPR-SMR proteins GUN1 and pTAC2 and other proteins with SMR domain were suggested to play roles in transcription (Fukui and Kuramitsu 2011; Koussevitzky et al. 2007; Pfalz et al. 2006). pTAC2 was isolated from transcriptional active chloroplast extracts and ptac2 mutants show an impaired accumulation and transcription of PEP derived transcripts (Pfalz et al. 2006). GUN1 has an unspecific DNA binding activity and shows sub-organellar co-localization with pTAC2 (Koussevitzky et al. 2007). To examine a possible involvement of SVR7 in chloroplast transcription, run-on labeling assays were carried out with svr7-2 and -3 mutant and wild-type chloroplasts (Fig. 6). Equal numbers of chloroplasts were used for in vitro labeling by transcriptional incorporation of [α-32P]-UTP. Labeled transcripts were hybridized to macro-arrays with probes for representative chloroplast genes encoding components of the ATP synthase, PSI, PSII, the Cyt b 6 f complex, Rubisco, and the translation apparatus (Fig. 6a, b). None of the 13 analyzed chloroplast genes was found to be affected in its transcription activity in svr7-2 and -3 mutants (Fig. 6c). Solely the 23S rRNA gene showed a ~30 % reduction in transcription activity in one of the two analyzed svr7 mutants, in the Landsberg background. However, the observed effects on protein accumulation, RNA processing, and translation caused by SVR7 mutations were consistently observed in both svr7 mutants (Figs. 3, 4, 5). Thus, it is unlikely that those defects are caused by an impaired transcription of the rrn23 gene. Furthermore, pronounced differences in transcription activities of several genes were observed between Arabidopsis plants of Landsberg erecta and Columbia accessions (Fig. 6c). Thus it is possible that the impaired chloroplast gene expression caused by SVR7 mutation leads to distinct secondary effects on transcription of the 23S rRNA gene in the analyzed Arabidopsis accessions.
Run-on analysis of chloroplast transcription activity in svr7 mutants. a Overview of probe arrangement on macroarrays with duplicate dots of DNA probes for plastid genes (as indicated), and control probes for the nuclear 18S-rRNA gene (rrn18), and the mitochondrial cytochrome c biogenesis orf203 gene (ccb203). b Equal amounts of isolated chloroplasts from wild-type and svr7 mutant plants were used for run-on transcription assays (10 min elongation). 32P-labeled transcripts were isolated and hybridized to macroarrays containing plastid and control gene probes as shown in a. c Signals from macroarray analyses shown in b and from one biological replicate were recorded using a Phosphor-Imager (Bio-Rad) and quantified using the Quantity One software (Bio-Rad). For each probe and genotype means of 4 replicates (two biological, and two technical) were calculated and plotted against the respective gene. Data for the representative dot blots shown in b may be slightly different from the averages of replicate experiments shown in c
SVR7 mutation does not disrupt Norflurazon induced retrograde signaling
GUN1, an SVR7 related chloroplast PPR-SMR protein, was shown to be involved in chloroplast to nucleus signaling (Koussevitzky et al. 2007). To test a similar function of SVR7 we examined the expression of a nuclear gene coding for a subunit of the chloroplast light harvesting complex (LhcB1.2). LhcB genes are under the control of retrograde signals and no longer expressed if chloroplast biogenesis is impaired (Oelmüller et al. 1986; Oelmüller and Mohr 1986). In GUN (genomes uncoupled) mutants this signaling is disrupted and LhcB expression still takes place despite the damage of chloroplast development (Mochizuki et al. 2001; Susek et al. 1993). We impaired chloroplast biogenesis by growing plants on Norflurazon (NF), which leads to an increased reactive oxygen stress and bleaching of plant tissue (Fig. 7). Under these conditions, expression of LhcB is not rescued by the SVR7 mutation, suggesting that SVR7 is not involved in transmitting at least the NF-generated retrograde signal.
Analysis of svr7 mutants for a genomes uncoupled (GUN) phenotype after Norflurazon treatment. a Arabidopsis wild-type (WT; Ler = Landsberg erecta) and svr7-2/-2 mutant plants were grown for 8 days on Murashige and Skoog medium supplemented with Sucrose (3 %) and with (+) or without (−) Norflurazon (NF, 5 μM). Norflurazon treated plants show albinotic phenotypes caused by inhibition of carotenoid biosynthesis. b 5 μg total RNA isolated from Norflurazon-treated or untreated plants were size-fractionated by agarose gel electrophoresis, transferred to Nylon membrane, and probed with a DNA-probe specific to an LhcB mRNA (light harvesting complex B). All lanes come from the same gel, irrelevant lanes are removed from the image (vertical lines). Methylene blue staining of total RNA is shown below the blot and serves as loading control. Norflurazon treatment causes the repression of LhcB expression in wild-type and svr7 mutant plants (Oelmüller et al. 1986; Oelmüller and Mohr 1986). An uncoupling of this feedback is typical for the GUN phenotype (Mochizuki et al. 2001; Susek et al. 1993), but was not observed in svr7 mutants. The assay was repeated once with the same result
SVR7 and its maize ortholog ATP4: evolutionary implications of conserved and diverged mutant defects
In the few instances in which orthologous proteins involved in chloroplast biogenesis have been studied in Arabidopsis and maize so far, their functions are largely conserved (Asakura and Barkan 2006, 2007; Asakura et al. 2008). In case of the orthologous PPR-SMR proteins SVR7 and ATP4 this appears to be only partially true. Both maize atp4 and Arabidopsis svr7 mutants exhibit impaired atpF transcript patterns, reduced accumulation of ATP synthase subunits and diminished ribosome association with the atpB/E mRNA. The observed common mutation effects together with the high degree in identity/similarity of the sequences strongly substantiate orthology (Zoschke et al. 2012). However, there are also remarkable differences in chloroplast gene expression between svr7 and atp4 mutants (Zoschke et al. 2012): Several atpF/-H and psaJ transcripts, which over-accumulate in svr7 mutants, are in fact not altered or even reduced in atp4 mutants. Additionally, the reduced accumulation of ATP synthase subunits is less pronounced in svr7 mutants compared to atp4 mutants and the translational defects caused by SVR7 mutation include also RbcL and thus appear to be broader than that of atp4 mutants.
Furthermore, the phenotypic consequences of SVR7 and ATP4 mutations are remarkably different. Whereas the knockout of ATP4 causes pale green phenotypes and seedling lethality, the mutations of SVR7 account only for slightly pale green and developmentally retarded but viable and fertile plants (Liu et al. 2010; Zoschke et al. 2012). These phenotypic variations could be explained by differences in the accumulation of ATP synthase subunits. In svr7 mutants, the ATP synthase is much less reduced than in atp4 mutants and it is known that even strongly reduced ATP synthase contents are sufficient for the plant’s energy supply by an upregulated activity of the remaining ATP synthase complexes (Rott et al. 2011). Consequently, reduced ATP synthase levels in svr7 mutants could be sufficient for viability of the plants whereas the severe ATP synthase reduction in atp4 mutants causes seedling lethality after exhaustion of the seed storage. Nevertheless, it is remarkable that the phenotypic effects of the Arabidopsis mutants are far less severe than the ones in maize, because in other cases, where knock-outs of PPR proteins have been investigated in orthologous genes between maize and Arabidopsis, the phenotypes in Arabidopsis were more drastic (e.g. Lurin et al. 2004; Schmitz-Linneweber and Small 2008; Tzafrir et al. 2004). For example, loss of PPR4 and -5 lead to albino seedlings in maize, while in Arabidopsis, plants die as early embryos (Beick et al. 2008; Cushing et al. 2005; Schmitz-Linneweber et al. 2006). These mutants have global defects in chloroplast translation, whereas ATP4/SVR7 mutations cause specific defects. Possibly, the ATP synthase activity is differently regulated in eudicotyledonous plants and thus less susceptible to changes in its expression levels than in monocotyledonous plants.
Alternatively, the protein functions of SVR7 and ATP4 maybe diverged during evolution. Finally, SVR7 functions in Arabidopsis might overlap with one or more other protein(s), which partially complement SVR7 mutation thus extenuating the effects compared to ATP4 mutations. Further studies are needed to discriminate between these alternatives.
Abbreviations
- Col:
-
Columbia (Arabidopsis accession)
- Cyt b 6 f :
-
Cytochrome b 6 f complex
- GUN:
-
Genomes uncoupled
- Ler:
-
Landsberg erecta (Arabidopsis accession)
- LHC:
-
Light harvesting complex
- NF:
-
Norflurazon
- PPR:
-
Pentatricopeptide repeat (protein domain)
- PSI/II:
-
Photosystem I/II
- RT-PCR:
-
Reverse transcription PCR
- SMR:
-
Small MutS related (protein domain)
- WT:
-
Wild-type
References
Asakura Y, Barkan A (2006) Arabidopsis orthologs of maize chloroplast splicing factors promote splicing of orthologous and species-specific group II introns. Plant Physiol 142:1656–1663
Asakura Y, Barkan A (2007) A CRM domain protein functions dually in group I and group II intron splicing in land plant chloroplasts. Plant Cell 19:3864–3875
Asakura Y, Bayraktar OA, Barkan A (2008) Two CRM protein subfamilies cooperate in the splicing of group IIB introns in chloroplasts. RNA 14:2319–2332
Barkan A (1993) Nuclear mutants of maize with defects in chloroplast polysome assembly have altered chloroplast RNA metabolism. Plant Cell 5:389–402
Barkan A (1998) Approaches to investigating nuclear genes that function in chloroplast biogenesis in land plants. Methods Enzymol 297:38–57
Barkan A (2011) Expression of plastid genes: organelle-specific elaborations on a prokaryotic scaffold. Plant Physiol 155:1520–1532
Barkan A, Walker M, Nolasco M, Johnson D (1994) A nuclear mutation in maize blocks the processing and translation of several chloroplast mRNAs and provides evidence for the differential translation of alternative mRNA forms. EMBO J 13:3170–3181
Beick S, Schmitz-Linneweber C, Williams-Carrier R, Jensen B, Barkan A (2008) The pentatricopeptide repeat protein PPR5 stabilizes a specific tRNA precursor in maize chloroplasts. Mol Cell Biol 28:5337–5347
Bock R, Timmis JN (2008) Reconstructing evolution: gene transfer from plastids to the nucleus. BioEssays 30:556–566
Boulouis A, Raynaud C, Bujaldon S, Aznar A, Wollman FA, Choquet Y (2011) The nucleus-encoded trans-acting factor MCA1 plays a critical role in the regulation of cytochrome f synthesis in Chlamydomonas chloroplasts. Plant Cell 23:333–349
Cai W, Okuda K, Peng L, Shikanai T (2011) PROTON GRADIENT REGULATION 3 recognizes multiple targets with limited similarity and mediates translation and RNA stabilization in plastids. Plant J 67:318–327
Cushing DA, Forsthoefel NR, Gestaut DR, Vernon DM (2005) Arabidopsis emb175 and other ppr knockout mutants reveal essential roles for pentatricopeptide repeat (PPR) proteins in plant embryogenesis. Planta 221:424–436
de Longevialle AF, Hendrickson L, Taylor NL, Delannoy E, Lurin C, Badger M, Millar AH, Small I (2008) The pentatricopeptide repeat gene OTP51 with two LAGLIDADG motifs is required for the cis-splicing of plastid ycf3 intron 2 in Arabidopsis thaliana. Plant J 56:157–168
Delannoy E, Stanley WA, Bond CS, Small ID (2007) Pentatricopeptide repeat (PPR) proteins as sequence-specificity factors in post-transcriptional processes in organelles. Biochem Soc Trans 35:1643–1647
Echeverria M, Lahmy S (1995) Identification of a 67 kDa protein that binds specifically to the pre-rRNA primary processing site in a higher plant. Nucleic Acids Res 23:4963–4970
Fukui K, Kuramitsu S (2011) Structure and function of the small MutS-related domain. Mol Biol Int 2011:691735
Gillman JD, Bentolila S, Hanson MR (2007) The petunia restorer of fertility protein is part of a large mitochondrial complex that interacts with transcripts of the CMS-associated locus. Plant J 49:217–227
Hammani K, des Francs-Small cc, Takenaka M, Tanz SK, Okuda k, Shikanai T, Brennicke A, Small I (2011) The pentatricopeptide repeat protein OTP87 is essential for RNA editing of nad7 and atp1 transcripts in Arabidopsis mitochondria. J Biol Chem 286:21361–21371
Hattori M, Miyake H, Sugita M (2007) A pentatricopeptide repeat protein is required for RNA processing of clpP pre-mRNA in moss chloroplasts. J Biol Chem 282:10773–10782
Kleine T, Maier UG, Leister D (2009) DNA transfer from organelles to the nucleus: the idiosyncratic genetics of endosymbiosis. Annu Rev Plant Biol 60:115–138
Koussevitzky S, Nott A, Mockler TC, Hong F, Sachetto-Martins G, Surpin M, Lim J, Mittler R, Chory J (2007) Signals from chloroplasts converge to regulate nuclear gene expression. Science 316:715–719
Lahmy S, Barneche F, Derancourt J, Filipowicz W, Delseny M, Echeverria M (2000) A chloroplastic RNA-binding protein is a new member of the PPR family. FEBS Lett 480:255–260
Liu X, Yu F, Rodermel S (2010) An Arabidopsis pentatricopeptide repeat protein, SUPPRESSOR OF VARIEGATION7, is required for FtsH-mediated chloroplast biogenesis. Plant Physiol 154:1588–1601
Lurin C, Andres C, Aubourg S, Bellaoui M, Bitton F, Bruyere C, Caboche M, Debast C, Gualberto J, Hoffmann B, Lecharny A, Le Ret M, Martin-Magniette ML, Mireau H, Peeters N, Renou JP, Szurek B, Taconnat L, Small I (2004) Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 16:2089–2103
Maiwald D, Dietzmann A, Jahns P, Pesaresi P, Joliot P, Joliot A, Levin JZ, Salamini F, Leister D (2003) Knock-out of the genes coding for the Rieske protein and the ATP-synthase delta-subunit of Arabidopsis. Effects on photosynthesis, thylakoid protein composition, and nuclear chloroplast gene expression. Plant Physiol 133:191–202
Malik Ghulam M, Zghidi-Abouzid O, Lambert E, Lerbs-Mache S, Merendino L (2012) Transcriptional organization of the large and the small ATP synthase operons, atpI/H/F/A and atpB/E, in Arabidopsis thaliana chloroplasts. Plant Mol Biol 79:259–272
Margulis L (1970) Origin of eukaryotic cells : evidence and research implications for a theory of the origin and evolution of microbial, plant, and animal cells on the Precambrian earth. Yale University Press, New Haven
Martin W (2003) Gene transfer from organelles to the nucleus: frequent and in big chunks. Proc Natl Acad Sci USA 100:8612–8614
Miyagi T, Kapoor S, Sugita M, Sugiura M (1998) Transcript analysis of the tobacco plastid operon rps2/atpI/H/F/A reveals the existence of a non-consensus type II (NCII) promoter upstream of the atpI coding sequence. Mol Gen Genet 257:299–307
Mochizuki N, Brusslan JA, Larkin R, Nagatani A, Chory J (2001) Arabidopsis genomes uncoupled 5 (GUN5) mutant reveals the involvement of Mg-chelatase H subunit in plastid-to-nucleus signal transduction. Proc Natl Acad Sci USA 98:2053–2058
Moreira D, Philippe H (1999) Smr: a bacterial and eukaryotic homologue of the C-terminal region of the MutS2 family. Trends Biochem Sci 24:298–300
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassay with tobacco tissue culture. Physiol Plant 15:473–497
Oelmüller R, Mohr H (1986) Photooxidative destruction of chloroplasts and its consequences for expression of nuclear genes. Planta 167:106–113
Oelmüller R, Levitan I, Bergfeld R, Rajasekhar VK, Mohr H (1986) Expression of nuclear genes as affected by treatments acting on the plastids. Planta 168:482–492
Okuda K, Nakamura T, Sugita M, Shimizu T, Shikanai T (2006) A pentatricopeptide repeat protein is a site recognition factor in chloroplast RNA editing. J Biol Chem 281:37661–37667
Pfalz J, Liere K, Kandlbinder A, Dietz KJ, Oelmuller R (2006) pTAC2, -6, and -12 are components of the transcriptionally active plastid chromosome that are required for plastid gene expression. Plant Cell 18:176–197
Pfalz J, Bayraktar OA, Prikryl J, Barkan A (2009) Site-specific binding of a PPR protein defines and stabilizes 5′ and 3′ mRNA termini in chloroplasts. EMBO J 28:2042–2052
Prikryl J, Rojas M, Schuster G, Barkan A (2011) Mechanism of RNA stabilization and translational activation by a pentatricopeptide repeat protein. Proc Natl Acad Sci USA 108:415–420
Rott M, Martins NF, Thiele W, Lein W, Bock R, Kramer DM, Schöttler MA (2011) ATP synthase repression in tobacco restricts photosynthetic electron transport, CO2 assimilation, and plant growth by overacidification of the thylakoid lumen. Plant Cell 23:304–321
Roy LM, Barkan A (1998) A SecY homologue is required for the elaboration of the chloroplast thylakoid membrane and for normal chloroplast gene expression. J Cell Biol 141:385–395
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schmitz-Linneweber C, Small I (2008) Pentatricopeptide repeat proteins: a socket set for organelle gene expression. Trends Plant Sci 13:663–670
Schmitz-Linneweber C, Williams-Carrier R, Barkan A (2005) RNA immunoprecipitation and microarray analysis show a chloroplast pentatricopeptide repeat protein to be associated with the 5′ region of mRNAs whose translation it activates. Plant Cell 17:2791–2804
Schmitz-Linneweber C, Williams-Carrier RE, Williams-Voelker PM, Kroeger TS, Vichas A, Barkan A (2006) A pentatricopeptide repeat protein facilitates the trans-splicing of the maize chloroplast rps12 pre-mRNA. Plant Cell 18:2650–2663
Schöttler MA, Flügel C, Thiele W, Stegemann S, Bock R (2007) The plastome-encoded PsaJ subunit is required for efficient photosystem I excitation, but not for plastocyanin oxidation in tobacco. Biochem J 403:251–260
Schweer J, Loschelder H, Link G (2006) A promoter switch that can rescue a plant sigma factor mutant. FEBS Lett 580:6617–6622
Small ID, Peeters N (2000) The PPR motif—a TPR-related motif prevalent in plant organellar proteins. Trends Biochem Sci 25:46–47
Stahl DJ, Rodermel SR, Bogorad L, Subramanian AR (1993) Co-transcription pattern of an introgressed operon in the maize chloroplast genome comprising four ATP synthase subunit genes and the ribosomal rps2. Plant Mol Biol 21:1069–1076
Stern DB, Goldschmidt-Clermont M, Hanson MR (2010) Chloroplast RNA metabolism. Annu Rev Plant Biol 61:125–155
Susek RE, Ausubel FM, Chory J (1993) Signal transduction mutants of Arabidopsis uncouple nuclear CAB and RBCS gene expression from chloroplast development. Cell 74:787–799
Suzuki JY, Ytterberg AJ, Beardslee TA, Allison LA, Wijk KJ, Maliga P (2004) Affinity purification of the tobacco plastid RNA polymerase and in vitro reconstitution of the holoenzyme. Plant J 40:164–172
Swiatecka-Hagenbruch M, Liere K, Börner T (2007) High diversity of plastidial promoters in Arabidopsis thaliana. Mol Genet Genom 277:725–734
Tasaki E, Hattori M, Sugita M (2010) The moss pentatricopeptide repeat protein with a DYW domain is responsible for RNA editing of mitochondrial ccmFc transcript. Plant J 62:560–570
Tzafrir I, Pena-Muralla R, Dickerman A, Berg M, Rogers R, Hutchens S, Sweeney TC, McElver J, Aux G, Patton D, Meinke D (2004) Identification of genes required for embryo development in Arabidopsis. Plant Physiol 135:1206–1220
Williams-Carrier R, Kroeger T, Barkan A (2008) Sequence-specific binding of a chloroplast pentatricopeptide repeat protein to its native group II intron ligand. RNA 14:1930–1941
Wobbe L, Schwarz C, Nickelsen J, Kruse O (2008) Translational control of photosynthetic gene expression in phototrophic eukaryotes. Physiol Plant 133:507–515
Woodson JD, Chory J (2008) Coordination of gene expression between organellar and nuclear genomes. Nat Rev Genet 9:383–395
Zehrmann A, Verbitskiy D, Hartel B, Brennicke A, Takenaka M (2011) PPR proteins network as site-specific RNA editing factors in plant organelles. RNA Biol 8:67–70
Zhelyazkova P, Sharma CM, Forstner KU, Liere K, Vogel J, Börner T (2012) The primary transcriptome of barley chloroplasts: numerous noncoding RNAs and the dominating role of the plastid-encoded RNA polymerase. Plant Cell 24:123–136
Zoschke R, Liere K, Börner T (2007) From seedling to mature plant: Arabidopsis plastidial genome copy number, RNA accumulation and transcription are differentially regulated during leaf development. Plant J 50:710–722
Zoschke R, Nakamura M, Liere K, Sugiura M, Börner T, Schmitz-Linneweber C (2010) An organellar maturase associates with multiple group II introns. Proc Natl Acad Sci USA 107:3245–3250
Zoschke R, Kroeger T, Belcher S, Schottler MA, Barkan A, Schmitz-Linneweber C (2012) The pentatricopeptide repeat-SMR protein ATP4 promotes translation of the chloroplast atpB/E mRNA. Plant J. doi:10.1111/j.1365-313X.2012.05081.x
Zubo YO, Kusnetsov VV, Börner T, Liere K (2011) Reverse protection assay: a tool to analyze transcriptional rates from individual promoters. Plant Methods 7:47
Zybailov B, Friso G, Kim J, Rudella A, Rodriguez VR, Asakura Y, Sun Q, van Wijk KJ (2009) Large scale comparative proteomics of a chloroplast Clp protease mutant reveals folding stress, altered protein homeostasis, and feedback regulation of metabolism. Mol Cell Proteom 8:1789–1810
Acknowledgments
The authors thank Reik Modrozynski and Simone Hardel for excellent technical support. We acknowledge Alice Barkan for providing the antibodies used in this study. We thank Michael Tillich for the introduction of Norflurazon assays. We acknowledge the Nottingham Arabidopsis Stock Centre and Rob Martienssen from the Cold Spring Harbor Laboratory for providing Arabidopsis seeds for insertion lines. We thank Alice Barkan, Ian Small, Kate Howell, Michael Tillich, and Karsten Liere for helpful discussions. This work was supported by the German Academic Exchange Service, and the German Research Foundation [Grant Numbers SCHM 1698/2-1, ZO 302/1].
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zoschke, R., Qu, Y., Zubo, Y.O. et al. Mutation of the pentatricopeptide repeat-SMR protein SVR7 impairs accumulation and translation of chloroplast ATP synthase subunits in Arabidopsis thaliana . J Plant Res 126, 403–414 (2013). https://doi.org/10.1007/s10265-012-0527-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-012-0527-1